Submitted:
27 December 2023
Posted:
09 January 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. MAPK-related genes are upregulated in MSC-induced MDSCs
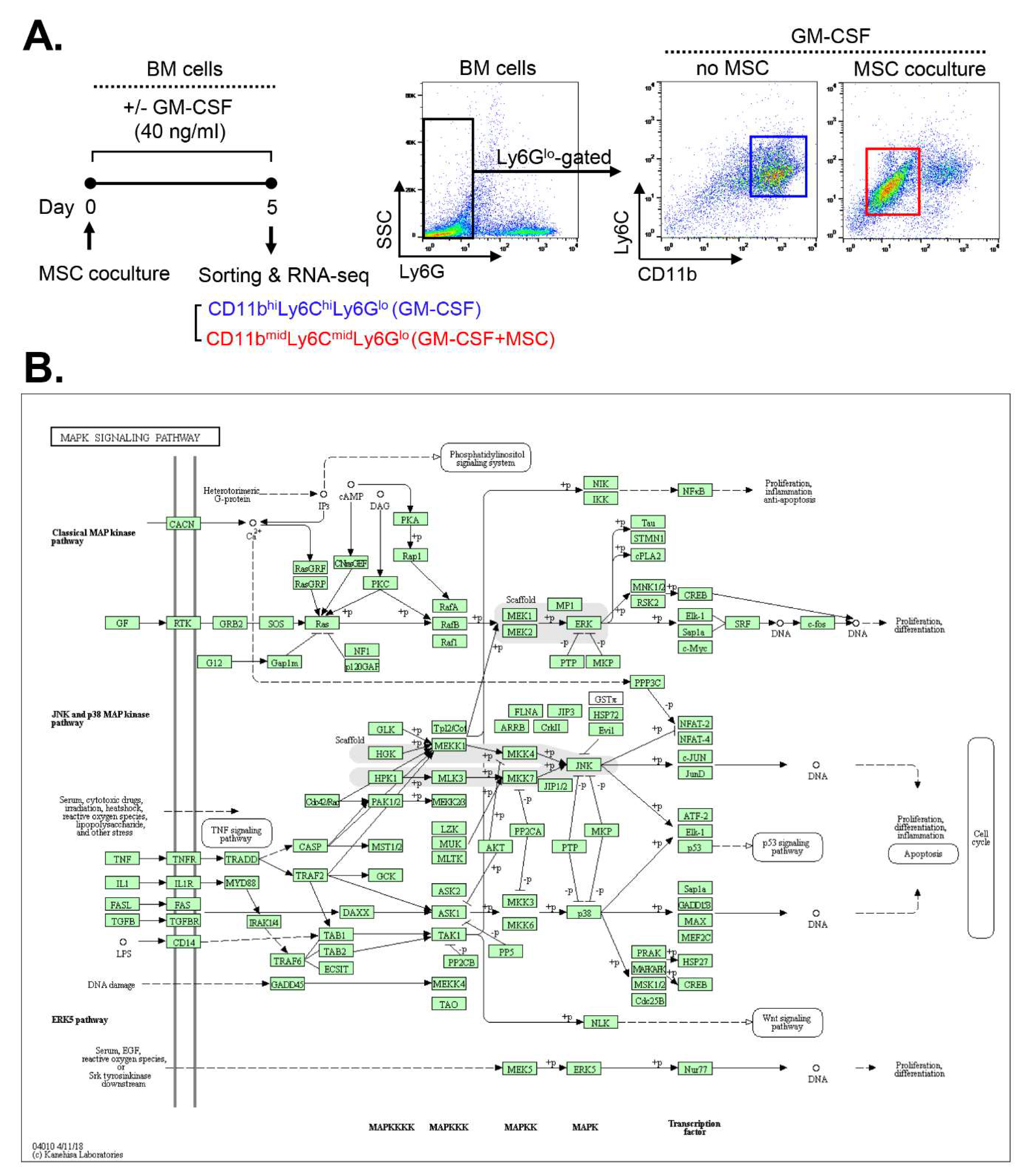
2.2. MSCs activate the JNK pathway in BM cells
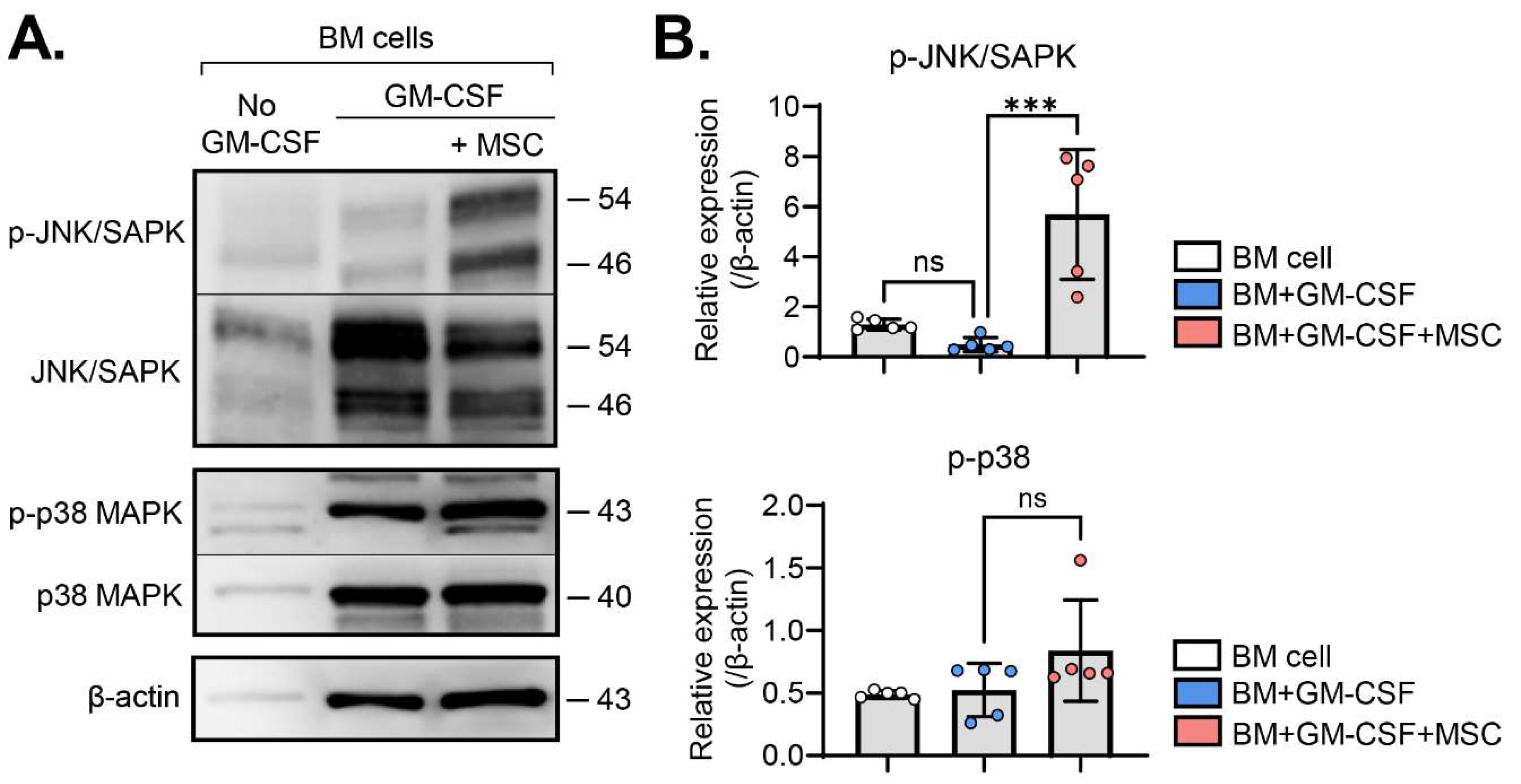
2.3. JNK inhibition abrogates MSC effects on MDSC induction
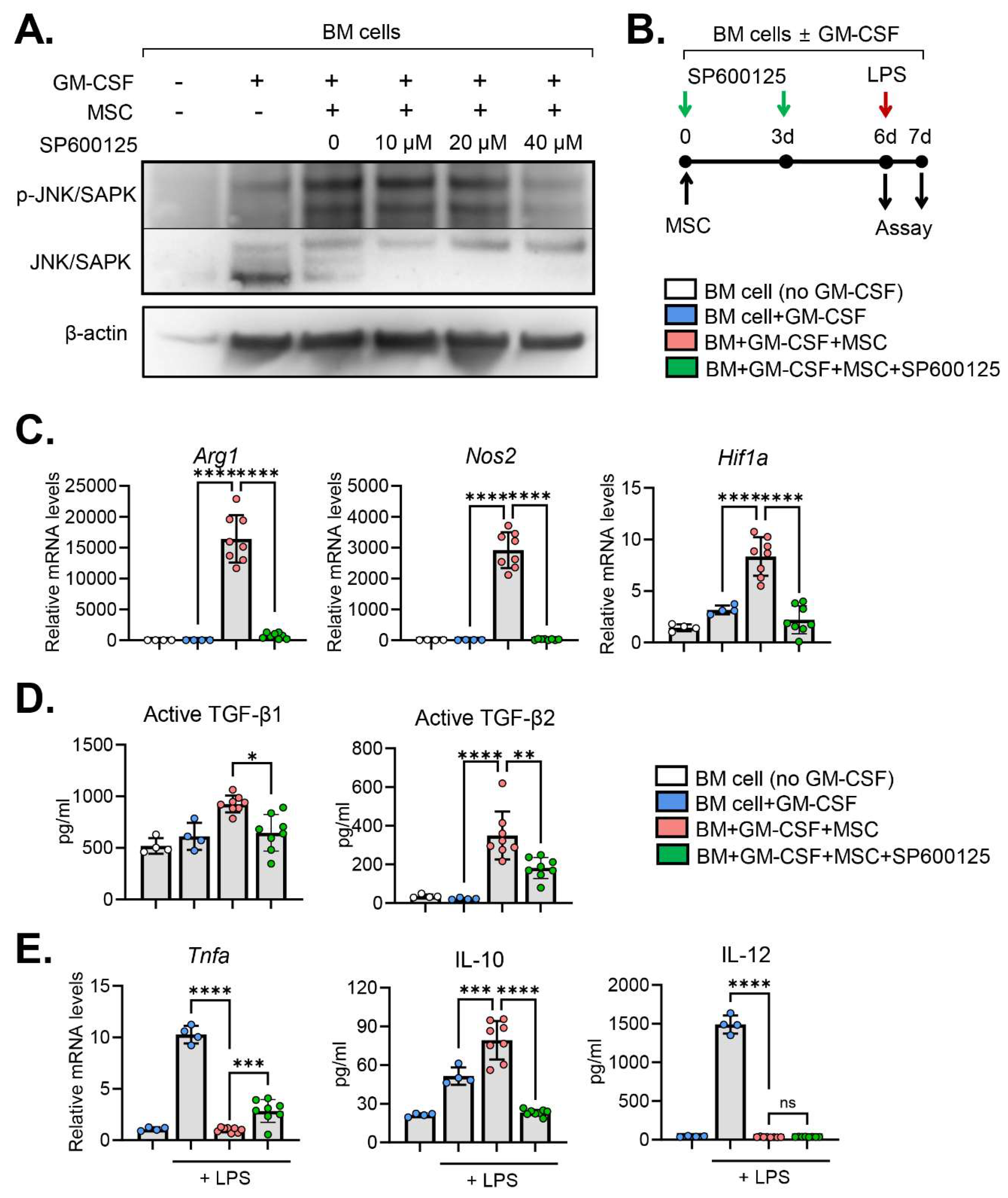
3. Discussion
4. Materials and Methods
4.1. Cell culture
4.2. RNA sequencing
4.3. Western blot assays
4.4. Quantitative real-time reverse-transcription PCR (qRT-PCR)
4.5. Enzyme-linked immunosorbent assay (ELISA)
4.6. Statistical analysis
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Galipeau, J. Macrophages at the nexus of mesenchymal stromal cell potency: The emerging role of chemokine cooperativity. Stem Cells. 2021, 39, 1145–1154. [Google Scholar] [CrossRef] [PubMed]
- Oh JY, Lee RH. Mesenchymal stromal cells for the treatment of ocular autoimmune diseases. Prog Retin Eye Res. 2021, 85, 100967. [CrossRef] [PubMed]
- Veglia, F.; Perego, M.; Gabrilovich, D. Myeloid-derived suppressor cells coming of age. Nat Immunol 2018, 19, 108–119. [Google Scholar] [CrossRef] [PubMed]
- Veglia F, Sanseviero E, Gabrilovich DI. Myeloid-derived suppressor cells in the era of increasing myeloid cell diversity. Nat Rev Immunol 2021, 21, 485–498. [CrossRef] [PubMed]
- Trikha P, Carson WE 3rd. Signaling pathways involved in MDSC regulation. Biochim Biophys Acta. 2014, 1846, 55–65. [CrossRef]
- Condamine T, Mastio J, Gabrilovich DI. Transcriptional regulation of myeloid-derived suppressor cells. J Leukoc Biol 2015, 98, 913–933. [CrossRef] [PubMed]
- Hegde S, Leader AM, Merad M. MDSC: Markers, development, states, and unaddressed complexity. Immunity 2021, 54, 875–884. [CrossRef] [PubMed]
- Lee HJ, Ko JH, Kim HJ, Jeong HJ, Oh JY. Mesenchymal stromal cells induce distinct myeloid-derived suppressor cells in inflammation. JCI Insight 2020, 18, e136059. [CrossRef]
- Zhang W, Liu HT. MAPK signal pathways in the regulation of cell proliferation in mammalian cells. Cell Res. 2002, 12, 9–18. [CrossRef] [PubMed]
- Bennett BL, Sasaki DT, Murray BW, O'Leary EC, Sakata ST, Xu W, Leisten JC, Motiwala A, Pierce S, Satoh Y, Bhagwat SS, Manning AM, Anderson DW. SP600125, an anthrapyrazolone inhibitor of Jun N-terminal kinase. Proc Natl Acad Sci U S A 2001, 98, 13681–13686. [CrossRef]
- Shalova IN, Lim JY, Chittezhath M, Zinkernagel AS, Beasley F, Hernández-Jiménez E, Toledano V, Cubillos-Zapata C, Rapisarda A, Chen J, Duan K, Yang H, Poidinger M, Melillo G, Nizet V, Arnalich F, López-Collazo E, Biswas SK. Human monocytes undergo functional re-programming during sepsis mediated by hypoxia-inducible factor-1α. Immunity 2015, 42, 484–498. [CrossRef]
- Dobreva ZG, Miteva LD, Stanilova SA. The inhibition of JNK and p38 MAPKs downregulates IL-10 and differentially affects c-Jun gene expression in human monocytes. Immunopharmacol Immunotoxicol. 2009, 31, 195–201. [CrossRef] [PubMed]
- Bayik D, Tross D, Haile LA, Verthelyi D, Klinman DM. Regulation of the maturation of human monocytes into immunosuppressive macrophages. Blood Adv. 2017, 1, 2510–2519. [CrossRef] [PubMed]
- TYu J, Li H, Zhang Z, Lin W, Wei X, Shao B. Targeting the MDSCs of Tumors In Situ With Inhibitors of the MAPK Signaling Pathway to Promote Tumor Regression. Front Oncol. 2021, 11, 647312. [CrossRef] [PubMed]
- Deng Y, Yang J, Qian J, Liu R, Huang E, Wang Y, Luo F, Chu Y. TLR1/TLR2 signaling blocks the suppression of monocytic myeloid-derived suppressor cell by promoting its differentiation into M1-type macrophage. Mol Immunol. 2019, 112, 266–273. [CrossRef] [PubMed]
- Liu L, Song H, Duan H, Chai J, Yang J, Li X, Yu Y, Zhang X, Hu X, Xiao M, Feng R, Yin H, Hu Q, Yang L, Du J, Li T. TSG-6 secreted by human umbilical cord-MSCs attenuates severe burn-induced excessive inflammation via inhibiting activations of P38 and JNK signaling. Sci Rep 2016, 6, 30121. [CrossRef]
- Valledor AF, Sánchez-Tilló E, Arpa L, Park JM, Caelles C, Lloberas J, Celada A. Selective roles of MAPKs during the macrophage response to IFN-gamma. J Immunol. 2008, 180, 4523–4529. [CrossRef] [PubMed]
- Galipeau J, Sensébé L. Mesenchymal Stromal Cells: Clinical Challenges and Therapeutic Opportunities. Cell Stem Cell. 2018, 22, 824–833. [CrossRef] [PubMed]
- Lee MJ, Park SY, Ko JH, Lee HJ, Ryu JS, Park JW, Khwarg SI, Yoon SO, Oh JY. Mesenchymal stromal cells promote B-cell lymphoma in lacrimal glands by inducing immunosuppressive microenvironment. Oncotarget. 2017, 8, 66281–66292. [CrossRef] [PubMed]
- Ren G, Zhao X, Wang Y, Zhang X, Chen X, Xu C, Yuan ZR, Roberts AI, Zhang L, Zheng B, Wen T, Han Y, Rabson AB, Tischfield JA, Shao C, Shi Y. CCR2-dependent recruitment of macrophages by tumor-educated mesenchymal stromal cells promotes tumor development and is mimicked by TNFα. Cell Stem Cell. 2012, 11, 812–824. [CrossRef]
| Gene Description | Gene symbol |
Fold Change |
|---|---|---|
| ELK1, member of ETS oncogene family | Elk1 | 13.873 |
| MAP kinase-activated protein kinase 5; similar to MK-5 type 2 | Mapkapk5 | 12.104 |
| RAS protein-specific guanine nucleotide-releasing factor 1 | Rasgrf1 | 11.334 |
| TNF receptor-associated factor 2 | Traf2 | 10.991 |
| activin A receptor, type IC | Acvr1c | 239.476 |
| brain derived neurotrophic factor | Bdnf | 134.205 |
| calcium channel, voltage-dependent, L type, alpha 1C subunit | Cacna1c | 925.475 |
| calcium channel, voltage-dependent, N type, alpha 1B subunit | Cacna1b | 27.199 |
| calcium channel, voltage-dependent, P/Q type, alpha 1A subunit | Cacna1a | 123.802 |
| calcium channel, voltage-dependent, T type, alpha 1G subunit | Cacna1g | 20.065 |
| calcium channel, voltage-dependent, alpha 2/delta subunit 2 | Cacna2d2 | 293.280 |
| calcium channel, voltage-dependent, alpha2/delta subunit 1 | Cacna2d1 | 129.935 |
| calcium channel, voltage-dependent, alpha2/delta subunit 3 | Cacna2d3 | 111.485 |
| calcium channel, voltage-dependent, beta 4 subunit | Cacnb4 | 20.455 |
| calcium channel, voltage-dependent, gamma subunit 2 | Cacng2 | 166.763 |
| dual specificity phosphatase 10 | Dusp10 | 119.544 |
| ecotropic viral integration site 1 | Mecom | 221.142 |
| fibroblast growth factor 12 | Fgf12 | 1521.701 |
| fibroblast growth factor 14 | Fgf14 | 112.772 |
| fibroblast growth factor 18 | Fgf18 | 37.001 |
| fibroblast growth factor 9 | Fgf19 | 99.477 |
| fibroblast growth factor receptor 1 | Fgfr1 | 29.610 |
| filamin, beta | Flnb | 559.332 |
| mitogen-activated protein kinase 13 | Mapk13 | 54.817 |
| mitogen-activated protein kinase 8 interacting protein 3 | Mapk8ip3 | 31.535 |
| mitogen-activated protein kinase kinase kinase 13 | Map3k13 | 131.863 |
| mitogen-activated protein kinase kinase kinase 4 | Map3k4 | 21.317 |
| mitogen-activated protein kinase kinase kinase kinase 3 | Map4k3 | 12.556 |
| neurofibromatosis 1 | Nf1 | 179.681 |
| nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 | Nfatc4 | 20.130 |
| phospholipase A2, group IID | Pla2g2d | 131.904 |
| phospholipase A2, group III | Pla2g3 | 33.243 |
| protein phosphatase 3, catalytic subunit, gamma isoform | Ppp3cc | 79.519 |
| related RAS viral (r-ras) oncogene homolog 2 | Rras2 | 21.369 |
| serine/threonine kinase 3 (Ste20, yeast homolog) | Stk3 | 98.308 |
| thymoma viral proto-oncogene 3 | Akt3 | 112.279 |
| transforming growth factor, beta 2 | Tgfb2 | 51.476 |
| transforming growth factor, beta 3 | Tgfb3 | 38.421 |
| v-crk sarcoma virus CT10 oncogene homolog (avian) | Crk | 8.968 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




