Submitted:
09 January 2024
Posted:
10 January 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
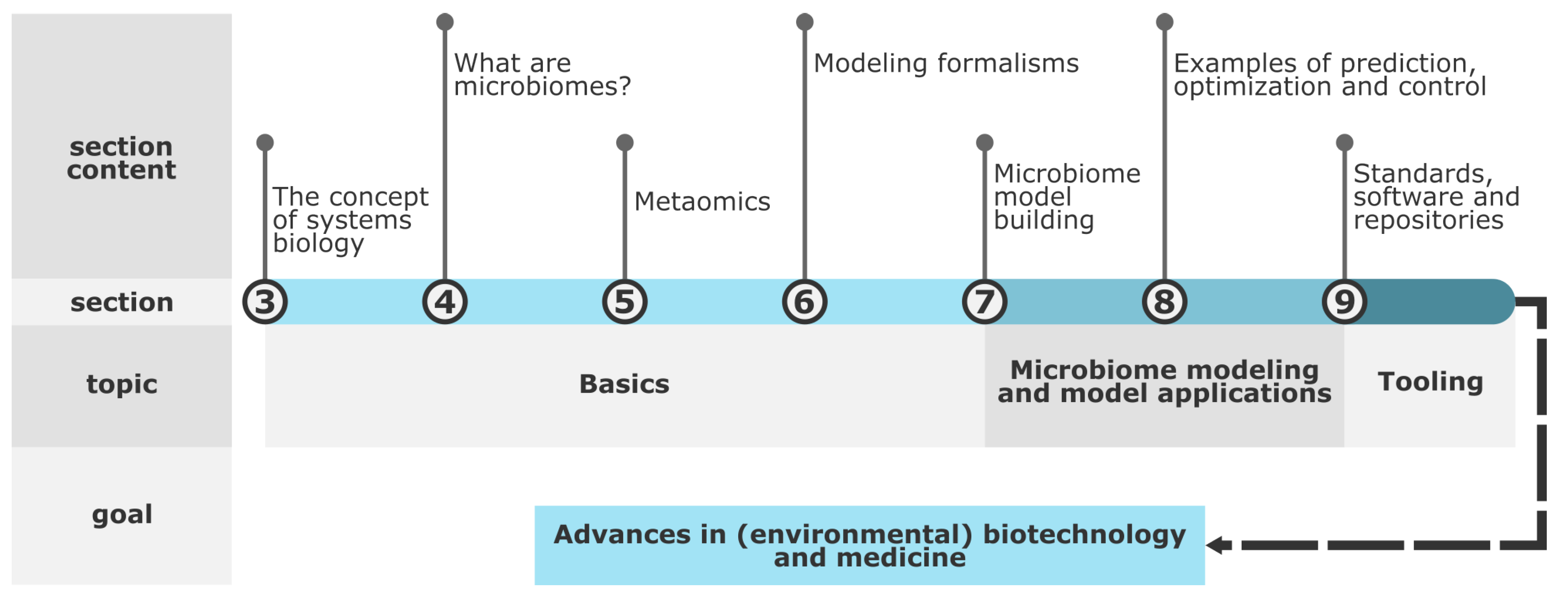
- First, the manuscript gives an interdisciplinary overview of microbiome modeling. To this end, the concept of systems biology (Section 3), microbiome properties (Section 4), metaomics methods (Section 5), and mathematical modeling (Section 6) are explained. In addition, mechanistic model building (Section 7) and their role in predicting, optimizing, and controlling microbiomes (Section 8) are covered. Finally, an overview of guidelines, software, and repositories for microbiome modeling is provided (Section 9).
- Second, metaomics and its peculiarities are explained. Metaomics methods based on liquid chromatography coupled with tandem mass spectrometry (LC-MS/MS) allow the determination of cellular phenotypes. Because it is difficult to cover every meta-omics method, metaproteomics is presented as an exemplary technology. In addition, extensive references to other omics technologies for microbiome analytics are given.
- Third, modeling concepts from metabolic modeling, as well as modeling of signaling and regulation are discussed. While metabolic models are standardized and can reach the scale of genomes [24], modeling of signaling and regulation is less uniform. To fully understand the interference of microbiomes and their hosts, highly resolved models of signaling and regulation are required. The building process of both model types is compared and formalisms that could facilitate genome-scale modeling of signaling and regulation are described.
- Fourth, guidelines facilitating reusability and reproducibility are introduced.
2. Methods
2.1. Targeted Literature Research Strategy
3. The Concept of Systems Biology
3.1. Summary – Section 3
4. What Are Microbiomes?
4.1. Microorganisms and Hosts Interact via Metabolism and Signaling
4.2. Microbiome Characteristics
4.3. Culturing Microorganisms to Model Microbiomes
4.4. Summary – Section 4
5. Meta-Omics Create Inventory Lists of Microbiomes
5.1. The Metaproteomics Workflow
5.2. Taxonomic and Functional Annotation of Protein Groups in Meta-Proteomics
5.3. Other Omics and Experimental Methods for Microbiome Analysis
5.4. Summary – Section 5
6. Mathematical Models Are Formalisms to Describe Biological Mechanisms
6.1. Statistical Models and Mechanistic Models
6.2. Scales of Mechanistic Models
6.3. Mathematical Modeling Frameworks
6.3.1. Graphs
6.3.2. Boolean Models
6.3.3. Models Based on Differential Equations
6.3.4. Constraint-Based Metabolic Models
6.3.5. Rule-Based Models and the Rxncon Language
6.3.6. Combining Model Formalisms
6.4. Summary – Section 6
7. Building and Adjusting Models to (meta)omics Data
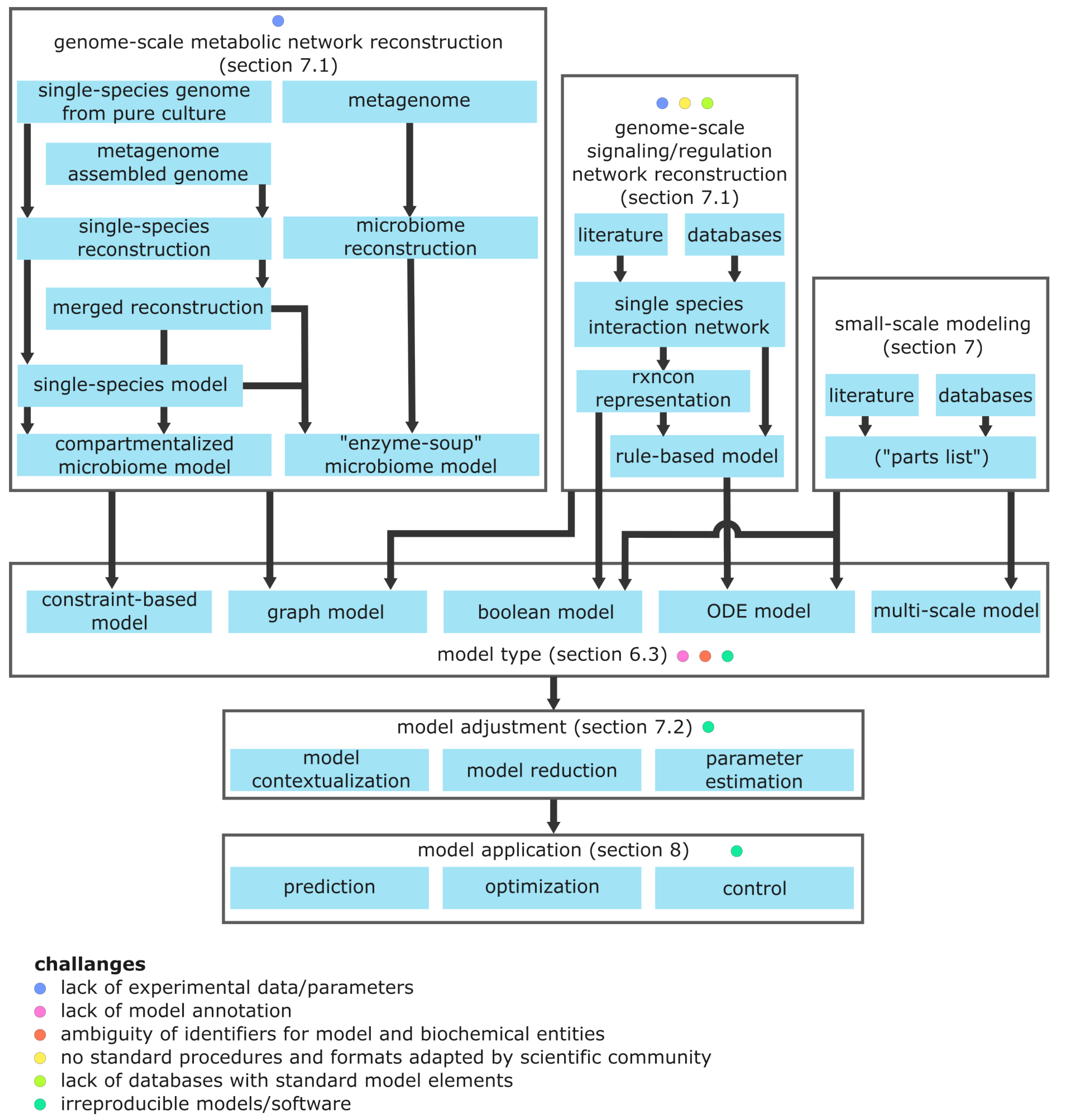
7.1. Reconstruction of Genome-Scale Biological Networks
7.1.1. Reconstruction of Single Species Metabolic Networks
- Draft Reconstruction: Starting point is a whole genome sequence of an organism. The genome is annotated, i.e., genes are linked to transcribed enzymes and transport proteins, which are associated with metabolic reactions. Biochemical databases (e.g., KEGG) can be used to annotate known genes. Genes and reactions are connected in Boolean expressions named gene protein reaction rules. These describe enzymatic subunits required to perform a reaction and facilitate in silico gene knockout analyses. The resulting “parts list” of genes and reactions is generated automatically and represents a draft reconstruction that needs further refinement.
- Refinement: Errors within the reconstruction, such as wrong stoichiometries, wrong cofactor usage, or falsely assigned reactions need to be resolved. This step often requires manual curation linked to extensive literature research and mining of organism-specific databases. Furthermore, processes such as non-growth associated maintenance, recovery of reducing agents, and biomass synthesis (i.e., cell growth) are typically lumped into respective model reactions and added to the reconstruction. For example, substrates of the biomass reaction (Section 6.3.4) are macromolecules or their precursors, whose stoichiometries are determined experimentally from the organism’s macromolecular composition (Section 4.2) [70] or adapted from other organisms. Beck et al. [69] reviewed and evaluated several lab procedures to obtain the macromolecular composition.
- Mathematical model implementation: Thirdly, the network reconstruction is converted to a constraint-based model, which involves creating the stoichiometric matrix, defining compartments, and specifying reaction directionalities.
- Model validation and refinement: The fourth step is a loop of model validation and refinement. The computational model is used to diminish flaws in the reconstruction, for example, missing pathways, or unreachable reactions and metabolites. Furthermore, model constraints can be fine-tuned to biological data (e.g., maximal uptake rates, growth, and non-growth-associated maintenance coefficients). It is ensured that biomass precursors can be synthesized and that the model reproduces relevant growth conditions. The primary reconstruction may contain network gaps, which can be closed by automated, optimization-based gap-filling algorithms. These gap-filling algorithms aim, for example, to identify a minimum set of reactions from a biochemical database enabling the model to simulate growth for different growth media [11]. Another gap-filling approach searches for reactions that support growth, biomass precursor synthesis, utilization of specified alternative energy sources, and metabolite production based on high genetic evidence [190]. After gap-filling, the model might contain blocked reactions (reactions unable to carry any flux), which can be identified using FVA (Section 6.3.4). Manual curation resolves these errors, for example, by adding further reactions. Growth and knockout screenings are used to validate the model output. Lastly, basic model properties, for example, the stoichiometric balance of reactions can be validated with the MEMOTE software [191].
7.1.2. Reconstruction of Microbiome Metabolism
7.1.3. Reconstruction of Signaling Networks
- Draft Reconstruction: Networks are reconstructed in the context of macroscopic behaviors in response to stimuli. Firstly, the inputs and outputs of interest are defined, which helps to restrict the scope of the reconstruction. Secondly, the molecules propagating signals from input to output are identified. In addition, information on the sequence of interactions should be collected [208]. Data on molecular interactions are determined experimentally [144] or predicted [209,210] and are available in interaction databases such as String [162] and scientific literature [42,208]. In non-specific interaction networks (e.g., if retrieved from a database), algorithms can determine potential connections between inputs and outputs [211]. For well-investigated processes, existing signaling networks are available in pathway databases [120,122,212,213,214] and can serve as templates [42]. The result of the first step is an interaction network specific to the defined scope.
- Rxncon model implementation: Firstly, the elemental reactions, involved molecules, and resulting states need to be defined. Secondly, the sequence of signaling events is implemented by defining the contingencies (i.e., conditions) for elemental reactions. Information from expression arrays, knock-out screenings, (meta)omics analyses, databases, and literature provide the required information [41,43,208]. The result of this step is a rxncon model, which is comparable to a metabolic network in the reconstruction state. It represents an interaction network with causal relationships and thus could be analyzed by graph methods [42].
- Boolean model implementation and
- Model validation: Rxncon models can be compiled into Boolean models, which can be validated on experimental data (e.g., reproduction of input-output behavior or activation of internal nodes). If model predictions are not consistent with data, model building is re-iterated from the first or second steps. Additionally, it is possible to compile a rxncon model into a rule-based model and subsequently to an ODE model [208].
7.1.4. A Perspective for Reconstruction of Signaling in Microbiomes
7.2. Parameter Estimation, Model Contextualization and Model Reduction
7.2.1. Parameter Estimation
7.2.2. Contextualization
7.2.3. Model Reduction
7.3. Summary – Section 7
8. Examples of Model-Based Microbiome Prediction, Optimization, and Control
8.1. Predicting and Understanding Microbiomes
8.2. Optimizing Microbiomes
8.3. Controlling Microbiomes
8.3.1. The Concept of Closed-Loop Control
8.3.2. System Inputs and System Outputs of Microbiomes
8.3.3. Control Algorithms and Model-Based Control
8.4. Summary - Section 8
9. Microbiome Modeling Requires Standards, Software and Repositories
9.1. FAIR Data
- Findability (“Datasets should be described, identified and registered or indexed in a clear and unequivocal manner”
- Accessibility (“Datasets should be accessible through a clearly defined access procedure, ideally using automated means. Metadata should always remain accessible.”)
- Interoperability (“Data and metadata are conceptualized, expressed and structured using common, published standards”)
- Reusability (“Characteristics of data and their provenance are described in detail according to domain-relevant community standards, with clear and accessible conditions for use”)
9.2. Initiatives and Community Guidelines
9.3. Languages for Modeling and Exchange Formats
9.4. Software
9.5. Repositories
9.6. Remarks on Languages and Software for Community Modeling
9.7. Summary - Section 9
10. Discussion
Supplementary Materials
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gilbert, J.A.; Blaser, M.J.; Caporaso, J.G.; Jansson, J.K.; Lynch, S.V.; Knight, R. Current understanding of the human microbiome. Nature Medicine 2018, 24, 392–400. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D. Human gut microbiome: hopes, threats and promises. Gut 2018, 67, 1716–1725. [Google Scholar] [CrossRef]
- Helmink, B.A.; Khan, M.A.W.; Hermann, A.; Gopalakrishnan, V.; Wargo, J.A. The microbiome, cancer, and cancer therapy. Nature Medicine 2019, 25, 377–388. [Google Scholar] [CrossRef] [PubMed]
- Jackson, R.B.; Saunois, M.; Bousquet, P.; Canadell, J.G.; Poulter, B.; Stavert, A.R.; Bergamaschi, P.; Niwa, Y.; Segers, A.; Tsuruta, A. Increasing anthropogenic methane emissions arise equally from agricultural and fossil fuel sources. Environmental Research Letters 2020, 15, 071002. [Google Scholar] [CrossRef]
- Naylor, D.; Sadler, N.; Bhattacharjee, A.; Graham, E.B.; Anderton, C.R.; McClure, R.; Lipton, M.; Hofmockel, K.S.; Jansson, J.K. Soil Microbiomes Under Climate Change and Implications for Carbon Cycling. Annual Review of Environment and Resources 2020, 45, 29–59. [Google Scholar] [CrossRef]
- Amann, R.I.; Ludwig, W.; Schleifer, K.H. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiological Reviews 1995, 59, 143–169. [Google Scholar] [CrossRef]
- Wade, W. Unculturable bacteria–the uncharacterized organisms that cause oral infections. JRSM 2002, 95, 81–83. [Google Scholar] [CrossRef]
- Qin, J.; Li, R.; Raes, J.; Arumugam, M.; Burgdorf, K.S.; Manichanh, C.; Nielsen, T.; Pons, N.; Levenez, F.; Yamada, T.; Mende, D.R.; Li, J.; Xu, J.; Li, S.; Li, D.; Cao, J.; Wang, B.; Liang, H.; Zheng, H.; Xie, Y.; Tap, J.; Lepage, P.; Bertalan, M.; Batto, J.M.; Hansen, T.; Le Paslier, D.; Linneberg, A.; Nielsen, H.B.; Pelletier, E.; Renault, P.; Sicheritz-Ponten, T.; Turner, K.; Zhu, H.; Yu, C.; Li, S.; Jian, M.; Zhou, Y.; Li, Y.; Zhang, X.; Li, S.; Qin, N.; Yang, H.; Wang, J.; Brunak, S.; Doré, J.; Guarner, F.; Kristiansen, K.; Pedersen, O.; Parkhill, J.; Weissenbach, J.; Bork, P.; Ehrlich, S.D.; Wang, J. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 2010, 464, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Aguiar-Pulido, V.; Huang, W.; Suarez-Ulloa, V.; Cickovski, T.; Mathee, K.; Narasimhan, G. Metagenomics, Metatranscriptomics, and Metabolomics Approaches for Microbiome Analysis: Supplementary Issue: Bioinformatics Methods and Applications for Big Metagenomics Data. Evolutionary Bioinformatics 2016, 12s1, EBO.S36436. [Google Scholar] [CrossRef]
- Heyer, R.; Schallert, K.; Zoun, R.; Becher, B.; Saake, G.; Benndorf, D. Challenges and perspectives of metaproteomic data analysis. Journal of Biotechnology 2017, 261, 24–36. [Google Scholar] [CrossRef]
- Henry, C.S.; DeJongh, M.; Best, A.A.; Frybarger, P.M.; Linsay, B.; Stevens, R.L. High-throughput generation, optimization and analysis of genome-scale metabolic models. Nature Biotechnology 2010, 28, 977–982. [Google Scholar] [CrossRef] [PubMed]
- Jünemann, S.; Kleinbölting, N.; Jaenicke, S.; Henke, C.; Hassa, J.; Nelkner, J.; Stolze, Y.; Albaum, S.P.; Schlüter, A.; Goesmann, A.; Sczyrba, A.; Stoye, J. Bioinformatics for NGS-based metagenomics and the application to biogas research. Journal of Biotechnology 2017, 261, 10–23. [Google Scholar] [CrossRef]
- Faust, K.; Raes, J. Microbial interactions: from networks to models. Nature Reviews Microbiology 2012, 10, 538–550. [Google Scholar] [CrossRef]
- Tobalina, L.; Bargiela, R.; Pey, J.; Herbst, F.A.; Lores, I.; Rojo, D.; Barbas, C.; Peláez, A.I.; Sánchez, J.; von Bergen, M.; Seifert, J.; Ferrer, M.; Planes, F.J. Context-specific metabolic network reconstruction of a naphthalene-degrading bacterial community guided by metaproteomic data. Bioinformatics 2015, 31, 1771–1779. [Google Scholar] [CrossRef]
- Machado, D.; Andrejev, S.; Tramontano, M.; Patil, K.R. Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Research 2018, 46, 7542–7553. [Google Scholar] [CrossRef]
- Aden, K.; Rehman, A.; Waschina, S.; Pan, W.H.; Walker, A.; Lucio, M.; Nunez, A.M.; Bharti, R.; Zimmerman, J.; Bethge, J.; Schulte, B.; Schulte, D.; Franke, A.; Nikolaus, S.; Schroeder, J.O.; Vandeputte, D.; Raes, J.; Szymczak, S.; Waetzig, G.H.; Zeuner, R.; Schmitt-Kopplin, P.; Kaleta, C.; Schreiber, S.; Rosenstiel, P. Metabolic Functions of Gut Microbes Associate With Efficacy of Tumor Necrosis Factor Antagonists in Patients With Inflammatory Bowel Diseases. Gastroenterology 2019, 157, 1279–1292.e11. [Google Scholar] [CrossRef] [PubMed]
- Marcelino, V.R.; Welsh, C.; Diener, C.; Gulliver, E.L.; Rutten, E.L.; Young, R.B.; Giles, E.M.; Gibbons, S.M.; Greening, C.; Forster, S.C. Disease-specific loss of microbial cross-feeding interactions in the human gut. Nature Communications 2023, 14. [Google Scholar] [CrossRef]
- García-Jiménez, B.; Torres-Bacete, J.; Nogales, J. Metabolic modelling approaches for describing and engineering microbial communities. Computational and Structural Biotechnology Journal 2021, 19, 226–246. [Google Scholar] [CrossRef] [PubMed]
- Curran, D.M.; Grote, A.; Nursimulu, N.; Geber, A.; Voronin, D.; Jones, D.R.; Ghedin, E.; Parkinson, J. Modeling the metabolic interplay between a parasitic worm and its bacterial endosymbiont allows the identification of novel drug targets. eLife 2020, 9. [Google Scholar] [CrossRef] [PubMed]
- Espinel-Ríos, S.; Bettenbrock, K.; Klamt, S.; Avalos, J.L.; Findeisen, R. Machine learning-supported cybergenetic modeling, optimization and control for synthetic microbial communities. In Computer Aided Chemical Engineering; Elsevier, 2023; pp. 2601–2606. [Google Scholar] [CrossRef]
- Espinel-Ríos, S.; Morabito, B.; Pohlodek, J.; Bettenbrock, K.; Klamt, S.; Findeisen, R. Toward a modeling, optimization, and predictive control framework for fed-batch metabolic cybergenetics. Biotechnology and Bioengineering 2023. [Google Scholar] [CrossRef]
- Xue, L.; Li, D.; Xi, Y. Nonlinear model predictive control of anaerobic digestion process based on reduced ADM1. In Proceedings of the 2015 10th Asian Control Conference (ASCC); IEEE, 2015; pp. 1–6. [Google Scholar] [CrossRef]
- Veenstra, T.D. Omics in Systems Biology: Current Progress and Future Outlook. PROTEOMICS 2021, 21, 2000235. [Google Scholar] [CrossRef] [PubMed]
- Thiele, I.; Palsson, B.Ø. A protocol for generating a high-quality genome-scale metabolic reconstruction. Nature Protocols 2010, 5, 93–121. [Google Scholar] [CrossRef]
- Sayers, E. Entrez programming utilities help. 2009; 17. Available online: http://www.ncbi.nlm.nih.gov/books/NBK25499.
- Bartel, J.; Krumsiek, J.; Theis, F.J. STATISTICAL METHODS FOR THE ANALYSIS OF HIGH-THROUGHPUT METABOLOMICS DATA. Computational and Structural Biotechnology Journal 2013, 4, e201301009. [Google Scholar] [CrossRef] [PubMed]
- Yamada, R.; Okada, D.; Wang, J.; Basak, T.; Koyama, S. Interpretation of omics data analyses. Journal of Human Genetics 2020, 66, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Arıkan, M.; Muth, T. Integrated multi-omics analyses of microbial communities: a review of the current state and future directions. Molecular Omics 2023. [Google Scholar] [CrossRef]
- Jiang, D.; Armour, C.R.; Hu, C.; Mei, M.; Tian, C.; Sharpton, T.J.; Jiang, Y. Microbiome Multi-Omics Network Analysis: Statistical Considerations, Limitations, and Opportunities. Frontiers in Genetics 2019, 10. [Google Scholar] [CrossRef]
- Reimand, J.; Isserlin, R.; Voisin, V.; Kucera, M.; Tannus-Lopes, C.; Rostamianfar, A.; Wadi, L.; Meyer, M.; Wong, J.; Xu, C.; Merico, D.; Bader, G.D. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nature Protocols 2019, 14, 482–517. [Google Scholar] [CrossRef]
- Salvato, F.; Hettich, R.L.; Kleiner, M. Five key aspects of metaproteomics as a tool to understand functional interactions in host-associated microbiomes. PLOS Pathogens 2021, 17, e1009245. [Google Scholar] [CrossRef] [PubMed]
- Gehlenborg, N.; O’Donoghue, S.I.; Baliga, N.S.; Goesmann, A.; Hibbs, M.A.; Kitano, H.; Kohlbacher, O.; Neuweger, H.; Schneider, R.; Tenenbaum, D.; Gavin, A.C. Visualization of omics data for systems biology. Nature Methods 2010, 7, S56–S68. [Google Scholar] [CrossRef]
- Bekiaris, P.S.; Klamt, S. Automatic construction of metabolic models with enzyme constraints. BMC Bioinformatics 2020, 21. [Google Scholar] [CrossRef]
- Machado, D.; Costa, R.S.; Rocha, M.; Ferreira, E.C.; Tidor, B.; Rocha, I. Modeling formalisms in Systems Biology. AMB Express 2011, 1, 45. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.M.; Tymoczko, J.L.; Stryer, L. Der Stoffwechsel: Konzepte und Grundmuster. In Stryer Biochemie; Springer: Berlin/Heidelberg, Germany, 2013; pp. 431–455. [Google Scholar] [CrossRef]
- Huang, J.; Zhang, P.; Solari, F.A.; Sickmann, A.; Garcia, A.; Jurk, K.; Heemskerk, J.W.M. Molecular Proteomics and Signalling of Human Platelets in Health and Disease. International Journal of Molecular Sciences 2021, 22, 9860. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.M.; Tymoczko, J.L.; Stryer, L. Signaltransduktionswege. In Stryer Biochemie; Springer: Berlin/Heidelberg, Germany, 2013; pp. 404–430. [Google Scholar] [CrossRef]
- Davidson, E.; Levin, M. Gene regulatory networks. Proceedings of the National Academy of Sciences 2005, 102, 4935–4935. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.M.; Tymoczko, J.L.; Stryer, L. Kontrolle der Genexpression bei Prokaryoten. In Stryer Biochemie; Springer: Berlin/Heidelberg, Germany, 2013; pp. 933–933. [Google Scholar] [CrossRef]
- Berg, J.M.; Tymoczko, J.L.; Stryer, L. Kontrolle der Genexpression bei Eukaryoten. In Stryer Biochemie; Springer: Berlin/Heidelberg, Germany, 2013; pp. 949–969. [Google Scholar] [CrossRef]
- Terfve, C.; Saez-Rodriguez, J. Modeling Signaling Networks Using High-throughput Phospho-proteomics. In Advances in Experimental Medicine and Biology; Springer: New York, 2011; pp. 19–57. [Google Scholar] [CrossRef]
- Samaga, R.; Klamt, S. Modeling approaches for qualitative and semi-quantitative analysis of cellular signaling networks. Cell Communication and Signaling 2013, 11, 43. [Google Scholar] [CrossRef] [PubMed]
- Papin, J.A.; Hunter, T.; Palsson, B.O.; Subramaniam, S. Reconstruction of cellular signalling networks and analysis of their properties. Nature Reviews Molecular Cell Biology 2005, 6, 99–111. [Google Scholar] [CrossRef] [PubMed]
- Sorrells, T.; Johnson, A. Making Sense of Transcription Networks. Cell 2015, 161, 714–723. [Google Scholar] [CrossRef] [PubMed]
- Panni, S.; Lovering, R.C.; Porras, P.; Orchard, S. Non-coding RNA regulatory networks. Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms 2020, 1863, 194417. [Google Scholar] [CrossRef] [PubMed]
- Popovic, M. Thermodynamic properties of microorganisms: determination and analysis of enthalpy, entropy, and Gibbs free energy of biomass, cells and colonies of 32 microorganism species. Heliyon 2019, 5, e01950. [Google Scholar] [CrossRef]
- Feijó Delgado, F.; Cermak, N.; Hecht, V.C.; Son, S.; Li, Y.; Knudsen, S.M.; Olcum, S.; Higgins, J.M.; Chen, J.; Grover, W.H.; Manalis, S.R. Intracellular Water Exchange for Measuring the Dry Mass, Water Mass and Changes in Chemical Composition of Living Cells. PLoS ONE 2013, 8, e67590. [Google Scholar] [CrossRef] [PubMed]
- Großkopf, T.; Soyer, O.S. Synthetic microbial communities. Current Opinion in Microbiology 2014, 18, 72–77. [Google Scholar] [CrossRef]
- Roell, G.W.; Zha, J.; Carr, R.R.; Koffas, M.A.; Fong, S.S.; Tang, Y.J. Engineering microbial consortia by division of labor. Microbial Cell Factories 2019, 18. [Google Scholar] [CrossRef] [PubMed]
- Schink, B. Energetics of syntrophic cooperation in methanogenic degradation. Microbiology and Molecular Biology Reviews 1997, 61, 262–280. [Google Scholar] [CrossRef]
- Heyer, R.; Kohrs, F.; Reichl, U.; Benndorf, D. Metaproteomics of complex microbial communities in biogas plants. Microbial Biotechnology 2015, 8, 749–763. [Google Scholar] [CrossRef]
- Muth, T.; Renard, B.Y.; Martens, L. Metaproteomic data analysis at a glance: advances in computational microbial community proteomics. Expert Review of Proteomics 2016, 13, 757–769. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef] [PubMed]
- Solano, C.; Echeverz, M.; Lasa, I. Biofilm dispersion and quorum sensing. Current Opinion in Microbiology 2014, 18, 96–104. [Google Scholar] [CrossRef] [PubMed]
- Quiza, L.; St-Arnaud, M.; Yergeau, E. Harnessing phytomicrobiome signaling for rhizosphere microbiome engineering. Frontiers in Plant Science 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Jameson, K.; Olson, C.; Kazmi, S.; Hsiao, E. Toward Understanding Microbiome-Neuronal Signaling. Molecular Cell 2020, 78, 577–583. [Google Scholar] [CrossRef]
- Shin, S.C.; Kim, S.H.; You, H.; Kim, B.; Kim, A.C.; Lee, K.A.; Yoon, J.H.; Ryu, J.H.; Lee, W.J. Drosophila Microbiome Modulates Host Developmental and Metabolic Homeostasis via Insulin Signaling. Science 2011, 334, 670–674. [Google Scholar] [CrossRef] [PubMed]
- Fischbach, M.A.; Segre, J.A. Signaling in Host-Associated Microbial Communities. Cell 2016, 164, 1288–1300. [Google Scholar] [CrossRef]
- The Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.M. Non-surface Attached Bacterial Aggregates: A Ubiquitous Third Lifestyle. Frontiers in Microbiology 2020, 11. [Google Scholar] [CrossRef]
- Rani, S.A.; Pitts, B.; Beyenal, H.; Veluchamy, R.A.; Lewandowski, Z.; Davison, W.M.; Buckingham-Meyer, K.; Stewart, P.S. Spatial Patterns of DNA Replication, Protein Synthesis, and Oxygen Concentration within Bacterial Biofilms Reveal Diverse Physiological States. Journal of Bacteriology 2007, 189, 4223–4233. [Google Scholar] [CrossRef]
- Kreft, J.U.; Plugge, C.M.; Prats, C.; Leveau, J.H.J.; Zhang, W.; Hellweger, F.L. From Genes to Ecosystems in Microbiology: Modeling Approaches and the Importance of Individuality. Frontiers in Microbiology 2017, 8. [Google Scholar] [CrossRef]
- Blum, W.E.; Zechmeister-Boltenstern, S.; Keiblinger, K.M. Does Soil Contribute to the Human Gut Microbiome? Microorganisms 2019, 7, 287. [Google Scholar] [CrossRef]
- Pasolli, E.; Asnicar, F.; Manara, S.; Zolfo, M.; Karcher, N.; Armanini, F.; Beghini, F.; Manghi, P.; Tett, A.; Ghensi, P.; Collado, M.C.; Rice, B.L.; DuLong, C.; Morgan, X.C.; Golden, C.D.; Quince, C.; Huttenhower, C.; Segata, N. Extensive Unexplored Human Microbiome Diversity Revealed by Over 150,000 Genomes from Metagenomes Spanning Age, Geography, and Lifestyle. Cell 2019, 176, 649–662.e20. [Google Scholar] [CrossRef] [PubMed]
- Almeida, A.; Mitchell, A.L.; Boland, M.; Forster, S.C.; Gloor, G.B.; Tarkowska, A.; Lawley, T.D.; Finn, R.D. A new genomic blueprint of the human gut microbiota. Nature 2019, 568, 499–504. [Google Scholar] [CrossRef]
- Louca, S.; Mazel, F.; Doebeli, M.; Parfrey, L.W. A census-based estimate of Earth’s bacterial and archaeal diversity. PLOS Biology 2019, 17, e3000106. [Google Scholar] [CrossRef] [PubMed]
- Overmann, J.; Abt, B.; Sikorski, J. Present and Future of Culturing Bacteria. Annual Review of Microbiology 2017, 71, 711–730. [Google Scholar] [CrossRef] [PubMed]
- Reimer, L.C.; Sardà Carbasse, J.; Koblitz, J.; Ebeling, C.; Podstawka, A.; Overmann, J. BacDive in 2022: the knowledge base for standardized bacterial and archaeal data. Nucleic Acids Research 2021, 50, D741–D746. [Google Scholar] [CrossRef] [PubMed]
- Beck, A.; Hunt, K.; Carlson, R. Measuring Cellular Biomass Composition for Computational Biology Applications. Processes 2018, 6, 38. [Google Scholar] [CrossRef]
- Lachance, J.C.; Lloyd, C.J.; Monk, J.M.; Yang, L.; Sastry, A.V.; Seif, Y.; Palsson, B.O.; Rodrigue, S.; Feist, A.M.; King, Z.A.; Jacques, P.É. BOFdat: Generating biomass objective functions for genome-scale metabolic models from experimental data. PLOS Computational Biology 2019, 15, e1006971. [Google Scholar] [CrossRef]
- Vos, T.; Hakkaart, X.D.V.; de Hulster, E.A.F.; van Maris, A.J.A.; Pronk, J.T.; Daran-Lapujade, P. Maintenance-energy requirements and robustness of Saccharomyces cerevisiae at aerobic near-zero specific growth rates. Microbial Cell Factories 2016, 15. [Google Scholar] [CrossRef]
- Zamboni, N.; Fendt, S.M.; Rühl, M.; Sauer, U. 13C-based metabolic flux analysis. Nature Protocols 2009, 4, 878–892. [Google Scholar] [CrossRef]
- Palazzotto, E.; Weber, T. Omics and multi-omics approaches to study the biosynthesis of secondary metabolites in microorganisms. Current Opinion in Microbiology 2018, 45, 109–116. [Google Scholar] [CrossRef]
- Winter, G.; Krömer, J.O. Fluxomics - connecting ‘omics analysis and phenotypes. Environmental Microbiology 2013, 15, 1901–1916. [Google Scholar] [CrossRef]
- Koch, S.; Benndorf, D.; Fronk, K.; Reichl, U.; Klamt, S. Predicting compositions of microbial communities from stoichiometric models with applications for the biogas process. Biotechnology for Biofuels 2016, 9. [Google Scholar] [CrossRef]
- Koch, S.; Kohrs, F.; Lahmann, P.; Bissinger, T.; Wendschuh, S.; Benndorf, D.; Reichl, U.; Klamt, S. RedCom: A strategy for reduced metabolic modeling of complex microbial communities and its application for analyzing experimental datasets from anaerobic digestion. PLOS Computational Biology 2019, 15, e1006759. [Google Scholar] [CrossRef] [PubMed]
- Schäpe, S.S.; Krause, J.L.; Engelmann, B.; Fritz-Wallace, K.; Schattenberg, F.; Liu, Z.; Müller, S.; Jehmlich, N.; Rolle-Kampczyk, U.; Herberth, G.; von Bergen, M. The Simplified Human Intestinal Microbiota (SIHUMIx) Shows High Structural and Functional Resistance against Changing Transit Times in In Vitro Bioreactors. Microorganisms 2019, 7, 641. [Google Scholar] [CrossRef] [PubMed]
- Hanreich, A.; Schimpf, U.; Zakrzewski, M.; Schlüter, A.; Benndorf, D.; Heyer, R.; Rapp, E.; Pühler, A.; Reichl, U.; Klocke, M. Metagenome and metaproteome analyses of microbial communities in mesophilic biogas-producing anaerobic batch fermentations indicate concerted plant carbohydrate degradation. Systematic and Applied Microbiology 2013, 36, 330–338. [Google Scholar] [CrossRef]
- Lui, L.M.; Majumder, E.L.W.; Smith, H.J.; Carlson, H.K.; von Netzer, F.; Fields, M.W.; Stahl, D.A.; Zhou, J.; Hazen, T.C.; Baliga, N.S.; Adams, P.D.; Arkin, A.P. Mechanism Across Scales: A Holistic Modeling Framework Integrating Laboratory and Field Studies for Microbial Ecology. Frontiers in Microbiology 2021, 12. [Google Scholar] [CrossRef]
- Petersen, C.; Hamerich, I.K.; Adair, K.L.; Griem-Krey, H.; Torres Oliva, M.; Hoeppner, M.P.; Bohannan, B.J.M.; Schulenburg, H. Host and microbiome jointly contribute to environmental adaptation. The ISME Journal 2023, 17, 1953–1965. [Google Scholar] [CrossRef] [PubMed]
- Muth, T.; Benndorf, D.; Reichl, U.; Rapp, E.; Martens, L. Searching for a needle in a stack of needles: challenges in metaproteomics data analysis. Mol. BioSyst. 2013, 9, 578–585. [Google Scholar] [CrossRef] [PubMed]
- Wilmes, P.; Bond, P.L. Metaproteomics: studying functional gene expression in microbial ecosystems. Trends in Microbiology 2006, 14, 92–97. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Valera, F. Environmental genomics, the big picture? FEMS Microbiology Letters 2004, 231, 153–158. [Google Scholar] [CrossRef]
- Kleiner, M. Metaproteomics: Much More than Measuring Gene Expression in Microbial Communities. 2019; 4. [Google Scholar] [CrossRef]
- Domenzain, I.; Sánchez, B.; Anton, M.; Kerkhoven, E.J.; Millán-Oropeza, A.; Henry, C.; Siewers, V.; Morrissey, J.P.; Sonnenschein, N.; Nielsen, J. Reconstruction of a catalogue of genome-scale metabolic models with enzymatic constraints using GECKO 2.0. Nature Communications 2022, 13. [Google Scholar] [CrossRef] [PubMed]
- Kohrs, F.; Heyer, R.; Magnussen, A.; Benndorf, D.; Muth, T.; Behne, A.; Rapp, E.; Kausmann, R.; Heiermann, M.; Klocke, M.; Reichl, U. Sample prefractionation with liquid isoelectric focusing enables in depth microbial metaproteome analysis of mesophilic and thermophilic biogas plants. Anaerobe 2014, 29, 59–67. [Google Scholar] [CrossRef] [PubMed]
- Aakko, J.; Pietilä, S.; Suomi, T.; Mahmoudian, M.; Toivonen, R.; Kouvonen, P.; Rokka, A.; Hänninen, A.; Elo, L.L. Data-Independent Acquisition Mass Spectrometry in Metaproteomics of Gut Microbiota—Implementation and Computational Analysis. Journal of Proteome Research 2019, 19, 432–436. [Google Scholar] [CrossRef] [PubMed]
- Pietilä, S.; Suomi, T.; Elo, L.L. Introducing untargeted data-independent acquisition for metaproteomics of complex microbial samples. ISME Communications 2022, 2. [Google Scholar] [CrossRef]
- Perkins, D.N.; Pappin, D.J.C.; Creasy, D.M.; Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 1999, 20, 3551–3567. [Google Scholar] [CrossRef]
- Craig, R.; Beavis, R.C. TANDEM: matching proteins with tandem mass spectra. Bioinformatics 2004, 20, 1466–1467. [Google Scholar] [CrossRef] [PubMed]
- Elias, J.E.; Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nature Methods 2007, 4, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Muth, T.; Behne, A.; Heyer, R.; Kohrs, F.; Benndorf, D.; Hoffmann, M.; Lehtevä, M.; Reichl, U.; Martens, L.; Rapp, E. The MetaProteomeAnalyzer: A Powerful Open-Source Software Suite for Metaproteomics Data Analysis and Interpretation. Journal of Proteome Research 2015, 14, 1557–1565. [Google Scholar] [CrossRef]
- Afgan, E.; Nekrutenko, A.; Grüning, B.A.; Blankenberg, D.; Goecks, J.; Schatz, M.C.; Ostrovsky, A.E.; Mahmoud, A.; Lonie, A.J.; Syme, A.; Fouilloux, A.; Bretaudeau, A.; Nekrutenko, A.; Kumar, A.; Eschenlauer, A.C.; DeSanto, A.D.; Guerler, A.; Serrano-Solano, B.; Batut, B.; Grüning, B.A.; Langhorst, B.W.; Carr, B.; Raubenolt, B.A.; Hyde, C.J.; Bromhead, C.J.; Barnett, C.B.; Royaux, C.; Gallardo, C.; Blankenberg, D.; Fornika, D.J.; Baker, D.; Bouvier, D.; Clements, D.; de Lima Morais, D.A.; Tabernero, D.L.; Lariviere, D.; Nasr, E.; Afgan, E.; Zambelli, F.; Heyl, F.; Psomopoulos, F.; Coppens, F.; Price, G.R.; Cuccuru, G.; Corguillé, G.L.; Kuster, G.V.; Akbulut, G.G.; Rasche, H.; Hotz, H.R.; Eguinoa, I.; Makunin, I.; Ranawaka, I.J.; Taylor, J.P.; Joshi, J.; Hillman-Jackson, J.; Goecks, J.; Chilton, J.M.; Kamali, K.; Suderman, K.; Poterlowicz, K.; Yvan, L.B.; Lopez-Delisle, L.; Sargent, L.; Bassetti, M.E.; Tangaro, M.A.; van den Beek, M.; Čech, M.; Bernt, M.; Fahrner, M.; Tekman, M.; Föll, M.C.; Schatz, M.C.; Crusoe, M.R.; Roncoroni, M.; Kucher, N.; Coraor, N.; Stoler, N.; Rhodes, N.; Soranzo, N.; Pinter, N.; Goonasekera, N.A.; Moreno, P.A.; Videm, P.; Melanie, P.; Mandreoli, P.; Jagtap, P.D.; Gu, Q.; Weber, R.J.M.; Lazarus, R.; Vorderman, R.H.P.; Hiltemann, S.; Golitsynskiy, S.; Garg, S.; Bray, S.A.; Gladman, S.L.; Leo, S.; Mehta, S.P.; Griffin, T.J.; Jalili, V.; Yves, V.; Wen, V.; Nagampalli, V.K.; Bacon, W.A.; de Koning, W.; Maier, W.; Briggs, P.J. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2022 update. Nucleic Acids Research 2022, 50, W345–W351. [Google Scholar] [CrossRef]
- Bossche, T.V.D.; Kunath, B.J.; Schallert, K.; Schäpe, S.S.; Abraham, P.E.; Armengaud, J.; Arntzen, M.Ø.; Bassignani, A.; Benndorf, D.; Fuchs, S.; Giannone, R.J.; Griffin, T.J.; Hagen, L.H.; Halder, R.; Henry, C.; Hettich, R.L.; Heyer, R.; Jagtap, P.; Jehmlich, N.; Jensen, M.; Juste, C.; Kleiner, M.; Langella, O.; Lehmann, T.; Leith, E.; May, P.; Mesuere, B.; Miotello, G.; Peters, S.L.; Pible, O.; Queiros, P.T.; Reichl, U.; Renard, B.Y.; Schiebenhoefer, H.; Sczyrba, A.; Tanca, A.; Trappe, K.; Trezzi, J.P.; Uzzau, S.; Verschaffelt, P.; von Bergen, M.; Wilmes, P.; Wolf, M.; Martens, L.; Muth, T. Critical Assessment of MetaProteome Investigation (CAMPI): a multi-laboratory comparison of established workflows. Nature Communications 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Delogu, F.; Kunath, B.J.; Evans, P.N.; Arntzen, M.Ø; Hvidsten, T.R.; Pope, P.B. Integration of absolute multi-omics reveals dynamic protein-to-RNA ratios and metabolic interplay within mixed-domain microbiomes. Nature Communications 2020, 11. [Google Scholar] [CrossRef]
- Sivanich, M.K.; Gu, T.; Tabang, D.N.; Li, L. Recent advances in isobaric labeling and applications in quantitative proteomics 2022. 22. [CrossRef]
- Ahrné, E.; Molzahn, L.; Glatter, T.; Schmidt, A. Critical assessment of proteome-wide label-free absolute abundance estimation strategies. PROTEOMICS 2013, 13, 2567–2578. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, B.J.; Zhang, C.; Nilsson, A.; Lahtvee, P.J.; Kerkhoven, E.J.; Nielsen, J. Improving the phenotype predictions of a yeast genome-scale metabolic model by incorporating enzymatic constraints. Molecular Systems Biology 2017, 13, 935. [Google Scholar] [CrossRef] [PubMed]
- Kirkpatrick, D.S.; Gerber, S.A.; Gygi, S.P. The absolute quantification strategy: a general procedure for the quantification of proteins and post-translational modifications. Methods 2005, 35, 265–273. [Google Scholar] [CrossRef]
- Starke, R.; Jehmlich, N.; Bastida, F. Using proteins to study how microbes contribute to soil ecosystem services: The current state and future perspectives of soil metaproteomics. Journal of Proteomics 2019, 198, 50–58. [Google Scholar] [CrossRef]
- Pratt, J.M.; Simpson, D.M.; Doherty, M.K.; Rivers, J.; Gaskell, S.J.; Beynon, R.J. Multiplexed absolute quantification for proteomics using concatenated signature peptides encoded by QconCAT genes. Nature protocols 2006, 1, 1029–1043. [Google Scholar] [CrossRef] [PubMed]
- Schallert, K.; Verschaffelt, P.; Mesuere, B.; Benndorf, D.; Martens, L.; Bossche, T.V.D. Pout2Prot: An Efficient Tool to Create Protein (Sub)groups from Percolator Output Files. Journal of Proteome Research 2022, 21, 1175–1180. [Google Scholar] [CrossRef]
- Zhang, X.; Figeys, D. Perspective and Guidelines for Metaproteomics in Microbiome Studies. Journal of Proteome Research 2019, 18, 2370–2380. [Google Scholar] [CrossRef] [PubMed]
- Mirzayi, C.; Renson, A.; Furlanello, C.; Sansone, S.A.; Zohra, F.; Elsafoury, S.; Geistlinger, L.; Kasselman, L.J.; Eckenrode, K.; van de Wijgert, J.; Loughman, A.; Marques, F.Z.; MacIntyre, D.A.; Arumugam, M.; Azhar, R.; Beghini, F.; Bergstrom, K.; Bhatt, A.; Bisanz, J.E.; Braun, J.; Bravo, H.C.; Buck, G.A.; Bushman, F.; Casero, D.; Clarke, G.; Collado, M.C.; Cotter, P.D.; Cryan, J.F.; Demmer, R.T.; Devkota, S.; Elinav, E.; Escobar, J.S.; Fettweis, J.; Finn, R.D.; Fodor, A.A.; Forslund, S.; Franke, A.; Furlanello, C.; Gilbert, J.; Grice, E.; Haibe-Kains, B.; Handley, S.; Herd, P.; Holmes, S.; Jacobs, J.P.; Karstens, L.; Knight, R.; Knights, D.; Koren, O.; Kwon, D.S.; Langille, M.; Lindsay, B.; McGovern, D.; McHardy, A.C.; McWeeney, S.; Mueller, N.T.; Nezi, L.; Olm, M.; Palm, N.; Pasolli, E.; Raes, J.; Redinbo, M.R.; Rühlemann, M.; Sartor, R.B.; Schloss, P.D.; Schriml, L.; Segal, E.; Shardell, M.; Sharpton, T.; Smirnova, E.; Sokol, H.; Sonnenburg, J.L.; Srinivasan, S.; Thingholm, L.B.; Turnbaugh, P.J.; Upadhyay, V.; Walls, R.L.; Wilmes, P.; Yamada, T.; Zeller, G.; Zhang, M.; Zhao, N.; Zhao, L.; Bao, W.; Culhane, A.; Devanarayan, V.; Dopazo, J.; Fan, X.; Fischer, M.; Jones, W.; Kusko, R.; Mason, C.E.; Mercer, T.R.; Sansone, S.A.; Scherer, A.; Shi, L.; Thakkar, S.; Tong, W.; Wolfinger, R.; Hunter, C.; Segata, N.; Huttenhower, C.; Dowd, J.B.; Jones, H.E.; Waldron, L. Reporting guidelines for human microbiome research: the STORMS checklist. Nature Medicine 2021, 27, 1885–1892. [Google Scholar] [CrossRef]
- Vizcaíno, J.A.; Walzer, M.; Jiménez, R.C.; Bittremieux, W.; Bouyssié, D.; Carapito, C.; Corrales, F.; Ferro, M.; Heck, A.J.; Horvatovich, P.; Hubalek, M.; Lane, L.; Laukens, K.; Levander, F.; Lisacek, F.; Novak, P.; Palmblad, M.; Piovesan, D.; Pühler, A.; Schwämmle, V.; Valkenborg, D.; van Rijswijk, M.; Vondrasek, J.; Eisenacher, M.; Martens, L.; Kohlbacher, O. A community proposal to integrate proteomics activities in ELIXIR. F1000Research 2017, 6, 875. [Google Scholar] [CrossRef]
- Van Den Bossche, T.; Arntzen, M.Ø; Becher, D.; Benndorf, D.; Eijsink, V.G.H.; Henry, C.; Jagtap, P.D.; Jehmlich, N.; Juste, C.; Kunath, B.J.; Mesuere, B.; Muth, T.; Pope, P.B.; Seifert, J.; Tanca, A.; Uzzau, S.; Wilmes, P.; Hettich, R.L.; Armengaud, J. The Metaproteomics Initiative: a coordinated approach for propelling the functional characterization of microbiomes. Microbiome 2021, 9. [Google Scholar] [CrossRef]
- Bateman, A.; Martin, M.J.; Orchard, S.; Magrane, M.; Ahmad, S.; Alpi, E.; Bowler-Barnett, E.H.; Britto, R.; Bye-A-Jee, H.; Cukura, A.; Denny, P.; Dogan, T.; Ebenezer, T.; Fan, J.; Garmiri, P.; da Costa Gonzales, L.J.; Hatton-Ellis, E.; Hussein, A.; Ignatchenko, A.; Insana, G.; Ishtiaq, R.; Joshi, V.; Jyothi, D.; Kandasaamy, S.; Lock, A.; Luciani, A.; Lugaric, M.; Luo, J.; Lussi, Y.; MacDougall, A.; Madeira, F.; Mahmoudy, M.; Mishra, A.; Moulang, K.; Nightingale, A.; Pundir, S.; Qi, G.; Raj, S.; Raposo, P.; Rice, D.L.; Saidi, R.; Santos, R.; Speretta, E.; Stephenson, J.; Totoo, P.; Turner, E.; Tyagi, N.; Vasudev, P.; Warner, K.; Watkins, X.; Zaru, R.; Zellner, H.; Bridge, A.J.; Aimo, L.; Argoud-Puy, G.; Auchincloss, A.H.; Axelsen, K.B.; Bansal, P.; Baratin, D.; Batista Neto, T.M.; Blatter, M.C.; Bolleman, J.T.; Boutet, E.; Breuza, L.; Gil, B.C.; Casals-Casas, C.; Echioukh, K.C.; Coudert, E.; Cuche, B.; de Castro, E.; Estreicher, A.; Famiglietti, M.L.; Feuermann, M.; Gasteiger, E.; Gaudet, P.; Gehant, S.; Gerritsen, V.; Gos, A.; Gruaz, N.; Hulo, C.; Hyka-Nouspikel, N.; Jungo, F.; Kerhornou, A.; Le Mercier, P.; Lieberherr, D.; Masson, P.; Morgat, A.; Muthukrishnan, V.; Paesano, S.; Pedruzzi, I.; Pilbout, S.; Pourcel, L.; Poux, S.; Pozzato, M.; Pruess, M.; Redaschi, N.; Rivoire, C.; Sigrist, C.J.A.; Sonesson, K.; Sundaram, S.; Wu, C.H.; Arighi, C.N.; Arminski, L.; Chen, C.; Chen, Y.; Huang, H.; Laiho, K.; McGarvey, P.; Natale, D.A.; Ross, K.; Vinayaka, C.R.; Wang, Q.; Wang, Y.; Zhang, J. UniProt: the Universal Protein Knowledgebase in 2023. Nucleic Acids Research 2022, 51, D523–D531. [Google Scholar] [CrossRef]
- Schiebenhoefer, H.; Bossche, T.V.D.; Fuchs, S.; Renard, B.Y.; Muth, T.; Martens, L. Challenges and promise at the interface of metaproteomics and genomics: an overview of recent progress in metaproteogenomic data analysis. Expert Review of Proteomics 2019, 16, 375–390. [Google Scholar] [CrossRef] [PubMed]
- Schoch, C.L.; Ciufo, S.; Domrachev, M.; Hotton, C.L.; Kannan, S.; Khovanskaya, R.; Leipe, D.; Mcveigh, R.; O’Neill, K.; Robbertse, B.; Sharma, S.; Soussov, V.; Sullivan, J.P.; Sun, L.; Turner, S.; Karsch-Mizrachi, I. NCBI Taxonomy: a comprehensive update on curation, resources and tools. Database 2020, 2020. [Google Scholar] [CrossRef]
- Parks, D.H.; Chuvochina, M.; Rinke, C.; Mussig, A.J.; Chaumeil, P.A.; Hugenholtz, P. GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Research 2021, 50, D785–D794. [Google Scholar] [CrossRef]
- Mesuere, B.; der Jeugt, F.V.; Willems, T.; Naessens, T.; Devreese, B.; Martens, L.; Dawyndt, P. High-throughput metaproteomics data analysis with Unipept: A tutorial. Journal of Proteomics 2018, 171, 11–22. [Google Scholar] [CrossRef]
- Huson, D.H.; Mitra, S.; Ruscheweyh, H.J.; Weber, N.; Schuster, S.C. Integrative analysis of environmental sequences using MEGAN4. Genome Research 2011, 21, 1552–1560. [Google Scholar] [CrossRef] [PubMed]
- Penzlin, A.; Lindner, M.S.; Doellinger, J.; Dabrowski, P.W.; Nitsche, A.; Renard, B.Y. Pipasic: similarity and expression correction for strain-level identification and quantification in metaproteomics. Bioinformatics 2014, 30, i149–i156. [Google Scholar] [CrossRef] [PubMed]
- Schiebenhoefer, H.; Schallert, K.; Renard, B.Y.; Trappe, K.; Schmid, E.; Benndorf, D.; Riedel, K.; Muth, T.; Fuchs, S. A complete and flexible workflow for metaproteomics data analysis based on MetaProteomeAnalyzer and Prophane. Nature Protocols 2020, 15, 3212–3239. [Google Scholar] [CrossRef]
- Mesuere, B.; Van der Jeugt, F.; Devreese, B.; Vandamme, P.; Dawyndt, P. The unique peptidome: Taxon-specific tryptic peptides as biomarkers for targeted metaproteomics. PROTEOMICS 2016, 16, 2313–2318. [Google Scholar] [CrossRef] [PubMed]
- Starke, R.; Fiore-Donno, A.M.; White, R.A.; Parente Fernandes, M.L.; Martinović, T.; Bastida, F.; Delgado-Baquerizo, M.; Jehmlich, N. Biomarker metaproteomics for relative taxa abundances across soil organisms. Soil Biology and Biochemistry 2022, 175, 108861. [Google Scholar] [CrossRef]
- Chang, A.; Jeske, L.; Ulbrich, S.; Hofmann, J.; Koblitz, J.; Schomburg, I.; Neumann-Schaal, M.; Jahn, D.; Schomburg, D. BRENDA, the ELIXIR core data resource in 2021: new developments and updates. Nucleic Acids Research 2020, 49, D498–D508. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; Harris, M.A.; Hill, D.P.; Issel-Tarver, L.; Kasarskis, A.; Lewis, S.; Matese, J.C.; Richardson, J.E.; Ringwald, M.; Rubin, G.M.; Sherlock, G. Gene Ontology: tool for the unification of biology. Nature Genetics 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Aleksander, S.A.; Balhoff, J.; Carbon, S.; Cherry, J.M.; Drabkin, H.J.; Ebert, D.; Feuermann, M.; Gaudet, P.; Harris, N.L.; et al. The Gene Ontology knowledgebase in 2023. Genetics 2023, 224, iyad031. [Google Scholar] [PubMed]
- Kanehisa, M.; Furumichi, M.; Sato, Y.; Kawashima, M.; Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Research 2022. [Google Scholar] [CrossRef]
- Karp, P.D.; Billington, R.; Caspi, R.; Fulcher, C.A.; Latendresse, M.; Kothari, A.; Keseler, I.M.; Krummenacker, M.; Midford, P.E.; Ong, Q.; Ong, W.K.; Paley, S.M.; Subhraveti, P. The BioCyc collection of microbial genomes and metabolic pathways. Briefings in Bioinformatics 2017, 20, 1085–1093. [Google Scholar] [CrossRef]
- Gillespie, M.; Jassal, B.; Stephan, R.; Milacic, M.; Rothfels, K.; Senff-Ribeiro, A.; Griss, J.; Sevilla, C.; Matthews, L.; Gong, C.; Deng, C.; Varusai, T.; Ragueneau, E.; Haider, Y.; May, B.; Shamovsky, V.; Weiser, J.; Brunson, T.; Sanati, N.; Beckman, L.; Shao, X.; Fabregat, A.; Sidiropoulos, K.; Murillo, J.; Viteri, G.; Cook, J.; Shorser, S.; Bader, G.; Demir, E.; Sander, C.; Haw, R.; Wu, G.; Stein, L.; Hermjakob, H.; D’Eustachio, P. The reactome pathway knowledgebase 2022. Nucleic Acids Research 2021, 50, D687–D692. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Chowdhury, D.; Zhang, Z.; Cheung, W.K.; Lu, A.; Bian, Z.; Zhang, L. A review of computational tools for generating metagenome-assembled genomes from metagenomic sequencing data. 2021, 19, 6301–6314. [CrossRef]
- Parks, D.H.; Rinke, C.; Chuvochina, M.; Chaumeil, P.A.; Woodcroft, B.J.; Evans, P.N.; Hugenholtz, P.; Tyson, G.W. Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nature Microbiology 2017, 2, 1533–1542. [Google Scholar] [CrossRef] [PubMed]
- Props, R.; Kerckhof, F.M.; Rubbens, P.; Vrieze, J.D.; Sanabria, E.H.; Waegeman, W.; Monsieurs, P.; Hammes, F.; Boon, N. Absolute quantification of microbial taxon abundances. The ISME Journal 2016, 11, 584–587. [Google Scholar] [CrossRef] [PubMed]
- Hatzenpichler, R.; Krukenberg, V.; Spietz, R.L.; Jay, Z.J. Next-generation physiology approaches to study microbiome function at single cell level. Nature Reviews Microbiology 2020, 18, 241–256. [Google Scholar] [CrossRef] [PubMed]
- Cesar, S.; Huang, K.C. Thinking big: the tunability of bacterial cell size. FEMS Microbiology Reviews 2017, 41, 672–678. [Google Scholar] [CrossRef] [PubMed]
- Bauermeister, A.; Mannochio-Russo, H.; Costa-Lotufo, L.V.; Jarmusch, A.K.; Dorrestein, P.C. Mass spectrometry-based metabolomics in microbiome investigations. Nature Reviews Microbiology 2021, 20, 143–160. [Google Scholar] [CrossRef]
- Liu, X.; Locasale, J.W. Metabolomics: A Primer. Trends in Biochemical Sciences 2017, 42, 274–284. [Google Scholar] [CrossRef] [PubMed]
- Zhang, A.; Sun, H.; Wang, P.; Han, Y.; Wang, X. Modern analytical techniques in metabolomics analysis. The Analyst 2012, 137, 293–300. [Google Scholar] [CrossRef]
- Bragg, L.; Tyson, G.W. Metagenomics Using Next-Generation Sequencing. In Methods in Molecular Biology; Humana Press, 2014; pp. 183–201. [Google Scholar] [CrossRef]
- Segata, N.; Boernigen, D.; Tickle, T.L.; Morgan, X.C.; Garrett, W.S.; Huttenhower, C. Computational meta’omics for microbial community studies. Molecular Systems Biology 2013, 9, 666. [Google Scholar] [CrossRef]
- Frioux, C.; Singh, D.; Korcsmaros, T.; Hildebrand, F. From bag-of-genes to bag-of-genomes: metabolic modelling of communities in the era of metagenome-assembled genomes. Computational and Structural Biotechnology Journal 2020, 18, 1722–1734. [Google Scholar] [CrossRef] [PubMed]
- Zorrilla, F.; Buric, F.; Patil, K.R.; Zelezniak, A. metaGEM: reconstruction of genome scale metabolic models directly from metagenomes. Nucleic Acids Research 2021, 49, e126–e126. [Google Scholar] [CrossRef] [PubMed]
- Bashiardes, S.; Zilberman-Schapira, G.; Elinav, E. Use of Metatranscriptomics in Microbiome Research. Bioinformatics and Biology Insights 2016, 10, BBI.S34610. [Google Scholar] [CrossRef] [PubMed]
- Gifford, S.M.; Sharma, S.; Rinta-Kanto, J.M.; Moran, M.A. Quantitative analysis of a deeply sequenced marine microbial metatranscriptome. The ISME Journal 2010, 5, 461–472. [Google Scholar] [CrossRef] [PubMed]
- Gosalbes, M.J.; Durbán, A.; Pignatelli, M.; Abellan, J.J.; Jiménez-Hernández, N.; Pérez-Cobas, A.E.; Latorre, A.; Moya, A. Metatranscriptomic Approach to Analyze the Functional Human Gut Microbiota. PLoS ONE 2011, 6, e17447. [Google Scholar] [CrossRef] [PubMed]
- Mijakovic, I.; Macek, B. Impact of phosphoproteomics on studies of bacterial physiology. FEMS Microbiology Reviews 2012, 36, 877–892. [Google Scholar] [CrossRef] [PubMed]
- Mashego, M.R.; Rumbold, K.; Mey, M.D.; Vandamme, E.; Soetaert, W.; Heijnen, J.J. Microbial metabolomics: past, present and future methodologies. Biotechnology Letters 2006, 29, 1–16. [Google Scholar] [CrossRef]
- Stitt, M.; Gibon, Y. Why measure enzyme activities in the era of systems biology? Trends in Plant Science 2014, 19, 256–265. [Google Scholar] [CrossRef] [PubMed]
- Wiechert, W. 13C Metabolic Flux Analysis. Metabolic Engineering 2001, 3, 195–206. [Google Scholar] [CrossRef]
- Wang, D.; Bodovitz, S. Single cell analysis: the new frontier in ‘omics’. Trends in Biotechnology 2010, 28, 281–290. [Google Scholar] [CrossRef]
- Duncan, K.D.; Fyrestam, J.; Lanekoff, I. Advances in mass spectrometry based single-cell metabolomics. The Analyst 2019, 144, 782–793. [Google Scholar] [CrossRef]
- Zhou, M.; Li, Q.; Wang, R. Current Experimental Methods for Characterizing Protein-Protein Interactions. ChemMedChem 2016, 11, 738–756. [Google Scholar] [CrossRef]
- Maier, R.M.; Pepper, I.L. Bacterial Growth. In Environmental Microbiology; Elsevier, 2015; pp. 37–56. [Google Scholar] [CrossRef]
- Oh, Y.K.; Palsson, B.O.; Park, S.M.; Schilling, C.H.; Mahadevan, R. Genome-scale Reconstruction of Metabolic Network in Bacillus subtilis Based on High-throughput Phenotyping and Gene Essentiality Data. Journal of Biological Chemistry 2007, 282, 28791–28799. [Google Scholar] [CrossRef]
- Noble, J.E.; Knight, A.E.; Reason, A.J.; Matola, A.D.; Bailey, M.J.A. A Comparison of Protein Quantitation Assays for Biopharmaceutical Applications. Molecular Biotechnology 2007, 37, 99–111. [Google Scholar] [CrossRef]
- Noble, J.E.; Bailey, M.J. Chapter 8 Quantitation of Protein. In Methods in Enzymology; Elsevier, 2009; pp. 73–95. [Google Scholar] [CrossRef]
- Stouthamer, A.; Bettenhaussen, C. Utilization of energy for growth and maintenance in continuous and batch cultures of microorganisms. Biochimica et Biophysica Acta (BBA) - Reviews on Bioenergetics 1973, 301, 53–70. [Google Scholar] [CrossRef]
- Motta, S.; Pappalardo, F. Mathematical modeling of biological systems. Briefings in Bioinformatics 2012, 14, 411–422. [Google Scholar] [CrossRef] [PubMed]
- Novère, N.L. Quantitative and logic modelling of molecular and gene networks. Nature Reviews Genetics 2015, 16, 146–158. [Google Scholar] [CrossRef] [PubMed]
- Bruggeman, F.J.; Westerhoff, H.V. The nature of systems biology. Trends in Microbiology 2007, 15, 45–50. [Google Scholar] [CrossRef]
- Baker, R.E.; Peña, J.M.; Jayamohan, J.; Jérusalem, A. Mechanistic models versus machine learning, a fight worth fighting for the biological community? Biology Letters 2018, 14, 20170660. [Google Scholar] [CrossRef] [PubMed]
- Bouwmeester, R.; Gabriels, R.; Bossche, T.V.D.; Martens, L.; Degroeve, S. The Age of Data-Driven Proteomics: How Machine Learning Enables Novel Workflows. PROTEOMICS 2020, 20, 1900351. [Google Scholar] [CrossRef]
- Pasolli, E.; Truong, D.T.; Malik, F.; Waldron, L.; Segata, N. Machine Learning Meta-analysis of Large Metagenomic Datasets: Tools and Biological Insights. PLOS Computational Biology 2016, 12, e1004977. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Mou, M.; Wang, Y.; Luo, Y.; Zhu, F. MetaFS: Performance assessment of biomarker discovery in metaproteomics. Briefings in Bioinformatics 2020, 22. [Google Scholar] [CrossRef]
- Sydor, S.; Dandyk, C.; Schwerdt, J.; Manka, P.; Benndorf, D.; Lehmann, T.; Schallert, K.; Wolf, M.; Reichl, U.; Canbay, A.; Bechmann, L.P.; Heyer, R. Discovering Biomarkers for Non-Alcoholic Steatohepatitis Patients with and without Hepatocellular Carcinoma Using Fecal Metaproteomics. International Journal of Molecular Sciences 2022, 23, 8841. [Google Scholar] [CrossRef]
- Ninfa, A.J.; Ballou, D.P.; Benore, M. Fundamental laboratory approaches for biochemistry and biotechnology; John Wiley & Sons, 2009. [Google Scholar]
- Suthers, P.F.; Foster, C.J.; Sarkar, D.; Wang, L.; Maranas, C.D. Recent advances in constraint and machine learning-based metabolic modeling by leveraging stoichiometric balances, thermodynamic feasibility and kinetic law formalisms. Metabolic Engineering 2021, 63, 13–33. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Yuan, L.; Lu, H.; Li, G.; Chen, Y.; Engqvist, M.K.M.; Kerkhoven, E.J.; Nielsen, J. Deep learning-based kcat prediction enables improved enzyme-constrained model reconstruction. Nature Catalysis 2022, 5, 662–672. [Google Scholar] [CrossRef]
- Koutrouli, M.; Karatzas, E.; Paez-Espino, D.; Pavlopoulos, G.A. A Guide to Conquer the Biological Network Era Using Graph Theory. Frontiers in Bioengineering and Biotechnology 2020, 8. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; Jensen, L.J.; von Mering, C. The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Research 2020, 49, D605–D612. [Google Scholar] [CrossRef] [PubMed]
- Walke, D.; Micheel, D.; Schallert, K.; Muth, T.; Broneske, D.; Saake, G.; Heyer, R. The importance of graph databases and graph learning for clinical applications. Database 2023, 2023, baad045. [Google Scholar] [CrossRef] [PubMed]
- Barbuti, R.; Gori, R.; Milazzo, P.; Nasti, L. A survey of gene regulatory networks modelling methods: from differential equations, to Boolean and qualitative bioinspired models. Journal of Membrane Computing 2020, 2, 207–226. [Google Scholar] [CrossRef]
- Wang, R.S.; Saadatpour, A.; Albert, R. Boolean modeling in systems biology: an overview of methodology and applications. Physical Biology 2012, 9, 055001. [Google Scholar] [CrossRef] [PubMed]
- Karlebach, G.; Shamir, R. Modelling and analysis of gene regulatory networks. Nature Reviews Molecular Cell Biology 2008, 9, 770–780. [Google Scholar] [CrossRef]
- Terzer, M.; Maynard, N.D.; Covert, M.W.; Stelling, J. Genome-scale metabolic networks. WIREs Systems Biology and Medicine 2009, 1, 285–297. [Google Scholar] [CrossRef] [PubMed]
- Aldridge, B.B.; Burke, J.M.; Lauffenburger, D.A.; Sorger, P.K. Physicochemical modelling of cell signalling pathways. Nature Cell Biology 2006, 8, 1195–1203. [Google Scholar] [CrossRef] [PubMed]
- Palsson, B.Ø. Systems biology: simulation of dynamic network states; Cambridge University Press, 2011. [Google Scholar]
- Reed, J.L.; Palsson, B.Ø. Thirteen Years of Building Constraint-Based In Silico Models of Escherichia coli. Journal of Bacteriology 2003, 185, 2692–2699. [Google Scholar] [CrossRef] [PubMed]
- Edwards, J.S.; Ibarra, R.U.; Palsson, B.O. In silico predictions of Escherichia coli metabolic capabilities are consistent with experimental data. Nature Biotechnology 2001, 19, 125–130. [Google Scholar] [CrossRef]
- Bonarius, H.P.; Schmid, G.; Tramper, J. Flux analysis of underdetermined metabolic networks: the quest for the missing constraints. Trends in Biotechnology 1997, 15, 308–314. [Google Scholar] [CrossRef]
- Orth, J.D.; Fleming, R.M.T.; Palsson, B.Ø. Reconstruction and Use of Microbial Metabolic Networks: the Core Escherichia coli Metabolic Model as an Educational Guide. EcoSal Plus 2010, 4. [Google Scholar] [CrossRef]
- Gudmundsson, S.; Thiele, I. Computationally efficient flux variability analysis. BMC Bioinformatics 2010, 11. [Google Scholar] [CrossRef]
- Lewis, N.E.; Nagarajan, H.; Palsson, B.O. Constraining the metabolic genotype–phenotype relationship using a phylogeny of in silico methods. Nature Reviews Microbiology 2012, 10, 291–305. [Google Scholar] [CrossRef]
- Bordbar, A.; Monk, J.M.; King, Z.A.; Palsson, B.O. Constraint-based models predict metabolic and associated cellular functions. Nature Reviews Genetics 2014, 15, 107–120. [Google Scholar] [CrossRef]
- Borisov, N.M.; Chistopolsky, A.S.; Faeder, J.R.; Kholodenko, B.N. Domain-oriented reduction of rule-based network models. IET systems biology 2008, 2, 342–351. [Google Scholar] [CrossRef]
- Münzner, U.; Klipp, E.; Krantz, M. A comprehensive, mechanistically detailed, and executable model of the cell division cycle in Saccharomyces cerevisiae. Nature Communications 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Faeder, J.R.; Blinov, M.L.; Hlavacek, W.S. Rule-Based Modeling of Biochemical Systems with BioNetGen. In Methods in Molecular Biology; Humana Press, 2009; pp. 113–167. [Google Scholar] [CrossRef]
- Romers, J.; Thieme, S.; Münzner, U.; Krantz, M. A scalable method for parameter-free simulation and validation of mechanistic cellular signal transduction network models. npj Systems Biology and Applications 2020, 6. [Google Scholar] [CrossRef]
- Dada, J.O.; Mendes, P. Multi-scale modelling and simulation in systems biology. Integrative Biology 2011, 3, 86. [Google Scholar] [CrossRef] [PubMed]
- Mahadevan, R.; Edwards, J.S.; Doyle, F.J. Dynamic Flux Balance Analysis of Diauxic Growth in Escherichia coli. Biophysical Journal 2002, 83, 1331–1340. [Google Scholar] [CrossRef]
- Sun, G.; Ahn-Horst, T.A.; Covert, M.W. The E. coli Whole-Cell Modeling Project. EcoSal Plus 2021, 9. [Google Scholar] [CrossRef] [PubMed]
- Feist, A.M.; Herrgård, M.J.; Thiele, I.; Reed, J.L.; Palsson, B.Ø. Reconstruction of biochemical networks in microorganisms. Nature Reviews Microbiology 2008, 7, 129–143. [Google Scholar] [CrossRef]
- Gu, C.; Kim, G.B.; Kim, W.J.; Kim, H.U.; Lee, S.Y. Current status and applications of genome-scale metabolic models. Genome Biology 2019, 20. [Google Scholar] [CrossRef]
- Heinken, A.; Basile, A.; Thiele, I. Advances in constraint-based modelling of microbial communities. Current Opinion in Systems Biology 2021, 27, 100346. [Google Scholar] [CrossRef]
- Garza, D.R.; Gonze, D.; Zafeiropoulos, H.; Liu, B.; Faust, K. Metabolic models of human gut microbiota: Advances and challenges. Cell Systems 2023, 14, 109–121. [Google Scholar] [CrossRef]
- do Rosario Martins Conde, P.; Sauter, T.; Pfau, T. Constraint Based Modeling Going Multicellular. Frontiers in Molecular Biosciences 2016, 3. [Google Scholar] [CrossRef]
- Mendes, P.; Hoops, S.; Sahle, S.; Gauges, R.; Dada, J.; Kummer, U. Computational Modeling of Biochemical Networks Using COPASI. In Systems Biology; Humana Press, 2009; pp. 17–59. [Google Scholar] [CrossRef]
- Zimmermann, J.; Kaleta, C.; Waschina, S. gapseq: informed prediction of bacterial metabolic pathways and reconstruction of accurate metabolic models. Genome Biology 2021, 22. [Google Scholar] [CrossRef] [PubMed]
- Lieven, C.; Beber, M.E.; Olivier, B.G.; Bergmann, F.T.; Ataman, M.; Babaei, P.; Bartell, J.A.; Blank, L.M.; Chauhan, S.; Correia, K.; Diener, C.; Dräger, A.; Ebert, B.E.; Edirisinghe, J.N.; Faria, J.P.; Feist, A.M.; Fengos, G.; Fleming, R.M.T.; García-Jiménez, B.; Hatzimanikatis, V.; van Helvoirt, W.; Henry, C.S.; Hermjakob, H.; Herrgård, M.J.; Kaafarani, A.; Kim, H.U.; King, Z.; Klamt, S.; Klipp, E.; Koehorst, J.J.; König, M.; Lakshmanan, M.; Lee, D.Y.; Lee, S.Y.; Lee, S.; Lewis, N.E.; Liu, F.; Ma, H.; Machado, D.; Mahadevan, R.; Maia, P.; Mardinoglu, A.; Medlock, G.L.; Monk, J.M.; Nielsen, J.; Nielsen, L.K.; Nogales, J.; Nookaew, I.; Palsson, B.O.; Papin, J.A.; Patil, K.R.; Poolman, M.; Price, N.D.; Resendis-Antonio, O.; Richelle, A.; Rocha, I.; Sánchez, B.J.; Schaap, P.J.; Sheriff, R.S.M.; Shoaie, S.; Sonnenschein, N.; Teusink, B.; Vilaça, P.; Vik, J.O.; Wodke, J.A.H.; Xavier, J.C.; Yuan, Q.; Zakhartsev, M.; Zhang, C. MEMOTE for standardized genome-scale metabolic model testing. Nature Biotechnology 2020, 38, 272–276. [Google Scholar] [CrossRef] [PubMed]
- Henry, C.S.; Bernstein, H.C.; Weisenhorn, P.; Taylor, R.C.; Lee, J.Y.; Zucker, J.; Song, H.S. Microbial Community Metabolic Modeling: A Community Data-Driven Network Reconstruction. Journal of Cellular Physiology 2016, 231, 2339–2345. [Google Scholar] [CrossRef] [PubMed]
- Greenblum, S.; Turnbaugh, P.J.; Borenstein, E. Metagenomic systems biology of the human gut microbiome reveals topological shifts associated with obesity and inflammatory bowel disease. Proceedings of the National Academy of Sciences 2011, 109, 594–599. [Google Scholar] [CrossRef] [PubMed]
- Klitgord, N.; Segre, D. The importance of compartmentalization in metabolic flux models: yeast as an ecosystem of organelles 2010. pp. 41–55.
- Biggs, M.B.; Medlock, G.L.; Kolling, G.L.; Papin, J.A. Metabolic network modeling of microbial communities. WIREs Systems Biology and Medicine 2015, 7, 317–334. [Google Scholar] [CrossRef] [PubMed]
- Zomorrodi, A.R.; Maranas, C.D. OptCom: A Multi-Level Optimization Framework for the Metabolic Modeling and Analysis of Microbial Communities. PLoS Computational Biology 2012, 8, e1002363. [Google Scholar] [CrossRef]
- Zelezniak, A.; Andrejev, S.; Ponomarova, O.; Mende, D.R.; Bork, P.; Patil, K.R. Metabolic dependencies drive species co-occurrence in diverse microbial communities. Proceedings of the National Academy of Sciences 2015, 112, 6449–6454. [Google Scholar] [CrossRef] [PubMed]
- Diener, C.; Gibbons, S.M.; Resendis-Antonio, O. MICOM: Metagenome-Scale Modeling To Infer Metabolic Interactions in the Gut Microbiota. mSystems 2020, 5. [Google Scholar] [CrossRef]
- Magnúsdóttir, S.; Heinken, A.; Kutt, L.; Ravcheev, D.A.; Bauer, E.; Noronha, A.; Greenhalgh, K.; Jäger, C.; Baginska, J.; Wilmes, P.; Fleming, R.M.T.; Thiele, I. Generation of genome-scale metabolic reconstructions for 773 members of the human gut microbiota. Nature Biotechnology 2016, 35, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Heinken, A.; Hertel, J.; Acharya, G.; Ravcheev, D.A.; Nyga, M.; Okpala, O.E.; Hogan, M.; Magnúsdóttir, S.; Martinelli, F.; Nap, B.; Preciat, G.; Edirisinghe, J.N.; Henry, C.S.; Fleming, R.M.T.; Thiele, I. Genome-scale metabolic reconstruction of 7,302 human microorganisms for personalized medicine. Nature Biotechnology 2023. [Google Scholar] [CrossRef] [PubMed]
- Geier, B.; Sogin, E.M.; Michellod, D.; Janda, M.; Kompauer, M.; Spengler, B.; Dubilier, N.; Liebeke, M. Spatial metabolomics of in situ host–microbe interactions at the micrometre scale. Nature Microbiology 2020, 5, 498–510. [Google Scholar] [CrossRef] [PubMed]
- Walke, D.; Schallert, K.; Ramesh, P.; Benndorf, D.; Lange, E.; Reichl, U.; Heyer, R. MPA_Pathway_Tool: User-Friendly, Automatic Assignment of Microbial Community Data on Metabolic Pathways. International Journal of Molecular Sciences 2021, 22, 10992. [Google Scholar] [CrossRef]
- Li, L.; Figeys, D. Proteomics and Metaproteomics Add Functional, Taxonomic and Biomass Dimensions to Modeling the Ecosystem at the Mucosal-luminal Interface. Molecular Cellular Proteomics 2020, 19, 1409–1417. [Google Scholar] [CrossRef] [PubMed]
- Rosario, D.; Boren, J.; Uhlen, M.; Proctor, G.; Aarsland, D.; Mardinoglu, A.; Shoaie, S. Systems Biology Approaches to Understand the Host–Microbiome Interactions in Neurodegenerative Diseases. Frontiers in Neuroscience 2020, 14. [Google Scholar] [CrossRef] [PubMed]
- Cermak, N.; Becker, J.W.; Knudsen, S.M.; Chisholm, S.W.; Manalis, S.R.; Polz, M.F. Direct single-cell biomass estimates for marine bacteria via Archimedes’ principle. The ISME Journal 2016, 11, 825–828. [Google Scholar] [CrossRef] [PubMed]
- Scott, W.T.; Benito-Vaquerizo, S.; Zimmermann, J.; Bajić, D.; Heinken, A.; Suarez-Diez, M.; Schaap, P.J. A structured evaluation of genome-scale constraint-based modeling tools for microbial consortia. PLOS Computational Biology 2023, 19, e1011363. [Google Scholar] [CrossRef] [PubMed]
- Saez-Rodriguez, J.; Simeoni, L.; Lindquist, J.A.; Hemenway, R.; Bommhardt, U.; Arndt, B.; Haus, U.U.; Weismantel, R.; Gilles, E.D.; Klamt, S.; Schraven, B. A Logical Model Provides Insights into T Cell Receptor Signaling. PLoS Computational Biology 2007, 3, e163. [Google Scholar] [CrossRef] [PubMed]
- Romers, J.; Thieme, S.; Münzner, U.; Krantz, M. Using rxncon to develop rule based models. Modeling Biomolecular Site Dynamics: Methods and Protocols 2018. [Google Scholar] [CrossRef]
- Andrighetti, T.; Bohar, B.; Lemke, N.; Sudhakar, P.; Korcsmaros, T. MicrobioLink: An Integrated Computational Pipeline to Infer Functional Effects of Microbiome–Host Interactions. Cells 2020, 9, 1278. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Beltrán, J.F.; Brito, I.L. Host-microbiome protein-protein interactions capture disease-relevant pathways. Genome Biology 2022, 23. [Google Scholar] [CrossRef] [PubMed]
- Ritz, A.; Poirel, C.L.; Tegge, A.N.; Sharp, N.; Simmons, K.; Powell, A.; Kale, S.D.; Murali, T. Pathways on demand: automated reconstruction of human signaling networks. npj Systems Biology and Applications 2016, 2. [Google Scholar] [CrossRef] [PubMed]
- Türei, D.; Korcsmáros, T.; Saez-Rodriguez, J. OmniPath: guidelines and gateway for literature-curated signaling pathway resources. Nature Methods 2016, 13, 966–967. [Google Scholar] [CrossRef]
- Kandasamy, K.; Mohan, S.; Raju, R.; Keerthikumar, S.; Kumar, G.S.S.; Venugopal, A.K.; Telikicherla, D.; Navarro, D.J.; Mathivanan, S.; Pecquet, C.; Gollapudi, S.K.; Tattikota, S.G.; Mohan, S.; Padhukasahasram, H.; Subbannayya, Y.; Goel, R.; Jacob, H.K.C.; Zhong, J.; Sekhar, R.; Nanjappa, V.; Balakrishnan, L.; Subbaiah, R.; l Ramachandra, Y.; Rahiman, A.; s Keshava Prasad, T.; Lin, J.X.; Houtman, J.C.D.; Desiderio, S.; Renauld, J.C.; Constantinescu, S.; Ohara, O.; Hirano, T.; Kubo, M.; Singh, S.; Khatri, P.; Draghici, S.; Bader, G.D.; Sander, C.; Leonard, W.J.; Pandey, A. NetPath: a public resource of curated signal transduction pathways. Genome Biology 2010, 11, R3. [Google Scholar] [CrossRef] [PubMed]
- Martens, M.; Ammar, A.; Riutta, A.; Waagmeester, A.; Slenter, D.N.; Hanspers, K.; Miller, R.A.; Digles, D.; Lopes, E.N.; Ehrhart, F.; Dupuis, L.J.; Winckers, L.A.; Coort, S.L.; Willighagen, E.L.; Evelo, C.T.; Pico, A.R.; Kutmon, M. WikiPathways: connecting communities. Nucleic Acids Research 2020, 49, D613–D621. [Google Scholar] [CrossRef] [PubMed]
- King, Z.A.; Lu, J.; Dräger, A.; Miller, P.; Federowicz, S.; Lerman, J.A.; Ebrahim, A.; Palsson, B.O.; Lewis, N.E. BiGG Models: A platform for integrating, standardizing and sharing genome-scale models. Nucleic Acids Research 2015, 44, D515–D522. [Google Scholar] [CrossRef]
- Seaver, S.M.D.; Liu, F.; Zhang, Q.; Jeffryes, J.; Faria, J.P.; Edirisinghe, J.N.; Mundy, M.; Chia, N.; Noor, E.; Beber, M.E.; Best, A.A.; DeJongh, M.; Kimbrel, J.A.; D’haeseleer, P.; McCorkle, S.R.; Bolton, J.R.; Pearson, E.; Canon, S.; Wood-Charlson, E.M.; Cottingham, R.W.; Arkin, A.P.; Henry, C.S. The ModelSEED Biochemistry Database for the integration of metabolic annotations and the reconstruction, comparison and analysis of metabolic models for plants, fungi and microbes. Nucleic Acids Research 2020, 49, D575–D588. [Google Scholar] [CrossRef]
- Ronda, C.; Wang, H.H. Engineering temporal dynamics in microbial communities. Current Opinion in Microbiology 2022, 65, 47–55. [Google Scholar] [CrossRef]
- Ashyraliyev, M.; Fomekong-Nanfack, Y.; Kaandorp, J.A.; Blom, J.G. Systems biology: parameter estimation for biochemical models. FEBS Journal 2009, 276, 886–902. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.M.; Tymoczko, J.L.; Gatto, G.J.; Stryer, L. Enzyme: Grundlegende Konzepte und Kinetik. In Stryer Biochemie; Springer: Berlin/Heidelberg, Germany, 2017; pp. 255–297. [Google Scholar] [CrossRef]
- Choi, K.; Medley, J.K.; König, M.; Stocking, K.; Smith, L.; Gu, S.; Sauro, H.M. Tellurium: An extensible python-based modeling environment for systems and synthetic biology. Biosystems 2018, 171, 74–79. [Google Scholar] [CrossRef] [PubMed]
- Albert, R.; Thakar, J. Boolean modeling: a logic-based dynamic approach for understanding signaling and regulatory networks and for making useful predictions. WIREs Systems Biology and Medicine 2014, 6, 353–369. [Google Scholar] [CrossRef] [PubMed]
- Hyduke, D.R.; Lewis, N.E.; Palsson, B.Ø. Analysis of omics data with genome-scale models of metabolism. Mol. BioSyst. 2013, 9, 167–174. [Google Scholar] [CrossRef] [PubMed]
- Agren, R.; Bordel, S.; Mardinoglu, A.; Pornputtapong, N.; Nookaew, I.; Nielsen, J. Reconstruction of Genome-Scale Active Metabolic Networks for 69 Human Cell Types and 16 Cancer Types Using INIT. PLoS Computational Biology 2012, 8, e1002518. [Google Scholar] [CrossRef] [PubMed]
- Agren, R.; Mardinoglu, A.; Asplund, A.; Kampf, C.; Uhlen, M.; Nielsen, J. Identification of anticancer drugs for hepatocellular carcinoma through personalized genome-scale metabolic modeling. Molecular Systems Biology 2014, 10. [Google Scholar] [CrossRef]
- Yizhak, K.; Benyamini, T.; Liebermeister, W.; Ruppin, E.; Shlomi, T. Integrating quantitative proteomics and metabolomics with a genome-scale metabolic network model. Bioinformatics 2010, 26, i255–i260. [Google Scholar] [CrossRef] [PubMed]
- Tian, M.; Reed, J.L. Integrating proteomic or transcriptomic data into metabolic models using linear bound flux balance analysis. Bioinformatics 2018, 34, 3882–3888. [Google Scholar] [CrossRef] [PubMed]
- Opdam, S.; Richelle, A.; Kellman, B.; Li, S.; Zielinski, D.C.; Lewis, N.E. A Systematic Evaluation of Methods for Tailoring Genome-Scale Metabolic Models. Cell Systems 2017, 4, 318–329.e6. [Google Scholar] [CrossRef]
- Kerkhoven, E.J. Advances in constraint-based models: methods for improved predictive power based on resource allocation constraints. Current Opinion in Microbiology 2022, 68, 102168. [Google Scholar] [CrossRef]
- Hädicke, O.; Klamt, S. EColiCore2: a reference network model of the central metabolism of Escherichia coli and relationships to its genome-scale parent model. Scientific Reports 2017, 7. [Google Scholar] [CrossRef]
- Robinson, J.L.; Kocabaş, P.; Wang, H.; Cholley, P.E.; Cook, D.; Nilsson, A.; Anton, M.; Ferreira, R.; Domenzain, I.; Billa, V.; Limeta, A.; Hedin, A.; Gustafsson, J.; Kerkhoven, E.J.; Svensson, L.T.; Palsson, B.O.; Mardinoglu, A.; Hansson, L.; Uhlén, M.; Nielsen, J. An atlas of human metabolism. Science Signaling 2020, 13. [Google Scholar] [CrossRef]
- Erdrich, P.; Steuer, R.; Klamt, S. An algorithm for the reduction of genome-scale metabolic network models to meaningful core models. BMC Systems Biology 2015, 9. [Google Scholar] [CrossRef] [PubMed]
- Danos, V.; Feret, J.; Fontana, W.; Harmer, R.; Krivine, J. Abstracting the Differential Semantics of Rule-Based Models: Exact and Automated Model Reduction. 2010 25th Annual IEEE Symposium on Logic in Computer Science. IEEE, 2010. [CrossRef]
- Wagner, D.; Schlüter, W. Vorhersage und Regelung der Methanproduktion durch maschinelles Lernen. Proceedings ASIM SST 2020. ARGESIM Publisher Vienna, 2020, Vol. 25. [CrossRef]
- Eng, A.; Borenstein, E. Microbial community design: methods, applications, and opportunities. Current Opinion in Biotechnology 2019, 58, 117–128. [Google Scholar] [CrossRef] [PubMed]
- Morgan, X.C.; Tickle, T.L.; Sokol, H.; Gevers, D.; Devaney, K.L.; Ward, D.V.; Reyes, J.A.; Shah, S.A.; LeLeiko, N.; Snapper, S.B.; Bousvaros, A.; Korzenik, J.; Sands, B.E.; Xavier, R.J.; Huttenhower, C. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biology 2012, 13, R79. [Google Scholar] [CrossRef]
- Batstone, D.; Keller, J.; Angelidaki, I.; Kalyuzhnyi, S.; Pavlostathis, S.; Rozzi, A.; Sanders, W.; Siegrist, H.; Vavilin, V. The IWA Anaerobic Digestion Model No 1 (ADM1). Water Science and Technology 2002, 45, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Batstone, D.; Keller, J.; Steyer, J. A review of ADM1 extensions, applications, and analysis: 2002–2005. Water Science and Technology 2006, 54, 1–10. [Google Scholar] [CrossRef]
- Weinrich, S.; Koch, S.; Bonk, F.; Popp, D.; Benndorf, D.; Klamt, S.; Centler, F. Augmenting Biogas Process Modeling by Resolving Intracellular Metabolic Activity. Frontiers in Microbiology 2019, 10. [Google Scholar] [CrossRef]
- Xu, X.; Zarecki, R.; Medina, S.; Ofaim, S.; Liu, X.; Chen, C.; Hu, S.; Brom, D.; Gat, D.; Porob, S.; Eizenberg, H.; Ronen, Z.; Jiang, J.; Freilich, S. Modeling microbial communities from atrazine contaminated soils promotes the development of biostimulation solutions. The ISME Journal 2018, 13, 494–508. [Google Scholar] [CrossRef] [PubMed]
- Stein, R.R.; Tanoue, T.; Szabady, R.L.; Bhattarai, S.K.; Olle, B.; Norman, J.M.; Suda, W.; Oshima, K.; Hattori, M.; Gerber, G.K.; Sander, C.; Honda, K.; Bucci, V. Computer-guided design of optimal microbial consortia for immune system modulation. eLife 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- van Leeuwen, P.T.; Brul, S.; Zhang, J.; Wortel, M.T. Synthetic microbial communities (SynComs) of the human gut: design, assembly, and applications. FEMS Microbiology Reviews 2023, 47. [Google Scholar] [CrossRef]
- Sorbara, M.T.; Pamer, E.G. Microbiome-based therapeutics. Nature Reviews Microbiology 2022, 20, 365–380. [Google Scholar] [CrossRef]
- Lee, T.A.; Steel, H. Cybergenetic control of microbial community composition. Frontiers in Bioengineering and Biotechnology 2022, 10, 1873. [Google Scholar] [CrossRef]
- Gutiérrez Mena, J.; Kumar, S.; Khammash, M. Dynamic cybergenetic control of bacterial co-culture composition via optogenetic feedback. Nature Communications 2022, 13. [Google Scholar] [CrossRef]
- Aditya, C.; Bertaux, F.; Batt, G.; Ruess, J. A light tunable differentiation system for the creation and control of consortia in yeast. Nature Communications 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Scott, S.R.; Hasty, J. Quorum Sensing Communication Modules for Microbial Consortia. ACS Synthetic Biology 2016, 5, 969–977. [Google Scholar] [CrossRef] [PubMed]
- Lu, T.K.; Collins, J.J. Dispersing biofilms with engineered enzymatic bacteriophage. Proceedings of the National Academy of Sciences 2007, 104, 11197–11202. [Google Scholar] [CrossRef]
- Buysschaert, B.; Kerckhof, F.; Vandamme, P.; De Baets, B.; Boon, N. Flow cytometric fingerprinting for microbial strain discrimination and physiological characterization. Cytometry Part A 2017, 93, 201–212. [Google Scholar] [CrossRef] [PubMed]
- Khesali Aghtaei, H.; Püttker, S.; Maus, I.; Heyer, R.; Huang, L.; Sczyrba, A.; Reichl, U.; Benndorf, D. Adaptation of a microbial community to demand-oriented biological methanation. Biotechnology for Biofuels and Bioproducts 2022, 15. [Google Scholar] [CrossRef] [PubMed]
- Bensmann, A.; Hanke-Rauschenbach, R.; Heyer, R.; Kohrs, F.; Benndorf, D.; Reichl, U.; Sundmacher, K. Biological methanation of hydrogen within biogas plants: A model-based feasibility study. Applied Energy 2014, 134, 413–425. [Google Scholar] [CrossRef]
- Simon, D. Kalman filtering. Embedded systems programming 2001, 14, 72–79. [Google Scholar]
- Waltemath, D.; Wolkenhauer, O. How Modeling Standards, Software, and Initiatives Support Reproducibility in Systems Biology and Systems Medicine. IEEE Transactions on Biomedical Engineering 2016, 63, 1999–2006. [Google Scholar] [CrossRef]
- Stanford, N.J.; Scharm, M.; Dobson, P.D.; Golebiewski, M.; Hucka, M.; Kothamachu, V.B.; Nickerson, D.; Owen, S.; Pahle, J.; Wittig, U.; Waltemath, D.; Goble, C.; Mendes, P.; Snoep, J. Data Management in Computational Systems Biology: Exploring Standards, Tools, Databases, and Packaging Best Practices. In Methods in Molecular Biology; Springer: New York, 2019; pp. 285–314. [Google Scholar] [CrossRef]
- Wilkinson, M.D.; Dumontier, M.; Aalbersberg, I.J.; Appleton, G.; Axton, M.; Baak, A.; Blomberg, N.; Boiten, J.W.; da Silva Santos, L.B.; Bourne, P.E.; Bouwman, J.; Brookes, A.J.; Clark, T.; Crosas, M.; Dillo, I.; Dumon, O.; Edmunds, S.; Evelo, C.T.; Finkers, R.; Gonzalez-Beltran, A.; Gray, A.J.; Groth, P.; Goble, C.; Grethe, J.S.; Heringa, J.; ’t Hoen, P.A.; Hooft, R.; Kuhn, T.; Kok, R.; Kok, J.; Lusher, S.J.; Martone, M.E.; Mons, A.; Packer, A.L.; Persson, B.; Rocca-Serra, P.; Roos, M.; van Schaik, R.; Sansone, S.A.; Schultes, E.; Sengstag, T.; Slater, T.; Strawn, G.; Swertz, M.A.; Thompson, M.; van der Lei, J.; van Mulligen, E.; Velterop, J.; Waagmeester, A.; Wittenburg, P.; Wolstencroft, K.; Zhao, J.; Mons, B. The FAIR Guiding Principles for scientific data management and stewardship. Scientific Data 2016, 3. [Google Scholar] [CrossRef]
- Boeckhout, M.; Zielhuis, G.A.; Bredenoord, A.L. The FAIR guiding principles for data stewardship: fair enough? European Journal of Human Genetics 2018, 26, 931–936. [Google Scholar] [CrossRef]
- Hughes, L.D.; Tsueng, G.; DiGiovanna, J.; Horvath, T.D.; Rasmussen, L.V.; Savidge, T.C.; Stoeger, T.; Turkarslan, S.; Wu, Q.; Wu, C.; Su, A.I.; and, L.P. Addressing barriers in FAIR data practices for biomedical data. Scientific Data 2023, 10. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, K.; Kananathan, S.; Roberts, M.G.; Meyer, J.P.; Shohan, M.U.S.; Xavier, A.; Maire, M.; Zyoud, A.; Men, J.; Ng, S.; Nguyen, T.V.N.; Glont, M.; Hermjakob, H.; Malik-Sheriff, R.S. Reproducibility in systems biology modelling. Molecular Systems Biology 2021, 17. [Google Scholar] [CrossRef]
- Wolstencroft, K.; Krebs, O.; Snoep, J.L.; Stanford, N.J.; Bacall, F.; Golebiewski, M.; Kuzyakiv, R.; Nguyen, Q.; Owen, S.; Soiland-Reyes, S.; Straszewski, J.; van Niekerk, D.D.; Williams, A.R.; Malmström, L.; Rinn, B.; Müller, W.; Goble, C. FAIRDOMHub: a repository and collaboration environment for sharing systems biology research. Nucleic Acids Research 2016, 45, D404–D407. [Google Scholar] [CrossRef] [PubMed]
- Hucka, M.; Nickerson, D.P.; Bader, G.D.; Bergmann, F.T.; Cooper, J.; Demir, E.; Garny, A.; Golebiewski, M.; Myers, C.J.; Schreiber, F.; Waltemath, D.; Novère, N.L. Promoting Coordinated Development of Community-Based Information Standards for Modeling in Biology: The COMBINE Initiative. Frontiers in Bioengineering and Biotechnology 2015, 3. [Google Scholar] [CrossRef] [PubMed]
- Waltemath, D.; Golebiewski, M.; Blinov, M.L.; Gleeson, P.; Hermjakob, H.; Hucka, M.; Inau, E.T.; Keating, S.M.; König, M.; Krebs, O.; Malik-Sheriff, R.S.; Nickerson, D.; Oberortner, E.; Sauro, H.M.; Schreiber, F.; Smith, L.; Stefan, M.I.; Wittig, U.; Myers, C.J. The first 10 years of the international coordination network for standards in systems and synthetic biology (COMBINE). Journal of Integrative Bioinformatics 2020, 17. [Google Scholar] [CrossRef] [PubMed]
- Naldi, A.; Monteiro, P.T.; Müssel, C.; Kestler, H.A.; Thieffry, D.; Xenarios, I.; Saez-Rodriguez, J.; Helikar, T.; and, C.C. Cooperative development of logical modelling standards and tools with CoLoMoTo. Bioinformatics 2015, 31, 1154–1159. [Google Scholar] [CrossRef] [PubMed]
- Tatka, L.T.; Smith, L.P.; Hellerstein, J.L.; Sauro, H.M. Adapting modeling and simulation credibility standards to computational systems biology. Journal of Translational Medicine 2023, 21. [Google Scholar] [CrossRef] [PubMed]
- Novère, N.L.; Finney, A.; Hucka, M.; Bhalla, U.S.; Campagne, F.; Collado-Vides, J.; Crampin, E.J.; Halstead, M.; Klipp, E.; Mendes, P.; Nielsen, P.; Sauro, H.; Shapiro, B.; Snoep, J.L.; Spence, H.D.; Wanner, B.L. Minimum information requested in the annotation of biochemical models (MIRIAM). Nature Biotechnology 2005, 23, 1509–1515. [Google Scholar] [CrossRef]
- Waltemath, D.; Adams, R.; Beard, D.A.; Bergmann, F.T.; Bhalla, U.S.; Britten, R.; Chelliah, V.; Cooling, M.T.; Cooper, J.; Crampin, E.J.; Garny, A.; Hoops, S.; Hucka, M.; Hunter, P.; Klipp, E.; Laibe, C.; Miller, A.K.; Moraru, I.; Nickerson, D.; Nielsen, P.; Nikolski, M.; Sahle, S.; Sauro, H.M.; Schmidt, H.; Snoep, J.L.; Tolle, D.; Wolkenhauer, O.; Novère, N.L. Minimum Information About a Simulation Experiment (MIASE). PLoS Computational Biology 2011, 7, e1001122. [Google Scholar] [CrossRef]
- Carey, M.A.; Dräger, A.; Beber, M.E.; Papin, J.A.; Yurkovich, J.T. Community standards to facilitate development and address challenges in metabolic modeling. Molecular Systems Biology 2020, 16. [Google Scholar] [CrossRef] [PubMed]
- Hucka, M.; Bergmann, F.T.; Chaouiya, C.; Dräger, A.; Hoops, S.; Keating, S.M.; König, M.; Novère, N.L.; Myers, C.J.; Olivier, B.G.; Sahle, S.; Schaff, J.C.; Sheriff, R.; Smith, L.P.; Waltemath, D.; Wilkinson, D.J.; Zhang, F. The Systems Biology Markup Language (SBML): Language Specification for Level 3 Version 2 Core Release 2. Journal of Integrative Bioinformatics 2019, 16. [Google Scholar] [CrossRef] [PubMed]
- Keating, S.M.; Waltemath, D.; König, M.; Zhang, F.; Dräger, A.; Chaouiya, C.; Bergmann, F.T.; Finney, A.; Gillespie, C.S.; Helikar, T.; Hoops, S.; Malik-Sheriff, R.S.; Moodie, S.L.; Moraru, I.I.; Myers, C.J.; Naldi, A.; Olivier, B.G.; Sahle, S.; Schaff, J.C.; Smith, L.P.; Swat, M.J.; Thieffry, D.; Watanabe, L.; Wilkinson, D.J.; Blinov, M.L.; Begley, K.; Faeder, J.R.; Gómez, H.F.; Hamm, T.M.; Inagaki, Y.; Liebermeister, W.; Lister, A.L.; Lucio, D.; Mjolsness, E.; Proctor, C.J.; Raman, K.; Rodriguez, N.; Shaffer, C.A.; Shapiro, B.E.; Stelling, J.; Swainston, N.; Tanimura, N.; Wagner, J.; Meier-Schellersheim, M.; Sauro, H.M.; Palsson, B.; Bolouri, H.; Kitano, H.; Funahashi, A.; Hermjakob, H.; Doyle, J.C.; Hucka, M.; Adams, R.R.; Allen, N.A.; Angermann, B.R.; Antoniotti, M.; Bader, G.D.; Červený, J.; Courtot, M.; Cox, C.D.; Pezze, P.D.; Demir, E.; Denney, W.S.; Dharuri, H.; Dorier, J.; Drasdo, D.; Ebrahim, A.; Eichner, J.; Elf, J.; Endler, L.; Evelo, C.T.; Flamm, C.; Fleming, R.M.; Fröhlich, M.; Glont, M.; Gonçalves, E.; Golebiewski, M.; Grabski, H.; Gutteridge, A.; Hachmeister, D.; Harris, L.A.; Heavner, B.D.; Henkel, R.; Hlavacek, W.S.; Hu, B.; Hyduke, D.R.; de Jong, H.; Juty, N.; Karp, P.D.; Karr, J.R.; Kell, D.B.; Keller, R.; Kiselev, I.; Klamt, S.; Klipp, E.; Knüpfer, C.; Kolpakov, F.; Krause, F.; Kutmon, M.; Laibe, C.; Lawless, C.; Li, L.; Loew, L.M.; Machne, R.; Matsuoka, Y.; Mendes, P.; Mi, H.; Mittag, F.; Monteiro, P.T.; Natarajan, K.N.; Nielsen, P.M.; Nguyen, T.; Palmisano, A.; Pettit, J.B.; Pfau, T.; Phair, R.D.; Radivoyevitch, T.; Rohwer, J.M.; Ruebenacker, O.A.; Saez-Rodriguez, J.; Scharm, M.; Schmidt, H.; Schreiber, F.; Schubert, M.; Schulte, R.; Sealfon, S.C.; Smallbone, K.; Soliman, S.; Stefan, M.I.; Sullivan, D.P.; Takahashi, K.; Teusink, B.; Tolnay, D.; Vazirabad, I.; von Kamp, A.; Wittig, U.; Wrzodek, C.; Wrzodek, F.; Xenarios, I.; Zhukova, A.; and, J.Z. SBML Level 3: an extensible format for the exchange and reuse of biological models. Molecular Systems Biology 2020, 16. [Google Scholar] [CrossRef] [PubMed]
- Lubitz, T.; Hahn, J.; Bergmann, F.T.; Noor, E.; Klipp, E.; Liebermeister, W. SBtab: a flexible table format for data exchange in systems biology. Bioinformatics 2016, 32, 2559–2561. [Google Scholar] [CrossRef]
- Köhn, D.; Novère, N.L. SED-ML – An XML Format for the Implementation of the MIASE Guidelines. In Computational Methods in Systems Biology; Springer: Berlin/Heidelberg, Germany, 2008; pp. 176–190. [Google Scholar] [CrossRef]
- Heirendt, L.; Arreckx, S.; Pfau, T.; Mendoza, S.N.; Richelle, A.; Heinken, A.; Haraldsdóttir, H.S.; Wachowiak, J.; Keating, S.M.; Vlasov, V.; Magnusdóttir, S.; Ng, C.Y.; Preciat, G.; Žagare, A.; Chan, S.H.J.; Aurich, M.K.; Clancy, C.M.; Modamio, J.; Sauls, J.T.; Noronha, A.; Bordbar, A.; Cousins, B.; Assal, D.C.E.; Valcarcel, L.V.; Apaolaza, I.; Ghaderi, S.; Ahookhosh, M.; Guebila, M.B.; Kostromins, A.; Sompairac, N.; Le, H.M.; Ma, D.; Sun, Y.; Wang, L.; Yurkovich, J.T.; Oliveira, M.A.P.; Vuong, P.T.; Assal, L.P.E.; Kuperstein, I.; Zinovyev, A.; Hinton, H.S.; Bryant, W.A.; Artacho, F.J.A.; Planes, F.J.; Stalidzans, E.; Maass, A.; Vempala, S.; Hucka, M.; Saunders, M.A.; Maranas, C.D.; Lewis, N.E.; Sauter, T.; Palsson, B.Ø.; Thiele, I.; Fleming, R.M.T. Creation and analysis of biochemical constraint-based models using the COBRA Toolbox v.3.0. Nature Protocols 2019, 14, 639–702. [Google Scholar] [CrossRef]
- Ebrahim, A.; Lerman, J.A.; Palsson, B.O.; Hyduke, D.R. COBRApy: COnstraints-Based Reconstruction and Analysis for Python. BMC Systems Biology 2013, 7. [Google Scholar] [CrossRef]
- Camborda, S.; Weder, J.N.; Töpfer, N. CobraMod: a pathway-centric curation tool for constraint-based metabolic models. Bioinformatics 2022, 38, 2654–2656. [Google Scholar] [CrossRef]
- Bauer, E.; Zimmermann, J.; Baldini, F.; Thiele, I.; Kaleta, C. BacArena: Individual-based metabolic modeling of heterogeneous microbes in complex communities. PLOS Computational Biology 2017, 13, e1005544. [Google Scholar] [CrossRef]
- Arkin, A.P.; Cottingham, R.W.; Henry, C.S.; Harris, N.L.; Stevens, R.L.; Maslov, S.; Dehal, P.; Ware, D.; Perez, F.; Canon, S.; Sneddon, M.W.; Henderson, M.L.; Riehl, W.J.; Murphy-Olson, D.; Chan, S.Y.; Kamimura, R.T.; Kumari, S.; Drake, M.M.; Brettin, T.S.; Glass, E.M.; Chivian, D.; Gunter, D.; Weston, D.J.; Allen, B.H.; Baumohl, J.; Best, A.A.; Bowen, B.; Brenner, S.E.; Bun, C.C.; Chandonia, J.M.; Chia, J.M.; Colasanti, R.; Conrad, N.; Davis, J.J.; Davison, B.H.; DeJongh, M.; Devoid, S.; Dietrich, E.; Dubchak, I.; Edirisinghe, J.N.; Fang, G.; Faria, J.P.; Frybarger, P.M.; Gerlach, W.; Gerstein, M.; Greiner, A.; Gurtowski, J.; Haun, H.L.; He, F.; Jain, R.; Joachimiak, M.P.; Keegan, K.P.; Kondo, S.; Kumar, V.; Land, M.L.; Meyer, F.; Mills, M.; Novichkov, P.S.; Oh, T.; Olsen, G.J.; Olson, R.; Parrello, B.; Pasternak, S.; Pearson, E.; Poon, S.S.; Price, G.A.; Ramakrishnan, S.; Ranjan, P.; Ronald, P.C.; Schatz, M.C.; Seaver, S.M.D.; Shukla, M.; Sutormin, R.A.; Syed, M.H.; Thomason, J.; Tintle, N.L.; Wang, D.; Xia, F.; Yoo, H.; Yoo, S.; Yu, D. KBase: The United States Department of Energy Systems Biology Knowledgebase. Nature Biotechnology 2018, 36, 566–569. [Google Scholar] [CrossRef]
- Klarner, H.; Streck, A.; Siebert, H. PyBoolNet: a python package for the generation, analysis and visualization of boolean networks. Bioinformatics 2016, 33, 770–772. [Google Scholar] [CrossRef]
- von Kamp, A.; Thiele, S.; Hädicke, O.; Klamt, S. Use of CellNetAnalyzer in biotechnology and metabolic engineering. Journal of Biotechnology 2017, 261, 221–228. [Google Scholar] [CrossRef] [PubMed]
- Müssel, C.; Hopfensitz, M.; Kestler, H.A. BoolNet—an R package for generation, reconstruction and analysis of Boolean networks. Bioinformatics 2010, 26, 1378–1380. [Google Scholar] [CrossRef] [PubMed]
- Helikar, T.; Kowal, B.; McClenathan, S.; Bruckner, M.; Rowley, T.; Madrahimov, A.; Wicks, B.; Shrestha, M.; Limbu, K.; Rogers, J.A. The Cell Collective: Toward an open and collaborative approach to systems biology. BMC Systems Biology 2012, 6, 96. [Google Scholar] [CrossRef] [PubMed]
- Matsuoka, Y.; Funahashi, A.; Ghosh, S.; Kitano, H. Modeling and Simulation Using CellDesigner. In Transcription Factor Regulatory Networks; Springer: New York, 2014; pp. 121–145. [Google Scholar] [CrossRef]
- Garny, A.; Hunter, P.J. OpenCOR: a modular and interoperable approach to computational biology. Frontiers in Physiology 2015, 6. [Google Scholar] [CrossRef]
- Hoops, S.; Sahle, S.; Gauges, R.; Lee, C.; Pahle, J.; Simus, N.; Singhal, M.; Xu, L.; Mendes, P.; Kummer, U. COPASI—a COmplex PAthway SImulator. Bioinformatics 2006, 22, 3067–3074. [Google Scholar] [CrossRef] [PubMed]
- Olivier, B.G.; Rohwer, J.M.; Hofmeyr, J.H.S. Modelling cellular systems with PySCeS. Bioinformatics 2004, 21, 560–561. [Google Scholar] [CrossRef] [PubMed]
- Lopez, C.F.; Muhlich, J.L.; Bachman, J.A.; Sorger, P.K. Programming biological models in Python using PySB. Molecular Systems Biology 2013, 9. [Google Scholar] [CrossRef] [PubMed]
- Starruß, J.; de Back, W.; Brusch, L.; Deutsch, A. Morpheus: a user-friendly modeling environment for multiscale and multicellular systems biology. Bioinformatics 2014, 30, 1331–1332. [Google Scholar] [CrossRef]
- Wittig, U.; Rey, M.; Weidemann, A.; Kania, R.; Müller, W. SABIO-RK: an updated resource for manually curated biochemical reaction kinetics. Nucleic Acids Research 2017, 46, D656–D660. [Google Scholar] [CrossRef]
- Mitchell, A.L.; Almeida, A.; Beracochea, M.; Boland, M.; Burgin, J.; Cochrane, G.; Crusoe, M.R.; Kale, V.; Potter, S.C.; Richardson, L.J.; Sakharova, E.; Scheremetjew, M.; Korobeynikov, A.; Shlemov, A.; Kunyavskaya, O.; Lapidus, A.; Finn, R.D. MGnify: the microbiome analysis resource in 2020. Nucleic Acids Research 2019. [Google Scholar] [CrossRef] [PubMed]
- Malik-Sheriff, R.S.; Glont, M.; Nguyen, T.V.N.; Tiwari, K.; Roberts, M.G.; Xavier, A.; Vu, M.T.; Men, J.; Maire, M.; Kananathan, S.; Fairbanks, E.L.; Meyer, J.P.; Arankalle, C.; Varusai, T.M.; Knight-Schrijver, V.; Li, L.; Dueñas-Roca, C.; Dass, G.; Keating, S.M.; Park, Y.M.; Buso, N.; Rodriguez, N.; Hucka, M.; Hermjakob, H. BioModels—15 years of sharing computational models in life science. Nucleic Acids Research 2019. [Google Scholar] [CrossRef] [PubMed]
- Moretti, S.; Tran, V.D.T.; Mehl, F.; Ibberson, M.; Pagni, M. MetaNetX/MNXref: unified namespace for metabolites and biochemical reactions in the context of metabolic models. Nucleic Acids Research 2020, 49, D570–D574. [Google Scholar] [CrossRef]
- Ankrah, N.Y.D.; Barker, B.E.; Song, J.; Wu, C.; McMullen, J.G.; Douglas, A.E. Predicted Metabolic Function of the Gut Microbiota of Drosophila melanogaster. mSystems 2021, 6. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.M.; Poline, J.B.; Dumas, G. Experimenting with reproducibility: a case study of robustness in bioinformatics. GigaScience 2018, 7. [Google Scholar] [CrossRef]
- Boettiger, C. An introduction to Docker for reproducible research. ACM SIGOPS Operating Systems Review 2015, 49, 71–79. [Google Scholar] [CrossRef]
- Pham, N.; van Heck, R.; van Dam, J.; Schaap, P.; Saccenti, E.; Suarez-Diez, M. Consistency, Inconsistency, and Ambiguity of Metabolite Names in Biochemical Databases Used for Genome-Scale Metabolic Modelling. Metabolites 2019, 9, 28. [Google Scholar] [CrossRef] [PubMed]
- Chindelevitch, L.; Stanley, S.; Hung, D.; Regev, A.; Berger, B. MetaMerge: scaling up genome-scale metabolic reconstructions, with application to Mycobacterium tuberculosis. Genome Biology 2012, 13, R6. [Google Scholar] [CrossRef]
- Ravikrishnan, A.; Raman, K. Critical assessment of genome-scale metabolic networks: the need for a unified standard. Briefings in Bioinformatics 2015, 16, 1057–1068. [Google Scholar] [CrossRef]
- Goodman, J.M.; Pletnev, I.; Thiessen, P.; Bolton, E.; Heller, S.R. InChI version 1.06: now more than 99.99. Journal of Cheminformatics 2021, 13. [Google Scholar] [CrossRef]
- Hastings, J.; Owen, G.; Dekker, A.; Ennis, M.; Kale, N.; Muthukrishnan, V.; Turner, S.; Swainston, N.; Mendes, P.; Steinbeck, C. ChEBI in 2016: Improved services and an expanding collection of metabolites. Nucleic Acids Research 2015, 44, D1214–D1219. [Google Scholar] [CrossRef]
- Vieira, L.S.; Laubenbacher, R.C. Computational models in systems biology: standards, dissemination, and best practices. Current Opinion in Biotechnology 2022, 75, 102702. [Google Scholar] [CrossRef]
- Papin, J.A.; Gabhann, F.M.; Sauro, H.M.; Nickerson, D.; Rampadarath, A. Improving reproducibility in computational biology research. PLOS Computational Biology 2020, 16, e1007881. [Google Scholar] [CrossRef] [PubMed]
- Wolf, M.; Schallert, K.; Knipper, L.; Sickmann, A.; Sczyrba, A.; Benndorf, D.; Heyer, R. Advances in the clinical use of metaproteomics. Expert Review of Proteomics 2023, 20, 71–86. [Google Scholar] [CrossRef] [PubMed]
- Thiele, I.; Clancy, C.M.; Heinken, A.; Fleming, R.M. Quantitative systems pharmacology and the personalized drug–microbiota–diet axis. Current Opinion in Systems Biology 2017, 4, 43–52. [Google Scholar] [CrossRef]
- Thiele, I.; Sahoo, S.; Heinken, A.; Hertel, J.; Heirendt, L.; Aurich, M.K.; Fleming, R.M. Personalized whole-body models integrate metabolism, physiology, and the gut microbiome. Molecular Systems Biology 2020, 16. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Li, F.; Sánchez, B.J.; Zhu, Z.; Li, G.; Domenzain, I.; Marcišauskas, S.; Anton, P.M.; Lappa, D.; Lieven, C.; Beber, M.E.; Sonnenschein, N.; Kerkhoven, E.J.; Nielsen, J. A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism. Nature Communications 2019, 10. [Google Scholar] [CrossRef] [PubMed]
| 1 | With the term "microbiomes", we include the terms "microbial community" or "microbiota" and refer to any community of microorganisms living with or without a eukaryotic host. |
| 2 |
https://www.destatis.de/DE/Presse/Pressemitteilungen/2023/03/PD23_090_43312.html, date of access: December 13, 2023 |
| 3 |
https://www.umweltbundesamt.de/daten/energie/erneuerbare-energien-vermiedene-treibhausgase#stromerzeugung, date of access: December 13, 2023 |
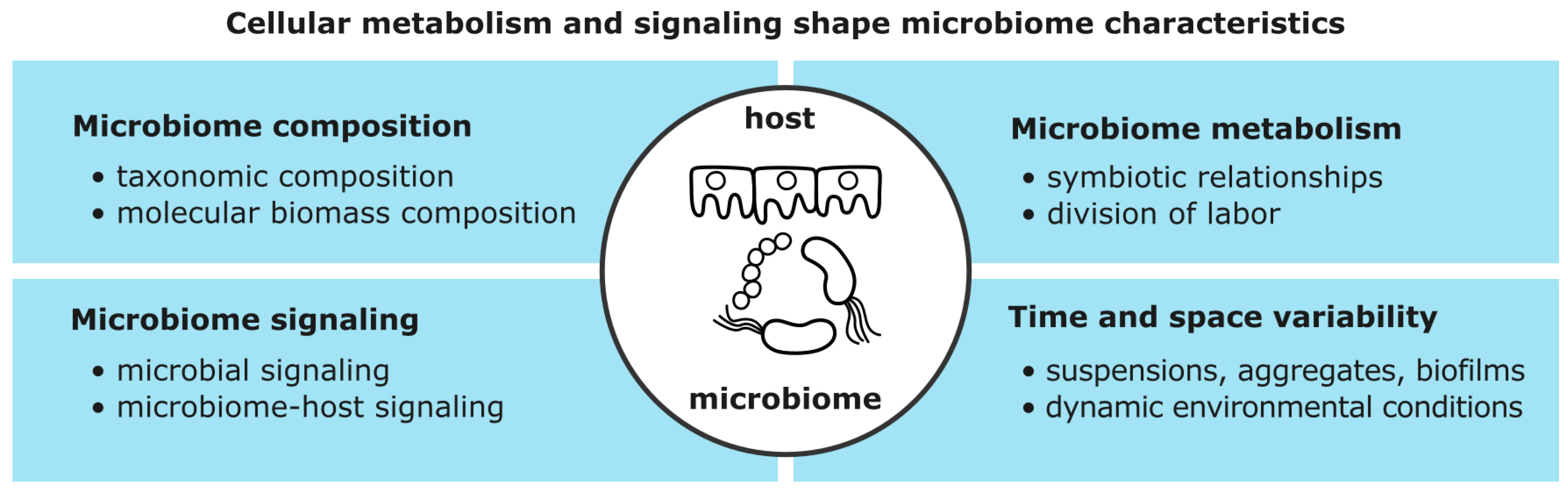
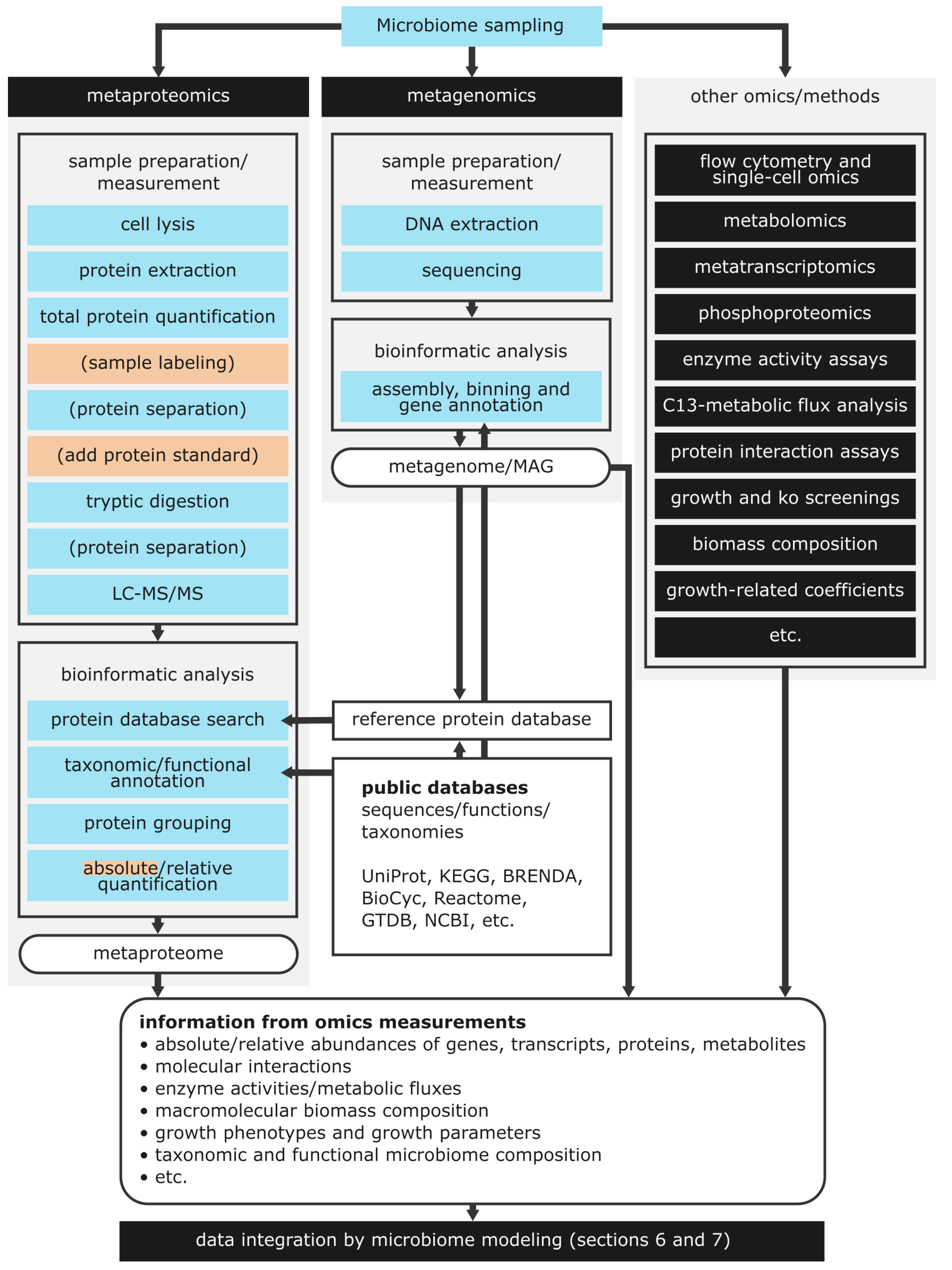
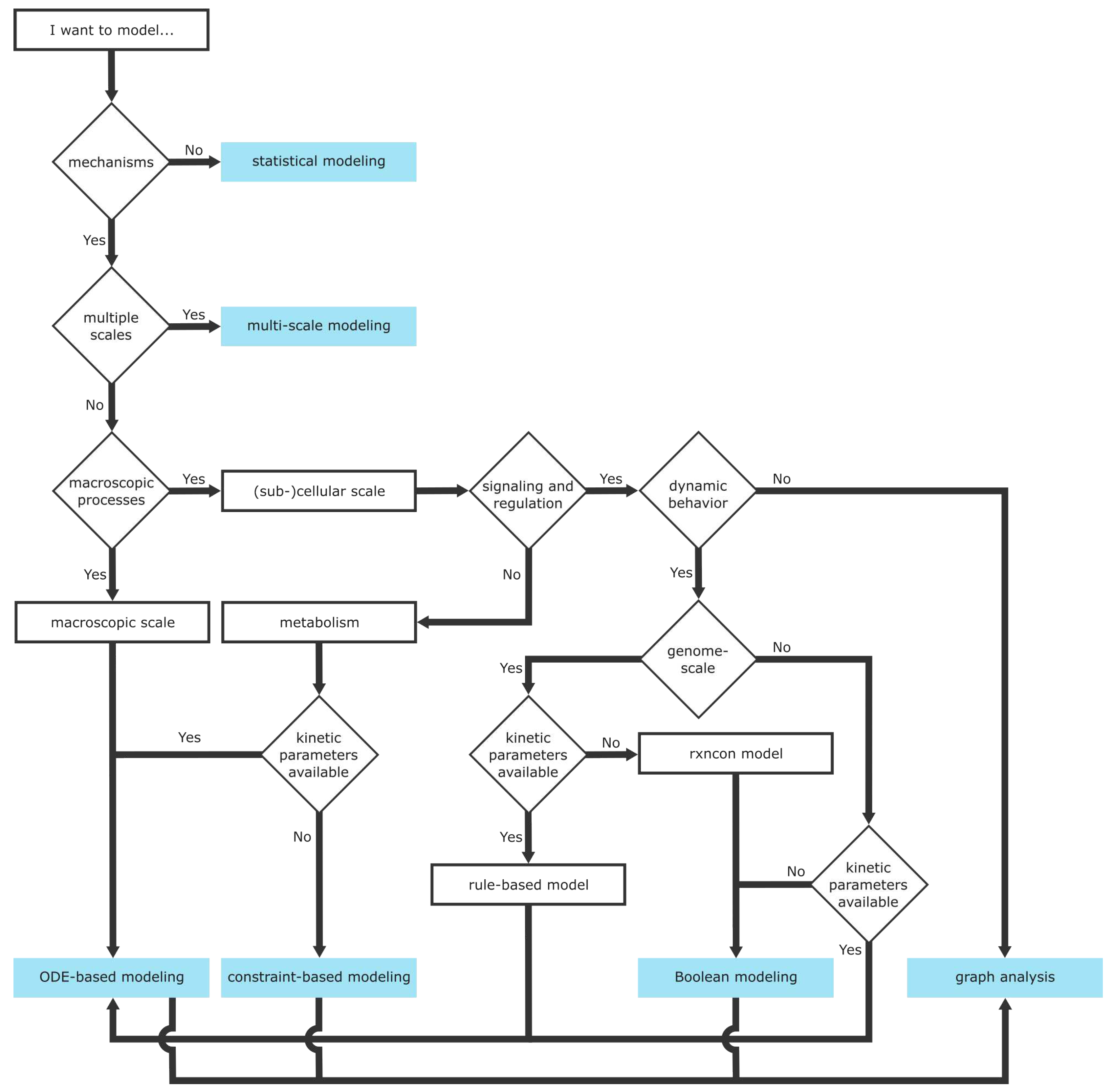
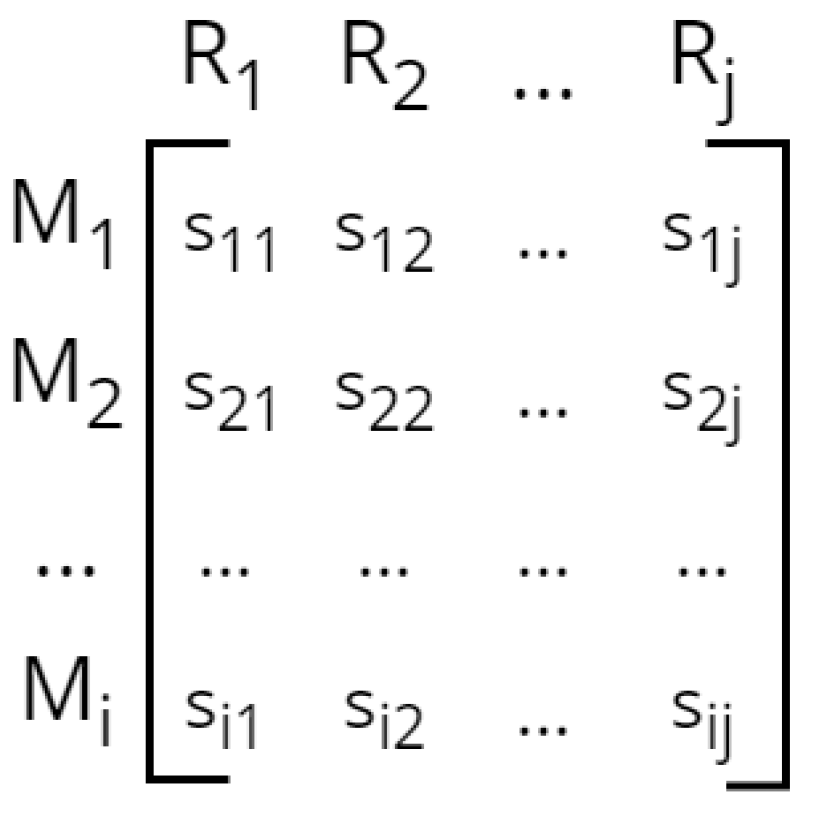
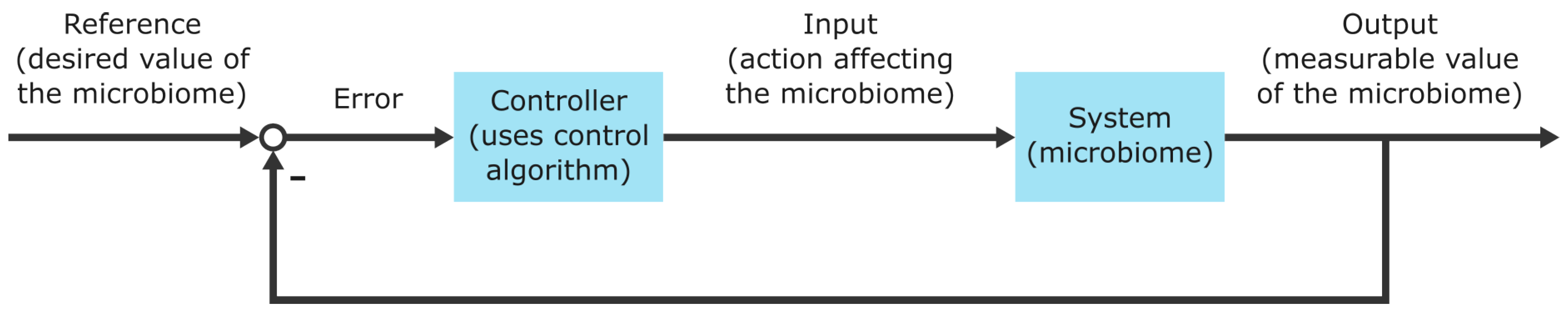
| section | Script query | date | Number of hits | Topics |
|---|---|---|---|---|
| 4 | (microbiome) AND (microbial community) | November 7, 2023 | 4465 | Role and properties of microbiomes |
| 5 | (metaproteomics) OR (metagenomics) OR (metaomics) | November 7, 2023 | 4200 | Metaomics methods, metaproteomics, bioinformatic challenges |
| 6 | (computational model) AND ((metabolism) OR (regulation) OR (signaling)) | November 7, 2023 | 3163 | model types, modeling approaches applicable to metabolism and signaling |
| 7.1 | (biological network reconstruction) AND ((microbiome) OR (microbial community)) | November 7, 2023 | 6531 | Reconstruction of metabolic and signaling networks |
| 7.2 | (computational model) AND ((parameter estimation) OR (contextualization) OR (reduction)) | November 7, 2023 | 1035 | parameter estimation, context-specific models, reduction of model size |
| 8.1 and 8.2 | (computational modeling) AND ((microbiome) OR (microbial community)) | November 7, 2023 | 1035 | examples of prediction, optimization |
| 8.3 | (control algorithm) AND ((microbiome) OR (microbial community)) | November 7, 2023 | 4313 | microbiome control |
| 9 | (network modeling) AND (guidelines OR software OR repository) | November 7, 2023 | 1671 | FAIR, initiatives, standards, languages, software, repositories |
| Data type/Method | References |
|---|---|
| WGS/amplicon | Bragg and Tyson [131], |
| Segata et al. [132], | |
| Frioux et al. [133], | |
| Jünemann et al. [12], | |
| Zorrilla et al. [134] | |
| metatranscriptomics | Bashiardes et al. [135], |
| Gifford et al. [136], | |
| Gosalbes et al. [137] | |
| phosphoproteomics | Terfve and Saez-Rodriguez [41], |
| Mijakovic and Macek [138] | |
| metabolomics | Mashego et al. [139], |
| Zhang et al. [130], | |
| Liu and Locasale [129], | |
| Bauermeister et al. [128] | |
| enzyme activity assays | Stitt and Gibon [140] |
| C13-metabolic flux analysis | Winter and Krömer [74], |
| Wiechert [141], | |
| Zamboni et al. [72] | |
| single-cell omics | Wang and Bodovitz [142], |
| Hatzenpichler et al. [126], | |
| Duncan et al. [143] | |
| protein interaction data | Zhou et al. [144] |
| growth screenings | Maier and Pepper [145], |
| Oh et al. [146] | |
| knock out screenings and gene essentiality data | Oh et al. [146] |
| biomass composition | Beck et al. [69], |
| Lachance et al. [70] | |
| total protein content | Noble et al. [147], |
| Noble and Bailey [148] | |
| maintenance coefficients | Stouthamer and Bettenhaussen [149], |
| Vos et al. [71] | |
| microscopy | Cesar and Huang [127] |
| flow cytometry | Hatzenpichler et al. [126], |
| Props et al. [125] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




