1. Introduction
The coronavirus disease 2019 (COVID-19) pandemic is caused by a highly transmissible and pathogenic positive-sense, single-stranded RNA coronavirus, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) [
1]. The virus primarily affects the respiratory system, causing sickness with symptoms such as fever, malaise, dry cough, shortness of breath and respiratory distress [
2]. SARS-CoV-2 infection is now known to manifest as systemic inflammation, leading to sepsis, acute cardiac injury, and heart failure and multi-organ dysfunction in patients at high risk [
3]. Globally, more than 768.04 million confirmed cases and 6.95 million deaths have been reported to WHO as of June 18, 2023.
SARS-CoV-2 genome undergoes thousands of mutations since the emergence of the virus [
4,
5,
6]. That leads to the surge of new viral strains, classified as variants of concern (VOCs), possessing increased virulence, transmissibility, and decreased response to available diagnostics, vaccines, and therapeutics [
7,
8]. The increasing prevalence of SARS-CoV-2 variants has raised serious concerns regarding possible increased infection severity or failure on the effectiveness of current vaccines thus giving rise to greater challenge to diagnostic and clinical management [
7]. Vaccination remains the most effective and cost-efficient means for prevention of infectious diseases [
9]. Many different types of vaccine have been developed, including some approved for immunization widely in human to control SARS-CoV-2 infection in the past three years. Although the authorized COVID-19 vaccines are still effective in preventing severe illness, mRNA and adenovirus vectored vaccines showed impaired effectiveness and rapid immunity waning against VOCs [
10]. Therefore, development of more effective, safer, and broad-spectrum vaccines is critically needed to respond to the evolution of SARS-CoV-2 and control COVID-19.
Virus-like particles (VLPs) are self-assembled structures from viral antigens that mimic the three-dimensional, morphological structure of authentic virions [
11,
12]. VLPs have been demonstrated to be very effective against a variety of viral pathogens in humans and thus are considered a highly potential vaccine platform [
13]. Different approaches have been reported for generation of SARS-CoV-2 VLPs [
13,
14,
15,
16]. Recombinant baculovirus-based vaccines are safe, and capable of inducing humoral and cellular responses [
17,
18,
19]. In addition, baculoviruses have been shown to enhance immunogenicity of the vaccine antigen, a similar function of an adjuvant [
20]. In this study, we generated a SARS-CoV-2 VLP using recombinant baculovirus BacMam and evaluated its immunogenicity in a mouse trial.
2. Materials and methods
2.1. Cells and culture media
Spodoptera frugiperda (Sf9) cells were propagated at 28℃ in Sf-900™ II SFM (ThermoFisher Scientific) supplemented with 100 units Penicillin and 100 µg Streptomycin/mL (MilliporeSigma) and 5% (vol/vol) Fetal Bovine Serum (MilliporeSigma). HEK-293 and HEK-293T cells were cultured and maintained in Dulbecco’s Modified Eagle Medium (DMEM, ThermoFisher Scientific) supplemented with 10% FBS and 100 units Penicillin and 100 µg Streptomycin/mL (MilliporeSigma) (designated D10 medium) at 37℃ in 5% CO2 atmosphere.
2.2. Plasmids construction
To generate baculoviruses (BacMam) capable of expressing a single and multiple proteins in mammalian cells thus could be used directly as an expression system in mammals [
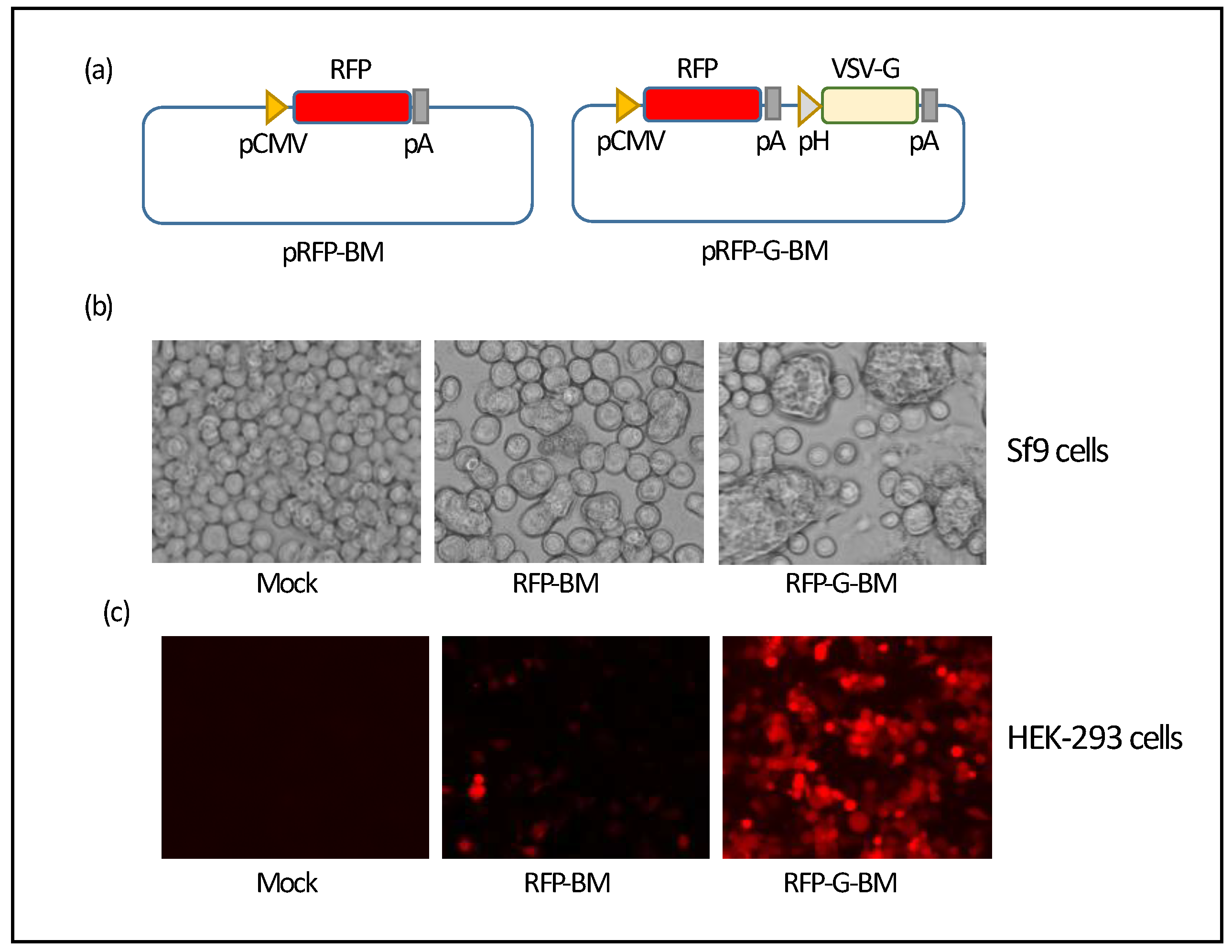
21], we first constructed a baculovirus donor plasmid carrying a reporter gene, red fluorescent protein (RFP). A DNA fragment containing CMV promoter followed by the RFP-coding sequence was amplified by PCR with pDsRed-Monomer-C1 (Clontech) as the template and primers CMV-RFP-F and CMV-RFP-R. PCR product was cloned into the pFastBac1 vector (ThermoFisher Scientific) for generation of an RFP-expressing BacMam (assigned as pRFP-BM) (
Figure 1a). To generate a BacMam with potential of increasing transduction efficiency in mammalian cells, an expression cassette containing vesicular stomatitis virus glycoprotein (VSV-G) coding sequence placed downstream of Polyhedrin promoter and followed by the SV40 poly A signal sequence was amplified by PCR with primer pairs (1) AvrII-pH-F and pH-VSV-G-R, (2) pH-VSV-G-F and VSV-G-SV40pA-R, (3) VSV-G-SV40pA-F and SV40pA-AvrII-R, and cloned into the pRFP-BM plasmid at the AvrII site (assigned as pRFP-G-BM) (
Figure 1a).
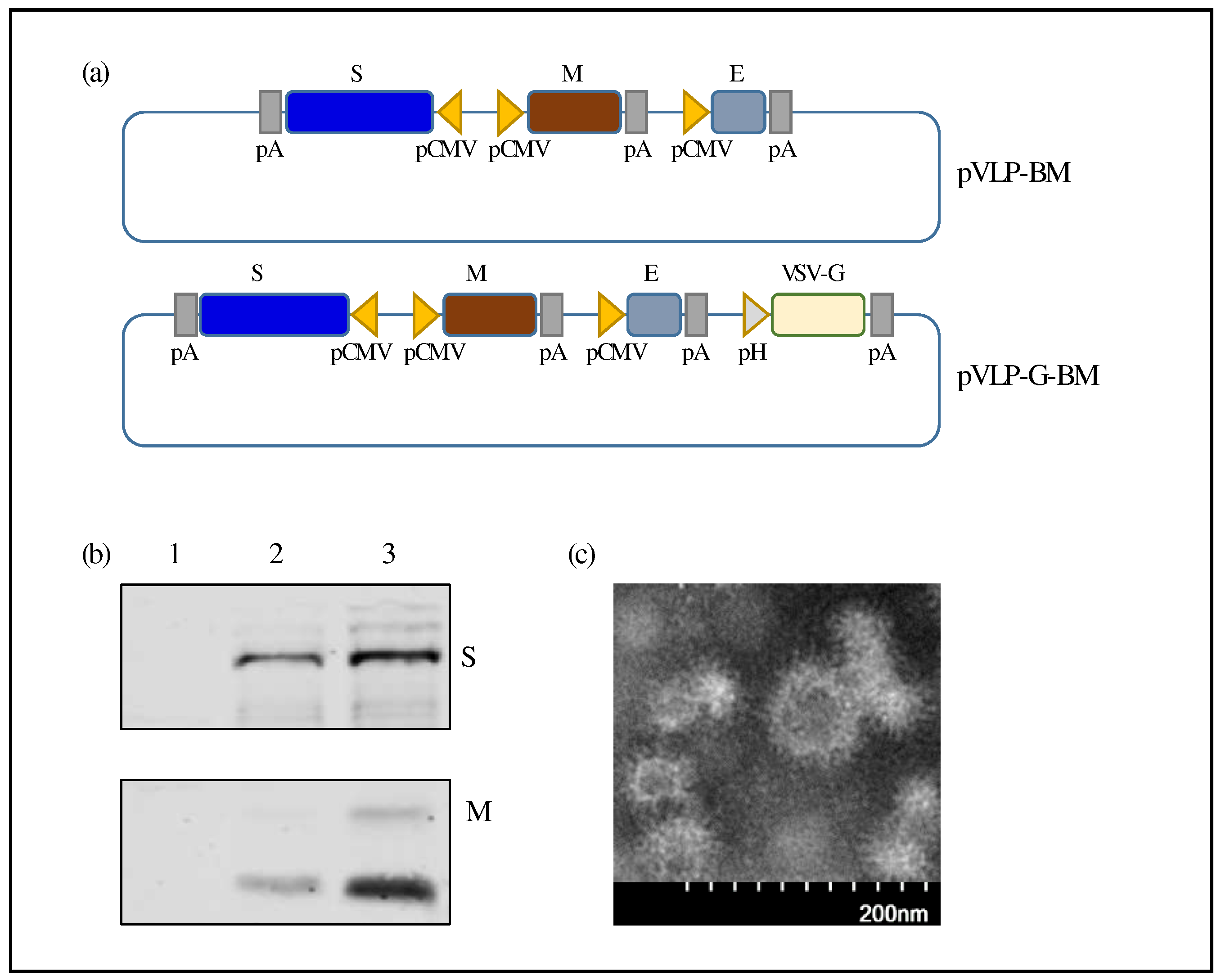
To construct recombinant donor plasmid for generation of a baculovirus expressing SARS-CoV-2 VLP, coding sequences for spike (S), membrane (M), and envelope (E) proteins of SARS-CoV-2 ancestral Wuhan strain (GenBank accession number NC_045512) were codon optimized for mammalian expression and synthesized (Bio Basic Canada). An intermediate plasmid (pS-BM) was first constructed by inserting the S gene into pFastBacTriple1 vector (ThermoFisher Scientific). Subsequently, four DNA fragments containing the M and E genes were amplified by PCR with 4 pairs of primers, (1) Rsr2-CMV-F and CMV-Cov2-M-R, (2) CMV-Cov2-M-F and pP10-CMV-R, (3) pP10-CMV-F and CMV-Cov2-E-R, (4) CMV-Cov2-E-F and MfeI-Cov2-E-R, then assembled into the pS-BM plasmid using GenBuilderᵀᴹ Plus Cloning Kit (Genscript) to generate pVLP-BM (
Figure 2a). Each gene was placed downstream of CMV promoter and followed by a poly A signal sequence. HSV TK poly A signal sequence was inserted downstream of the S and M genes, while SV40 poly A signal sequence was cloned downstream of the E gene. To generate a second version of SARS-CoV-2 VLP BacMam, a donor plasmid was constructed by assembling PCR fragments containing the VSV-G-expressing cassette as described above into the pVLP-BM plasmid and assigned as pVLP-G-BM (
Figure 2a). H
uman angiotensin-converting enzyme 2 (ACE2) cDNA in a plasmid was obtained from Sino Biological Inc. ACE2 coding sequence with a carboxyl V5-polyhistidine tag was cloned into the pEB4.6 episomal vector with a puromycin-resistant gene to allow selection of cells that stably express ACE2 protein after transfection [
22]. pHAGE-CMV-Luc2-IRES-ZsGreen-W is a lentiviral backbone plasmid expressing both luciferase and GFP reporters was obtained from BEI Resources (NR-52516) [
23]. A packaging plasmid psPAX2 was a gift from Didier Trono (Addgene plasmid #12260). Plasmid expressing the SARS-CoV-2 D614G spike protein with a C-terminal 19 amino acid deletion pcDNA-SARS-CoV-2-D614G-D19 WT was kindly provided by Dr. Bo Meng [
24]. Phusion™ High-Fidelity DNA Polymerase (ThermoFisher Scientific) was used for PCR amplification. Primers and plasmids used in this study are listed in
Table 1 and
Table 2, respectively. All constructed plasmids were verified by PCR targeting the insertion junctions, restriction enzyme digestion, and DNA sequencing.
2.3. Generation of recombinant BacMam viruses
To generate recombinant baculoviruses, bacmid DNA was extracted from E. coli DH10Bac (ThermoFisher Scientific) that were transformed with the donor plasmids. Sf9 cells in a T25 flask were transfected with 3 µg of the bacmid DNA using Cellfectin™ II Reagent (ThermoFisher Scientific). Culture media containing the viruses were harvested at 4 - 6 days post-transfection (dpt). Viruses were amplified by infecting Sf9 in suspension culture at an MOI of 0.1 and harvested at 4 days post-infection (dpi). To purify the virus from infected cell culture, cell debris was removed by centrifugation at 2,000 rpm for 10 minutes. The supernatant was then layered over a 27% sucrose cushion in TSE buffer and centrifuged at 24,000 RPM for 75 minutes. The virus pellet was re-suspended in phosphate-buffered saline (PBS, pH 7.5) and centrifuged at 27,000 RPM for 150 minutes then the final pellet was resuspended in PBS. Purified virus was stored at -80°C and titrated with the BacPAK™ Baculovirus Rapid Titer Kit (Takara).
2.4. Transduction and fluorescent microscopy
HEK-293 cells were pre-seeded overnight in 12-well plate to reach 60 - 70% confluence then incubated with the BacMam viruses in PBS or DMEM for 2 - 3 h at 28℃. The transduction mixtures were subsequently replaced with 1 mL of D10 medium supplemented with 5 mM sodium butyrate. The cells were observed under a fluorescent microscope at 48 h post-transduction and/or harvested in SDS sample buffer for protein identification by Western blotting.
2.5. Western blotting
Cell lysates were subjected to sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto nitrocellulose membranes (Millipore). The membranes were blocked with 3% BSA in PBST (PBS + 0.1% Tween 20) at room temperature (RT) for 1 h and incubated with a primary antibody at 4℃ overnight. After five washes with PBST, the membranes were subsequently incubated with the appropriate infrared dye-labeled secondary antibodies for 1 h at room temperature, followed by five washes with PBST then was scanned with Odyssey CLx Imaging System (Li-Cor Biosciences). Rabbit anti-SARS-CoV-2 Spike Protein S1/S2 (ThermoFisher Scientific), rabbit anti-SARS-CoV Membrane (M) Protein (Rockland), mouse anti-His-tag (Qiagen), and mouse anti-β-actin (Cell Signaling Technology) antibodies were used as primary antibodies. IRDye 680 goat anti-rabbit IgG (Li-Cor Biosciences) and IRDye 800 goat anti-mouse IgG (Li-Cor Biosciences) were utilized as secondary antibodies.
2.6. Transmission electron microscopy (TEM)
Purified SARS-CoV-2 VLPs were subjected to negative staining and examined by transmission electron microscopy. Specifically, a 400-mesh copper grid (Electron Microscopy Sciences) coated with formvar and carbon was placed on a drop of the purified VLP sample and incubated at RT for 1-2 min for the particles to adhere, prior to be transferred to a droplet of water for 20 seconds. Subsequently, the grid was placed on a drop of 0.5% PTA stain solution for 1 min, wicked away the excessive stain and dried. The grid was then analyzed by using a transmission electron microscope, HT7700 (Hitachi High-Tech), and images were acquired through a CCD camera, XR16 (AMT) at the University of Saskatchewan WCVM Imaging Centre.
2.7. Mouse immunogenicity trial
To examine the immunogenicity of SARS-CoV-2 VLP BacMam, BALB/c mice (5-6 weeks old, 8 per group) were intramuscularly injected with 50 µL of PBS or 2.25 x 108 ifu of VLP-G-BM. A second dose of the same vaccine was given after two weeks. Vaccinated animals were observed for general health status and reaction at the injection sites. Blood samples were collected before and 14 days post-vaccination (dpv). On day 14 after the second immunization, all mice were euthanized to collect bronchoalveolar lavage (BAL) and blood samples for measuring antibody titers and neutralization test.
2.8. ELISA
Clear round-bottom Immuno 96-well plates (ThermoFisher Scientific) were coated with 100 μl of purified SARS-CoV-2 S1 protein (2 µg/mL) at 4℃ overnight. The plates were blocked with 150 μl of 1% Bovine Serum Albumin (MilliporeSigma) in TBST (Tris-buffered saline + 0.1% Tween 20) for 1 h. Subsequently, 100 μl of two-fold diluted mouse serum at an initial dilution of 1:200 were added to the coated wells and incubated for 2 h. Then Biotin-labeled goat anti-mouse IgG (ThermoFisher Scientific) was added and incubated for 1 h. Subsequently, the wells were incubated with Streptavidin Alkaline Phosphatase (Jackson Immunoresearch Laboratories) for 1 h at RT. The plates were washed five times with TBST between incubation steps. Finally, p-Nitrophenyl Phosphate (ThermoFisher Scientific) diluted in DE buffer (10.5 mM Diethanolamine, 0.5 mM Magnesium Chloride) was added for colored product development and absorbance was measured at 450 nm with a subtracted reference at 490 nm using a SpectraMax microplate reader (Molecular Devices).
2.9. Creation of ACE2-expressing HEK-293 cells
HEK-293 cells in six-well plate were transfected with 2.5 µg of the ACE2-V5-polyhistine expressing plasmid using JetPEI (Polyplus) and the culture medium was replaced after 4 h. On the next day, 2 µg/mL Puromycin was added to the cell culture medium and concentration of the antibiotic was gradually increased to 10 µg/mL after four days of transfection and onwards. The cells survived under the selection medium were sub-cultured and ACE2 protein expression was confirmed by Western blotting using an anti-His-tag antibody.
2.10. Generation of SARS-CoV-2 spike pseudotyped lentivirus, confocal microscopy, and luciferase assay
Pseudotyped lentivirus with both luciferase and green fluorescent protein (GFP) reporters was generated as described previously [
23,
25]. In brief, HEK-293T cells in six-well plate were co-transfected with 0.34 µg of spike-expressing plasmid pcDNA-SARS-CoV-2-D614G-S-D19 WT [
24], 1 µg of lentiviral backbone plasmid pHAGE-CMV-Luc2-IRES-ZsGreen-W [
23] and 0.66 µg of packaging plasmid psPAX2 plasmids using the JetPEI transfection reagent. Cell culture medium was replaced at 16 hpt and supernatants containing SARS-CoV-2 pseudovirus were collected at 48 and 72 hpt, filtered through a 0.45 µm filter, aliquoted, and stored at -80 ℃.
HEK-293-ACE2 cells seeded in 96-well cell-culture plate (~5 x 10
4 cells/well) pre-treated with Poly-L-lysine (MilliporeSigma) were infected with the SARS-CoV-2 pseudovirus in the presence of 5 µg/mL Polybrene (MilliporeSigma) to minimize charge-repulsion between the virus and cells [
26]. The lentiviral pseudovirus stock volume was used to sufficiently produce >2 x 10
6 relative light units (RLUs) per well in a 96-well plate for achieving a signal of 10
4-fold above the background luciferase activity of mock-infected controls to give a dynamic range sufficient for use in neutralization assay [
27]. The infected cells were harvested at 48 h post-infection (hpi) and luciferase assay was performed using Luciferase™ Reporter Assay System (Promega) in a GloMax 20/20 Luminometer according to the manufacturer’s instructions.
HEK-293-ACE2 cells cultured in 2-well chamber slides (ThermoFisher Scientific) were similarly infected with the SARS-CoV-2 pseudovirus. At 48 hpi, the infected cells were fixed, and the nuclei were stained with 300 nM DAPI (MilliporeSigma) for 10 min at RT. After applying a mounting medium (ProLong™ Diamond Antifade Mountant, ThermoFisher Scientific), the slides were covered with a coverslip, and dried in the dark for 24 h before examination of GFP expression by confocal microscopy.
2.11. Pseudotype virus neutralization assay
Neutralization assay was performed as described previously with minor modifications [
23,
28,
29]. In brief, pooled serum samples of VLP-G BM- and PBS-injected mice at 28 dpv diluted 1:20 in D10 medium were incubated with pseudovirus for 1 h at 37 °C, then utilized to infect the HEK-293-ACE2 cells in 96-well cell-culture plate. The cells were observed under a fluorescence microscope (Zeiss Axioplan 2) with appropriate settings for GFP expression at 24 - 48 hpi and then lyzed with the Passive Lysis buffer (Promega) at 48 hpi for 30 min at RT for luciferase assay. Neutralization activity was determined by the RLUs per well in 96-well plate of the vaccination group relative to the control group.
2.12. Statistical analysis
All experiments in cell culture were done in triplicates and data were expressed as mean ± SEM values. Data analysis was performed using GraphPad Prism 9 and statistical differences were determined by the Student’s t-test. Statistical significance was demonstrated as follows: * if P < 0.05, ** if P < 0.01, *** if P < 0.001, **** if P < 0.0001, and NS if not significant.
3. Results
3.1. Generation and characterization of SARS-CoV-2 VLP BacMam
To explore the potential of a baculovirus-based VLP vaccine against SARS-CoV-2, we first generated recombinant baculoviruses expressing an easily detectable reporter for testing the capability of expressing foreign proteins and transduction/infection efficiency of this system
in vitro. To this end, we first constructed two baculovirus donor plasmids containing RFP gene under the control of CMV promoter followed by a poly A sequence with and without incorporation of VSV-G-coding sequence under the control of Polyhedrin promoter assigned as pRFP-BM and pRFP-G-BM, respectively (
Figure 1a). Recombinant baculoviruses expressing RFP protein in mammalian cells were successfully generated by transfecting Sf9 cells with the bacmids. Typical cytopathic effects (CPEs) in the form of cell to cell fusion and cell swelling were clearly observed in Sf9 cells after two days of infection with these two BacMam viruses and this phenotype was stronger in the RFP-G-BM-infected cells possibly due to membrane-fusion activity of VSV-G protein (
Figure 1b). Expression of RFP was clearly observed two days post-transduction with noticeably higher number of RFP-positive HEK-293 cells transduced with the RFP-G-BM containing VSV-G (
Figure 1c). Our results confirmed that incorporation of VSV-G in recombinant baculoviruses significantly enhances transduction efficiency of the virus into mammalian cells and results in a significant improvement on protein expression [
30,
31]. In addition, our recombinant BacMam viruses showed great capability of infecting/transducing several mammalian cell lines, including HEK-293, HEK-293T, Vero 76, Vero E6, and Huh-7, as evidenced by RFP expression observed under a fluorescent microscope (data not shown).
We then utilized this BacMam system to produce SARS-CoV-2 VLPs. Two BacMam viruses expressing SARS-CoV-2 S, M, and E proteins, a minimal requirement for coronavirus VLP formation [
32], with and without VSV-G were generated, assigned as VLP-BM and VLP-G-BM, respectively. The arrangement of SARS-CoV-2 structural protein-coding sequences in the BacMam donor constructs was shown in Fig 2a. The expression of SARS-CoV-2 S and M proteins was confirmed by Western blotting using protein-specific antibodies. Higher levels of SARS-CoV-2 S and M protein expression in HEK-293 cells transduced with the VLP-G-BM was observed compared with those transduced with the VLP-BM (
Figure 2b). Formation of SARS-CoV-2 VLPs in HEK-293 cells transduced with the VLP-G-BM virus was demonstrated by transmission electron microscopy with negative staining (
Figure 2c). Our VLPs displayed corona-like morphology with
round shape, spike-like structures decorated on the surface, and the average diameter fell around 85.2 ± 19.8 nm
, which is in accordance with previously reported SARS-CoV-2 VLPs [
15] and SARS-CoV-2 particles [
33]
. These results showed that transduction of our VLP-G-BM virus led to the formation of SARS-CoV-2 VLP in mammalian cells.
3.2. Production of SARS-CoV-2 VLP BacMam virus
For production of the VLP BacMam, we first adapted Sf9 cells in suspension culture and then infected the cells with VLP-G-BM at an MOI of 1. The culture medium of infected cells was harvested 3 - 4 days post-infection and the recombinant VLP BacMam was purified and concentrated by ultra-centrifugation and stored at -80℃. The titre of the purified VLP BacMam virus was 1.25 x 1010 ifu/mL.
3.3. Immunogenicity of SARS-CoV-2 VLP BacMam virus in mice
To evaluate the immunogenicity of our baculovirus-based VLP vaccine in mice, VLP-G-BM at 2.5 x 10
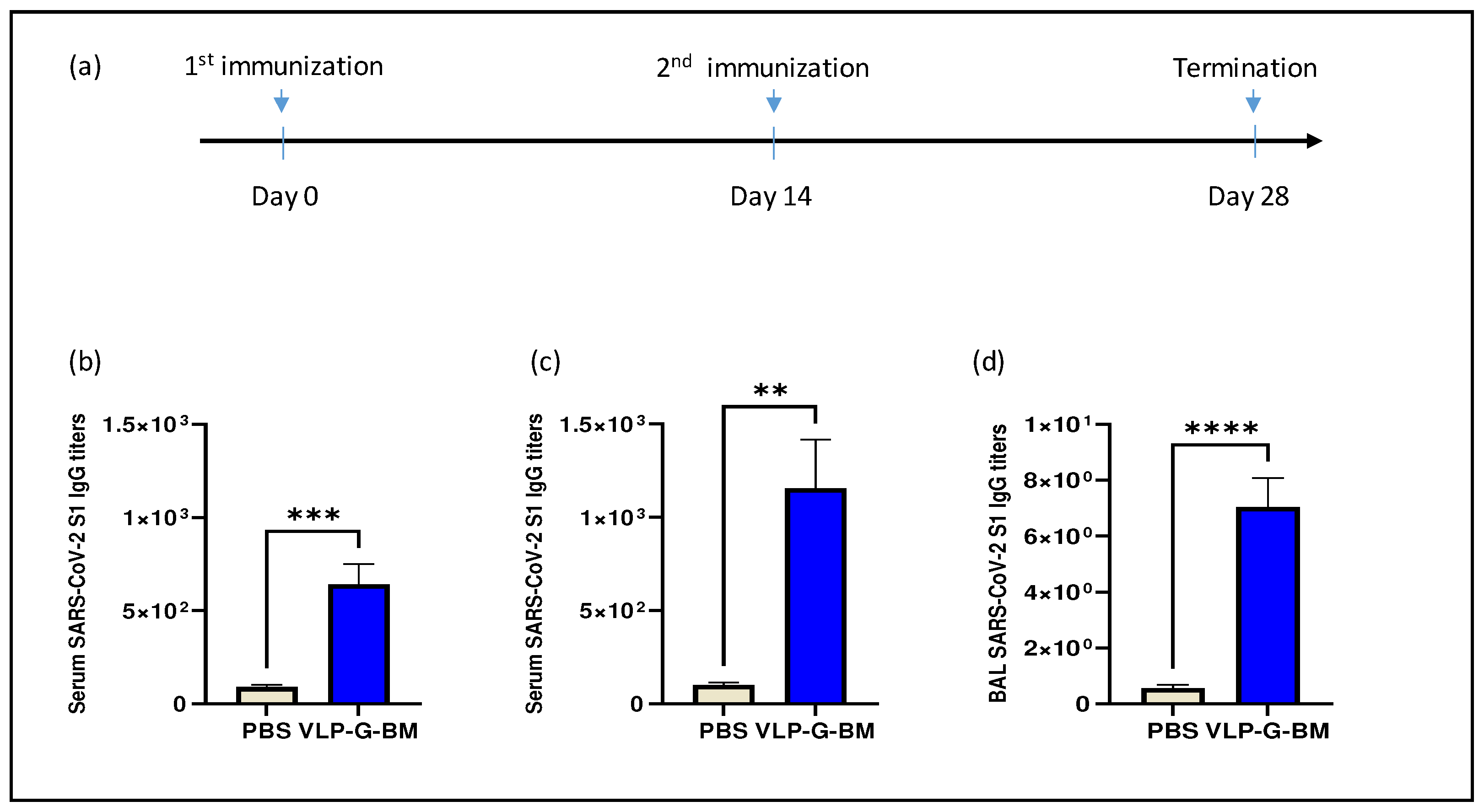
8 ifu per mouse was injected intramuscularly and boosted with the same dose of vaccine at two weeks post-primary immunization (
Figure 3a). No overt adverse clinical events were observed in the animals throughout the experiment following vaccination, demonstrating the safety of the vaccine.
The immune response to SARS-CoV-2 spike S1 protein was examined in sera and bronchoalveolar lavage (BAL) after 14 and 28 days post-vaccination of the VLP BacMam vaccine. Mice vaccinated with the VLP BacMam showed significantly higher serum IgG levels on day 14 after primary immunization (
Figure 3b) and further increased on day 28 after the boosting shot (
Figure 3c). In addition, the VLP-G-BM induced IgG antibody production in the BAL of vaccinated animals on 28 dpv (
Figure 3d). These results demonstrated that the VLP BacMam vaccine was safe and capable of inducing spike-specific immune responses in sera and in the lung after intramuscular immunization.
3.4. VLP BacMam induced neutralization against SARS-CoV-2
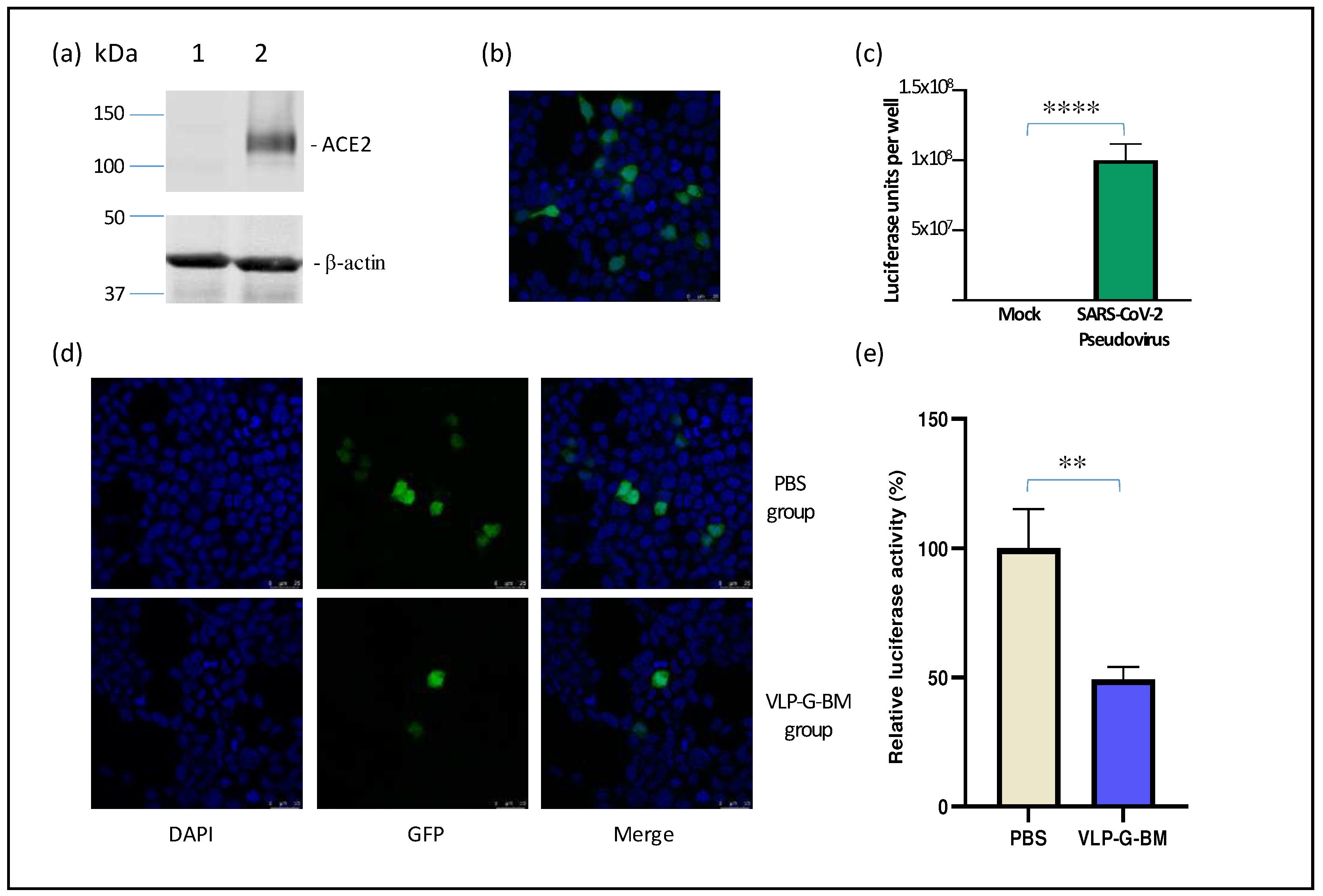
To evaluate the capability of inducing neutralization against SARS-CoV-2 of our VLP BacMam vaccine, we produced a pseudotyped lentivirus bearing SARS-CoV-2 spike protein with GFP and luciferase reporters. For efficient infection of SARS-CoV-2 pseudotyped virus, we also generated a ACE2-expressing cell line (HEK-293-ACE2). The stable expression of ACE2 protein in HEK-293-ACE2 cells was confirmed by Western blotting (
Figure 4a). HEK-293-ACE2 cells infected with the SARS-CoV-2 pseudotyped virus displayed both GFP determined by confocal microscopy (
Figure 4b) and luciferase activity by luciferase assay (
Figure 4c), indicating successful transduction. Importantly, we observed similar neutralization titres of a reference positive serum sample using both the pseudotyped virus and authentic SARS-CoV-2 virus (data not shown), demonstrating the reliability of the pseudotyped virus for use in neutralization assay.
In comparison to PBS-vaccinated animals, sera of VLP BacMam-vaccinated animals showed lower number of GFP positive cells (
Figure 4d) and approximately 50% reduction in luciferase activity in HEK-293-ACE2 cells infected with the SARS-CoV-2 spike psuedotyped virus pre-treated with the VLP BacMam-vaccinated mouse sera. These results demonstrated that vaccination with our VLP BacMam induced neutralizing antibodies, which could be capable of providing protection against SARS-CoV-2 infection.
4. Discussion
Many infectious diseases have emerged or re-emerged over the past few decades. These diseases have led to millions of deaths and caused enormous slowdown of the global economy [
34]. The COVID-19
pandemic, caused by highly transmissible SARS-CoV-2, is
the most recent example of those deadly infections and has spread all over the globe within a short period, infecting hundreds of millions of people and resulting in millions of deaths. The global spread and impacts of COVID-19 had provoked an urgent need for preventative approaches through vaccinations [
35], which has greatly contributed to the overall success in controlling the disease globally so far. However, similar with other RNA viruses, SARS-CoV-2 constantly changes through mutation and lead to the emergence of new variants over time [
36]. Therefore, tremendous efforts have been put into improving the efficacy of existing vaccines and developing newer vaccines.
Recombinant baculoviruses have been reported previously as a safe and effective platform for gene delivery and vaccine development against important viral pathogens [
17,
18,
19,
30,
37]. Engineering mammalian promoters into the baculovirus genome (BacMam) has allowed protein expression in mammalian cells as an additional feature of the platform [
38]. In this work, we constructed a recombinant BacMam expressing SARS-CoV-2 structural proteins spike, membrane, and envelope. Transmission electron microscopy experiment showed the formation of VLPs in mammalian cells after BacMam transduction. Immunogenicity study showed that the VLP BacMam elicited the production of SARS-COV-2 spike-specific IgG antibody after intramuscular injections in mouse sera and in mouse lung. More importantly, we showed that the sera of VLP BacMam vaccinated mice inhibited the transduction of a SARS-CoV-2 spike pseudotyped lentivirus, indicating the presence of neutralizing antibodies.
The spike protein is the main component of the approved vaccines due primarily to its ability of inducing neutralizing antibodies [
39]. Unfortunately, however, there have been constant mutations in the spike protein, resulting in altered antigenicity and thus reduced efficacy and rapid immunity waning of the vaccines developed against previous spike sequences [
40]. Therefore, inclusion of more conserved viral components has been proposed to provide broader protection [
39]. Another consideration when it comes to improving the existing vaccines is to induce cross-protective T cell responses, which have been shown to be important for preventing severe COVID-19 disease [
40]. In this regard, the M and E proteins have become good candidates due to their high-level conservation and the identification of multiple T cell epitopes in these two proteins [
39]. The VLP constructed in our work expresses both M and E proteins. It would be interesting to characterize the immune responses to these two proteins in future research.
In conclusion, we demonstrated the formation of SARS-CoV-2 VLP by a recombinant BacMam virus and its capability of inducing immune responses in sera and in the lung in mice after immunization. These results warrant further investigation to evaluate the protective efficacy of the VLP vaccine against viral challenge.
Author Contributions
Conceptualization, Q.L.; methodology, H.T.N. and Q.L.; validation, H.T.N. and Q.L.; formal analysis, H.T.N. and Q.L.; investigation, H.T.N.; resources, D.F. and Q.L.; writing—original draft preparation, H.T.N.; writing—review and editing, Q.L. and D.F.; supervision, Q.L.; funding acquisition, Q.L., D.F., and V.G.. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by grants from Canadian Institutes of Health Research (CIHR) DC0190GP, Saskatchewan Agriculture Development Fund 2018-0101, Natural Sciences and Engineering Research Council of Canada (NSERC) RGPIN-2018-04138, and VIDO. VIDO receives operational funding from the Government of Saskatchewan through Innovation Saskatchewan and the Ministry of Agriculture and from the Canada Foundation for Innovation through the Major Science Initiatives.
Institutional Review Board Statement
The animal study protocol was approved by the Animal Research Ethics Board, University of Saskatchewan (Protocol code 20200016, August 31, 2020).
Data Availability Statement
The original Western blotting images presented in the study are included in the supplementary material.
Acknowledgments
We would like to thank the animal care staff at VIDO for their assistance in the animal trial. We also would like to thank Mingmin Liao, Erin Scruten, Ken Lai, Jocelyne Lew, and WCVM Imaging Centre for their help. The following reagent was obtained through BEI Resources, NIAID, NIH: SARS-related Coronavirus 2, Wuhan-Hu-1 Spike-Pseudotyped Lentivirus Kit, NR-52948. This article is published with the permission of the Director of VIDO, journal series no. 1034.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Shereen, M.A., et al., COVID-19 infection: Origin, transmission, and characteristics of human coronaviruses. J Adv Res, 2020. 24: p. 91-98.
- Hui, D.S., et al., The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health - The latest 2019 novel coronavirus outbreak in Wuhan, China. International Journal of Infectious Diseases, 2020. 91: p. 264-266. [CrossRef]
- Chen, T., et al., Clinical characteristics of 113 deceased patients with coronavirus disease 2019: retrospective study. Bmj-British Medical Journal, 2020. 368. [CrossRef]
- Cosar, B., et al., SARS-CoV-2 Mutations and their Viral Variants. Cytokine & Growth Factor Reviews, 2022. 63: p. 10-22.
- Chen, J.H., et al., Mutations Strengthened SARS-CoV-2 Infectivity. Journal of Molecular Biology, 2020. 432(19): p. 5212-5226.
- Wang, R., et al., Decoding SARS-CoV-2 Transmission and Evolution and Ramifications for COVID-19 Diagnosis, Vaccine, and Medicine. Journal of Chemical Information and Modeling, 2020. 60(12): p. 5853-5865.
- Khateeb, J., Y.C. Li, and H.B. Zhang, Emerging SARS-CoV-2 variants of concern and potential intervention approaches. Critical Care, 2021. 25(1).
- Sanyaolu, A., et al., The emerging SARS-CoV-2 variants of concern. Therapeutic Advances in Infectious Disease, 2021. 8.
- Wen, Z.S., et al., Chitosan Nanoparticles Act as an Adjuvant to Promote both Th1 and Th2 Immune Responses Induced by Ovalbumin in Mice. Marine Drugs, 2011. 9(6): p. 1038-1055. [CrossRef]
- Tada, T., et al., Comparison of Neutralizing Antibody Titers Elicited by mRNA and Adenoviral Vector Vaccine against SARS-CoV-2 Variants. bioRxiv, 2021.
- Grgacic, E.V.L. and D.A. Anderson, Virus-like particles: Passport to immune recognition. Methods, 2006. 40(1): p. 60-65. [CrossRef]
- Kushnir, N., S.J. Streatfield, and V. Yusibov, Virus-like particles as a highly efficient vaccine platform: Diversity of targets and production systems and advances in clinical development. Vaccine, 2012. 31(1): p. 58-83. [CrossRef]
- Hemmati, F., et al., Plant-derived VLP: a worthy platform to produce vaccine against SARS-CoV-2. Biotechnology Letters, 2022. 44(1): p. 45-57.
- Geng, Q., et al., Novel virus-like nanoparticle vaccine effectively protects animal model from SARS-CoV-2 infection. PLoS Pathog, 2021. 17(9): p. e1009897.
- Xu, R.D., et al., Construction of SARS-CoV-2 Virus-Like Particles by Mammalian Expression System (vol 8, 862, 2020). Frontiers in Bioengineering and Biotechnology, 2020. 8.
- Lu, J., et al., A COVID-19 mRNA vaccine encoding SARS-CoV-2 virus-like particles induces a strong antiviral-like immune response in mice. Cell Research, 2020. 30(10): p. 936-939.
- Argilaguet, J.M., et al., BacMam immunization partially protects pigs against sublethal challenge with African swine fever virus. Antiviral Research, 2013. 98(1): p. 61-65. [CrossRef]
- Keil, G.M., et al., BacMam Platform for Vaccine Antigen Delivery. Vaccine Technologies for Veterinary Viral Diseases, 2016. 1349: p. 105-119.
- Zhang, J., et al., BacMam virus-based surface display of the infectious bronchitis virus (IBV) S1 glycoprotein confers strong protection against virulent IBV challenge in chickens. Vaccine, 2014. 32(6): p. 664-670. [CrossRef]
- Hervas-Stubbs, S., et al., Insect baculoviruses strongly potentiate adaptive immune responses by inducing type I IFN. J Immunol, 2007. 178(4): p. 2361-9.
- Kost, T.A., J.P. Condreay, and D.L. Jarvis, Baculovirus as versatile vectors for protein expression in insect and mammalian cells. Nat Biotechnol, 2005. 23(5): p. 567-575.
- Nguyen, H.T., et al., Construction of a Noninfectious SARS-CoV-2 Replicon for Antiviral-Drug Testing and Gene Function Studies. Journal of Virology, 2021. 95(18).
- Crawford, K.H.D., et al., Protocol and Reagents for Pseudotyping Lentiviral Particles with SARS-CoV-2 Spike Protein for Neutralization Assays. Viruses-Basel, 2020. 12(5).
- Meng, B., et al., Recurrent emergence of SARS-CoV-2 spike deletion H69/V70 and its role in the Alpha variant B.1.1.7. Cell Reports, 2021. 35(13).
- Shi, J., et al., Effective vaccination strategy using SARS-CoV-2 spike cocktail against Omicron and other variants of concern. Npj Vaccines, 2022. 7(1).
- Denning, W., et al., Optimization of the Transductional Efficiency of Lentiviral Vectors: Effect of Sera and Polycations. Molecular Biotechnology, 2013. 53(3): p. 308-314. [CrossRef]
- Thakur, N., et al., Production of Recombinant Replication-defective Lentiviruses Bearing the SARS-CoV or SARS-CoV-2 Attachment Spike Glycoprotein and Their Application in Receptor Tropism and Neutralisation Assays. Bio Protoc, 2021. 11(21): p. e4249.
- Liu, X.L., et al., Enhanced elicitation of potent neutralizing antibodies by the SARS-CoV-2 spike receptor binding domain Fc fusion protein in mice. Vaccine, 2020. 38(46): p. 7205-7212.
- Yi, C.Y., et al., Key residues of the receptor binding motif in the spike protein of SARS-CoV-2 that interact with ACE2 and neutralizing antibodies. Cellular & Molecular Immunology, 2020. 17(6): p. 621-630.
- Tang, X.C., H.R. Lu, and T.M. Ross, Baculovirus-Produced Influenza Virus-like Particles in Mammalian Cells Protect Mice from Lethal Influenza Challenge. Viral Immunology, 2011. 24(4): p. 311-319. [CrossRef]
- Barsoum, J., et al., Efficient transduction of mammalian cells by a recombinant baculovirus having the vesicular stomatitis virus G glycoprotein. Human Gene Therapy, 1997. 8(17): p. 2011-2018. [CrossRef]
- Swann, H., et al., Minimal system for assembly of SARS-CoV-2 virus like particles (vol 10, 21877, 2020). Scientific Reports, 2021. 11(1).
- Prasad, S., et al., Transmission electron microscopy imaging of SARS-CoV-2. Indian Journal of Medical Research, 2020. 151(2-3): p. 241-243.
- Pati, R., M. Shevtsov, and A. Sonawane, Nanoparticle Vaccines Against Infectious Diseases. Frontiers in Immunology, 2018. 9. [CrossRef]
- Kim, C., J.D. Kim, and S.U. Seo, Nanoparticle and virus-like particle vaccine approaches against SARS-CoV-2. Journal of Microbiology, 2022. 60(3): p. 335-346.
- Walensky, R.P., H.T. Walke, and A.S. Fauci, SARS-CoV-2 Variants of Concern in the United States-Challenges and Opportunities. Jama-Journal of the American Medical Association, 2021. 325(11): p. 1037-1038.
- Kost, T.A., et al., Implementation of BacMam virus gene delivery technology in a drug discovery setting. Drug Discovery Today, 2007. 12(9-10): p. 396-403. [CrossRef]
- Kost, T.A. and J.P. Condreay, Recombinant baculoviruses as mammalian cell gene-delivery vectors. Trends Biotechnol, 2002. 20(4): p. 173-80. [CrossRef]
- Dai, L. and G.F. Gao, Viral targets for vaccines against COVID-19. Nat Rev Immunol, 2021. 21(2): p. 73-82. [CrossRef]
- Carabelli, A.M., et al., SARS-CoV-2 variant biology: immune escape, transmission and fitness. Nat Rev Microbiol, 2023. 21(3): p. 162-177.
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).