Submitted:
23 January 2024
Posted:
24 January 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Methods:
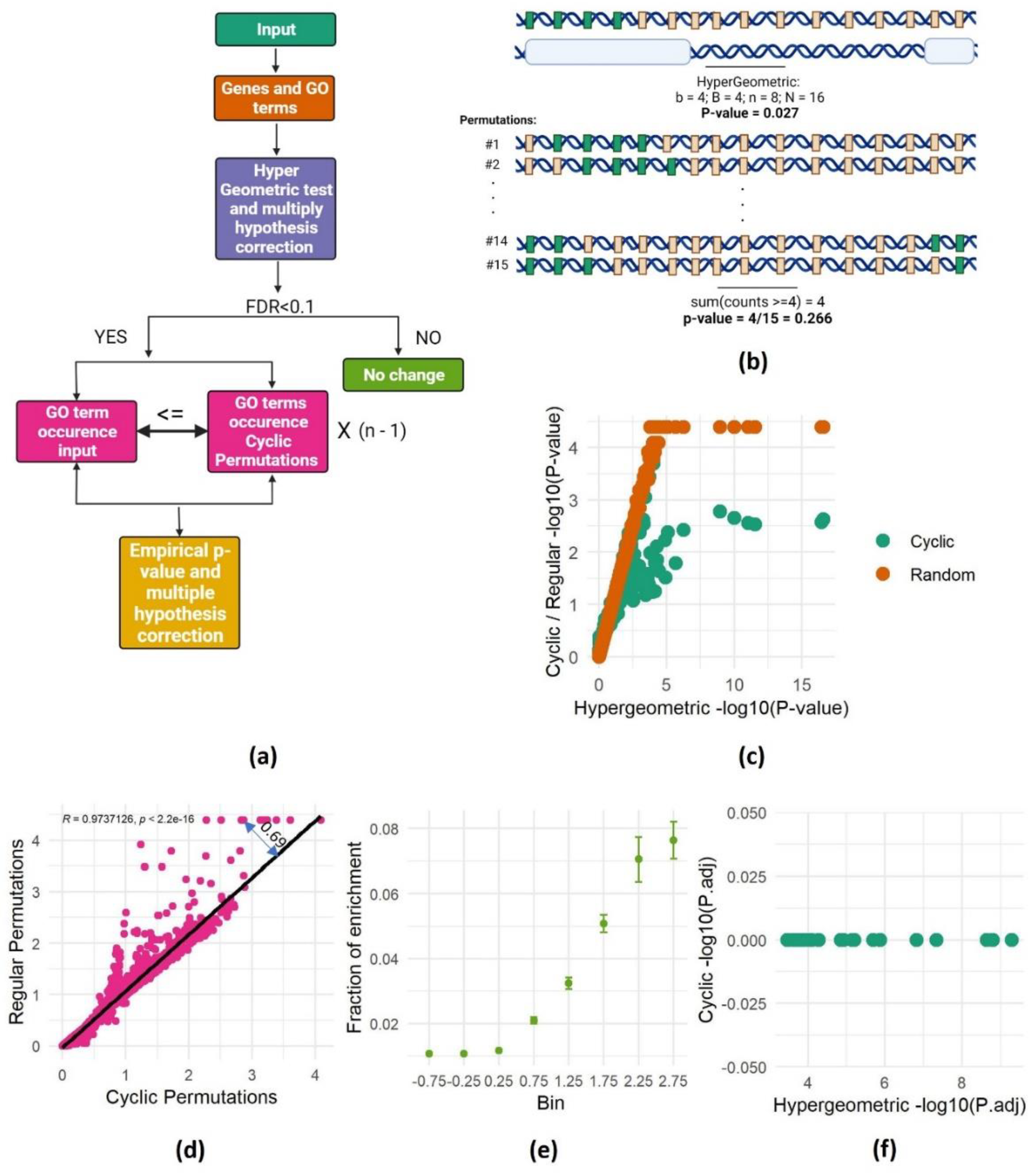
SAGO Pipeline
Hypergeometric Test for GO Terms Enrichment:
Cyclic and Random Permutations:
Linear Regression Analysis:
Random intervals analysis
Data Sources and Processing
Results
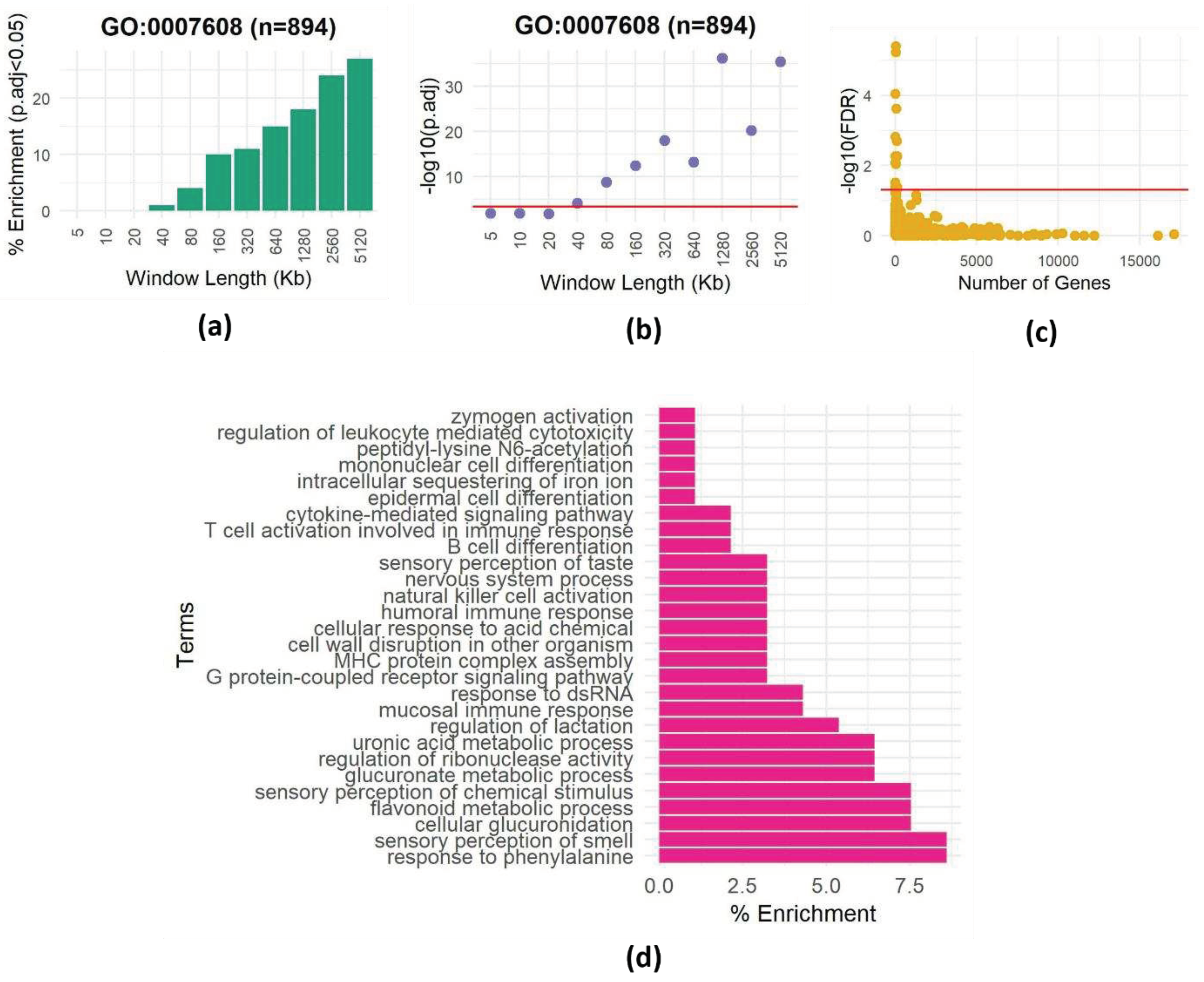
Spatial dependencies affect enrichment analyses.
Multiple hypothesis corrections
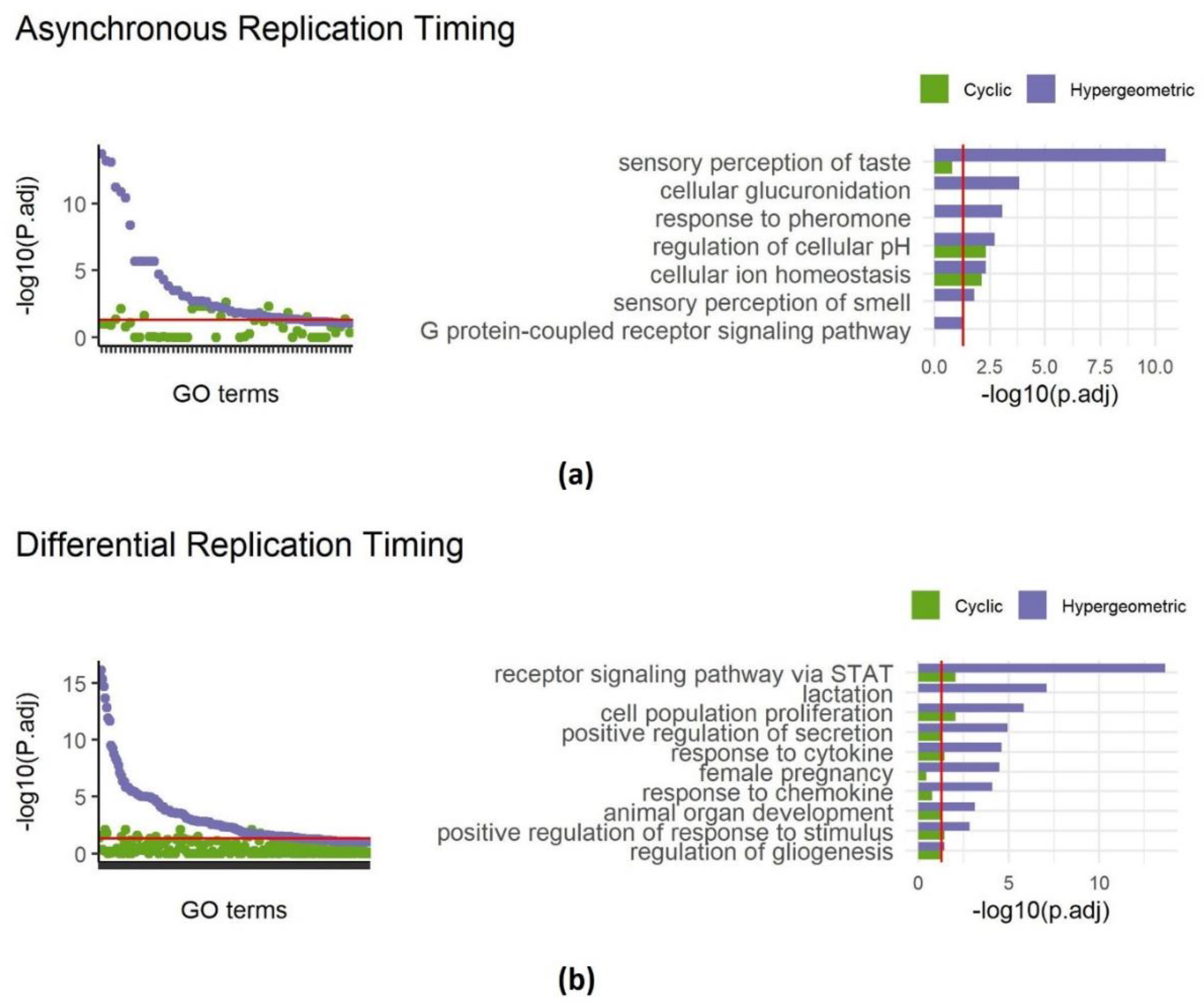
Applying SAGO to replication timing data
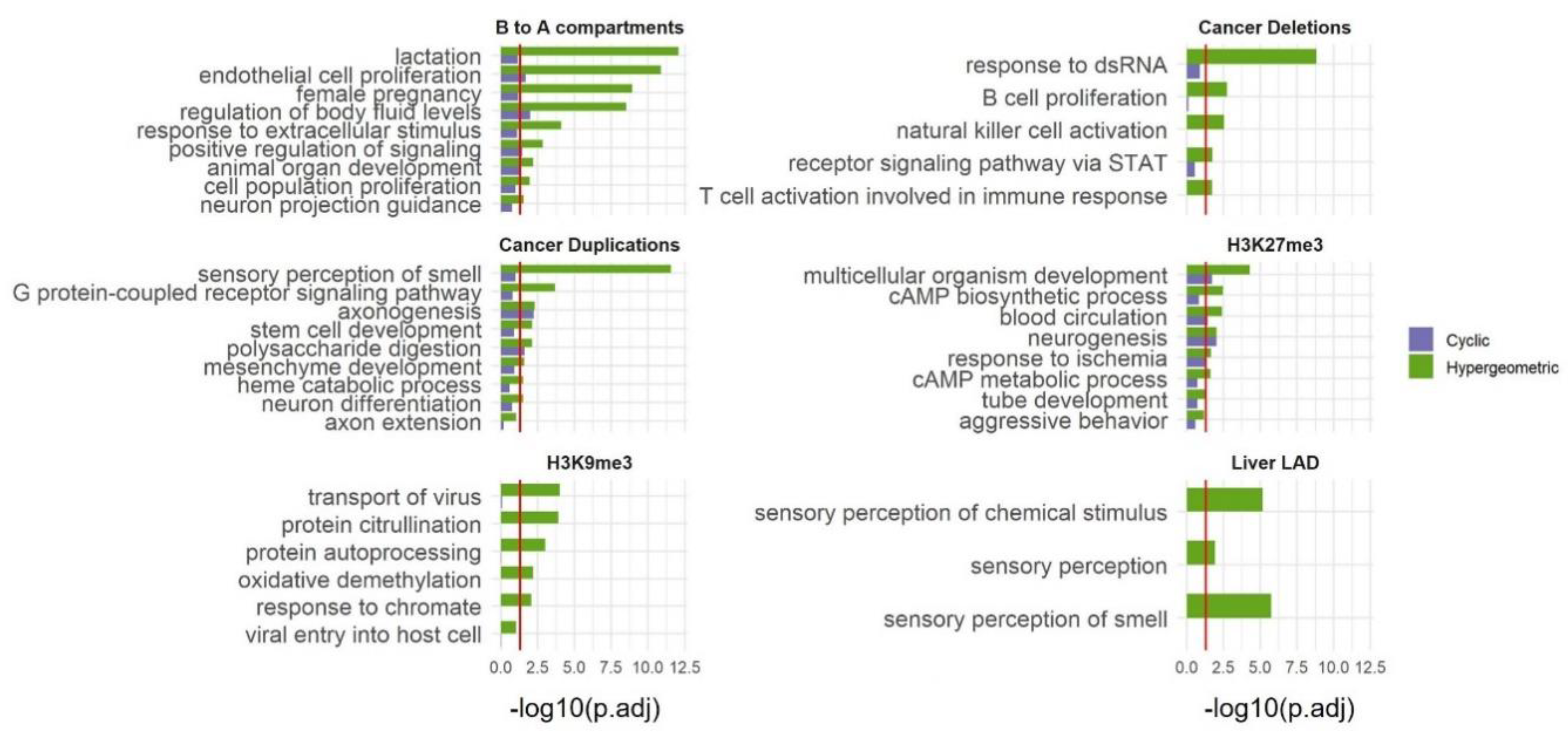
Expanding the use of SAGO to additional types of data
Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ashburner, M., et al., Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet, 2000. 25(1): p. 25-9. [CrossRef]
- Mooney, M.A. and B. Wilmot, Gene set analysis: A step-by-step guide. Am J Med Genet B Neuropsychiatr Genet, 2015. 168(7): p. 517-27.
- Eden, E., et al., GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics, 2009. 10: p. 48. [CrossRef]
- Rivals, I., et al., Enrichment or depletion of a GO category within a class of genes: which test? Bioinformatics, 2007. 23(4): p. 401-7.
- Li, W., et al., Beyond standard pipeline and p < 0.05 in pathway enrichment analyses. Comput Biol Chem, 2021. 92: p. 107455.
- Takebayashi, S.I., M. Ogata, and K. Okumura, Anatomy of Mammalian Replication Domains. Genes (Basel), 2017. 8(4). [CrossRef]
- Poulet, A., et al., RT States: systematic annotation of the human genome using cell type-specific replication timing programs. Bioinformatics, 2019. 35(13): p. 2167-2176. [CrossRef]
- Du, Q., et al., Replication timing and epigenome remodelling are associated with the nature of chromosomal rearrangements in cancer. Nat Commun, 2019. 10(1): p. 416. [CrossRef]
- Kosak, S.T. and M. Groudine, Gene order and dynamic domains. Science, 2004. 306(5696): p. 644-7. [CrossRef]
- Hurst, L.D., C. Pal, and M.J. Lercher, The evolutionary dynamics of eukaryotic gene order. Nat Rev Genet, 2004. 5(4): p. 299-310. [CrossRef]
- Michalak, P., Coexpression, coregulation, and cofunctionality of neighboring genes in eukaryotic genomes. Genomics, 2008. 91(3): p. 243-8. [CrossRef]
- Ben-Elazar, S., Z. Yakhini, and I. Yanai, Spatial localization of co-regulated genes exceeds genomic gene clustering in the Saccharomyces cerevisiae genome. Nucleic Acids Res, 2013. 41(4): p. 2191-201. [CrossRef]
- Elizondo, L.I., et al., Gene clusters, molecular evolution and disease: a speculation. Curr Genomics, 2009. 10(1): p. 64-75. [CrossRef]
- Singer, G.A., et al., Clusters of co-expressed genes in mammalian genomes are conserved by natural selection. Mol Biol Evol, 2005. 22(3): p. 767-75. [CrossRef]
- Lercher, M.J., A.O. Urrutia, and L.D. Hurst, Clustering of housekeeping genes provides a unified model of gene order in the human genome. Nat Genet, 2002. 31(2): p. 180-3. [CrossRef]
- Lee, J.M. and E.L. Sonnhammer, Genomic gene clustering analysis of pathways in eukaryotes. Genome Res, 2003. 13(5): p. 875-82. [CrossRef]
- Tiirikka, T., M. Siermala, and M. Vihinen, Clustering of gene ontology terms in genomes. Gene, 2014. 550(2): p. 155-64. [CrossRef]
- Cabrera, C.P., et al., Uncovering networks from genome-wide association studies via circular genomic permutation. G3 (Bethesda), 2012. 2(9): p. 1067-75. [CrossRef]
- Gel, B., et al., regioneR: an R/Bioconductor package for the association analysis of genomic regions based on permutation tests. Bioinformatics, 2016. 32(2): p. 289-91. [CrossRef]
- Zang, C., Y. Wang, and W. Peng, RECOGNICER: A coarse-graining approach for identifying broad domains from ChIP-seq data. Quant Biol, 2020. 8(4): p. 359-368. [CrossRef]
- Chakraborty, A., J.G. Wang, and F. Ay, dcHiC detects differential compartments across multiple Hi-C datasets. Nat Commun, 2022. 13(1): p. 6827.
- Yehuda, Y., et al., Germline DNA replication timing shapes mammalian genome composition. Nucleic Acids Res, 2018. 46(16): p. 8299-8310. [CrossRef]
- Malnic, B., P.A. Godfrey, and L.B. Buck, The human olfactory receptor gene family. Proc Natl Acad Sci U S A, 2004. 101(8): p. 2584-9.
- Wen, S.H., et al., A two-stage design for multiple testing in large-scale association studies. J Hum Genet, 2006. 51(6): p. 523-532. [CrossRef]
- Consortium, E.P., An integrated encyclopedia of DNA elements in the human genome. Nature, 2012. 489(7414): p. 57-74.
- Luo, Y., et al., New developments on the Encyclopedia of DNA Elements (ENCODE) data portal. Nucleic Acids Res, 2020. 48(D1): p. D882-D889. [CrossRef]
- Sloan, C.A., et al., ENCODE data at the ENCODE portal. Nucleic Acids Res, 2016. 44(D1): p. D726-32. [CrossRef]
- Bonev, B., et al., Multiscale 3D Genome Rewiring during Mouse Neural Development. Cell, 2017. 171(3): p. 557-572 e24. [CrossRef]
- Shah, P.P., et al., An atlas of lamina-associated chromatin across twelve human cell types reveals an intermediate chromatin subtype. Genome Biol, 2023. 24(1): p. 16. [CrossRef]
- Labani, M., et al., PeakCNV: A multi-feature ranking algorithm-based tool for genome-wide copy number variation-association study. Comput Struct Biotechnol J, 2022. 20: p. 4975-4983. [CrossRef]
- Conesa, A., et al., A survey of best practices for RNA-seq data analysis. Genome Biol, 2016. 17: p. 13. [CrossRef]
- Eden, E., et al., Discovering motifs in ranked lists of DNA sequences. PLoS Comput Biol, 2007. 3(3): p. e39. [CrossRef]
- Lazar, N.H., et al., High-resolution genome-wide mapping of chromosome-arm-scale truncations induced by CRISPR-Cas9 editing. bioRxiv, 2023: p. 2023.04.15.537038.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



