Submitted:
22 January 2024
Posted:
23 January 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Context
Method
Sample Collection and Sequencing
Genome Assembly, Annotation, and Assessment
Results
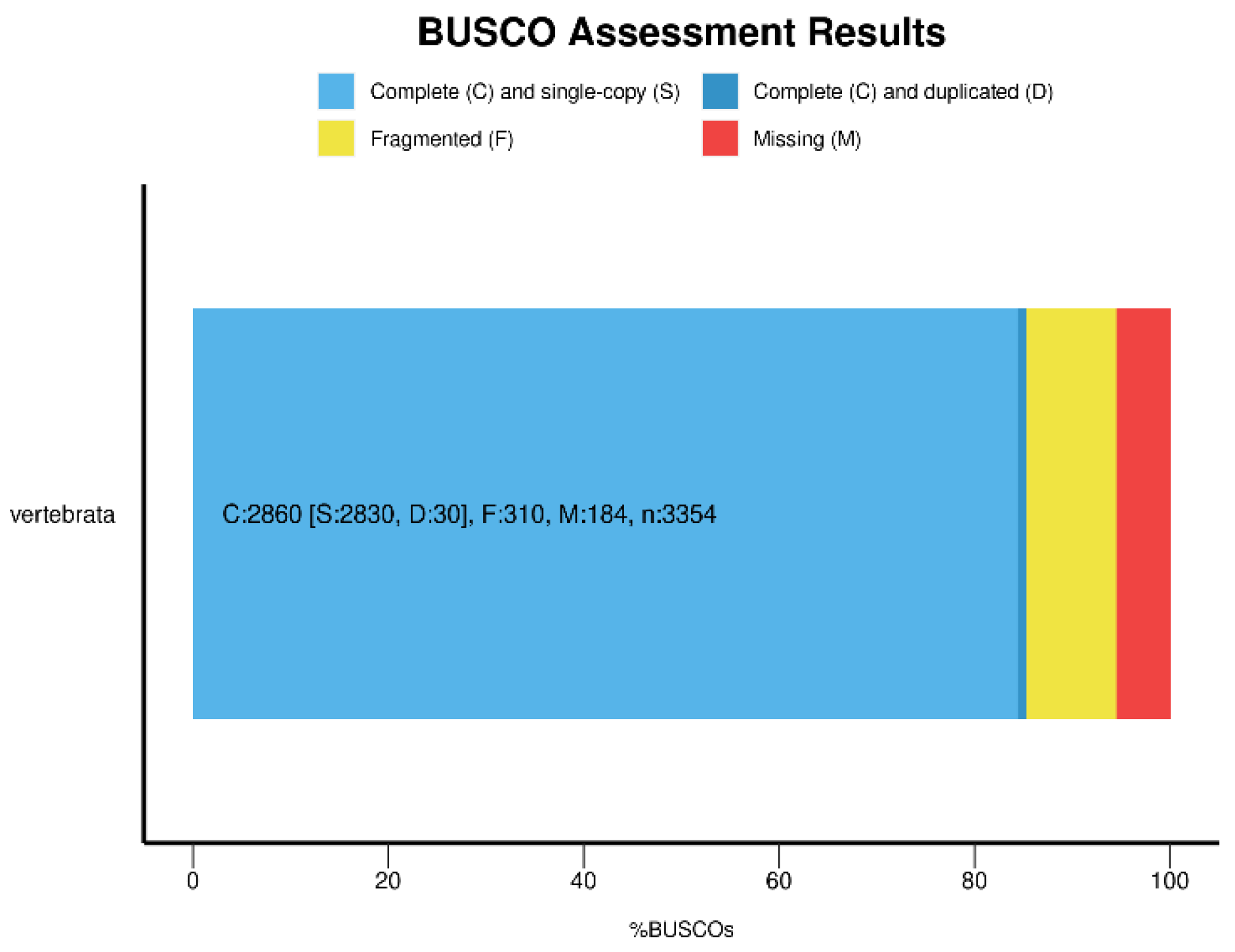
Data Validation and Quality Control
Reuse Potential
Author Contributions
Data Availability Statement
Editor’s note
Ethics approval
Acknowledgments
Competing Interests
Abbrevitions
References
- Gray JE. Synopsis of the species of Rattle snakes, or Family of Crotalidae. Zoological Miscellany 2: 47–51. http://repositorio.fciencias.unam.mx:8080/xmlui/handle/11154/158063.
- Leviton A, Wogan G, Koo M, Zug G, Lucas R, Vindum J. The Dangerously Venomous Snakes of Myanmar. Illustrated Key and Checklist. 542003; https://repository.si.edu/handle/10088/4542.
- Zhu F, Liu Q, Che J, Zhang L, Chen X, Yan F, et al.. Molecular phylogeography of white-lipped tree viper (Trimeresurus; Viperidae). Zoologica Scripta. 2016;. [CrossRef]
- Cockram CS, Chan JC, Chow KY. Bites by the white-lipped pit viper (Trimeresurus albolabris) and other species in Hong Kong. A survey of 4 years’ experience at the Prince of Wales Hospital. J Trop Med Hyg. 93:79–86 1990;
- Liew JL, Tan NH, Tan CH. Proteomics and preclinical antivenom neutralization of the mangrove pit viper (Trimeresurus purpureomaculatus, Malaysia) and white-lipped pit viper (Trimeresurus albolabris, Thailand) venoms. Acta Tropica. 2020;. [CrossRef]
- Jangprasert P, Rojnuckarin P. Molecular cloning, expression and characterization of albolamin: A type P-IIa snake venom metalloproteinase from green pit viper (Cryptelytrops albolabris). Toxicon. 2014;. [CrossRef]
- Pinyachat A, Rojnuckarin P, Muanpasitporn C, Singhamatr P, Nuchprayoon S. Albocollagenase, a novel recombinant P-III snake venom metalloproteinase from green pit viper (Cryptelytrops albolabris), digests collagen and inhibits platelet aggregation. Toxicon. 2011;. [CrossRef]
- Tan CH. Snake Venomics: Fundamentals, Recent Updates, and a Look to the Next Decade. Toxins. Multidisciplinary Digital Publishing Institute; 2022;. [CrossRef]
- Olaoba OT, Karina Dos Santos P, Selistre-de-Araujo HS, Ferreira de Souza DH. Snake Venom Metalloproteinases (SVMPs): A structure-function update. Toxicon X. 2020;. [CrossRef]
- Zhu F, Chen L, Guo P, Xu Y, Liu Q. Sexual Dimorphism and Geographic Variation of the White-lipped Pit Viper (Trimeresurus albolabris) in China. Current Herpetology. 2022;. [CrossRef]
- Malhotra A, Thorpe RS. A Phylogeny of the Trimeresurus Group of Pit Vipers: New Evidence from a Mitochondrial Gene Tree. Molecular Phylogenetics and Evolution. 2000;. [CrossRef]
- Casewell NR, Wüster W, Vonk FJ, Harrison RA, Fry BG. Complex cocktails: the evolutionary novelty of venoms. Trends in Ecology & Evolution. 2013;. [CrossRef]
- Calvete JJ. Venomics: integrative venom proteomics and beyond*. Biochemical Journal. 2017;. [CrossRef]
- Rao W, Kalogeropoulos K, Allentoft ME, Gopalakrishnan S, Zhao W, Workman CT, et al.. The rise of genomics in snake venom research: recent advances and future perspectives. GigaScience. 2022;. [CrossRef]
- Song T, Zhang C, Zhang L, Huang X, Hu C, Xue C, et al.. Complete mitochondrial genome of Trimeresurus albolabris (Squamata: Viperidae: Crotalinae). Mitochondrial DNA. 2015;. [CrossRef]
- Wang O, Chin R, Cheng X, Wu MKY, Mao Q, Tang J, et al.. Efficient and unique cobarcoding of second-generation sequencing reads from long DNA molecules enabling cost-effective and accurate sequencing, haplotyping, and de novo assembly. Genome Res. 2019;. [CrossRef]
- Liu B, Cui L, Deng Z, Ma Y, Yang D, Gong Y, et al.. The annotation pipeline for the genome of a snake. protocols.io. 2023;. [CrossRef]
- Weisenfeld NI, Kumar V, Shah P, Church DM, Jaffe DB. Direct determination of diploid genome sequences. Genome Res. 2017;. [CrossRef]
- Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J, et al.. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience. 2012;. [CrossRef]
- Pryszcz LP, Gabaldón T. Redundans: an assembly pipeline for highly heterozygous genomes. Nucleic Acids Res. 2016;. [CrossRef]
- Benson G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Research. 1999;. [CrossRef]
- Xu Z, Wang H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Research. 2007;. [CrossRef]
- Smit AFA, Hubley R, Green P. RepeatModeler Open-1.0. 2008–2015. Seattle, USA: Institute for Systems Biology. http://www.repeatmasker.org, Last Accessed May. 1:20182015;
- Tarailo-Graovac M, Chen N. Using RepeatMasker to identify repetitive elements in genomic sequences Curr. Protoc. Bioinformatics, 2009; 25: 4.10.1–4.10.14. [CrossRef]
- Tempel S. Using and understanding RepeatMasker. Mobile genetic elements: protocols and genomic applications. Methods Mol. Biol., 2012; 859: 29–51. [CrossRef]
- Stanke M, Steinkamp R, Waack S, Morgenstern B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Research. 2004;. [CrossRef]
- Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;. [CrossRef]
- Haas BJ, Papanicolaou A, Yassour M, Grabherr M, Blood PD, Bowden J, et al.. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc. 2013;. [CrossRef]
- Haas BJ, Salzberg SL, Zhu W, Pertea M, Allen JE, Orvis J, et al.. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 2008;. [CrossRef]
- Mount DW. Using the Basic Local Alignment Search Tool (BLAST). Cold Spring Harb Protoc. 2007;. [CrossRef]
- Birney E, Clamp M, Durbin R. GeneWise and Genomewise. Genome Res. 2004;. [CrossRef]
- Campbell MS, Holt C, Moore B, Yandell M. Genome Annotation and Curation Using MAKER and MAKER-P. Current Protocols in Bioinformatics. 2014;. [CrossRef]
- Manni M, Berkeley MR, Seppey M, Simão FA, Zdobnov EM. BUSCO Update: Novel and Streamlined Workflows along with Broader and Deeper Phylogenetic Coverage for Scoring of Eukaryotic, Prokaryotic, and Viral Genomes. Mol Biol Evol. 2021;. [CrossRef]
- Emms DM, Kelly S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biology. 2015;. [CrossRef]
- Yin W; Wang Z; Li Q; Lian J; Zhou Y; Lu B; Jin L; Qiu P; Zhang P; Zhu W; Wen B; Huang Y; Lin Z; Qiu B; Su X; Zhang G; Yan G; Zhou Q (2016): Supporting data for "Evolutionary trajectories of snake genes and genomes revealed by comparative analyses of five-pacer viper" GigaScience Database. [CrossRef]
- Margres MJ, Rautsaw RM, Strickland JL, Mason AJ, Schramer TD, Hofmann EP, et al.. The Tiger Rattlesnake genome reveals a complex genotype underlying a simple venom phenotype. Proceedings of the National Academy of Sciences. Proceedings of the National Academy of Sciences; 2021;. [CrossRef]
- Jones P, Binns D, Chang H-Y, Fraser M, Li W, McAnulla C, et al.. InterProScan 5: genome-scale protein function classification. Bioinformatics. 2014;. [CrossRef]
- Kanehisa M. The KEGG Database. ‘In Silico’ Simulation of Biological Processes. John Wiley & Sons, Ltd; [CrossRef]
- Bairoch A, Apweiler R. The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Research. 2000;. [CrossRef]
- Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics. 2015;. [CrossRef]
- Guo X, Chen F, Gao F, Li L, Liu K, You L, et al. CNSA: a data repository for archiving omics data. Database. 2020;. [CrossRef]
- Chen FZ, You LJ, Yang F, Wang LN, Guo XQ, Gao F, et al.. CNGBdb: China National GeneBank DataBase. Yi Chuan (Hereditas,). 2020;. [CrossRef]
- Niu X; Lv Y; Chen J; Feng Y; Cui Y; Lu H; Liu H (2024): Supporting data for "The genome assembly and annotation of the White-Lipped Tree viper Trimeresurus albolabris" GigaScience Database. [CrossRef]
- Snake Genomes. GigaByte. 2023; [CrossRef]


| WGS-1 | WGS-2 | WGS-3 | ||||
| fq1 | fq2 | fq1 | fq2 | fq1 | fq2 | |
| %Q20 | 96.98 | 97.81 | 97.74 | 94.96 | 95.69 | 97.59 |
| %Q30 | 90.79 | 90.6 | 92.83 | 84.27 | 84.46 | 89.81 |
| %GC | 40.37 | 40.21 | 41.02 | 40.81 | 40.36 | 40.47 |
| %ErrorRate | 0.351809 | 0.233019 | 0.264481 | 0.540272 | 0.449364 | 0.257064 |
| TotalReads | 492,445,828 | 425,689,572 | 104,911,172 | |||
| TotalBases | 98,489,165,600 | 85,137,914,400 | 20,982,234,400 | |||
| stLFR-1 | stLFR-2 | RNA-seq | ||||
| fq1 | fq2 | fq1 | fq2 | fq1 | fq2 | |
| %Q20 | 96.53 | 95.59 | 96.41 | 96.3 | 98.3 | 98.19 |
| %Q30 | 89.83 | 87.26 | 87.98 | 86.37 | 94.3 | 93.71 |
| %GC | 39.34 | 42.16 | 39.28 | 42.15 | 44.11 | 44.07 |
| %ErrorRate | 0.403065 | 0.525486 | 0.442228 | 0.392415 | 0.194523 | 0.205665 |
| TotalReads | 633,976,833 | 161,105,172 | 50,828,075 | |||
| TotalBases | 145,814,671,590 | 37,054,189,560 | 10,165,615,000 | |||
| contigs | contigs >(1,000bp) | contigs >(10,000bp) | |
| Total number (>) | 71,131 | 46608 | 10,016 |
| Total length of (bp) | 1,513,852,334 | 1,501,212,553 | 1,355,102,082 |
| N50 Length (bp) | 381,553 | ||
| N75 Length (bp) | 115,212 | ||
| GC content is (%) | 39.97 | ||
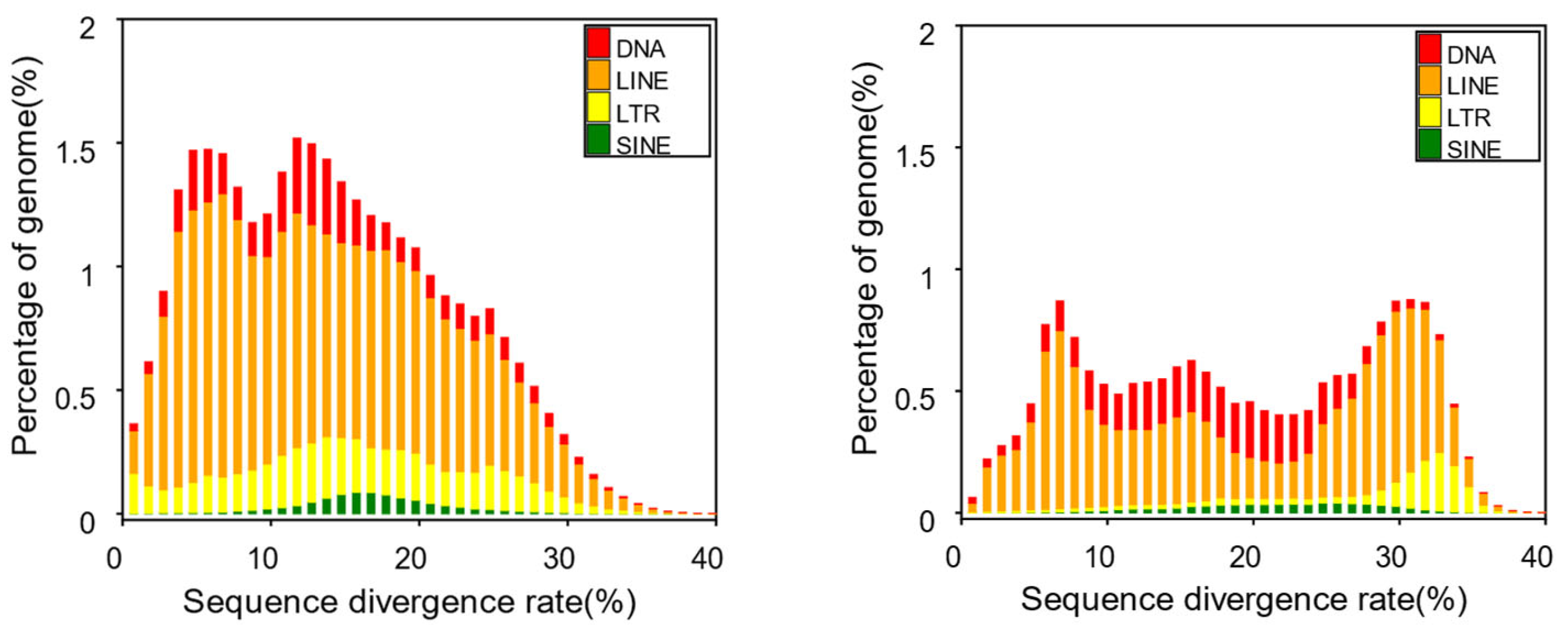
| Type | Repeat Size | % of genome |
| Trf | 47,767,541 | 3.155363 |
| Repeatmasker | 252,985,952 | 16.711402 |
| Proteinmask | 185,792,360 | 12.272819 |
| De novo | 498,353,737 | 32.919574 |
| Total | 581,568,803 | 38.416482 |
| Type | Repbase TEs | TE protiens | De novo | Combined TEs | ||||
| Length (Bp) | % in genome | Length (Bp) | % in genome | Length (Bp) | % in genome | Length (Bp) | % in genome | |
| DNA | 51,357,881 | 3.392529 | 2,032,636 | 0.134269 | 64,605,374 | 4.267614 | 104,513,127 | 6.903786 |
| LINE | 184,866,441 | 12.211656 | 157,414,659 | 10.398284 | 294,829,469 | 19.475444 | 362,351,919 | 23.935751 |
| SINE | 9,622,825 | 0.635651 | 0 | 0 | 13,144,889 | 0.868307 | 18,769,499 | 1.23985 |
| LTR | 23,685,560 | 1.564589 | 26,413,572 | 1.744792 | 74,868,256 | 4.945546 | 88,305,953 | 5.833195 |
| Other | 77,658 | 0.00513 | 141 | 0.000009 | 0 | 0 | 77,799 | 0.005139 |
| Unknown | 0 | 0 | 0 | 0 | 98,895,691 | 6.532717 | 98,895,691 | 6.532717 |
| Total | 252,985,952 | 16.711402 | 185,792,360 | 12.272819 | 496,342,637 | 32.786727 | 566,552,754 | 37.424572 |
| Type | Copy(w) | Average length(bp) | Total length(bp) | % of genome |
| miRNA | 250 | 98.992 | 24,748 | 0.001635 |
| tRNA | 179 | 75.70949721 | 13,552 | 0.000895 |
| rRNA | 104 | 137.6057692 | 14,311 | 0.000945 |
| snRNA | 301 | 115.1229236 | 34,652 | 0.002289 |
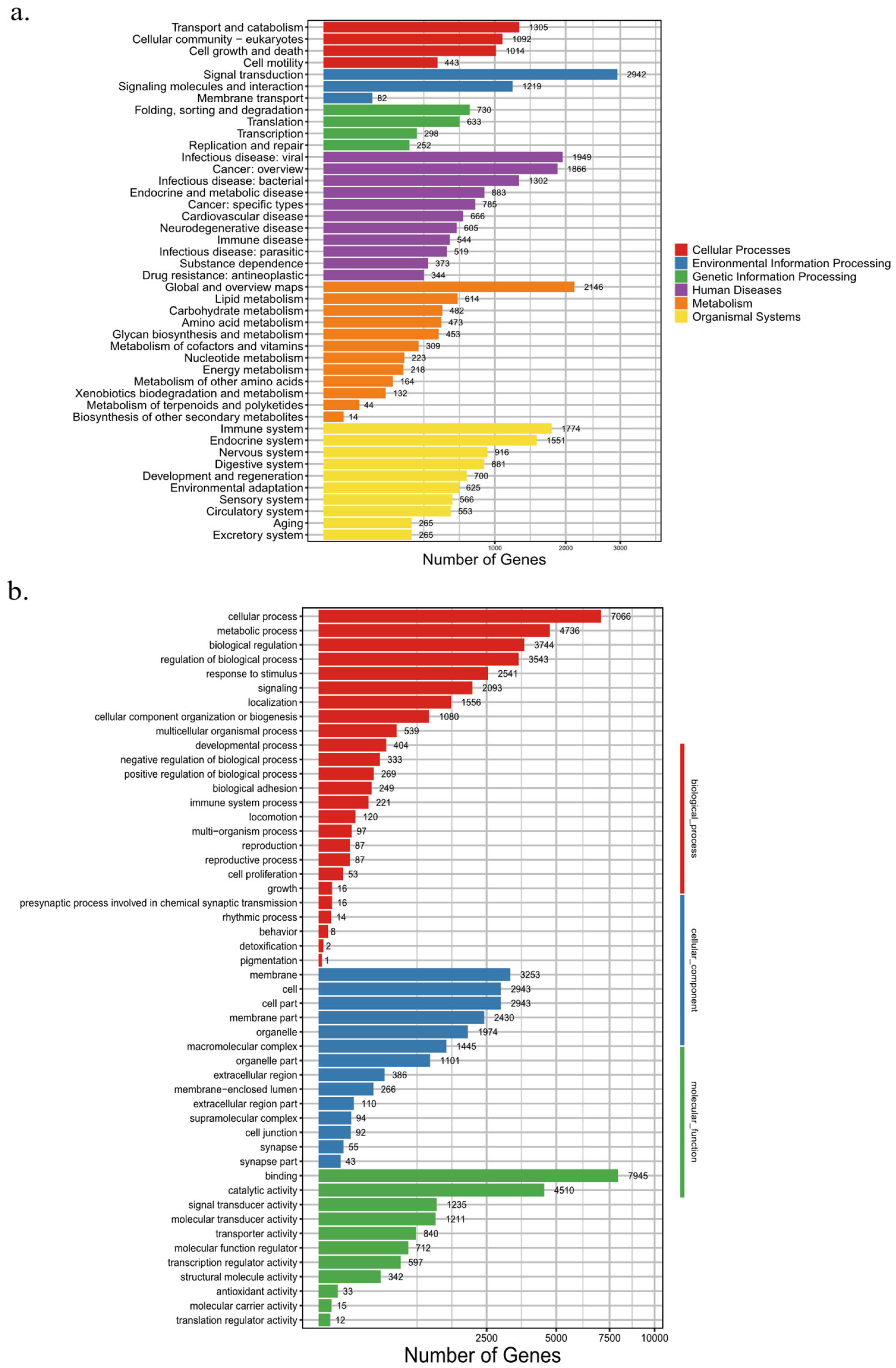
| Values | Total | Swissprot-Annotated | KEGG-Annotated | TrEMBL-Annotated | Interpro-Annotated | GO-Annotated | Overall |
| Number | 21,695 | 20,240 | 19,216 | 21,134 | 21,019 | 14,786 | 21,516 |
| Percentage | 100% | 93.29% | 88.57% | 97.41% | 96.88% | 68.15% | 99.17% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




