Submitted:
15 December 2023
Posted:
18 December 2023
You are already at the latest version
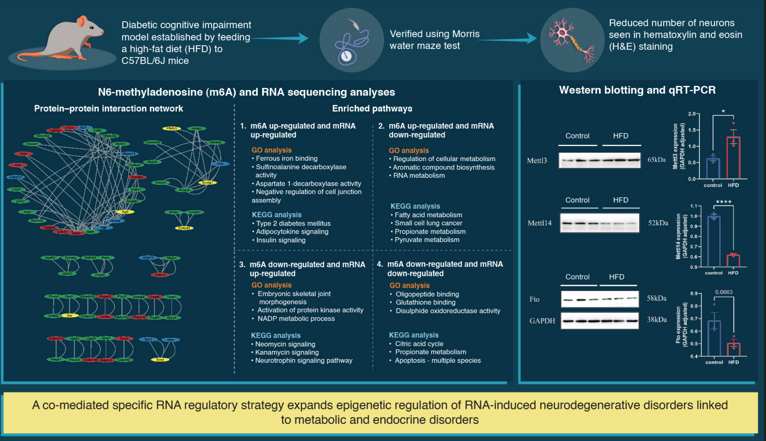
Abstract
Keywords:
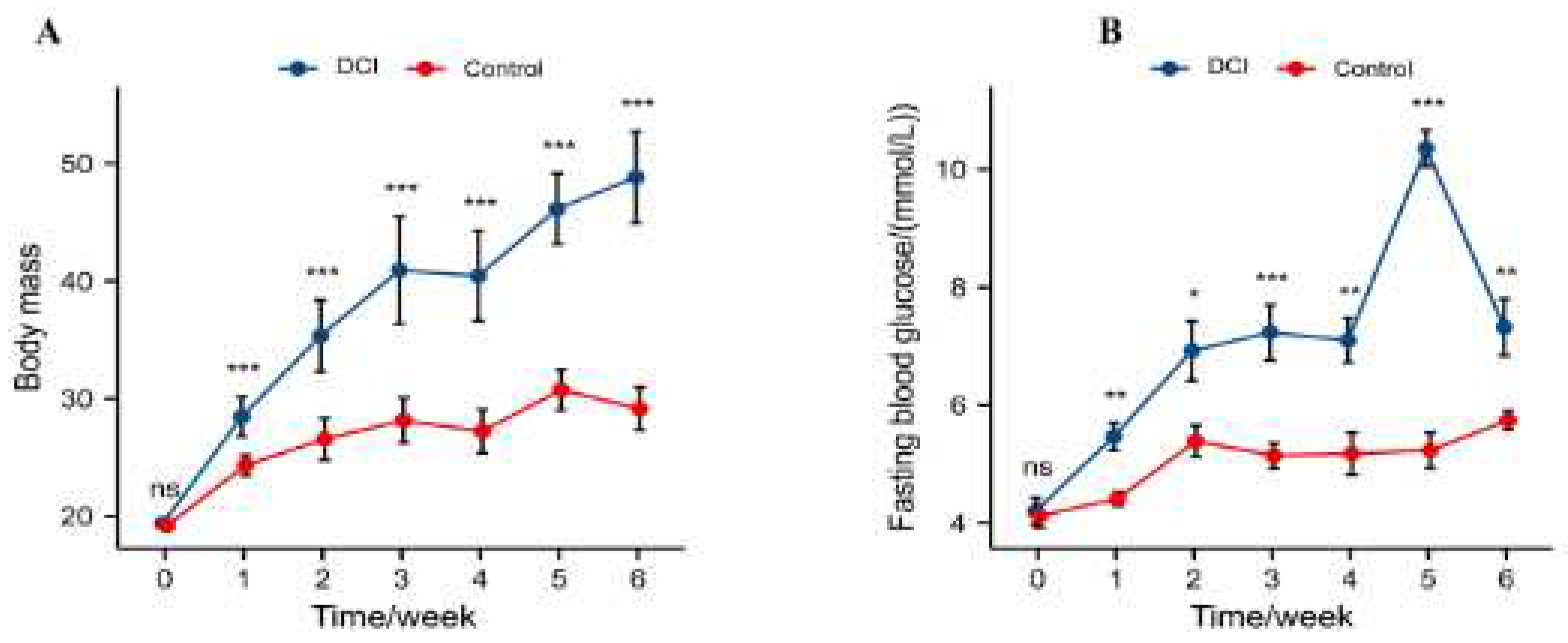
1. Introduction
2. Results
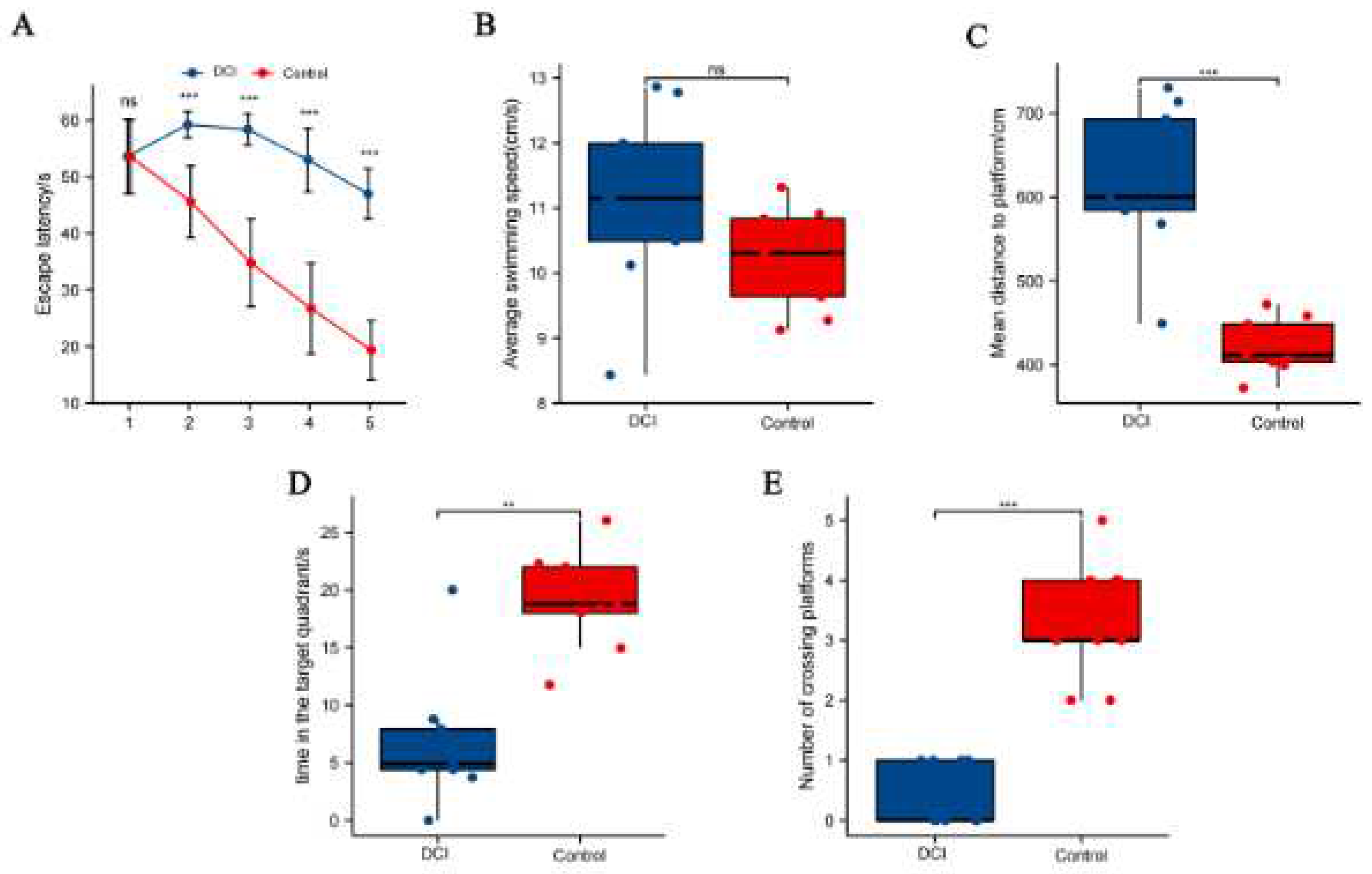
Results of the Morris water maze test
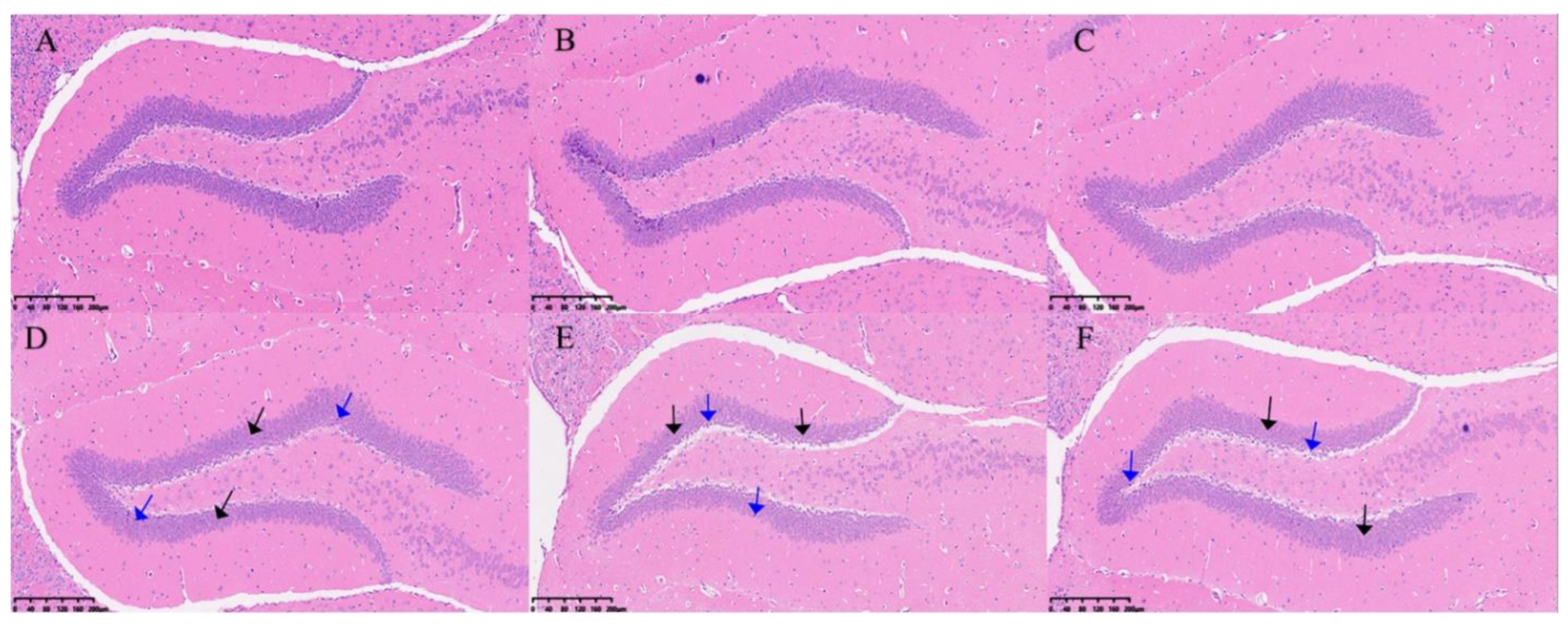
H&E staining of the hippocampus in the two groups of mice
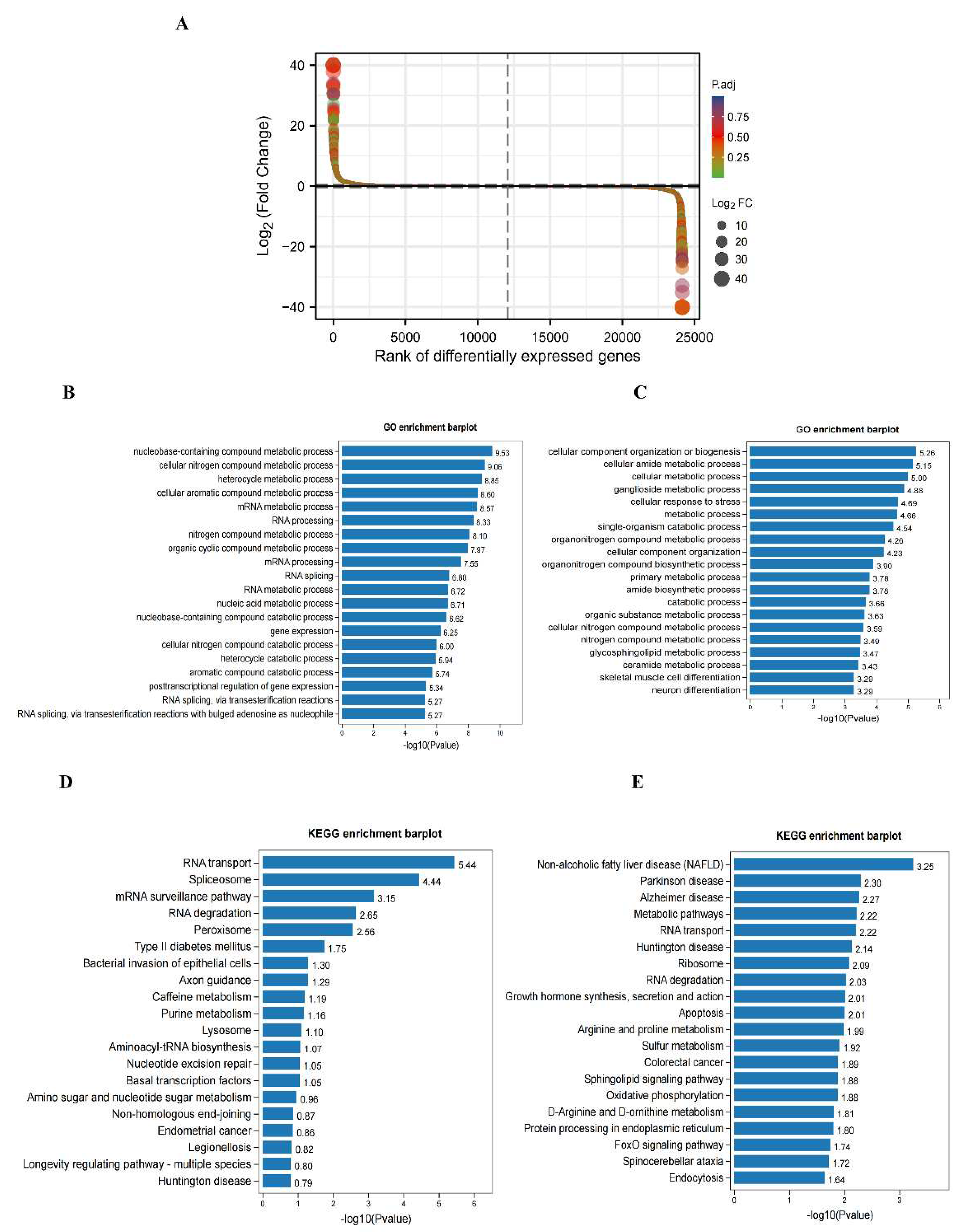
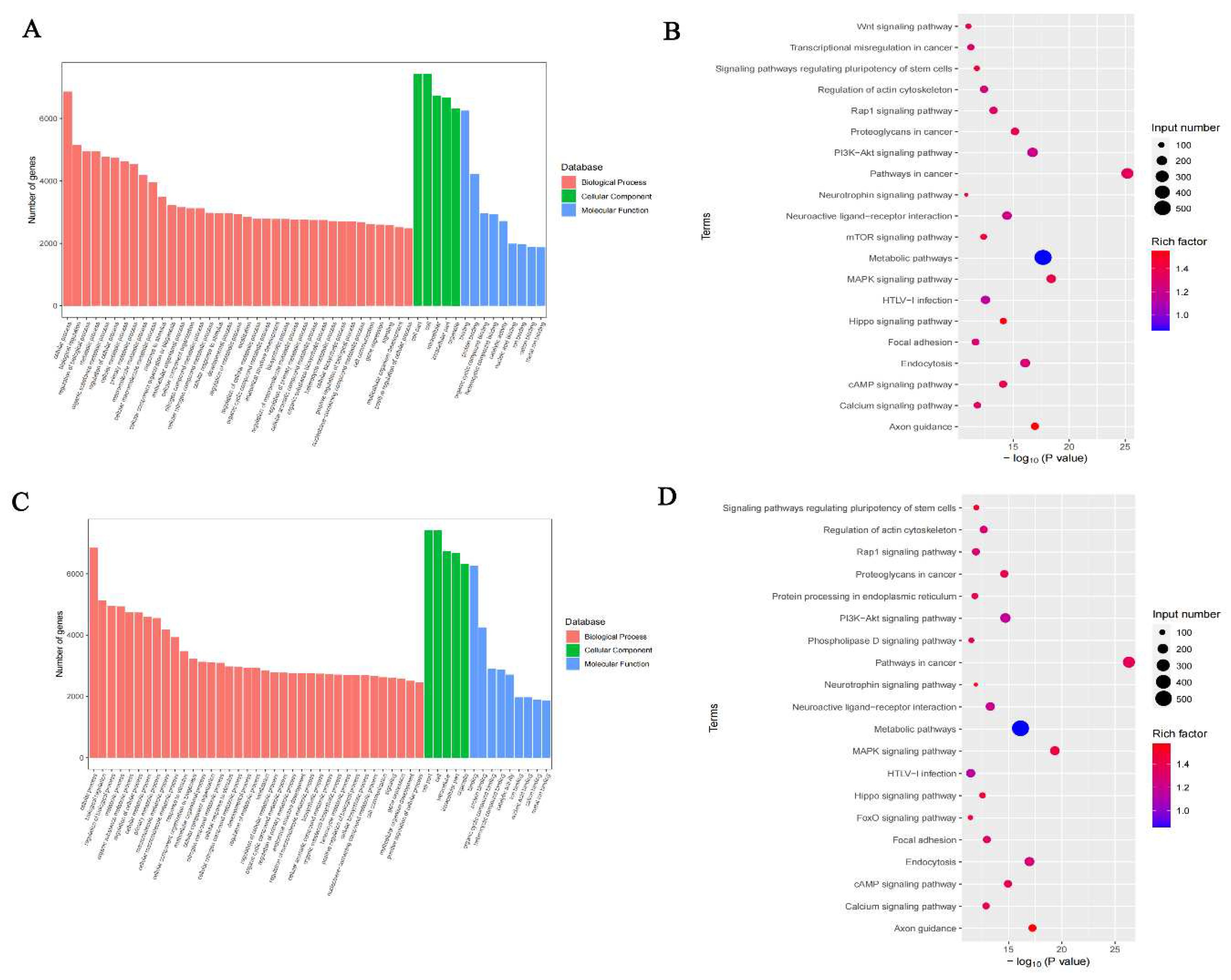
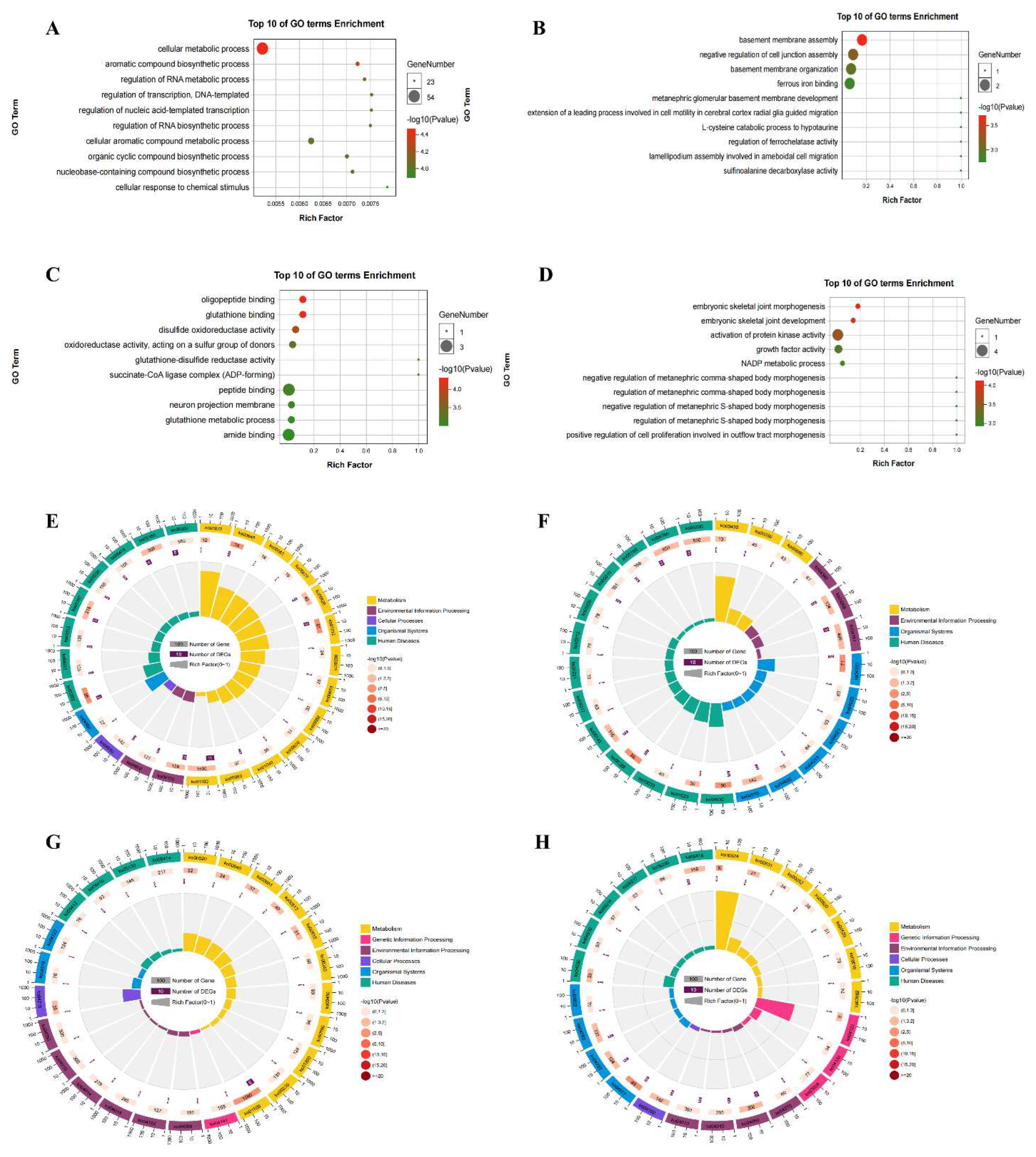
GO and KEGG pathway analyses of DEGs in the hippocampus of DCI mice
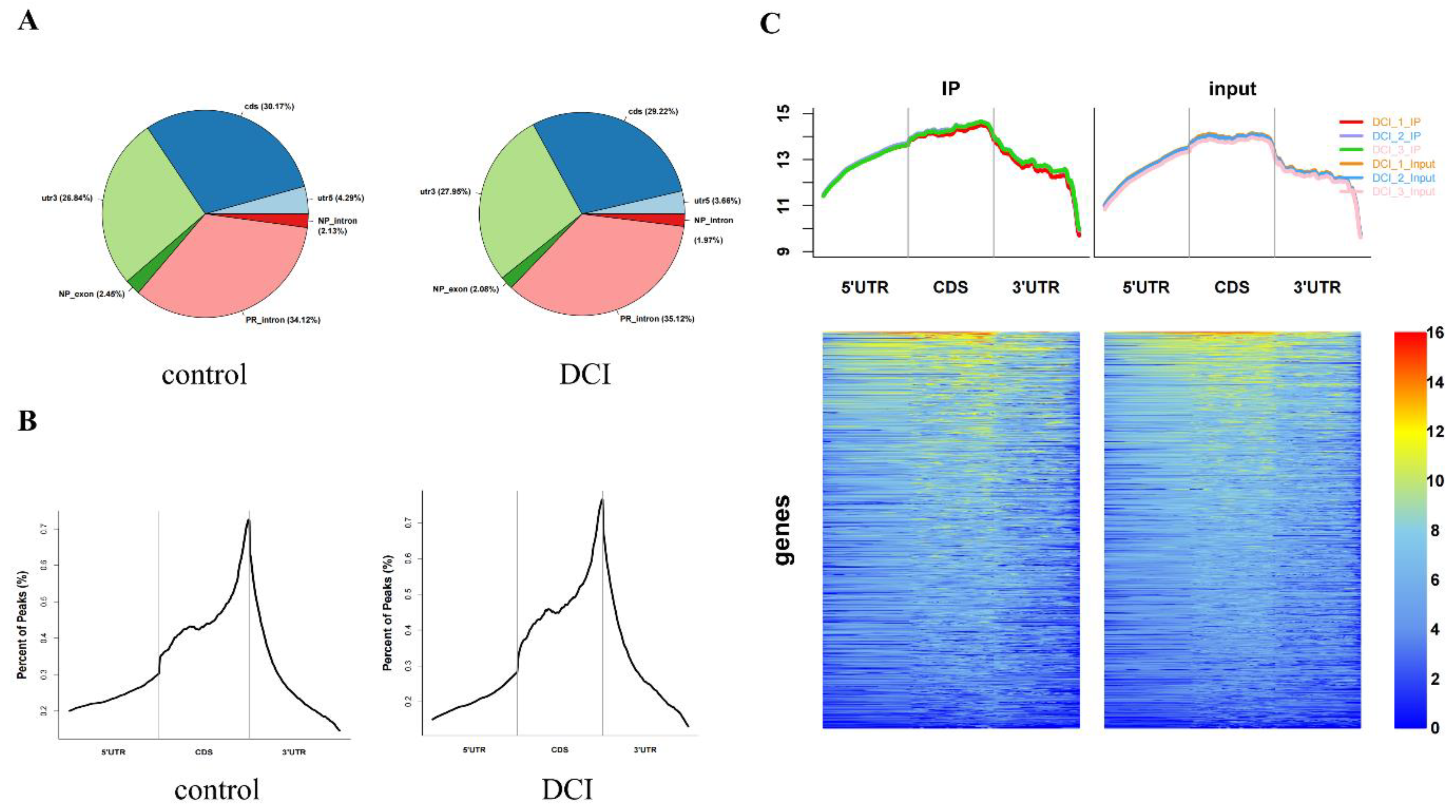
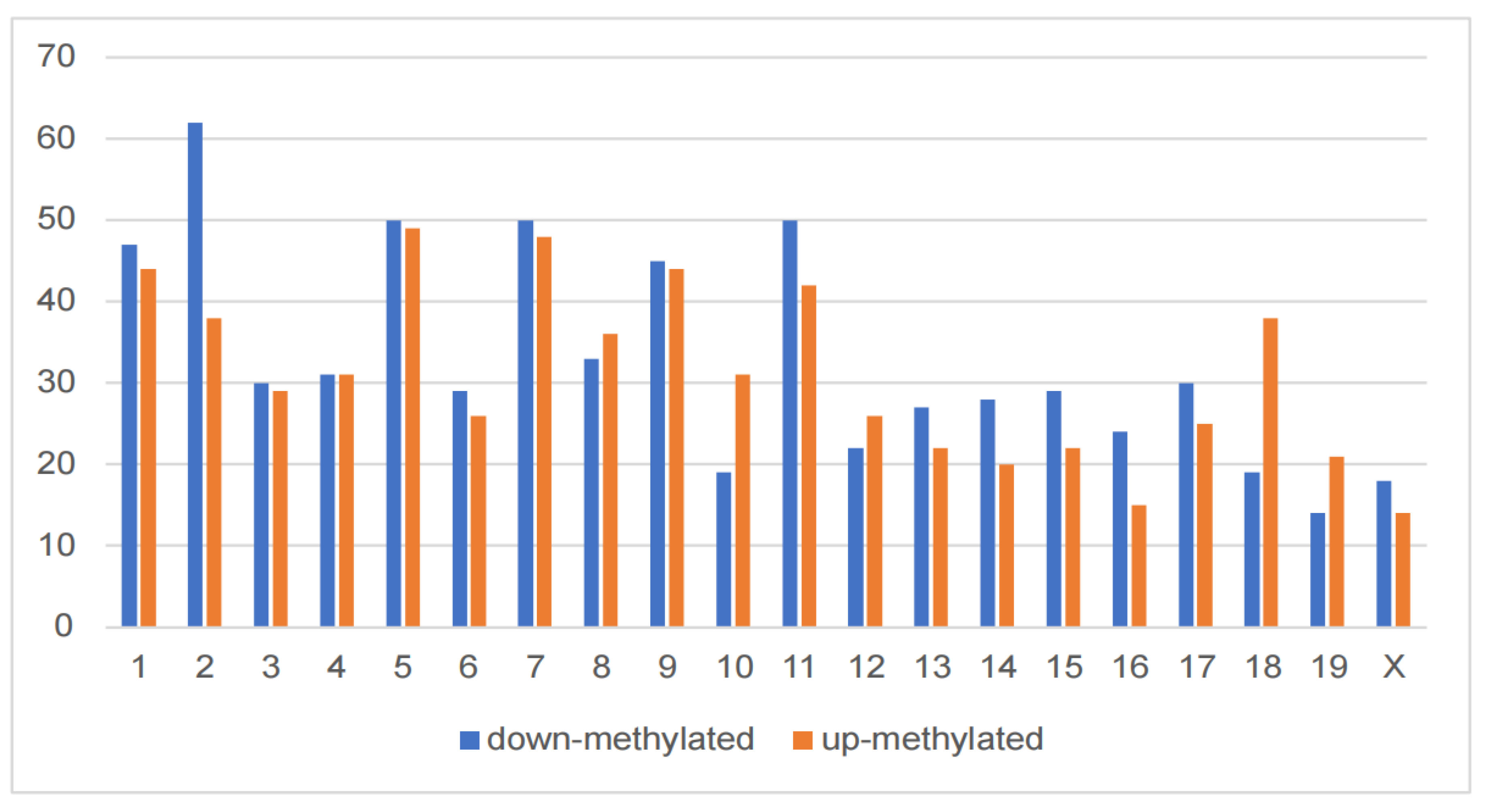
Altered m6A modification of genes in the hippocampus of DCI mice
Differential m6A-modification and expression of genes in the cerebral cortex result from altered m6A methyltransferase and demethylase levels.
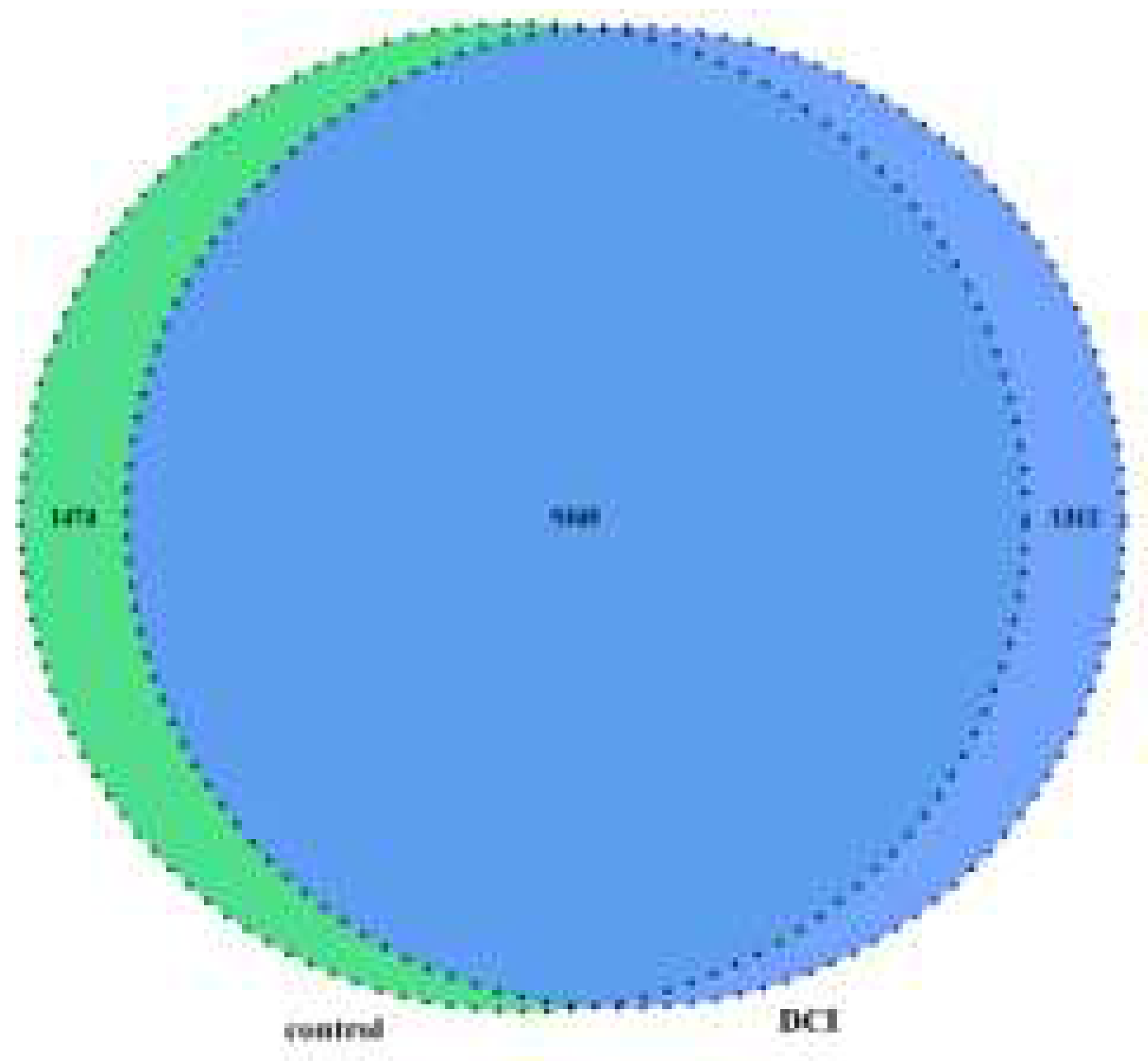
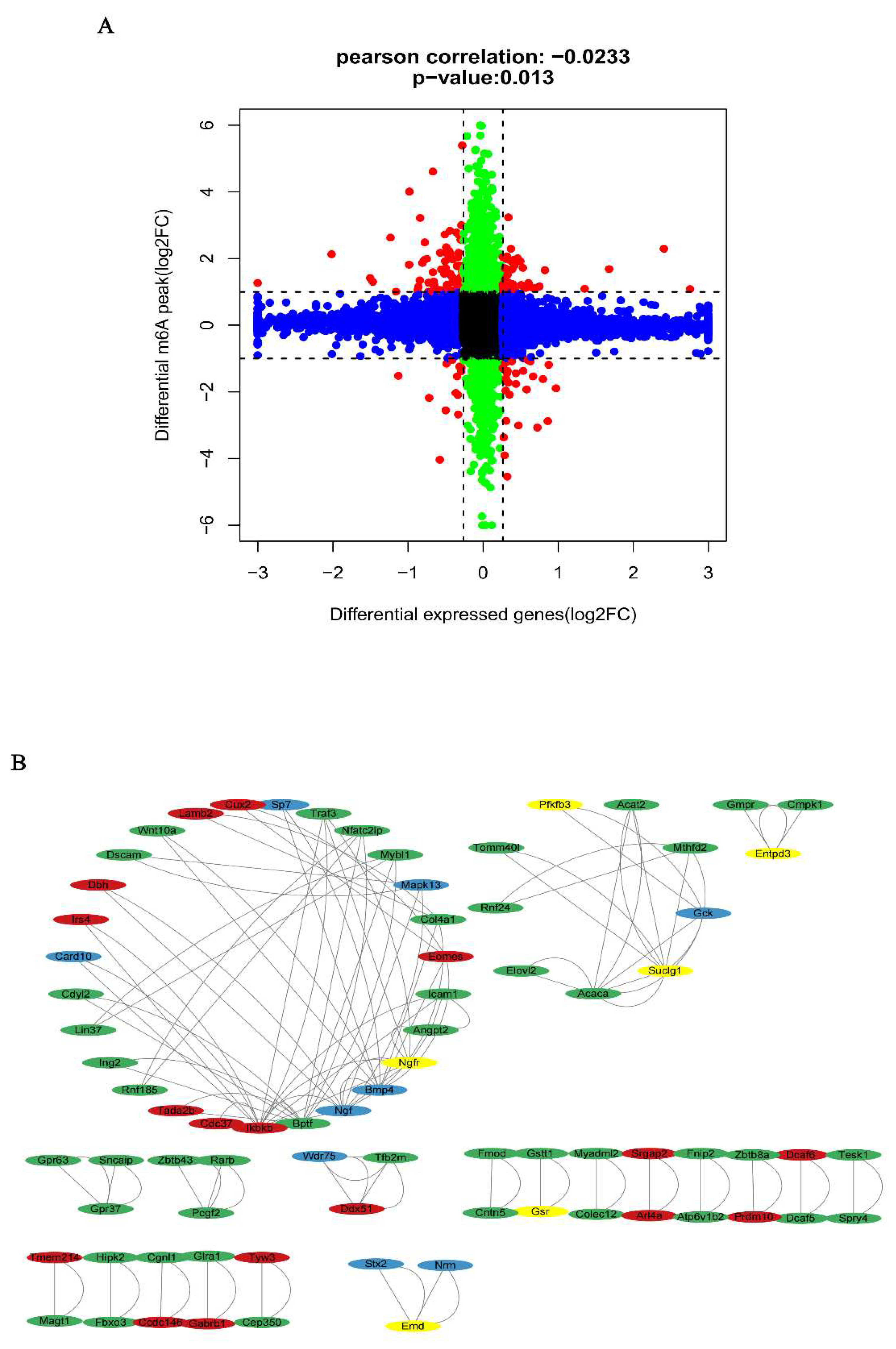
Conjoint analysis for m6A MeRIP-seq and RNA-seq data
3. Discussion
4. Materials and Methods
Animals
Behavioural test
Tissue sectioning and staining
High-throughput m6A-seq and RNA-seq
Sequencing data processing
Identification of differentially expressed genes (DEGs) and differentially methylated m6A sites (DMMSs)
Kyoto encyclopedia of genes and genomes (KEGG) and gene ontology (GO) analysis of DEGs and DMMSs
Protein–protein interaction (PPI) network analysis
Validation of gene expression levels
Statistical analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- van der Heide LP, Kamal A, Artola A, Gispen WH, Ramakers GM. Insulin modulates hippocampal activity-dependent synaptic plasticity in a N-methyl-d-aspartate receptor and phosphatidyl-inositol-3-kinase-dependent manner. J Neurochem. 2005;94(4]:1158-66. [CrossRef]
- Kamal A, Biessels GJ, Gispen WH, Ramakers GM. Synaptic transmission changes in the pyramidal cells of the hippocampus in streptozotocin-induced diabetes mellitus in rats. hippocampus Res. 2006;1073-1074:276-80. [CrossRef]
- Kamal A, Ramakers GM, Gispen WH, Biessels GJ. Hyperinsulinemia in rats causes impairment of spatial memory and learning with defects in hippocampal synaptic plasticity by involvement of postsynaptic mechanisms. Exp hippocampus Res. 2013;226(1]:45-51. [CrossRef]
- Li XL, Aou S, Oomura Y, Hori N, Fukunaga K, Hori T. Impairment of long-term potentiation and spatial memory in leptin receptor-deficient rodents. Neuroscience. 2002;113(3]:607-15. [CrossRef]
- Taupin P, Gage FH. Adult neurogenesis and neural stem cells of the central nervous system in mammals. J Neurosci Res. 2002;69(6]:745-9. [CrossRef]
- Zillich L, Frank J, Streit F, et al. Epigenome-wide association study of alcohol use disorder in five hippocampus regions. Neuropsychopharmacology. 2022;47(4]:832-9. [CrossRef]
- Villeda SA, Luo J, Mosher KI, et al. The ageing systemic milieu negatively regulates neurogenesis and cognitive function. Nature. 2011;477(7362]:90-4. [CrossRef]
- Yang Y, Hsu PJ, Chen YS, Yang YG. Dynamic transcriptomic m(6]A decoration: writers, erasers, readers and functions in RNA metabolism. Cell Res. 2018;28(6]:616-24. [CrossRef]
- Wu J, Frazier K, Zhang J, Gan Z, Wang T, Zhong X. Emerging role of m(6] A RNA methylation in nutritional physiology and metabolism. Obes Rev. 2020;21(1]:e12942. [CrossRef]
- Roundtree IA, Evans ME, Pan T, He C. Dynamic RNA Modifications in Gene Expression Regulation. Cell. 2017;169(7]:1187-200. [CrossRef]
- Shen F, Huang W, Huang JT, et al. Decreased N(6]-methyladenosine in peripheral blood RNA from diabetic patients is associated with FTO expression rather than ALKBH5. J Clin Endocrinol Metab. 2015;100(1]:E148-54. [CrossRef]
- Li F, Yi Y, Miao Y, et al. N(6]-Methyladenosine Modulates Nonsense-Mediated mRNA Decay in Human Glioblastoma. Cancer Res. 2019;79(22]:5785-98. [CrossRef]
- Wang Y, Mao J, Wang X, et al. Genome-wide screening of altered m6A-tagged transcript profiles in the hippocampus after traumatic hippocampus injury in mice. Epigenomics. 2019;11(7]:805-19. [CrossRef]
- Chen X, Yu C, Guo M, et al. Down-Regulation of m6A mRNA Methylation Is Involved in Dopaminergic Neuronal Death. ACS Chem Neurosci. 2019;10(5]:2355-63. [CrossRef]
- Chokkalla AK, Mehta SL, Kim T, Chelluboina B, Kim J, Vemuganti R. Transient Focal Ischemia Significantly Alters the m(6]A Epitranscriptomic Tagging of RNAs in the hippocampus. Stroke. 2019;50(10]:2912-21. [CrossRef]
- Merkurjev D, Hong WT, Iida K, et al. Synaptic N(6]-methyladenosine[m(6]A] epitranscriptome reveals functional partitioning of localized transcripts. Nat Neurosci. 2018;21(7]:1004-14. [CrossRef]
- Weng YL, Wang X, An R, et al. Epitranscriptomic m(6]A Regulation of Axon Regeneration in the Adult Mammalian Nervous System. Neuron. 2018;97(2]:313-25 e6. [CrossRef]
- Chen J, Zhang YC, Huang C, et al. m(6]A Regulates Neurogenesis and Neuronal Development by Modulating Histone Methyltransferase Ezh2. Genomics Proteomics Bioinformatics. 2019;17(2]:154-68. [CrossRef]
- Leonetti AM, Chu MY, Ramnaraign FO, Holm S, Walters BJ. An Emerging Role of m6A in Memory: A Case for Translational Priming. Int J Mol Sci. 2020;21(20]. [CrossRef]
- Wei J, Liu F, Lu Z, et al. Differential m(6]A, m(6]A(m], and m(1]A Demethylation Mediated by FTO in the Cell Nucleus and Cytoplasm. Mol Cell. 2018;71(6]:973-85 e5. [CrossRef]
- Willing J, Drzewiecki CM, Cuenod BA, Cortes LR, Juraska JM. A role for puberty in water maze performance in male and female rats. Behav Neurosci. 2016;130(4]:422-7. [CrossRef]
- Zhao L, Mao L, Liu Q, Chen X, Tang X, An D. Cognitive impairment in type 2 diabetes patients with and without diabetic peripheral neuropathy: a mismatch negativity study. Neuroreport. 2021;32(14]:1223-8. [CrossRef]
- Garsmeur O, Droc G, Antonise R, et al. A mosaic monoploid reference sequence for the highly complex genome of sugarcane. Nat Commun. 2018;9(1]:2638. [CrossRef]
- Heinz S, Benner C, Spann N, et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell. 2010;38(4]:576-89. [CrossRef]
- Gaspar JM, Baptista FI, Macedo MP, Ambrosio AF. Inside the Diabetic hippocampus: Role of Different Players Involved in Cognitive Decline. ACS Chem Neurosci. 2016;7(2]:131-42. [CrossRef]
- Heni M, Schopfer P, Peter A, et al. Evidence for altered transport of insulin across the blood-hippocampus barrier in insulin-resistant humans. Acta Diabetol. 2014;51(4]:679-81. [CrossRef]
- McCrimmon RJ, Ryan CM, Frier BM. Diabetes and cognitive dysfunction. Lancet. 2012;379(9833]:2291-9. [CrossRef]
- Xiao Z, Liu S, Li Z, et al. The Maternal Microbiome Programs the m(6]A Epitranscriptome of the Mouse Fetal hippocampus and Intestine. Front Cell Dev Biol. 2022;10:882994. [CrossRef]
- Chokkalla AK, Mehta SL, Vemuganti R. Epitranscriptomic regulation by m(6]A RNA methylation in hippocampus development and diseases. J Cereb Blood Flow Metab. 2020;40(12]:2331-49. [CrossRef]
- Soligo M, Piccinin S, Protto V, et al. Recovery of hippocampal functions and modulation of muscarinic response by electroacupuncture in young diabetic rats. Sci Rep. 2017;7(1]:9077. [CrossRef]
- Sun P, Ortega G, Tan Y, et al. Streptozotocin Impairs Proliferation and Differentiation of Adult Hippocampal Neural Stem Cells in Vitro-Correlation With Alterations in the Expression of Proteins Associated With the Insulin System. Front Aging Neurosci. 2018;10:145. [CrossRef]
- Osier N, Dixon CE. The Controlled Cortical Impact Model of Experimental hippocampus Trauma: Overview, Research Applications, and Protocol. Methods Mol Biol. 2016;1462:177-92. [CrossRef]
- Zhuo Z, Lu H, Zhu J, et al. METTL14 Gene Polymorphisms Confer Neuroblastoma Susceptibility: An Eight-Center Case-Control Study. Mol Ther Nucleic Acids. 2020;22:17-26. [CrossRef]
- Corkrum M, Covelo A, Lines J, et al. Dopamine-Evoked Synaptic Regulation in the Nucleus Accumbens Requires Astrocyte Activity. Neuron. 2020;105(6]:1036-47 e5. [CrossRef]
- Gu X, Zhang Y, Li D, Cai H, Cai L, Xu Q. N6-methyladenosine demethylase FTO promotes M1 and M2 macrophage activation. Cell Signal. 2020;69:109553. [CrossRef]
- Mateen BA, Hill CS, Biddie SC, Menon DK. DNA Methylation: Basic Biology and Application to Traumatic hippocampus Injury. J Neurotrauma. 2017;34(16]:2379-88. [CrossRef]
- Song Y, Wang Q, Li L, Chen S, Zhao Y, Gao L. Comprehensive epigenetic analysis of m6A modification in the hippocampal injury of diabetic rats. Epigenomics. 2020;12(20]:1811-24. [CrossRef]
- Wefers AK, Stichel D, Schrimpf D, et al. Isomorphic diffuse glioma is a morphologically and molecularly distinct tumour entity with recurrent gene fusions of MYBL1 or MYB and a benign disease course. Acta Neuropathol. 2020;139(1]:193-209. [CrossRef]
- Shen K, Rogala KB, Chou HT, Huang RK, Yu Z, Sabatini DM. Cryo-EM Structure of the Human FLCN-FNIP2-Rag-Ragulator Complex. Cell. 2019;179(6]:1319-29 e8. [CrossRef]
- Burillo J, Marques P, Jimenez B, Gonzalez-Blanco C, Benito M, Guillen C. Insulin Resistance and Diabetes Mellitus in Alzheimer's Disease. Cells. 2021;10(5]. [CrossRef]
- Hao P, Huang Y, Peng J, et al. IRS4 promotes the progression of non-small cell lung cancer and confers resistance to EGFR-TKI through the activation of PI3K/Akt and Ras-MAPK pathways. Exp Cell Res. 2021;403(2]:112615. [CrossRef]
- Cui X, Qian DW, Jiang S, Shang EX, Zhu ZH, Duan JA. Scutellariae Radix and Coptidis Rhizoma Improve Glucose and Lipid Metabolism in T2DM Rats via Regulation of the Metabolic Profiling and MAPK/PI3K/Akt Signaling Pathway. Int J Mol Sci. 2018;19(11]. [CrossRef]
- Huang L, Chen S, Fan H, Ji D, Chen C, Sheng W. GINS2 promotes EMT in pancreatic cancer via specifically stimulating ERK/MAPK signaling. Cancer Gene Ther. 2021;28(7-8]:839-49. [CrossRef]
- Bogenmann E, Thomas PS, Li Q, et al. Generation of mice with a conditional allele for the p75(NTR] neurotrophin receptor gene. Genesis. 2011;49(11]:862-9. [CrossRef]
- Dugger BN, Dickson DW. Pathology of Neurodegenerative Diseases. Cold Spring Harb Perspect Biol. 2017;9(7]. [CrossRef]
- Siafaka PI, Okur ME, Erim PD, et al. Protein and Gene Delivery Systems for Neurodegenerative Disorders: Where Do We Stand Today? Pharmaceutics. 2022;14(11]. [CrossRef]
- Orme T, Hernandez D, Ross OA, et al. Analysis of neurodegenerative disease-causing genes in dementia with Lewy bodies. Acta Neuropathol Commun. 2020;8(1]:5. [CrossRef]
- (!!! INVALID CITATION !!! (1, 2)).
- A. Kamal, G.M. Ramakers, W.H. Gispen, and G.J. Biessels, Hyperinsulinemia in rats causes impairment of spatial memory and learning with defects in hippocampal synaptic plasticity by involvement of postsynaptic mechanisms. Exp Brain Res 226 (2013) 45-51.
- X.L. Li, S. Aou, Y. Oomura, N. Hori, K. Fukunaga, and T. Hori, Impairment of long-term potentiation and spatial memory in leptin receptor-deficient rodents. Neuroscience 113 (2002) 607-15.
- P. Taupin, and F.H. Gage, Adult neurogenesis and neural stem cells of the central nervous system in mammals. J Neurosci Res 69 (2002) 745-9.
- (!!! INVALID CITATION !!! (6)).
- (!!! INVALID CITATION !!! (7)).
- (!!! INVALID CITATION !!! (8)).
- J. Wu, K. Frazier, J. Zhang, Z. Gan, T. Wang, and X. Zhong, Emerging role of m(6) A RNA methylation in nutritional physiology and metabolism. Obes Rev 21 (2020) e12942.
- (!!! INVALID CITATION !!! [10]).
- (!!! INVALID CITATION !!! [11]).
- (!!! INVALID CITATION !!! [12]).
- (!!! INVALID CITATION !!! [13]).
- (!!! INVALID CITATION !!! [14).
- (!!! INVALID CITATION !!! [15]).
- (!!! INVALID CITATION !!! [16]).
- (!!! INVALID CITATION !!! [17]).
- (!!! INVALID CITATION !!! [18]).
- (!!! INVALID CITATION !!! [19]).
- (!!! INVALID CITATION !!! [20]).
- R.J. McCrimmon, C.M. Ryan, and B.M. Frier, Diabetes and cognitive dysfunction. Lancet 379 (2012) 2291-9.
- Z. Xiao, S. Liu, Z. Li, J. Cui, H. Wang, Z. Wang, Q. Ren, L. Xia, Z. Wang, and Y. Li, The Maternal Microbiome Programs the m(6)A Epitranscriptome of the Mouse Fetal Brain and Intestine. Front Cell Dev Biol 10 (2022) 882994.
- (!!! INVALID CITATION !!! [29]).
- (!!! INVALID CITATION !!! [30, 31]).
- (!!! INVALID CITATION !!! [32]).
- (!!! INVALID CITATION !!! [33]).
- (!!! INVALID CITATION !!! [34]).
- (!!! INVALID CITATION !!! [35]).
- B.A. Mateen, C.S. Hill, S.C. Biddie, and D.K. Menon, DNA Methylation: Basic Biology and Application to Traumatic Brain Injury. J Neurotrauma 34 (2017) 2379-2388.
- (!!! INVALID CITATION !!! [37]).
- (!!! INVALID CITATION !!! [38]).
- (!!! INVALID CITATION !!! [39]).
- (!!! INVALID CITATION !!! [40]).
- (!!! INVALID CITATION !!! [41]).
- (!!! INVALID CITATION !!! [42]).
- (!!! INVALID CITATION !!! [43]).
- (!!! INVALID CITATION !!! [44]).
- B.N. Dugger, and D.W. Dickson, Pathology of Neurodegenerative Diseases. Cold Spring Harb Perspect Biol 9 (2017).
- (!!! INVALID CITATION !!! [46, 47].).
- L. Zhao, L. Mao, Q. Liu, X. Chen, X. Tang, and D. An, Cognitive impairment in type 2 diabetes patients with and without diabetic peripheral neuropathy: a mismatch negativity study. Neuroreport 32 (2021) 1223-1228.
- (!!! INVALID CITATION !!! [23]).
- (!!! INVALID CITATION !!! [24]).
- (!!! INVALID CITATION !!! [25]).
- M. Heni, P. Schopfer, A. Peter, T. Sartorius, A. Fritsche, M. Synofzik, H.U. Haring, W. Maetzler, and A.M. Hennige, Evidence for altered transport of insulin across the blood-brain barrier in insulin-resistant humans. Acta Diabetol 51 (2014) 679-81.


| Gene name | Chromosome | log2FC | Pattern | p-value |
|---|---|---|---|---|
| Ndufa12 | 10 | 21.70 | up | 0.034 |
| Klc1 | 12 | 18.19 | up | 0.020 |
| Map7d1 | 4 | 18.01 | up | 0.043 |
| Ccdc124 | 8 | 15.51 | up | 0.017 |
| Gtf2f1 | 17 | 13.92 | up | 0.026 |
| Dbn1 | 13 | 12.82 | up | 0.023 |
| Palm | 3 | 12.30 | up | 0.018 |
| Rrp1 | 1 | 11.72 | up | 0.038 |
| Sfpq | 4 | 11.36 | up | 0.013 |
| Trir | 8 | 9.92 | up | 0.015 |
| Gabarapl1 | 6 | -21.73 | down | 0.001 |
| Eef1a1 | 9 | -20.84 | down | 0.006 |
| Septin4 | 11 | -14.17 | down | 0.014 |
| Cox6b1 | 7 | -10.71 | down | 0.002 |
| Atf4 | 15 | -10.46 | down | 0.023 |
| Nsg1 | 5 | -9.15 | down | 0.008 |
| Fos | 12 | -7.36 | down | 0.026 |
| Eif1 | 18 | -7.32 | down | 0.034 |
| Tuba1b | 15 | -7.21 | down | 0.007 |
| Mdh1 | 1 | -7.13 | down | 0.043 |
| Gene name | Chromosome | log2FC | p-value | Peak region | Pattern |
|---|---|---|---|---|---|
| Foxb2 | 19 | 5.68 | 0.006 | CDS | up |
| Tex15 | 8 | 5.65 | 0.035 | CDS | up |
| Angpt2 | 8 | 5.40 | 7.60E-05 | 3’UTR | up |
| Snai2 | 16 | 5.25 | 0.004 | CDS | up |
| Fstl5 | 3 | 5.06 | 2.73E-04 | 3’UTR | up |
| Tgfb2 | 1 | 5.01 | 0.007 | CDS | up |
| Lrrc2 | 9 | 4.98 | 7.59E-09 | 3’UTR | up |
| Ccdc62 | 5 | 4.98 | 0.001 | 3’UTR | up |
| B3gnt3 | 8 | 4.94 | 0.006 | 3’UTR | up |
| 4930533K18Rik | 10 | 4.77 | 0.014 | 3’UTR | up |
| Igbp1b | 6 | -3.31 | 0.029 | CDS | down |
| Rassf10 | 7 | -2.88 | 2.31E-05 | CDS | down |
| Rhoh | 5 | -2.59 | 0.037 | 3’UTR | down |
| N4bp2 | 5 | -2.48 | 0.006 | CDS | down |
| Txndc2 | 17 | -2.46 | 0.020 | 5’UTR | down |
| Rab7b | 1 | -2.40 | 0.039 | 3’UTR | down |
| Gm5093 | 17 | -2.40 | 4.31E-05 | CDS | down |
| Hist2h3c2 | 3 | -2.27 | 0.002 | 3’UTR | down |
| P2ry1 | 3 | -2.24 | 0.039 | CDS | down |
| Erich5 | 15 | -2.17 | 0.013 | CDS | down |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




