Submitted:
07 January 2024
Posted:
08 January 2024
Read the latest preprint version here
Abstract
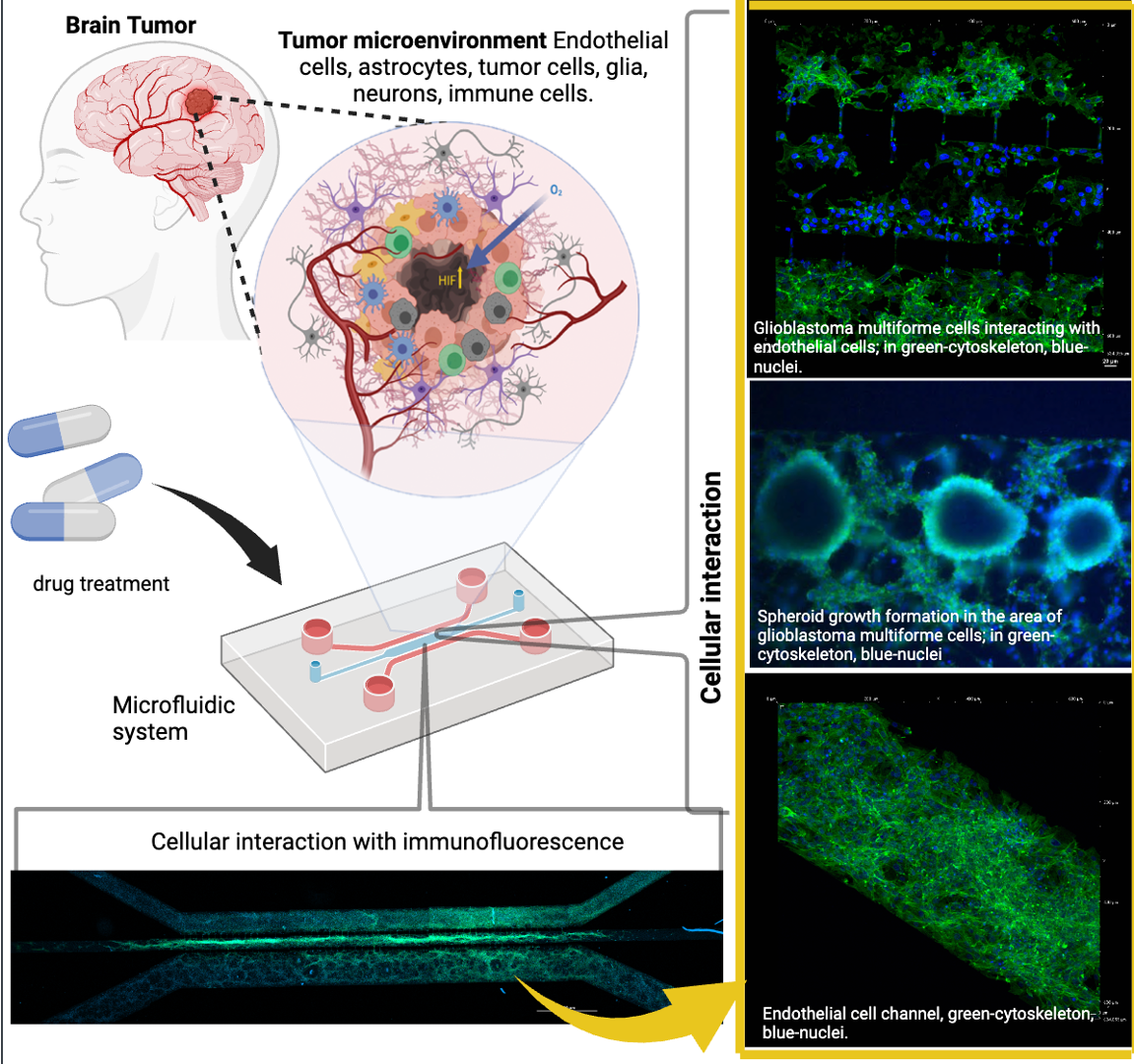
Keywords:
1. Introduction
2. Materials and Methods
2.1. Design and Fabrication of Microfluidic System
2.2. Characterization of the Microfluidic System
2.3. Immunophenotype Cells
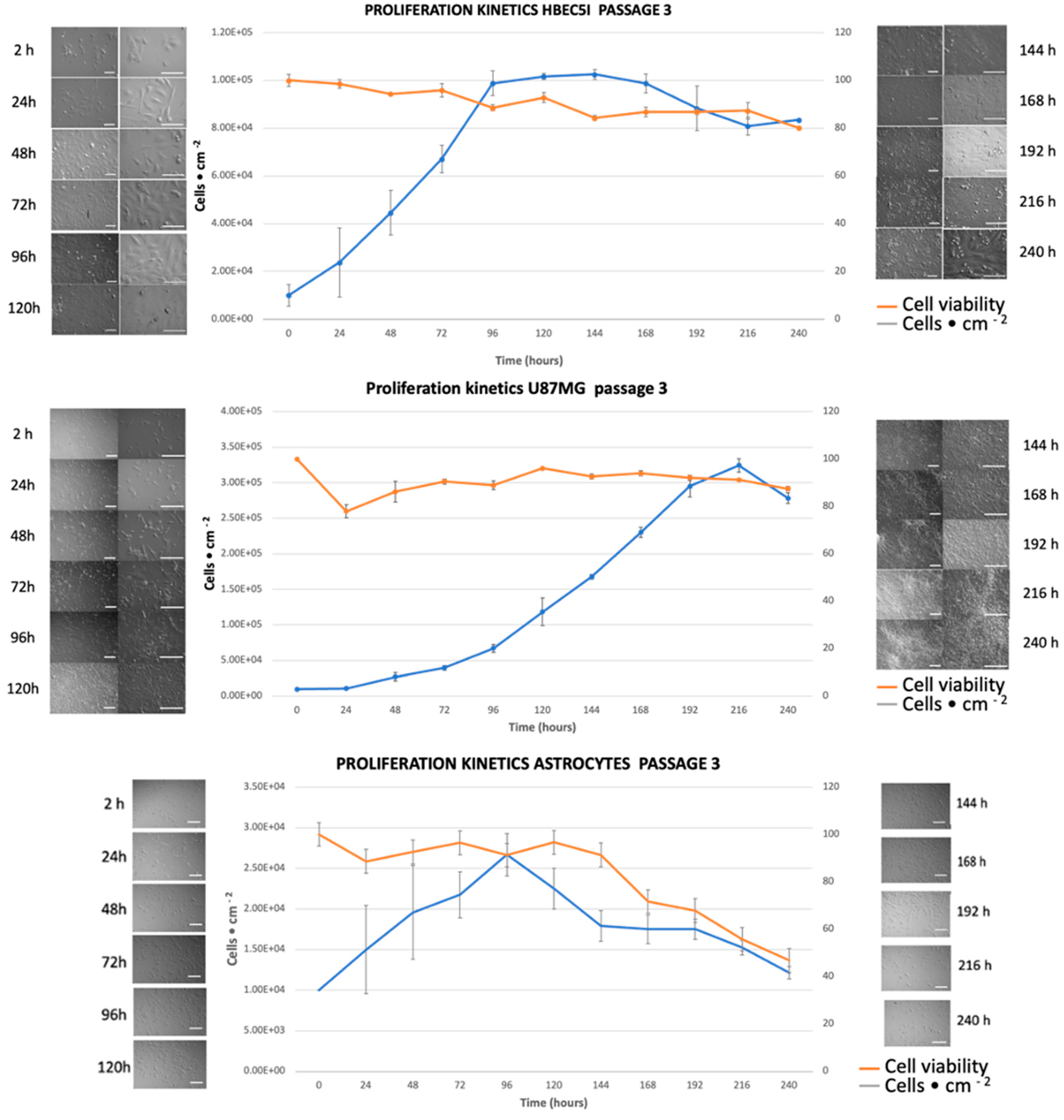
2.4. Cell Proliferation Kinetics
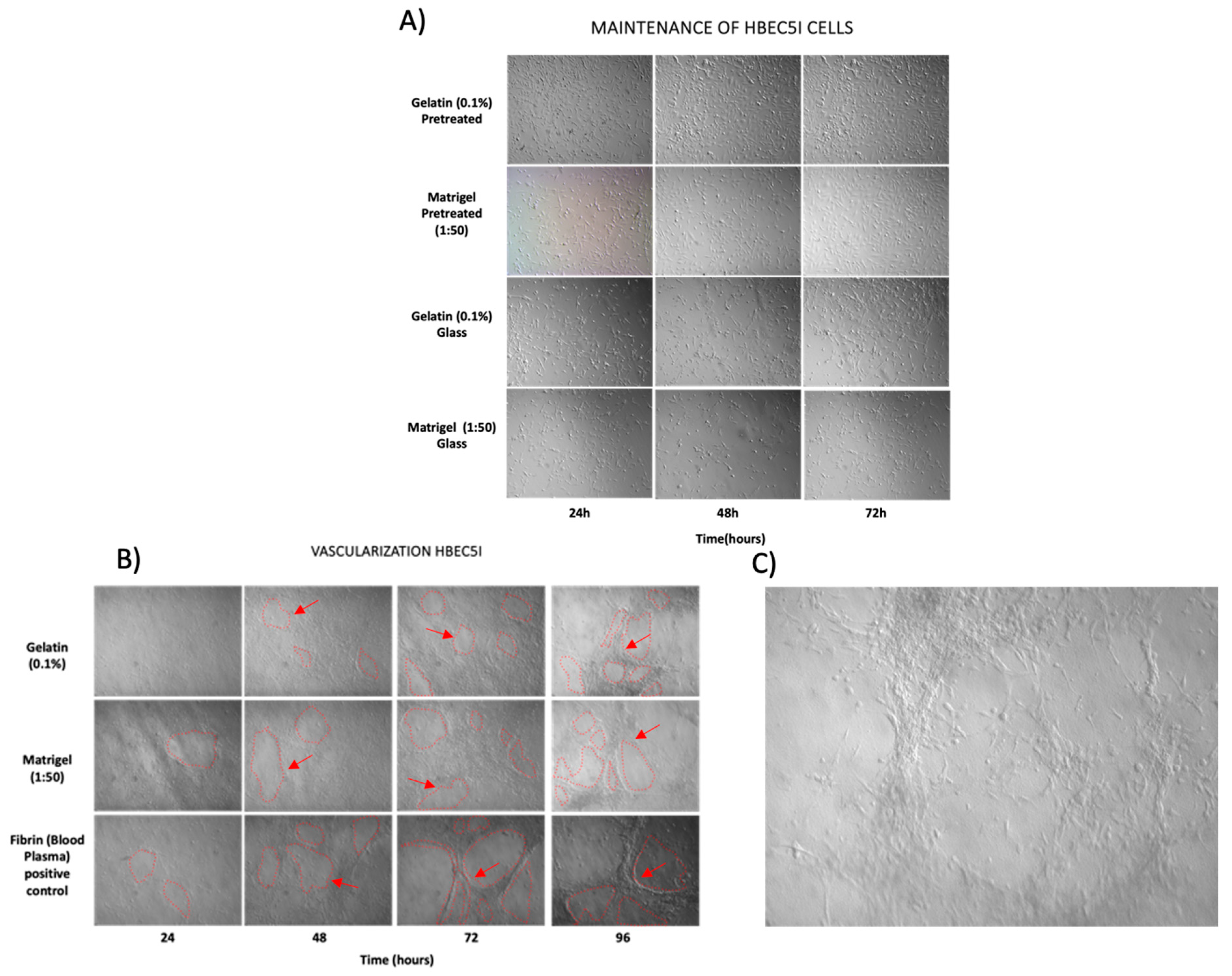
2.5. Protein Matrix Test for HBEC5i Endothelial Cells
2.6. Endothelial Vascularization Test
2.7. Epifluorescence Microscopy
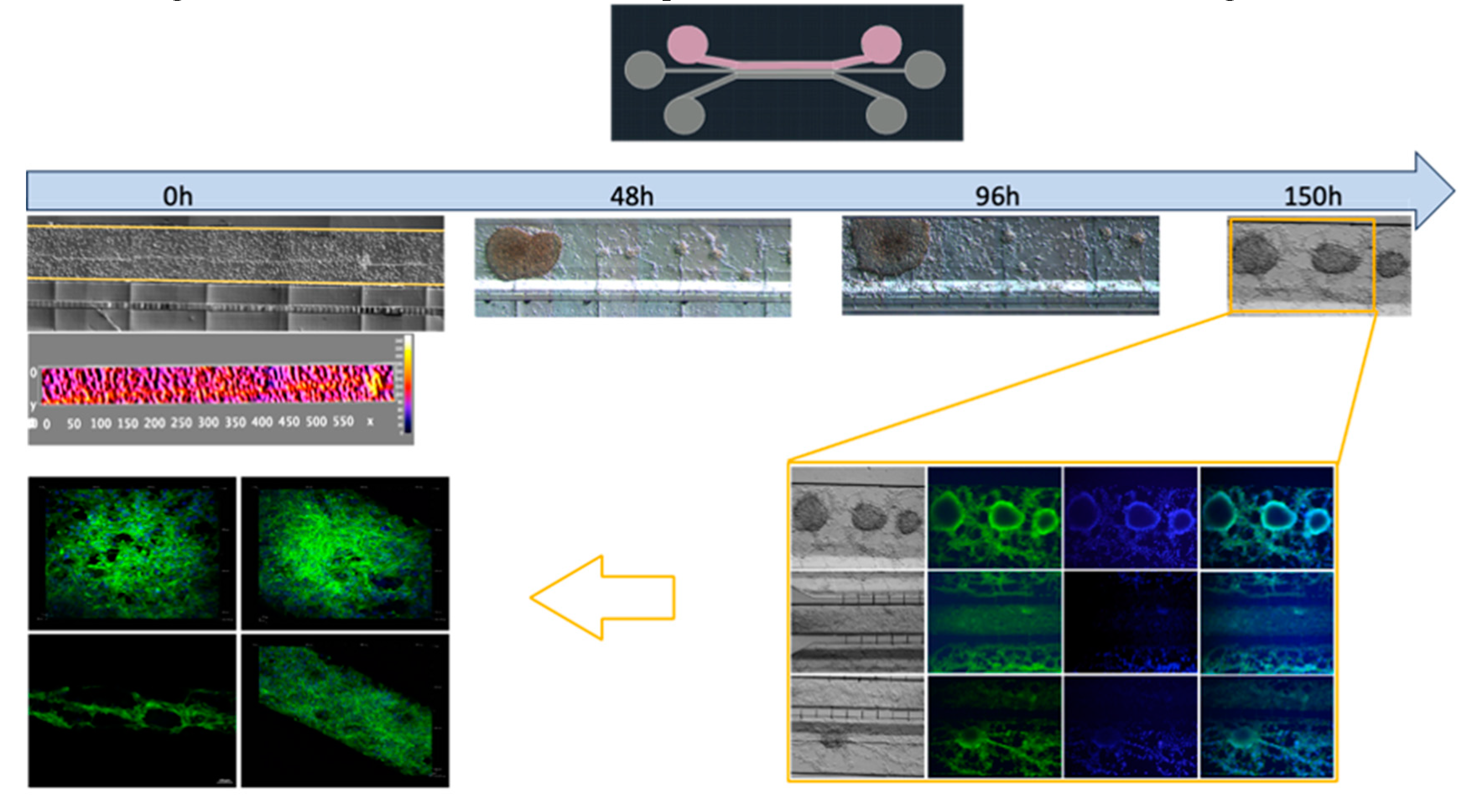
2.8. Cell Line Cultures in the Microfluidic System
2.9. Microenvironment of Hematoencefalic Barrier
2.10. Confocal Microscopy
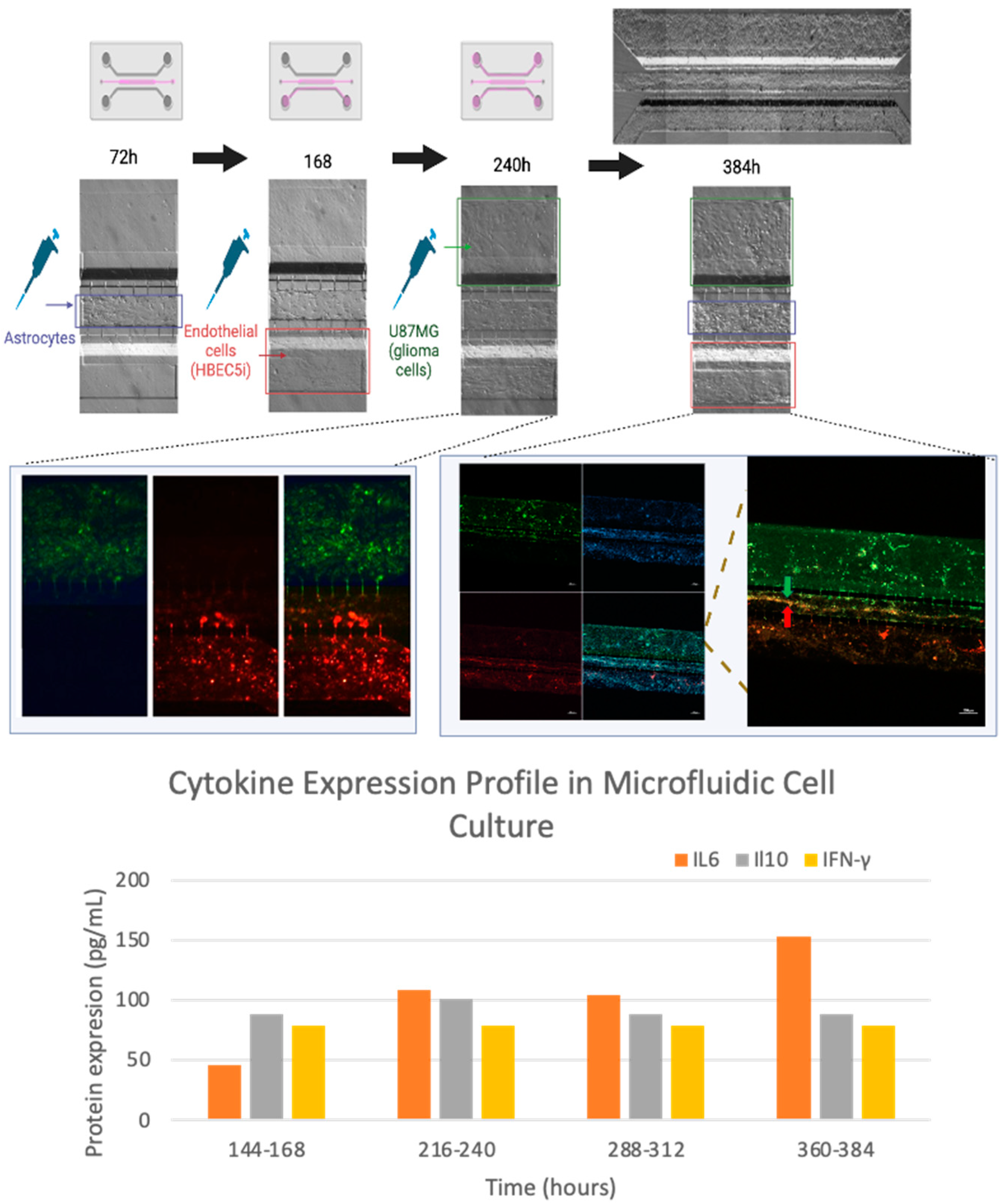
2.11. Cytokine Expression Assay
3. Results
3.1. The Microfluidic System: Device Design
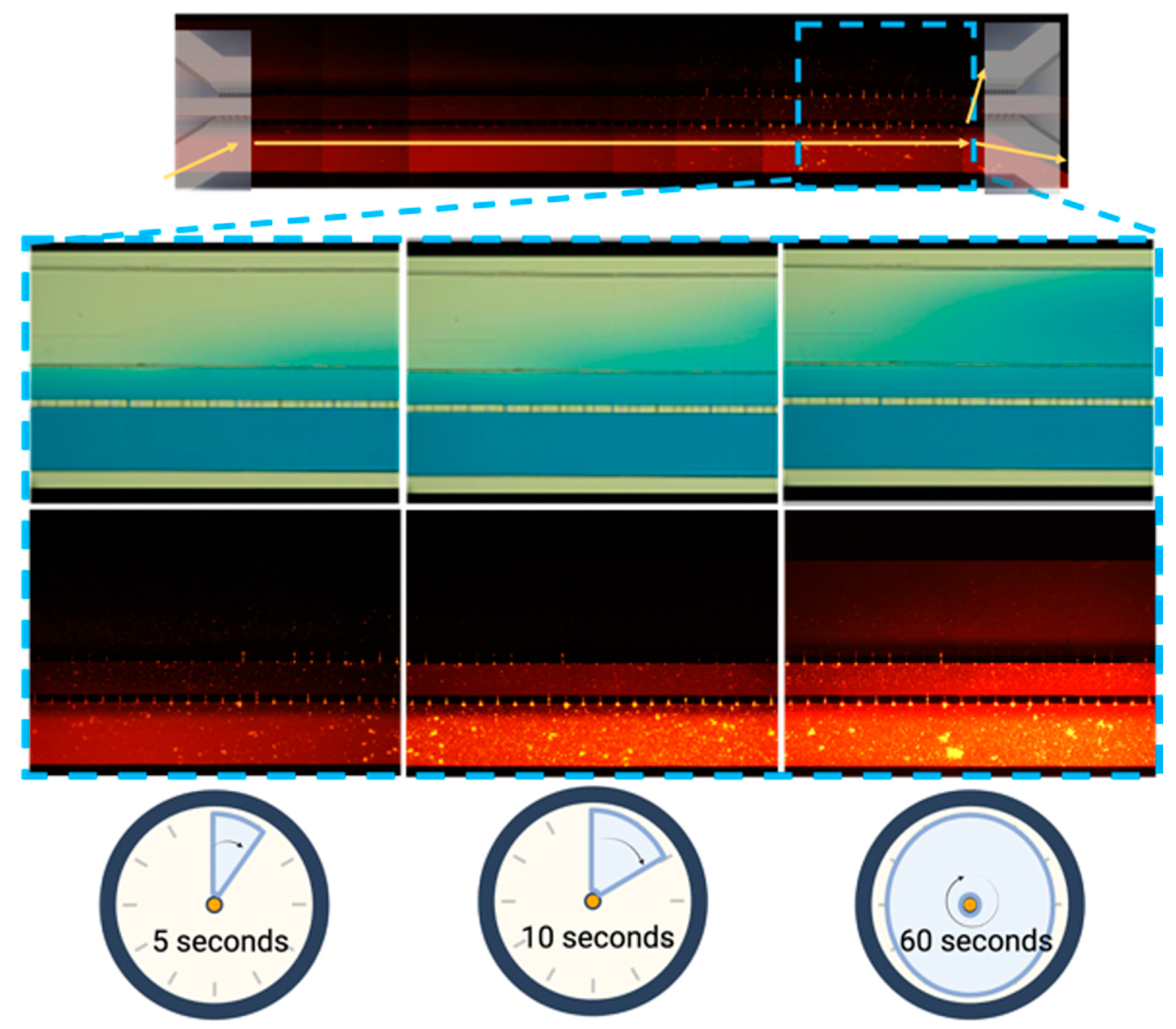
3.2. The Microfluidic System Permeability Test
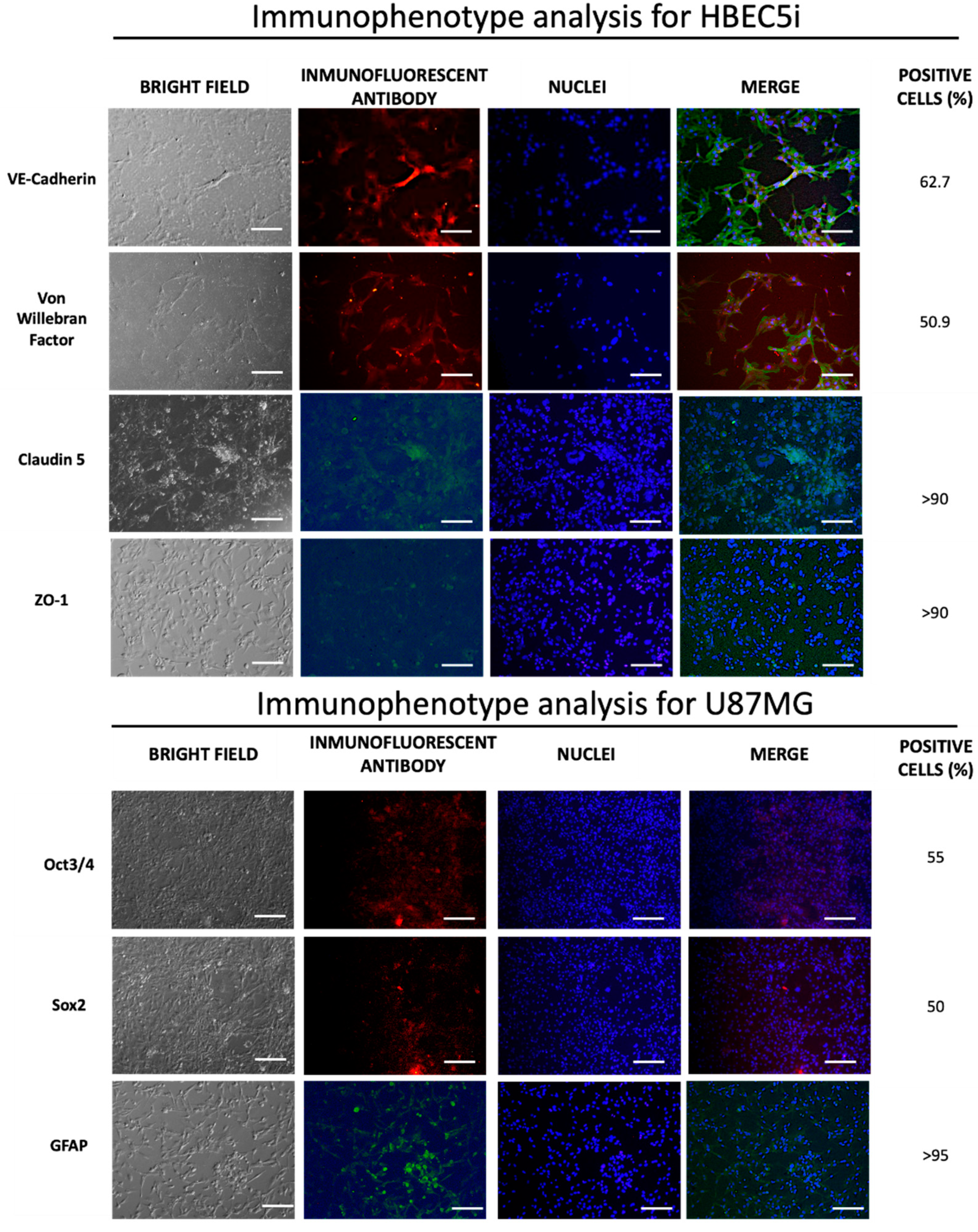
3.3. Characterization of Cellular Population
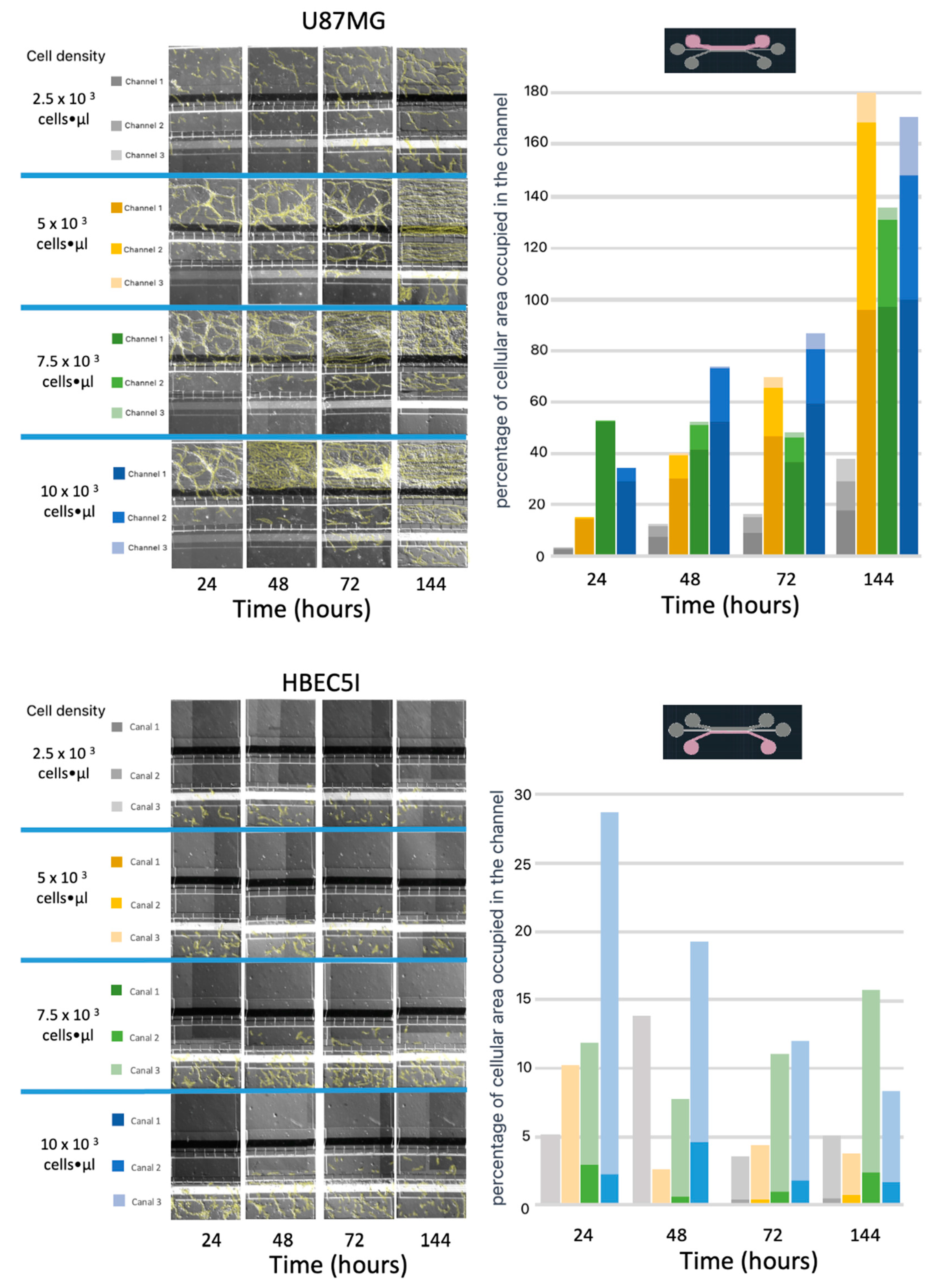
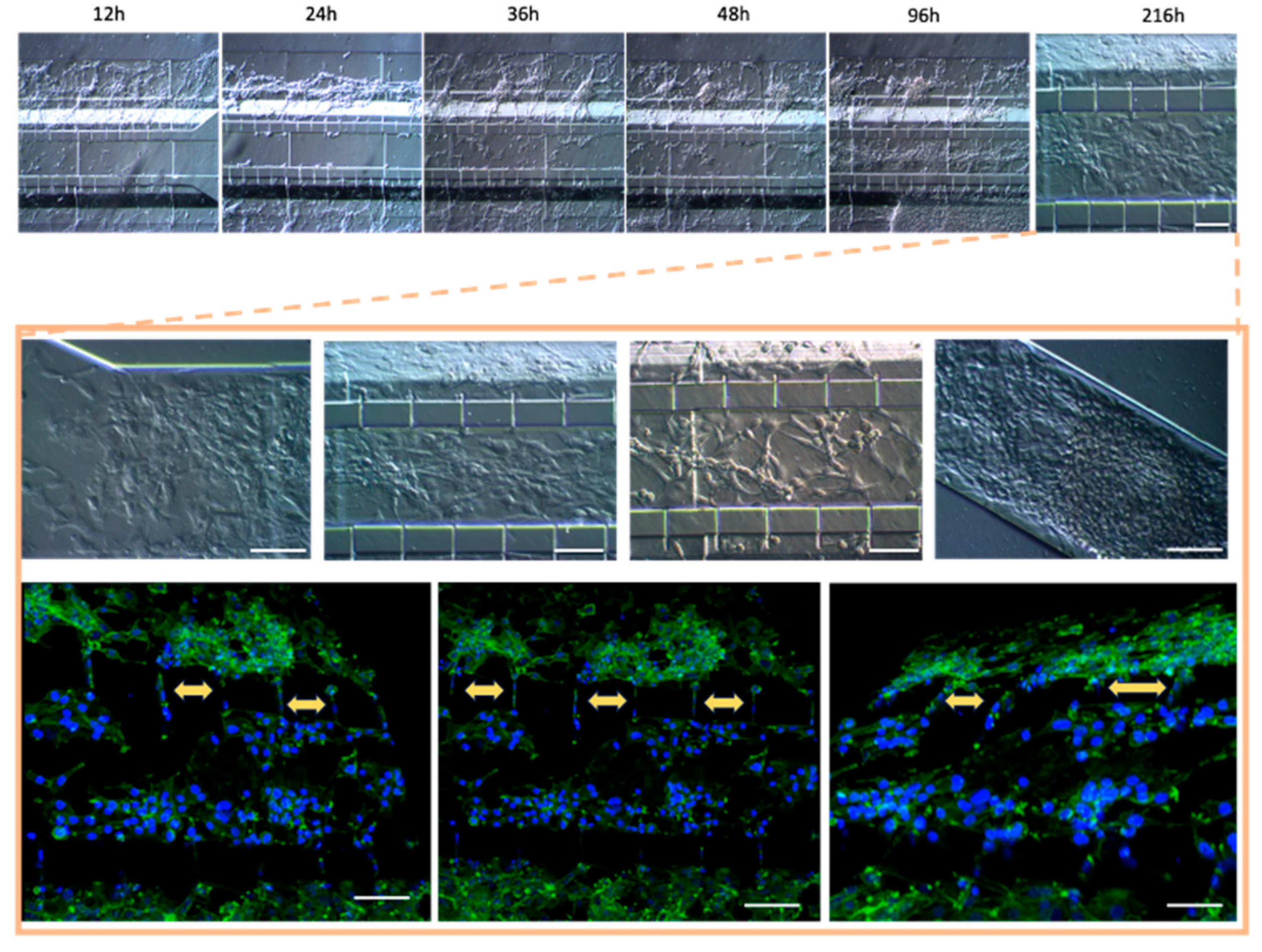
3.4. Optimization of Inoculation Cell Density in the Microfluidic System
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Furnari, F.B.; Fenton, T.; Bachoo, R.M.; Mukasa, A.; Stommel, J.M.; Stegh, A.; Hahn, W.C.; Ligon, K.L.; Louis, D.N.; Brennan, C. Malignant astrocytic glioma: genetics, biology, and paths to treatment. Genes Dev 2017, 21, 2683–710. [Google Scholar] [CrossRef] [PubMed]
- Junttila, M.R.; De Sauvage, F.J. Influence of tumour micro-environment heterogeneity on therapeutic response. Nature 2013, 501, 346–354. [Google Scholar] [CrossRef] [PubMed]
- Becker, J.C.; Andersen, M.H.; Schrama, D.; thor Straten, P. Immune-suppressive properties of the tumor microenvironment. Cancer immunology, immunotherapy 2013, 62, 1137–1148. [Google Scholar] [CrossRef] [PubMed]
- Charles, N.A.; Holland, E.C.; Gilbertson, R.; Glass, R.; Kettenmann, H. The brain tumor microenvironment. Glia 2012, 60, 502–514. [Google Scholar] [CrossRef]
- Meacham, C.E.; Morrison, S.J. Tumour heterogeneity and cancer cell plasticity. Nature 2013, 501, 328–337. [Google Scholar] [CrossRef] [PubMed]
- Hacking, R.; Hunt, K. Cerebral abscess: A review of its pathophysiology, diagnosis and management. British Journal of Neuroscience Nursing 2007, 3, 400–403. [Google Scholar] [CrossRef]
- Lecuyer, M.A.; Kebir, H.; Prat, A. Glial influences on BBB functions and molecular players in immune cell trafficking. Biochimica et Biophysica Acta (BBA)-Molecular Basis of Disease 2016, 1862, 472–482. [Google Scholar] [CrossRef]
- Vatine, G.D.; Barrile, R.; Workman, M.J.; Sances, S.; Barriga, B.K.; Rahnama, M.; Svendsen, C.N. Human iPSC-derived blood-brain barrier chips enable disease modeling and personalized medicine applications. Cell stem cell 2019, 24, 995–1005. [Google Scholar] [CrossRef]
- Wang, X.; Hou, Y.; Ai, X.; Sun, J.; Xu, B.; Meng, X.; Zhang, S. Potential applications of microfluidics-based blood-brain barrier (BBB)-on-chips for in vitro drug development. Biomedicine & Pharmacotherapy 2020, 132, 110822. [Google Scholar]
- Gronseth, E.; Wang, L.; Harder, D.R.; Ramchandran, R. The Role of Astrocytes in Tumor Growth and Progression. Astrocyte-Physiology and Pathology 2018. [Google Scholar]
- Lee, G.; Hall, I.I.I.; RR; Ahmed, A.U. Cancer stem cells: cellular plasticity, niche, and its clinical relevance. Journal of stem cell research & therapy 2016, 6. Ahmed, A.U. Cancer stem cells: cellular plasticity, niche.
- Abou-Antoun, T.J.; Hale, J.S.; Lathia, J.D.; Dombrowski, S.M. Brain cancer stem cells in adults and children: cell biology and therapeutic implications. Neurotherapeutics 2017, 14, 372–384. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.H.V.; Middleton, K.; You, L.; Sun, Y. A review of microfluidic approaches for investigating cancer extravasation during metastasis. Microsystems & Nanoengineering 2018, 4, 17104. [Google Scholar]
- Ruiz-Garcia, H.; Alvarado-Estrada, K.; Schiapparelli, P.; Quinones-Hinojosa, A.; Trifiletti, D.M. Engineering three- dimensional tumor models to study glioma cancer stem cells and tumor microenvironment. Frontiers in Cellular Neuroscience 2020, 14. [Google Scholar] [CrossRef] [PubMed]
- Yissachar, N.; Zhou, Y.; Ung, L.; Lai, N.Y.; Mohan, J.F.; Ehrlicher, A.; Benoist, C. An intestinal organ culture system uncovers a role for the nervous system in microbe-immune crosstalk. Cell 2017, 168, 1135–1148. [Google Scholar] [CrossRef]
- Basma, E.; KS, B.R. A dynamic perfusion based blood-brain barrier model for cytotoxicity testing and drug permeation. Scientific Reports (Nature Publisher Group) 2020, 10. [Google Scholar]
- van Duinen, V.; Trietsch, S.J.; Joore, J.; Vulto, P.; Hankemeier, T. Microfluidic 3D cell culture: from tools to tissue models. Current opinion in biotechnology 2015, 35, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Plummer, S.; Wallace, S.; Ball, G.; Lloyd, R.; Schiapparelli, P.; Quiñones-Hinojosa, A.; Pamies, D. A Human iPSC-derived 3D platform using primary brain cancer cells to study drug development and personalized medicine. Scientific reports 2019, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Tsai, H.F.; Trubelja, A.; Shen, A.Q.; Bao, G. Tumour-on-a-chip: microfluidic models of tumour morphology, growth and microenvironment. Journal of the Royal Society Interface 2017, 14, 20170137. [Google Scholar] [CrossRef]
- Meng, F.; Meyer, C.M.; Joung, D.; Vallera, D.A.; McAlpine, M.C.; Panoskaltsis-Mortari, A. 3D Bioprinted In Vitro Metastatic Models via Reconstruction of Tumor Microenvironments. Advanced Materials 2019, 1806899. [Google Scholar] [CrossRef]
- Zhang, H.; Zhu, Y.; Shen, Y. Microfluidics for cancer nanomedicine: from fabrication to evaluation. Small 2018, 14, 1800360. [Google Scholar] [CrossRef]
- Logun, M.; Zhao, W.; Mao, L.; Karumbaiah, L. Microfluidics in malignant glioma research and precision medicine. Advanced biosystems 2018, 2, 170022. [Google Scholar] [CrossRef]
- Deosarkar, S.P.; Prabhakarpandian, B.; Wang, B.; Sheffield, J.B.; Krynska, B.; Kiani, M.F. A novel dynamic neonatal blood-brain barrier on a chip. PloS one 2015, 10, e0142725. [Google Scholar] [CrossRef]
- Sontheimer-Phelps, A.; Hassell, B.A.; Ingber, D.E. Modelling cancer in microfluidic human organs-on-chips. Nature Reviews Cancer. 2019. [CrossRef]
- Kamei, K.I.; Ohashi, M.; Gschweng, E.; Ho, Q.; Suh, J.; Tang, J.; Tseng, H.R. Microfluidic image cytometry for quantitative single-cell profiling of human pluripotent stem cells in chemically defined conditions. Lab on a Chip 2010, 10, 1113–1119. [Google Scholar] [CrossRef]
- Lee, H.; Kim, N.; Rheem, H.B.; Kim, B.J.; Park, J.H.; Choi, I.S. A Decade of Advances in Single-Cell Nanocoating for Mammalian Cells. Advanced Healthcare Materials 2021, 10, 2100347. [Google Scholar] [CrossRef]
- Arvanitis, C.D.; Ferraro, G.B.; Jain, R.K. The blood–brain barrier and blood–tumour barrier in brain tumours and metastases. Nature Reviews Cancer 2020, 20, 26–41. [Google Scholar] [CrossRef]
- Tovar-Lopez, F.; Thurgood, P.; Gilliam, C.; Nguyen, N.; Pirogova, E.; Khoshmanesh, K.; Baratchi, S. A microfluidic system for studying the effects of disturbed flow on endothelial cells. Frontiers in bioengineering and biotechnology 2019, 7, 81. [Google Scholar] [CrossRef] [PubMed]
- Adriani, G.; Ma, D.; Pavesi, A.; Kamm, R.D.; Goh, E.L. A 3D neurovascular microfluidic model consisting of neurons, astrocytes and cerebral endothelial cells as a blood–brain barrier. Lab on a Chip 2017, 17, 448–459. [Google Scholar] [CrossRef] [PubMed]
- May, S.; Evans, S.; Parry, L. Organoids, organs-on-chips and other systems, and microbiota. Emerging Topics in Life Sciences 2017, 1. [Google Scholar]
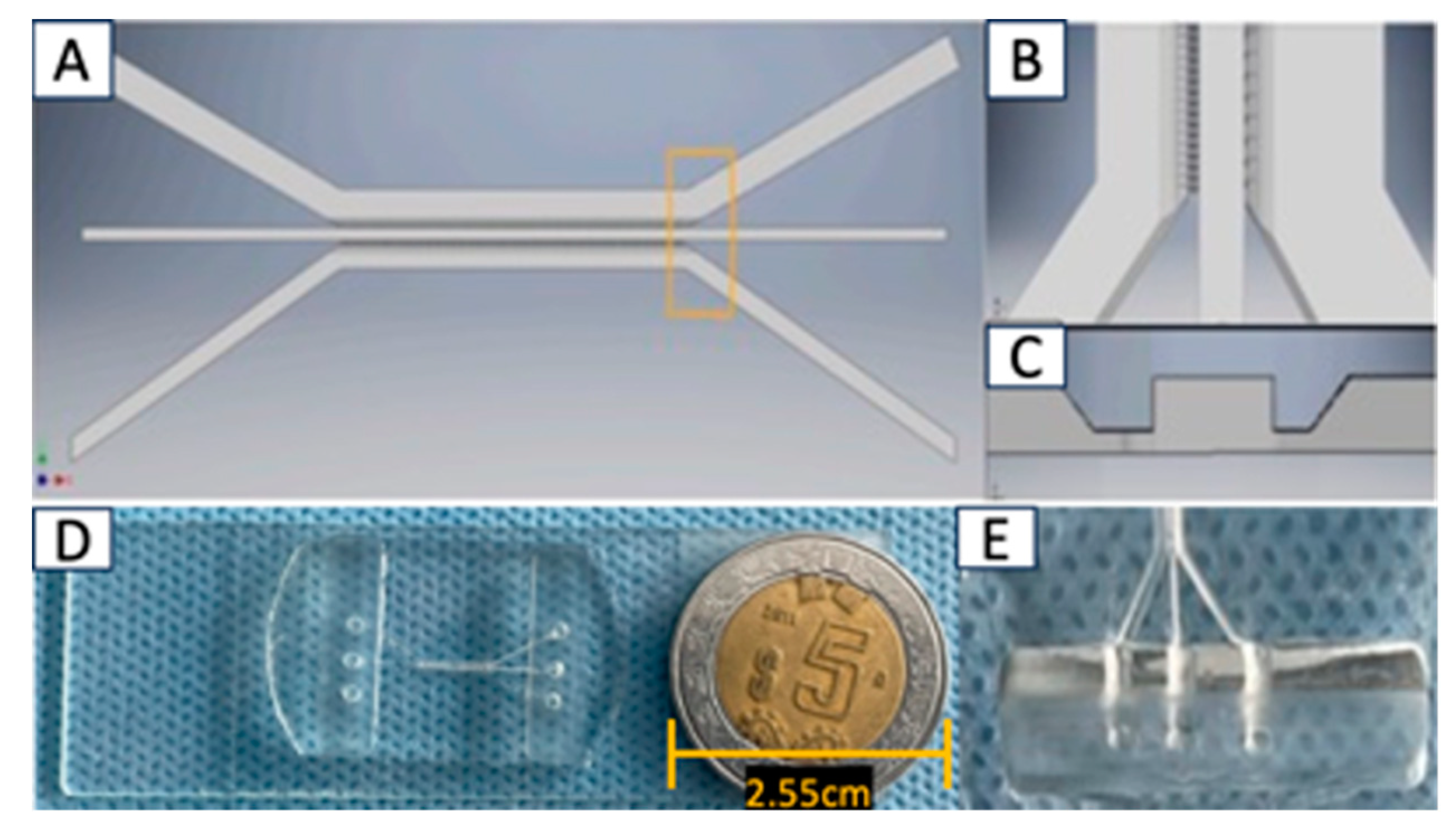

|
Antibody |
Primary antibody | Secondary antibody |
Target description |
| Anti Ve-cadherin | 1:200 (Ab33168) | 1:1000 (Ab6718) | The Ve-cadherin is an adhesion protein present in endothelial cells that form the inner lining of blood vessels |
| Anti Von Willebrand | 1:200 (Ab6994) | 1:1000 (Ab6718) | The Von Willebrand protein is mainly found in endothelial cells that line the blood vessels. |
| Anti VEGFR | 1:200 (ab9530) | 1:1000 (ab6786) | The anti-VEGFR antibody is a specific antibody that targets the vascular endothelial growth factor receptor (VEGFR), a protein that plays a crucial role in angiogenesis and blood vessel growth. |
| Anti Oct3/4 | 1:200(C10-Z210Ms) | 1:1000 (ab6786) | The expression of Oct-3/4 in glioblastoma cells, such as U87MG, is an indication of these tumor cells' ability to acquire more undifferentiated and aggressive characteristics. |
| Anti Sox2 | 1:200 (ZM57-Z236MS) | 1:1000 (ab6786) | Sox-2 expression in glioblastoma cells, including U87MG, signals the existence of stem cells capable of self-renewal and generating differentiated tumor cell subpopulations. This phenomenon contributes to therapy resistance and tumor recurrence |
| Anti GFAP | 1:200 (G3893) | 1:1000 (ab6785) | GFAP (glial fibrillary acidic protein) is a valuable tool for identifying and locating astrocytes in cell cultures, brain tissues, and histological sections. |
| Anti Falloidin-Alexa Fluor 488 | 1:100 (A12379) | --------------------- | Cells and tissues. By using the anti-phalloidin antibody labeled with a fluorescent substance or an enzyme, it is possible to visualize the distribution, organization, and dynamics of actin filaments in living cells or fixed samples |
| Claudin5 | 1:50 (Ab15106) | 1:1000 (ab6785) | Claudin 5 is a protein expressed in the tight junctions of endothelial cells, playing a crucial role in the formation and maintenance of the blood-brain barrier. This membrane protein has a vital function in contributing to the regulation of capillary permeability in the brain. |
| ZO1 | 1:50 (Ab221547) | 1:1000 (Ab6717) | Zonula occludens-1, commonly referred to as ZO-1, is a tight junction protein that plays a key role in the formation and maintenance of cellular tight junctions. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




