Submitted:
25 December 2023
Posted:
26 December 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
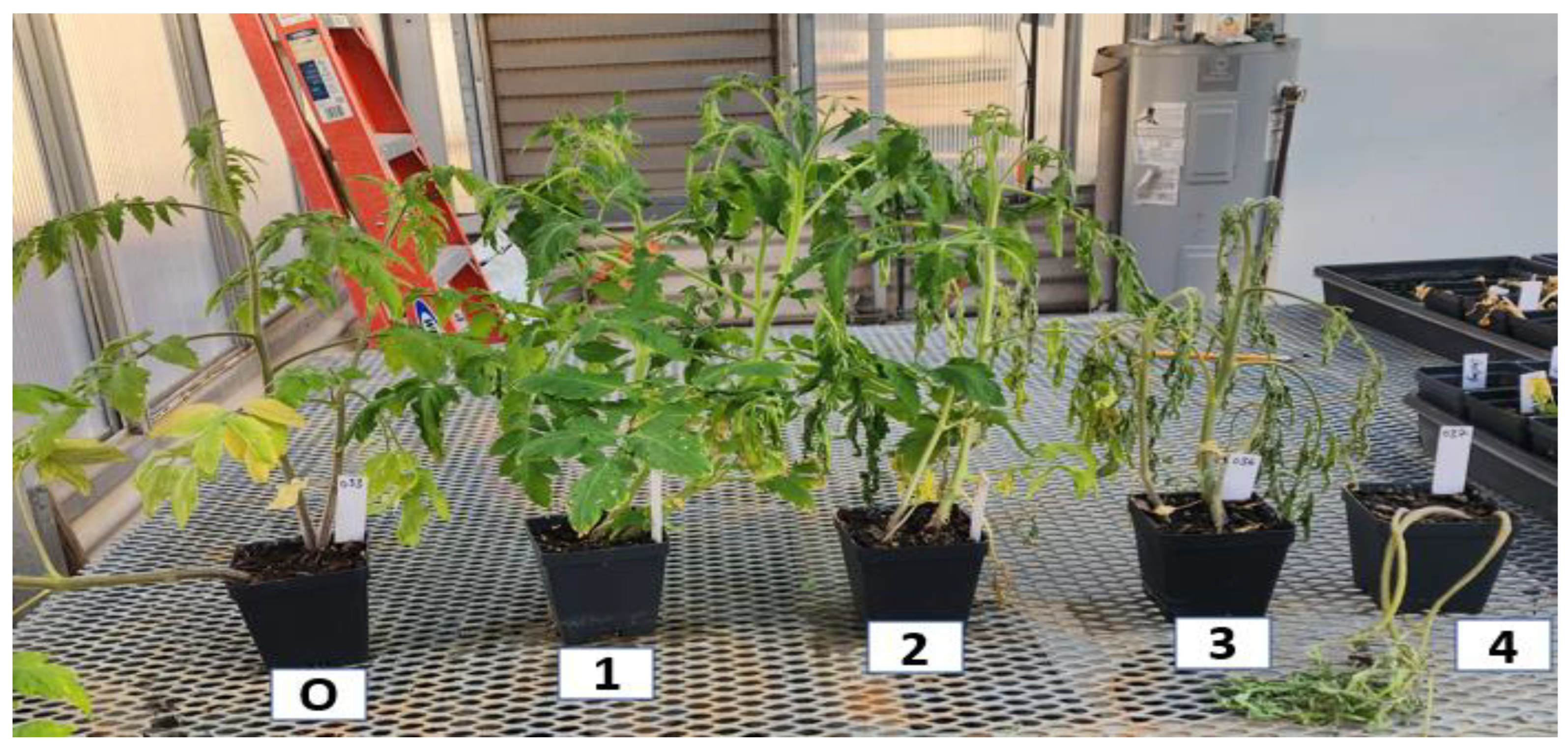
2.2. Plant materials and phenotyping
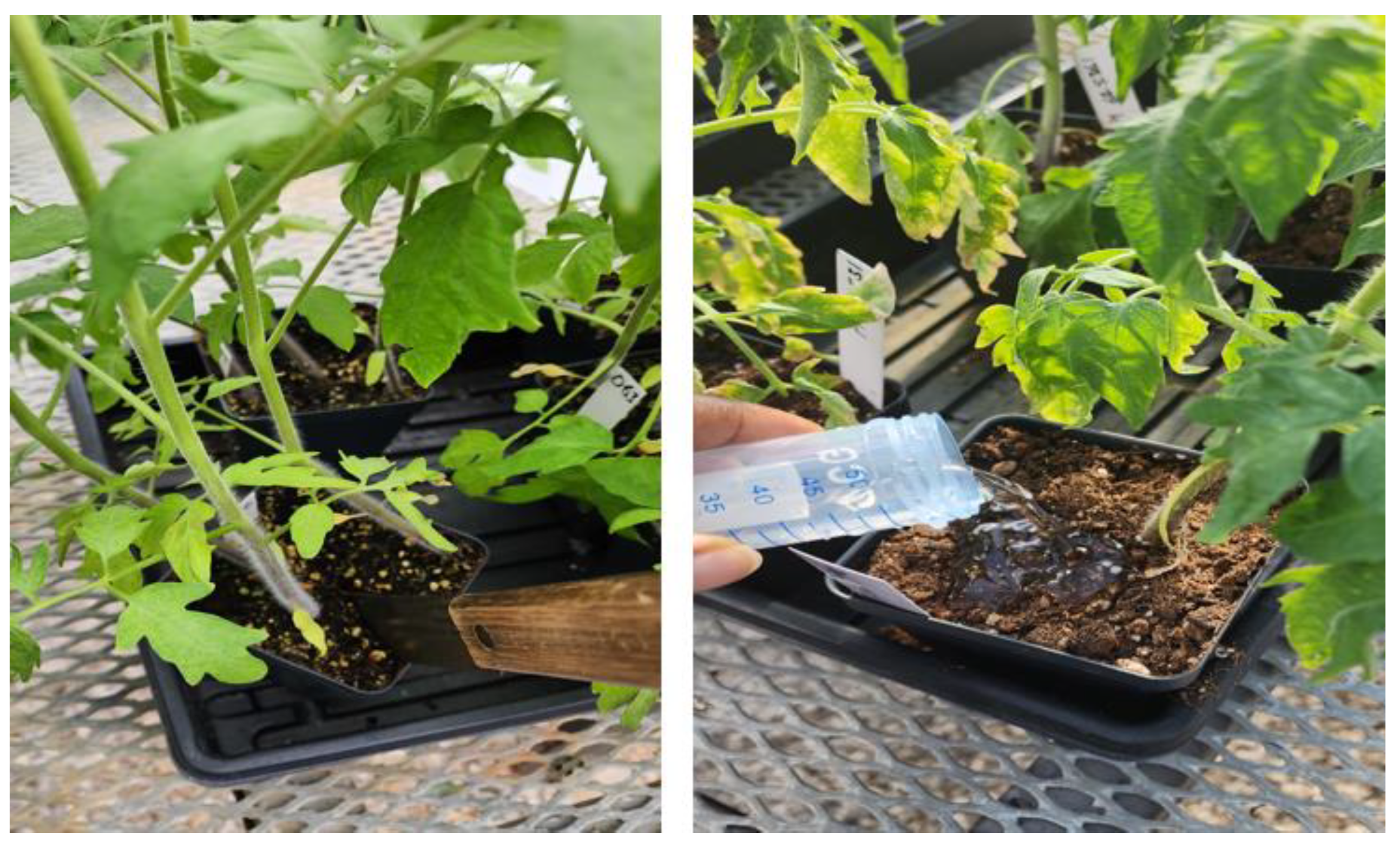
2.3. Greenhouse experiment for pathogen tests
3. Data analysis
3.1. Statistical model
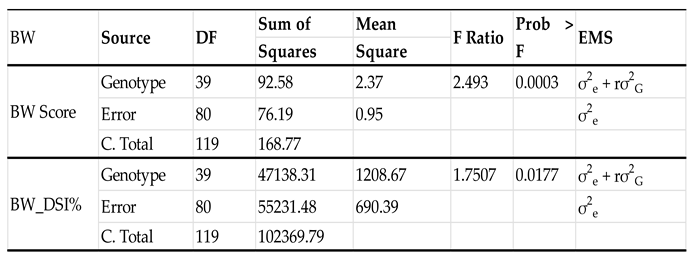
3.2. ANOVA, distribution, descriptive statistics, Pearson’s correlation, and broad-sense heritability
3.3. DNA extraction: genotyping by sequencing (GBS) and SNP discovery.
3.4. Principal Component Analysis (PCA) and Genetic Diversity
4. Results
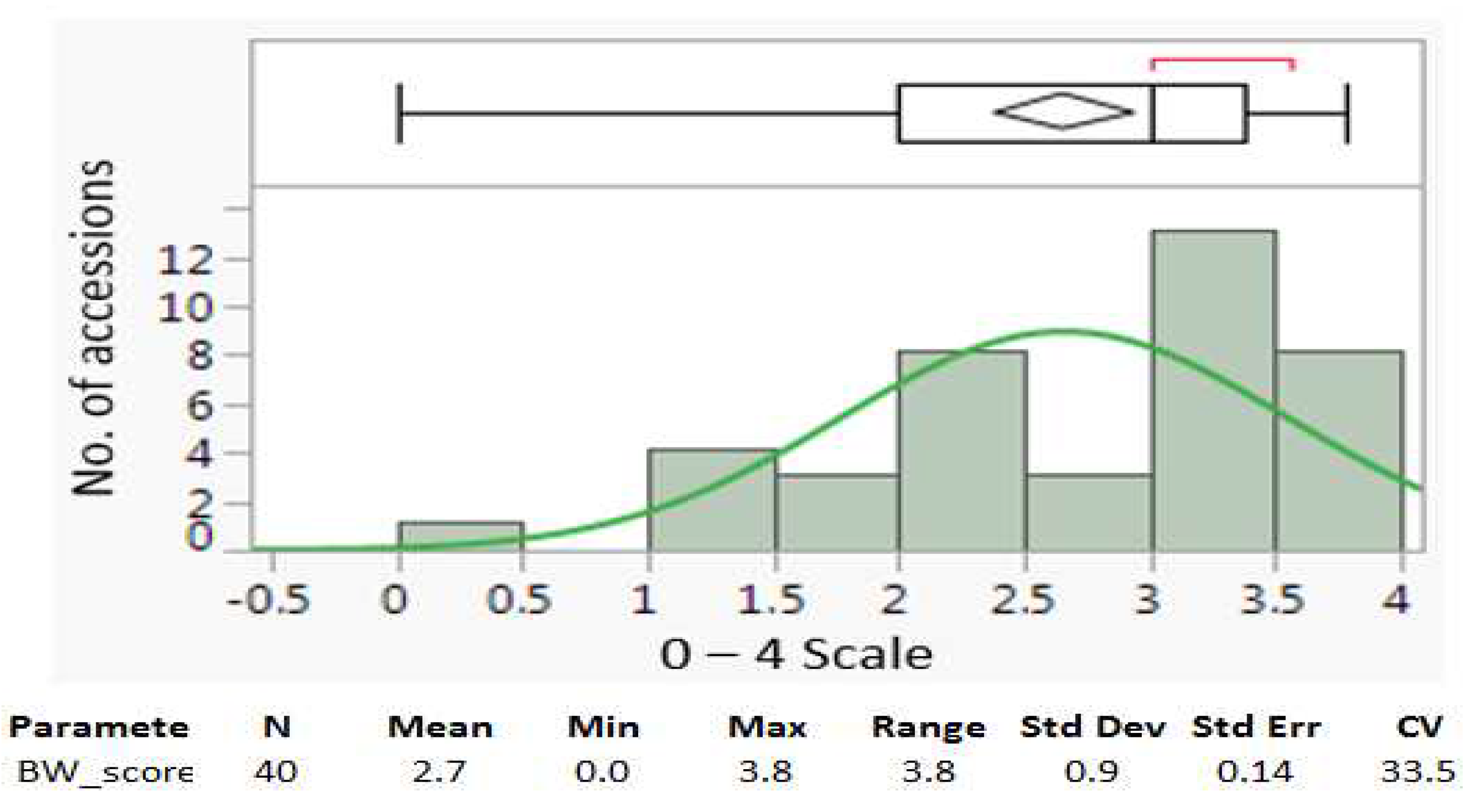
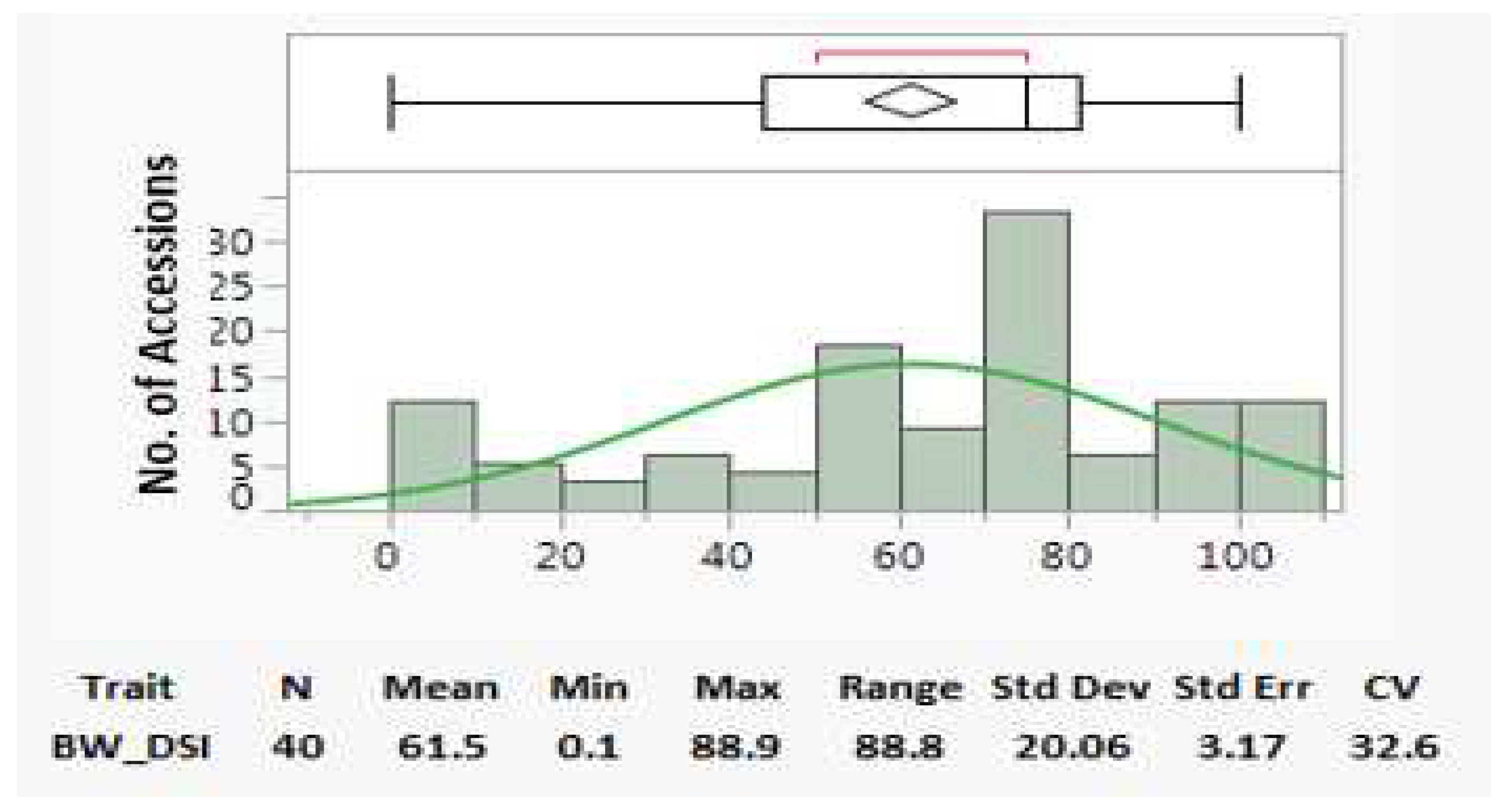
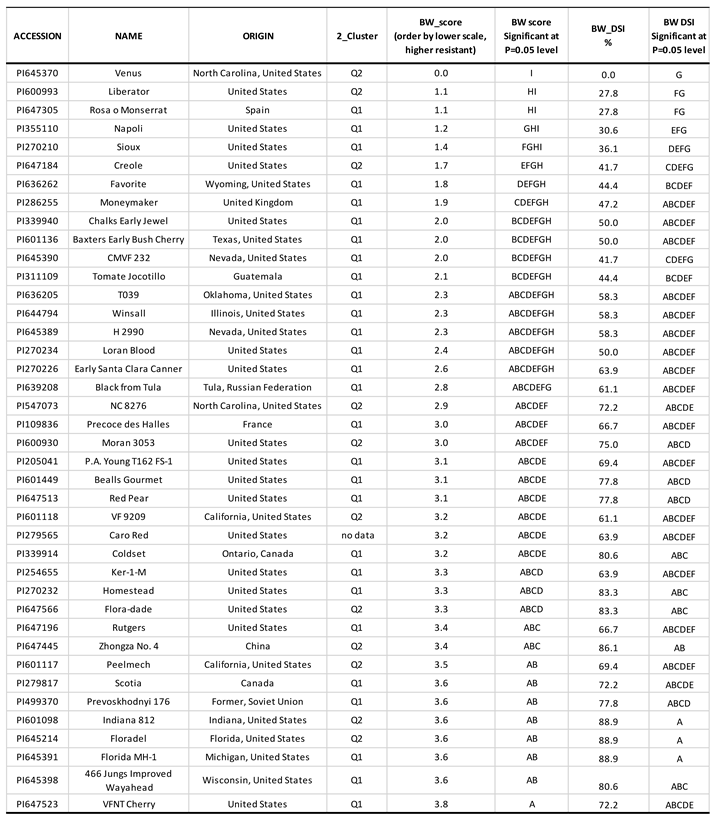
4.1. Parameters and distributions of bacterial wilt resistance
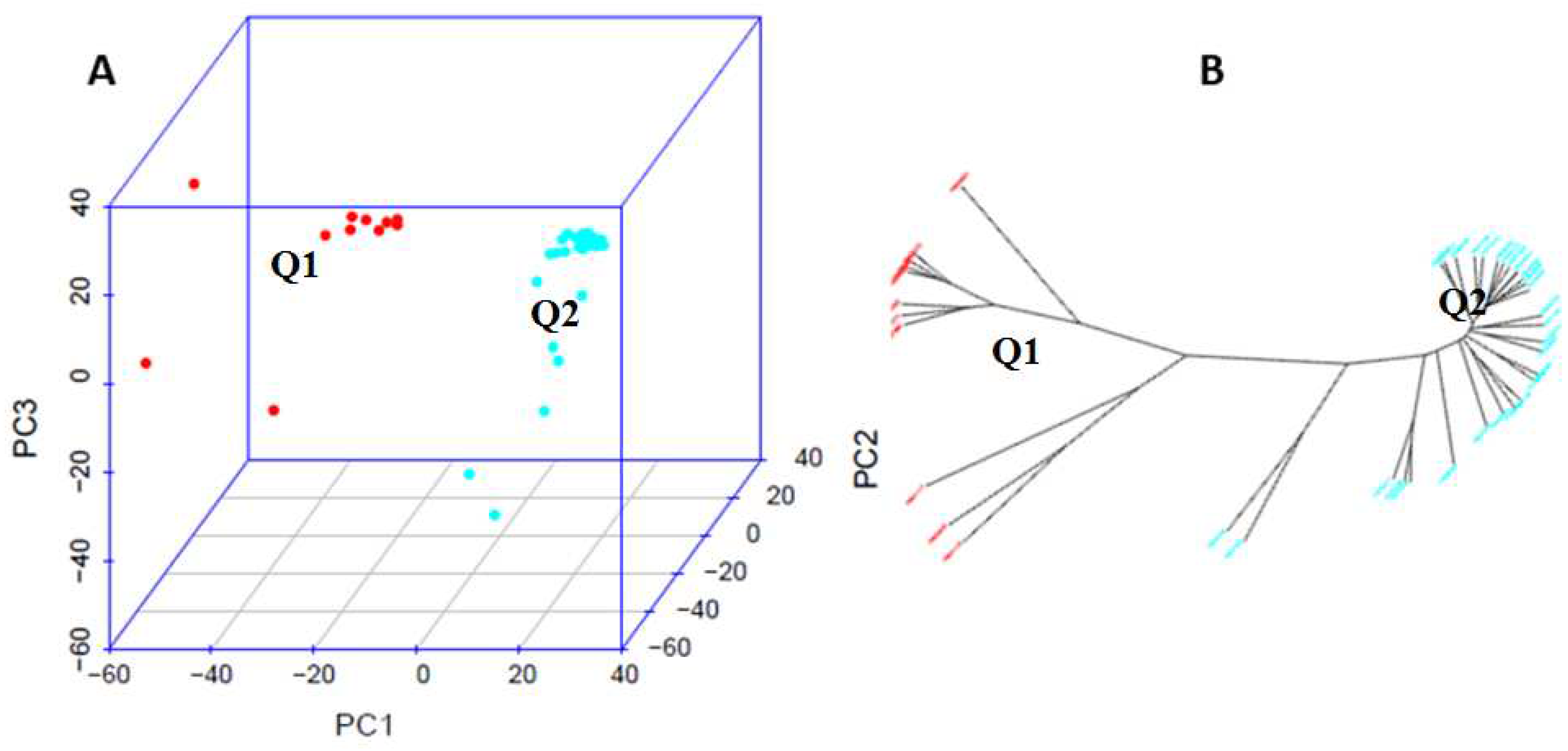
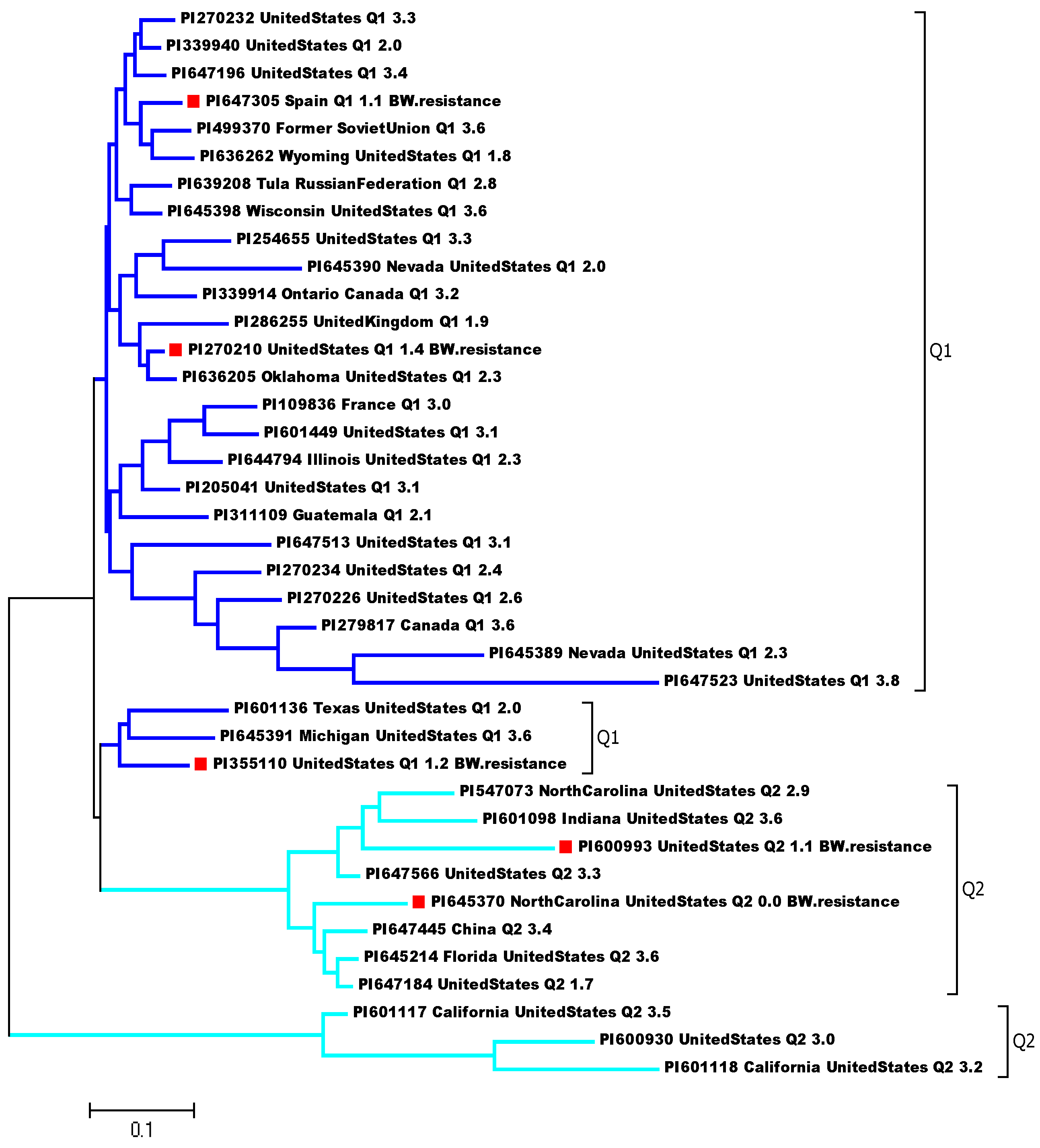
4.2. Principal Component Analysis (PCA) and Genetic Diversity Analysis
5. Discussion
6. Conclusion
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Álvarez, B., López, M. M., & Biosca, E. G. (2007). Influence of native microbiota on survival of Ralstonia solanacearum phylotype II in river water microcosms. Applied and Environmental Microbiology, 73(22), 7210–7217. [CrossRef]
- Aslam, M. N., Mukhtar, T., Hussain, M. A., & Raheel, M. (2017). Assessment of resistance to bacterial wilt incited by Ralstonia solanacearum in tomato germplasm. Journal of Plant Diseases and Protection, 124(6), 585–590. [CrossRef]
- Barik, S., Reddy, A. C., Ponnam, N., Kumari, M., C, A. G., Reddy D C, L., Petikam, S., & Gs, S. (2020). Breeding for bacterial wilt resistance in eggplant (Solanum melongena L.): Progress and prospects. Crop Protection, 137, 105270. [CrossRef]
- Bi-Hao, C., Jian-Jun, L., Yong, W., & Guo-Ju, C. (2009). Inheritance and identification of SCAR marker linked to bacterial wilt-resistance in eggplant. African Journal of Biotechnology, 8(20), 5201–5207. http://www.academicjournals.org/AJB.
- Bindal, S., & Srivastava, S. (2019). Bacterial wilt of solanaceous crops: Sign, symptoms and management. Agrica, 8(2), 134. [CrossRef]
- Bita, C. E., & Gerats, T. (2013). Plant tolerance to high temperature in a changing environment: Scientific fundamentals and production of heat stress-tolerant crops. Frontiers in Plant Science (Vol. 4, Issue JUL). Frontiers Research Foundation. [CrossRef]
- Bocsanczy, A. M., Espindola, A. S., & Norman, D. J. (2019). Whole-genome sequences of Ralstonia solanacearum strains, emerging pathogens of blueberry in Florida. [CrossRef]
- Champoiseau, P. G., Jones, J. B., & Allen, C. (2009). Ralstonia solanacearum Race 3 Biovar 2 causes tropical losses and temperate anxieties. Plant Health Progress, 10(1). [CrossRef]
- Chellemi, D. O., Dankers, H. A., Olson, S. M., Hodge, N. C., & Scott, J. W. (1994). Evaluating bacterial wilt-resistant tomato genotypes using a regional approach. J. AMER. SOC. HORT. SCI (Vol. 119, Issue 2).
- Chiang, K. S., Liu, H. I., & Bock, C. H. (2017). A discussion on disease severity index values. Part I: warning on inherent errors and suggestions to maximize accuracy. Annals of Applied Biology, 171(2), 139–154. [CrossRef]
- Collins, E. J., Bowyer, C., Tsouza, A., & Chopra, M. (2022). Tomatoes: an extensive review of the associated health impacts of tomatoes and factors that can affect their cultivation. Biology (Vol. 11, Issue 2). MDPI. [CrossRef]
- Deslandes, L., Olivier, J., Theulières, F., Hirsch, J., Feng, D. X., Bittner-Eddy, P., Beynon, J., & Marco, Y. (2002). Resistance to Ralstonia solanacearum in Arabidopsis thaliana is conferred by the recessive rrs1-r gene, a member of a novel family of resistance genes. Proceedings of the National Academy of Sciences of the United States of America, 99(4), 2404. [CrossRef]
- Du, H., Wen, C., Zhang, X., Xu, X., Yang, J., Chen, B., & Geng, S. (2019). Identification of a major qtl (Qrrs-10.1) that confers resistance to Ralstonia solanacearum in pepper (capsicum annuum) using slaf-bsa and qtl mapping. International Journal of Molecular Sciences, 20(23). [CrossRef]
- Elshire, R. J., Glaubitz, J. C., Sun, Q., Poland, J. A., Kawamoto, K., Buckler, E. S., & Mitchell, S. E. (2011). A robust, simple genotyping-by-sequencing (GBS) approach for high-diversity species. PLOS ONE, 6(5). [CrossRef]
- Fock, I., Collonnier, C., Purwito, A., Luisetti, J., Souvannavong, V., Vedel, F., Servaes, A., Ambroise, A., Kodja, H., Ducreux, G., & Sihachakr, D. (2000). Resistance to bacterial wilt in somatic hybrids between Solanum tuberosum and Solanum phureja. Plant Science, 160(1), 165–176. [CrossRef]
- Glaubitz, J. C., Casstevens, T. M., Lu, F., Harriman, J., Elshire, R. J., Sun, Q., & Buckler, E. S. (2014). TASSEL-GBS: A high-capacity genotyping by sequencing analysis pipeline. PLOS ONE, 9(2). [CrossRef]
- Grimault, V., Prior, P., & Anaïs, G. (1995). A monogenic dominant resistance of tomato to bacterial wilt in Hawaii 7996 is associated with plant colonization by pseudomonas solanacearum. Journal of Phytopathology, 143(6), 349–352. [CrossRef]
- Hong Hai, T. T., Esch, E., & Wang, J. F. (2008a). Resistance to Taiwanese race 1 strains of Ralstonia solanacearum in wild tomato germplasm. European Journal of Plant Pathology, 122(4), 471–479. [CrossRef]
- Hong Hai, T. T., Esch, E., & Wang, J. F. (2008b). Resistance to Taiwanese race 1 strains of Ralstonia solanacearum in wild tomato germplasm. European Journal of Plant Pathology, 122(4), 471–479. [CrossRef]
- Hong, J. C., Norman, D. J., Reed, D. L., Timur Momol, M., & Jones, J. B. (2012). Diversity among Ralstonia solanacearum strains isolated from the southeastern United States. Phytopathology, 102(10), 924–936. [CrossRef]
- Hong, J. K., Jang, S. J., Lee, Y. H., Jo, Y. S., Yun, J. G., Jo, H., Park, C. J., & Kim, H. J. (2018). Reduced bacterial wilt in tomato plants by bactericidal peroxyacetic acid mixture treatment. Plant Pathology Journal, 34(1), 78–84. [CrossRef]
- Huang, M., Liu, X., Zhou, Y., Summers, R. M., & Zhang, Z. (2019). BLINK: A package for the next level of genome-wide association studies with both individuals and markers in the millions. GigaScience, 8(2). [CrossRef]
- Huet, G. (2014). Breeding for resistance to Ralstonia solanacearum. Frontiers in Plant Science (Vol. 5, Issue DEC). Frontiers Media S.A. [CrossRef]
- Ji, P., Allen, C., Sanchez-Perez, A., Yao, J., Elphinstone, J. G., Jones, J. B., & Momol, M. T. (2007). New diversity of Ralstonia solanacearum strains associated with vegetable and ornamental crops in Florida. Plant Disease, 91(2), 195–203. [CrossRef]
- Jung, E. J., Joo, H. J., Choi, S. Y., Lee, S. Y., Jung, Y. H., Lee, M. H., Kong, H. G., & Lee, S.-W. (2014). Resistance evaluation of tomato germplasm against bacterial wilt by Ralstonia solanacearum. Research in Plant Disease, 20(4), 253–258. [CrossRef]
- Kalpage, M. D., & De Costa, D. M. (2014). Isolation of bacteriophages and determination of their efficiency in controlling Ralstonia solanacearum causing bacterial wilt of tomato. Tropical Agricultural Research (Vol. 26, Issue 1).
- Kim, S. G., Hur, O. S., Ro, N. Y., Ko, H. C., Rhee, J. H., Sung, J. S., Ryu, K. Y., Lee, S. Y., & Baek, H. J. (2016a). Evaluation of resistance to Ralstonia solanacearum in tomato genetic resources at seedling stage. Plant Pathology Journal, 32(1), 58–64. [CrossRef]
- Kim, S. G., Hur, O. S., Ro, N. Y., Ko, H. C., Rhee, J. H., Sung, J. S., Ryu, K. Y., Lee, S. Y., & Baek, H. J. (2016b). Evaluation of resistance to Ralstonia solanacearum in tomato genetic resources at seedling stage. Plant Pathology Journal, 32(1), 58–64. [CrossRef]
- Kumar, S., Ramanjini Gowda, P. H., Saikia, B., Debbarma, J., Velmurugan, N., & Chikkaputtaiah, C. (2018). Screening of tomato genotypes against bacterial wilt (Ralstonia solanacearum) and validation of resistance-linked DNA markers. Australasian Plant Pathology, 47(4), 365–374. [CrossRef]
- Kumar, S., Stecher, G., & Tamura, K. (2016). MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution, 33(7), 1870–1874. [CrossRef]
- Kwon, J. S., Nam, J. Y., Yeom, S. I., & Kang, W. H. (2021). Leaf-to-whole plant spread bioassay for pepper and Ralstonia solanacearum interaction determines the inheritance of resistance to bacterial wilt for further breeding. International Journal of Molecular Sciences, 22(5), 1–14. [CrossRef]
- Lebeau, A., Daunay, M. C., Frary, A., Palloix, A., Wang, J. F., Dintinger, J., Chiroleu, F., Wicker, E., & Prior, P. (2011). Bacterial wilt resistance in tomato, pepper, and eggplant: Genetic resources respond to diverse strains in the Ralstonia solanacearum species complex. Phytopathology, 101(1), 154–165. [CrossRef]
- Lee, J. H., Jang, K. S., Choi, Y. H., Kim, J.-C., & Choi, G. J. (2015). Development of an efficient screening system for resistance of tomato cultivars to Ralstonia solanacearum. Research in Plant Disease, 21(4), 290–296. [CrossRef]
- Lu, Y., Rao, S., Huang, F., Cai, Y., Wang, G., & Cai, K. (2016). Effects of biochar amendment on tomato bacterial wilt resistance and soil microbial amount and activity. International Journal of Agronomy, 2016. [CrossRef]
- Maji A. T. (2012a). Application of principal component analysis for rice germplasm characterization and evaluation. Journal of Plant Breeding and Crop Science, 4(6). [CrossRef]
- Maji A. T. (2012b). Application of principal component analysis for rice germplasm characterization and evaluation. Journal of Plant Breeding and Crop Science, 4(6). [CrossRef]
- Mcavoy, T., Freeman, J. H., Rideout, S. L., Olson, S. M., & Paret, M. L. (2012). Evaluation of grafting using hybrid rootstocks for management of bacterial wilt in field tomato production. HORTSCIENCE (Vol. 47, Issue 5).
- Mohan, V., Gupta, S., Thomas, S., Mickey, H., Charakana, C., Chauhan, V. S., Sharma, K., Kumar, R., Tyagi, K., Sarma, S., Gupta, S. K., Kilambi, H. V., Nongmaithem, S., Kumari, A., Gupta, P., Sreelakshmi, Y., & Sharma, R. (2016). Tomato fruits show wide phenomic diversity but fruit developmental genes show low genomic diversity. PLOS ONE, 11(4). [CrossRef]
- Morel, A., Peeters, N., Vailleau, F., Barberis, P., Jiang, G., Berthomé, R., & Guidot, A. (2018). Plant pathogenicity phenotyping of Ralstonia solanacearum strains. Methods in Molecular Biology, 1734, 223–239. [CrossRef]
- Muthoni, J., Shimelis, H., & Melis, R. (2012). Management of bacterial wilt [Rhalstonia solanacearum Yabuuchi et al., 1995] of potatoes: An opportunity for host resistance in Kenya. Journal of Agricultural Science, 4(9). [CrossRef]
- Newton B-Mensah, I., Osei, K., & Prempeh, R. N. A. (2021). Screening tomato genotypes for bacterial wilt disease (Ralstonia solanacearum) resistance in Ghana. European Journal of Agriculture and Food Sciences, 3(5), 1–8. [CrossRef]
- Newton Boakye-Mensah, I. (2020). Evaluation of tomato (solanum lycopersicum l) genotypes for bacterial wilt (Ralstonia solanacearum) resistance in Ghana. https://ir.ucc.edu.gh/xmlui.
- Patil, V. U., Gopal, J., & Singh, B. P. (2012). Improvement for bacterial wilt resistance in potatoes by conventional and biotechnological approaches. Agricultural Research (Vol. 1, Issue 4, pp. 299–316). Springer. [CrossRef]
- Perveen, R., Suleria, H. A. R., Anjum, F. M., Butt, M. S., Pasha, I., & Ahmad, S. (2015). Tomato (solanum lycopersicum) carotenoids and lycopene chemistry; metabolism, absorption, nutrition, and allied health claims—a comprehensive review. Critical Reviews in Food Science and Nutrition, 55(7), 919–929. [CrossRef]
- Prior, P. (2005). How complex is the “Ralstonia solanacearum species complex? https://www.researchgate.net/publication/37628297.
- Ritchie, H., Rosado, P., & Roser, M. (2023). Agricultural production. Our World in Data. https://ourworldindata.org/agricultural-production.
- Rochette, N. C., Rivera-Colón, A. G., & Catchen, J. M. (2019). Stacks 2: Analytical methods for paired-end sequencing improve RADseq-based population genomics. Molecular Ecology, 28(21), 4737–4754. [CrossRef]
- Roy, S. J., Tucker, E. J., & Tester, M. (2011). Genetic analysis of abiotic stress tolerance in crops. Current Opinion in Plant Biology, 14(3), 232–239. [CrossRef]
- S. K. Kaushik. (2011). Genetics of fruit yield and its contributing characteristics in tomato (Solanum lycopersicom). Journal of Agricultural Biotechnology and Sustainable Development, 3(10). [CrossRef]
- Schmidt, P., Hartung, J., Bennewitz, J., & Hans-Peter, P. (2019). Heritability in plant breeding on a genotype-difference basis. Genetics, 212(4), 991–1008. [CrossRef]
- Shaheen, M. R., Ayyub, C. M., Amjad, M., & Waraich, E. A. (2016). Morpho-physiological evaluation of tomato genotypes under high-temperature stress conditions. Journal of the Science of Food and Agriculture, 96(8), 2698–2704. [CrossRef]
- Singh, S., Gautam, R. K., Singh, D. R., Sharma, T. V. R. S., Sakthivel, K., & Roy, S. D. (2015). Genetic approaches for mitigating losses caused by bacterial wilt of tomato in tropical islands. European Journal of Plant Pathology (Vol. 143, Issue 2, pp. 205–221). Kluwer Academic Publishers. [CrossRef]
- Wang, G., Kong, J., Cui, D., Zhao, H., Niu, Y., Xu, M., Jiang, G., Zhao, Y., & Wang, W. (2019). Resistance against Ralstonia solanacearum in tomatoes depends on the methionine cycle and the γ-aminobutyric acid metabolic pathway. Plant Journal, 97(6), 1032–1047. [CrossRef]
- Yadav, R. K., Tripathi, M. K., Tiwari, S., Tripathi, N., Asati, R., Patel, V., Sikarwar, R. S., & Payasi, D. K. (2023). Breeding and genomic approaches towards the development of fusarium wilt resistance in chickpea. Life (Vol. 13, Issue 4). MDPI. [CrossRef]
- Yuliar, Asi Nion, Y., & Toyota, K. (2015). Recent trends in control methods for bacterial wilt diseases caused by Ralstonia solanacearum. Microbes and Environments (Vol. 30, Issue 1, pp. 1–11). Japanese Society of Microbial Ecology. [CrossRef]
- Zhou, R., Wu, Z., Wang, X., Rosenqvist, E., Wang, Y., Zhao, T., & Ottosen, C. O. (2018). Evaluation of temperature stress tolerance in cultivated and wild tomatoes using photosynthesis and chlorophyll fluorescence. Horticulture Environment and Biotechnology, 59(4), 499–509. [CrossRef]
- Zohoungbogbo, H., Quenum, A., Honfoga, J., Chen, J. R., Achigan-Dako, E., Kenyon, L., & Hanson, P. (2021). Evaluation of resistance sources of tomato (Solanum lycopersicum L.) to phylotype I strain of Ralstonia solanacearum species complex in Benin. Agronomy, 11(8). [CrossRef]


 |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




