1. Introduction
Deoxyuridine triphosphate nucleotidohydrolases (dUTPases or Duts) are produced by most living organisms as part of their pyrimidine biosynthesis networks [
1]. Their role is to eliminate excess dUTP, which can be mistakenly incorporated into DNA by DNA polymerases since most of them cannot distinguish between dUTP and dTTP. In addition, dUTPases provide the dTTP precursor – dUMP [
2]. The indispensability of dUTPase genes has been demonstrated in various organisms, including bacteria [
3,
4], yeast [
5] and mice [
6]. Moreover, many viruses and phages also possess the genes for encoding dUTPases.
Based on their oligomerization state, dUTPases can be classified into three families: monomeric [
7], homodimeric [
8], and homotrimeric [
9] enzymes. The homotrimeric family is the largest and includes dUTPases found in various organisms such as plants, animals, fungi, bacteria, and certain viruses like adenoviruses, poxviruses, and retroviruses [
10].
It has been described in the publications that dUTPases can have extra functions in addition to their dUTPase activity. For instance, trimeric dUTPases encoded by staphylococcal phages have been shown to induce the transfer of staphylococcal pathogenicity islands (SaPI) by interacting with the SaPI-encoded Stl repressor [
11,
12,
13]. This mechanism relies on the presence of dUTP and is reminiscent of the nucleotide-dependent signaling mechanism seen in eukaryotic G proteins. Interestingly, this function is not related to the dUTPase enzymatic activity itself but is determined by an extended segment (loop) of the polypeptide chain up to 40 amino acids in length in the protein structure. In contrast, the
Staphylococcus epidermidis phage PH15 dUTPase, which lacks this loop, is unable to induce SaPI transfer. Another example is the dUTPase of
Mycobacterium smegmatis, where the deletion of a mycobacteria-specific loop near the C-terminal region does not significantly affect its dUTPase enzymatic properties but proves to be lethal for the cell [
4]. This suggests the existence of an essential additional function associated with that loop.
Our study focuses on investigating the involvement of the homotrimeric dUTPase, which is synthesized during bacteriophage T5 infection of
Escherichia coli, in the process of phage development. In the study by Warner H.B.
et al. [
14], it was shown that during T5 phage infection of
E.coli cells, the overall level of dUTPase activity increases. As a consequence, they obtained a viable mutant phage through random mutagenesis in which this activity was not observed during its development in host cells. This is interesting because viral genomes typically follow the pattern of occupying as little space as possible, suggesting that each gene serves a purpose and is often vital for efficient phage development. This raises the question of why does the T5 phage possesses its own dUTPase if its primary function is not essential for its life cycle. We aim to uncover the additional function performed by T5 dUTPase during phage development that has allowed it to be preserved throughout evolution, and determine which structural element is responsible for this function.
Here we conducted a comparison of the dUTPases from T5 phage, T5-like phages, and the host enzyme dUTPase of
E.coli (
Figure S1). Although there is limited sequence similarity between the T5 phage dUTPase (T5Dut) and the
E.coli dUTPase, the comparison of their spatial structures revealed a high degree of similarity, except for the presence of a specific loop in the T5 and T5-like phages that is absent in the bacterial protein. We presume that this role is facilitated by the structural element unique to T5 and T5-like phages, which enables the phage dUTPase to carry out its additional function that is not compensated by the presence of the
E.coli dUTPase.
2. Results
2.1. Extra Function of T5 Dut in Addition to Their dUTPase Activity
As previously mentioned, there is the evidence that dUTPase of T5-like phages, besides its primary role in DNA repair, may also play an additional role in the development of phages within host cells [
11,
12,
13,
15]. To investigate this fact widely, a mutant T5 phage with an amber mutation in the
dut gene was generated using site-directed mutagenesis. Subsequently, phage progeny derived from crossing were analyzed, and several plaques exhibiting phenotypic differences (smaller phage colonies were formed) when grown on the
E.coli XAC (Su0) nonsuppressing strain were selected.
Dut gene sequencing of these phages confirmed the presence of amber mutations, and they were designated as T5
dut-am. Furthermore, it was observed that the amber mutation in the T5
dut-am phage genome could be effectively complemented by the wild-type
dut gene nucleotide sequence when the mutant phage was plated on the
E.coli XAC strain containing the pBADex1/T5dut plasmid. Conversely, no complementation effect was observed when the mutant (T5
dut-am) phage was plated on the
E.coli XAC strain containing the pBADex1/Ecdut plasmid consisted the
E.coli dUTPase gene. These findings strongly support the importance of this enzyme in the development of T5 bacteriophage within
E.coli cells and suggest the existence of an additional function inherent to phage dUTPase.
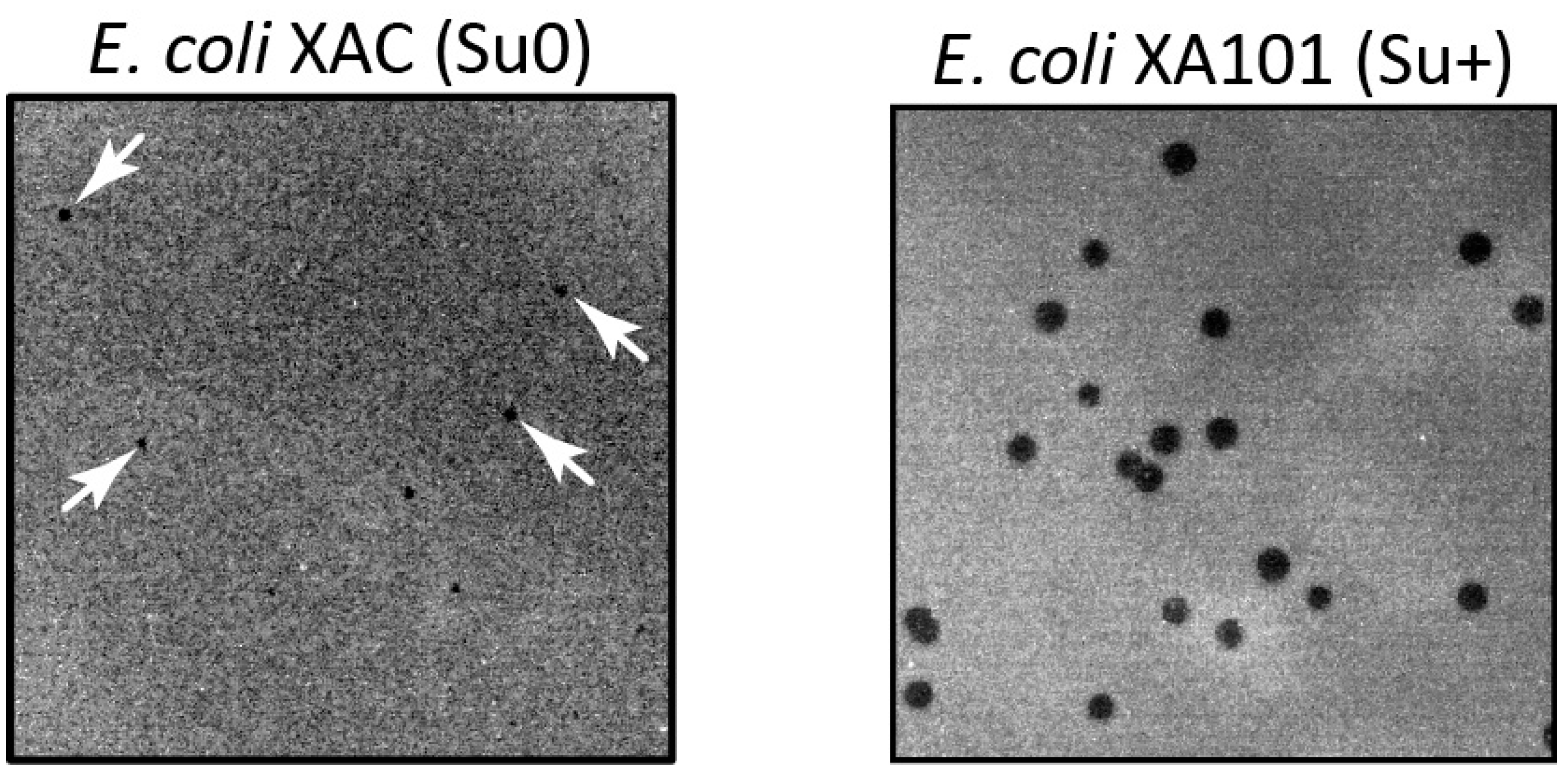
It is worth noting that the manifestation of the
dut gene mutation in T5 phage has some peculiarities. When the mutant T5 phage is plated on the
E.coli XAC (Su0) strain, very small and slightly visible colonies are formed (
Figure 1), and their quantity is approximately three times less compared to plating on the
E.coli XA101 (Su+) suppressing strain (
Table 1). Additionally, experiments aimed at determining the burst size of the phage have demonstrated that this specific mutation results in a nearly tenfold decrease in phage yield.
2.2. The Additional Structural Element of the T5 Phage dUTPase Is Responsible for Carrying out the Novel Function
In the previously observed cases of double dUTPase function, the specific structural elements associated with the supplemental action were identified [
12,
13]. The comparative analysis of dUTPase sequences from bacteriophage T5 and the host organism
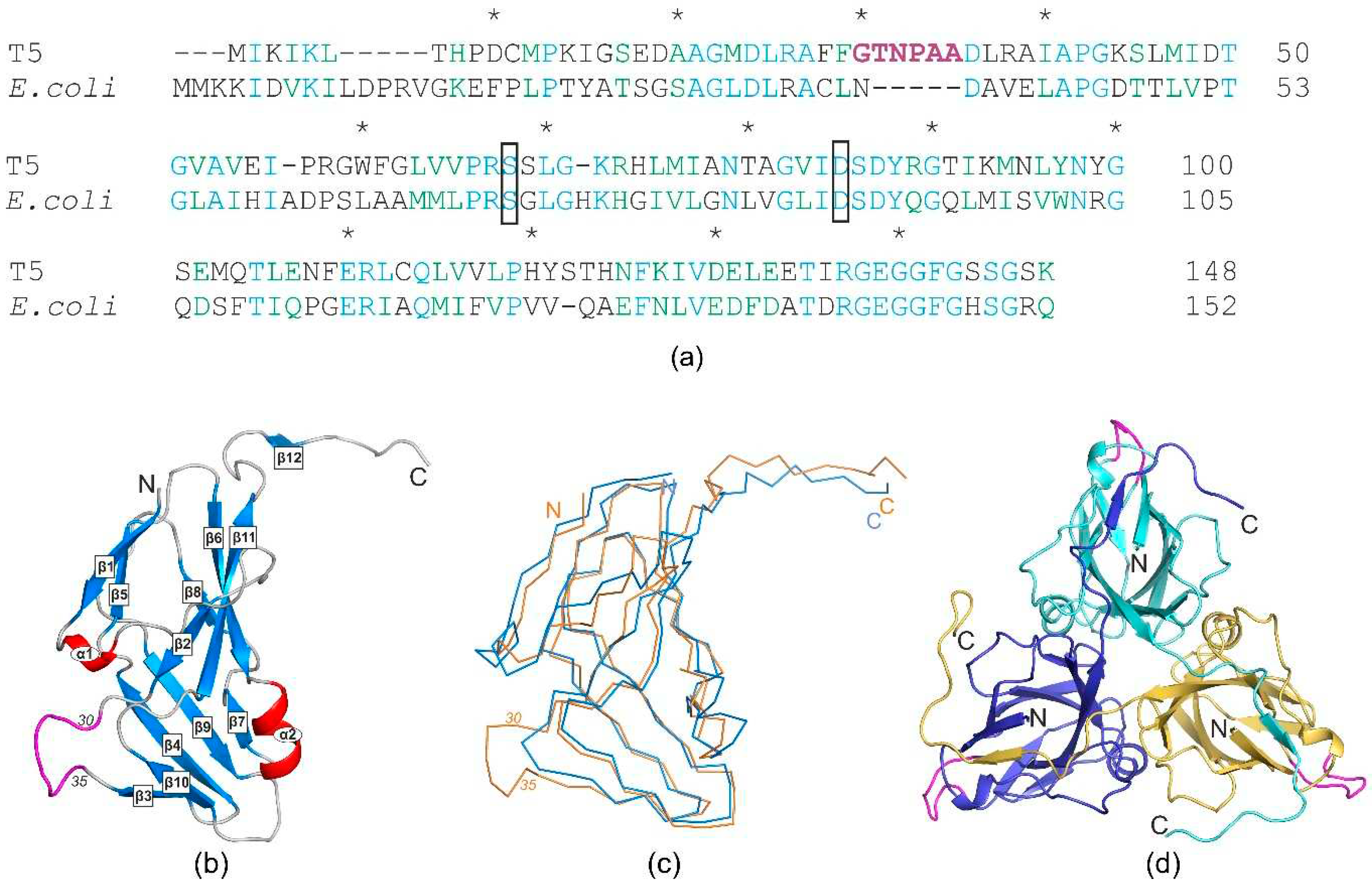
E.coli revealed a low identity of only 34% (
Figure 2a). Additionally, both gaps and insertions were observed, making it difficult to confidently identify the presumed regions responsible for performing the additional function. To assess the significance of these differences in the spatial structures of the given proteins, the crystal structure of T5Dut was obtained at a resolution of 2.0 Å, pdb id 8QKY (
Table 2). Despite the low identity, the overall tertiary and quaternary dUTPase structures of phage and host were found to be very similar, with root mean square deviation (RMSD) of 1.3 Å when aligning the Cα atoms of the monomers. The structure shows the typical trimeric dUTPase antiparallel β
-pleated fold of the subunits (
Figure 2b). Three subunits are attached to each other in the same manner as in other trimeric dUTPases (
Figure 2d), forming three active sites per enzyme molecule in the region of monomer interaction.
At the same time, the analysis of the amino acid sequence and the obtained structure allowed for the identification of an additional loop, a short polypeptide segment six amino acid residues long (residues 30-35 in pdb 8QKY) located on the surface of each subunit of the trimer, between β-strands β2 and β3 (
Figure 2), which is presumably responsible for the new function of T5 Dut. To test our suggestion, the next mutant T5 phage T5
dut(Δloop) with a dUTPase sequence 28-FFGTNPAADL-37 reduced to 28-FFNDL-32, was obtained. The mutant phage T5
dut(Δloop), did not differ from the previously obtained mutant T5
dut-am in terms of main growth characteristics. To confirm that the loop reduction remains overall folding without global changes we obtained the mutant form T5DutΔloop structure (2.1 Å) (
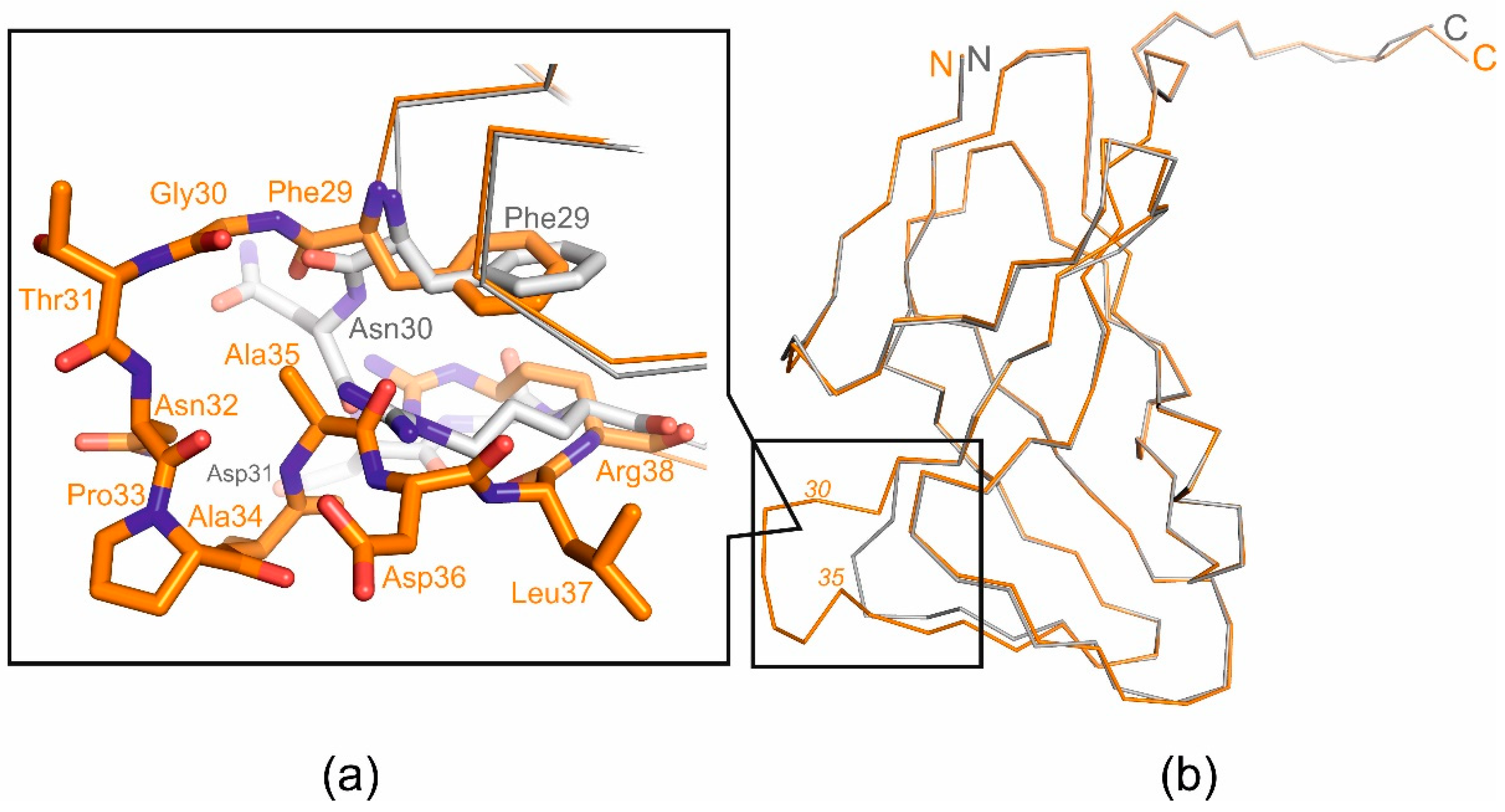
Table 1) Comparison of the crystal structures of wild-type T5 Dut and its mutant form revealed no significant changes in the overall folding of the protein, except for the mutated region. Superimposition of the wild-type and mutant forms, with a RMSD of 0.31 Å for the Cα atoms shows a nearly identical folding of these proteins, with the only difference observed in the region of interest (
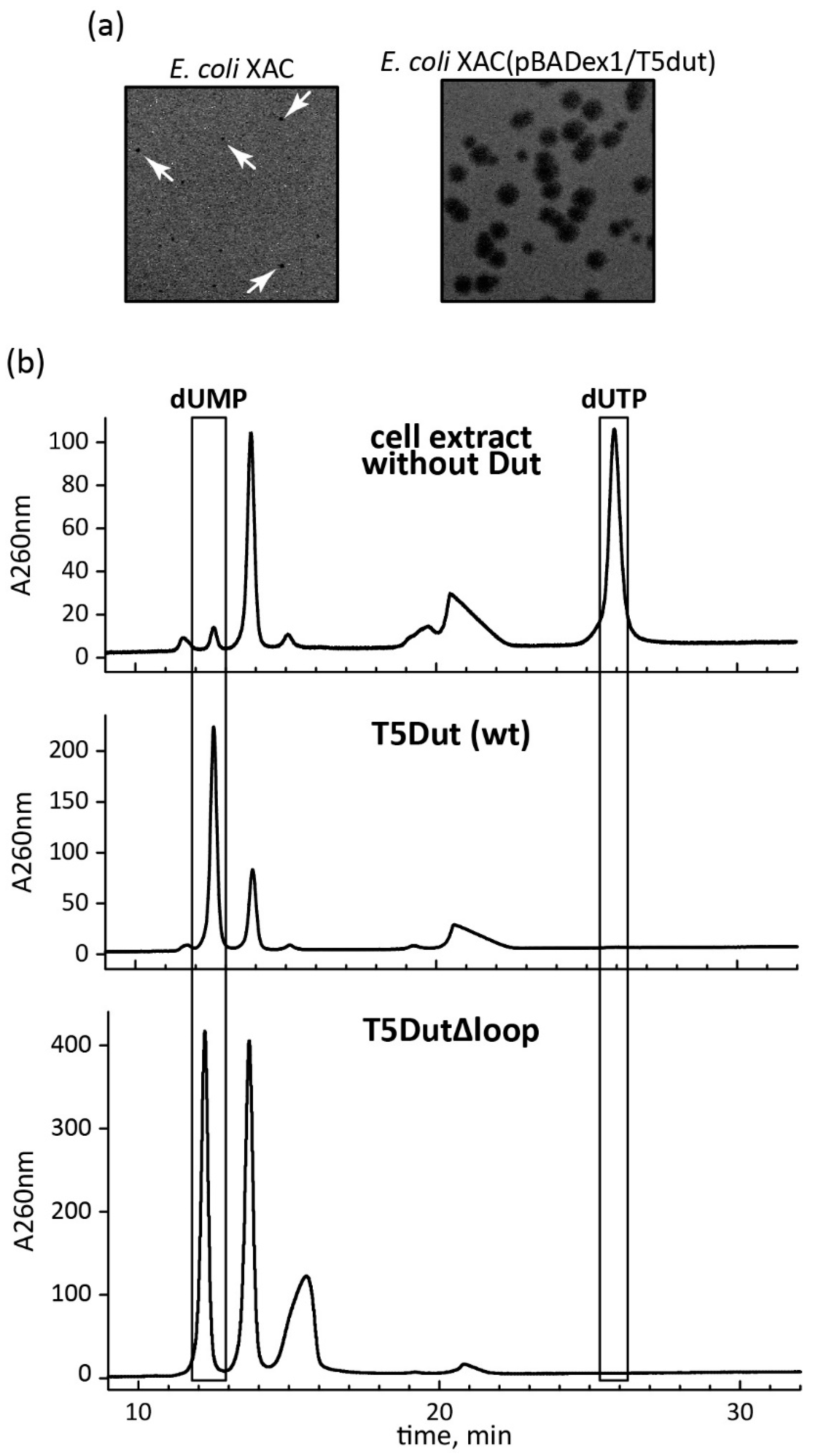
Figure 3). This change in the amino acid sequence of T5 phage Dut also had no impact on its enzymatic activity, specifically the hydrolysis of dUTP, as shown in the curve (
Figure 4). The results of these experiments clearly demonstrate that the short segment of the polypeptide chain, consisting of 6 amino acid residues (30-GTNPAA-35), is responsible for carrying out an additional function of T5Dut.
2.3. The Canonical Function of T5 Phage dUTPase Is Not Crucial for Performing the Additional Function
In the study of the mutant T5 phage lacking phage-specific dUTPase activity during development in
E.coli cells, no mapping or description of the mutation had unfortunately been performed [
14].
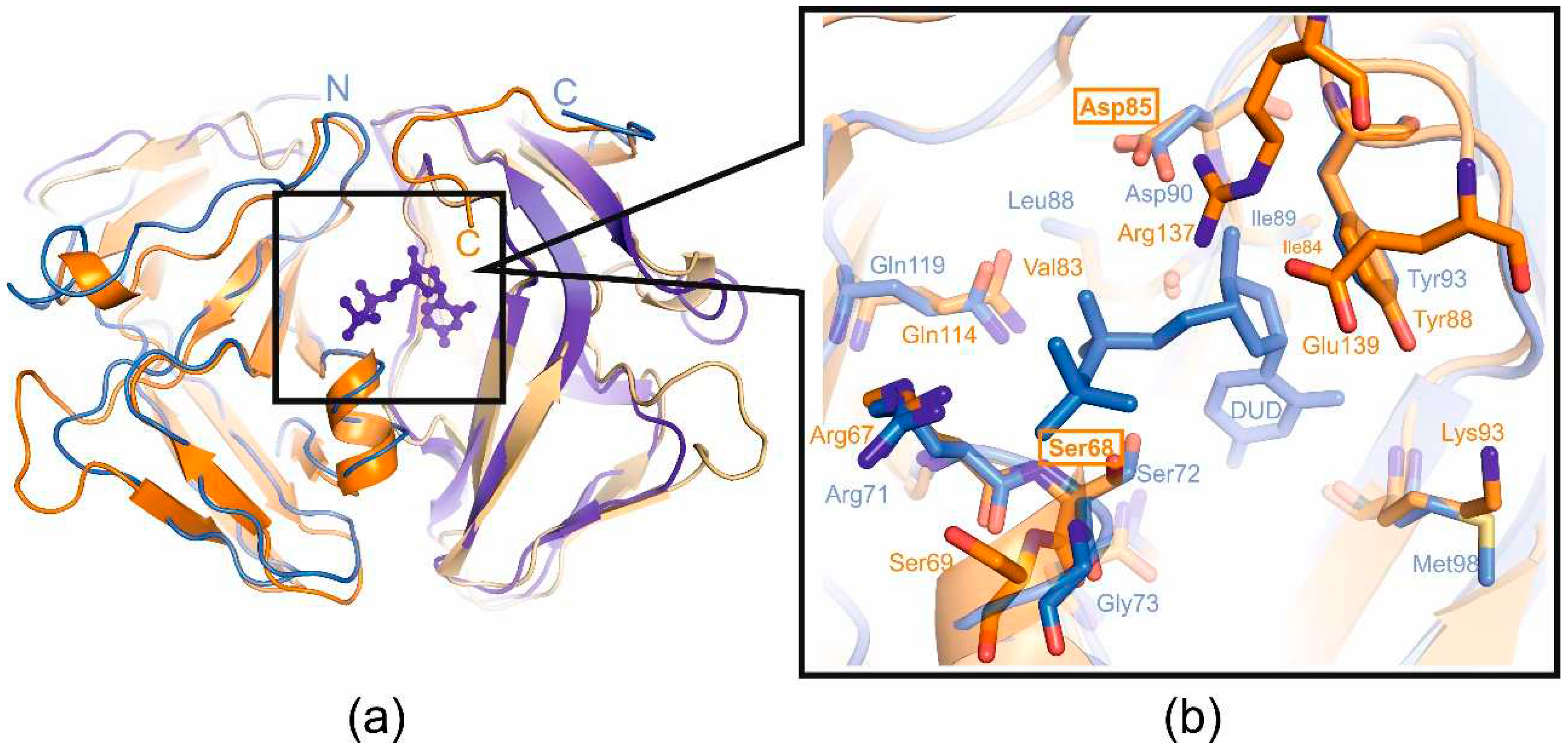
In order to answer the question of whether enzymatic activity plays a role in the additional function of T5 phage dUTPase, the mutant forms of T5Dut that do not possess enzymatic activity were obtained. Comparison of the amino acid sequence and structure of T5Dut phage with other characterized enzymes allowed us to identify two conserved amino acid residues (Ser68 and Asp85) (
Figure 2a) within the active site that are directly involved in the enzymatic reaction in T5 Dut (
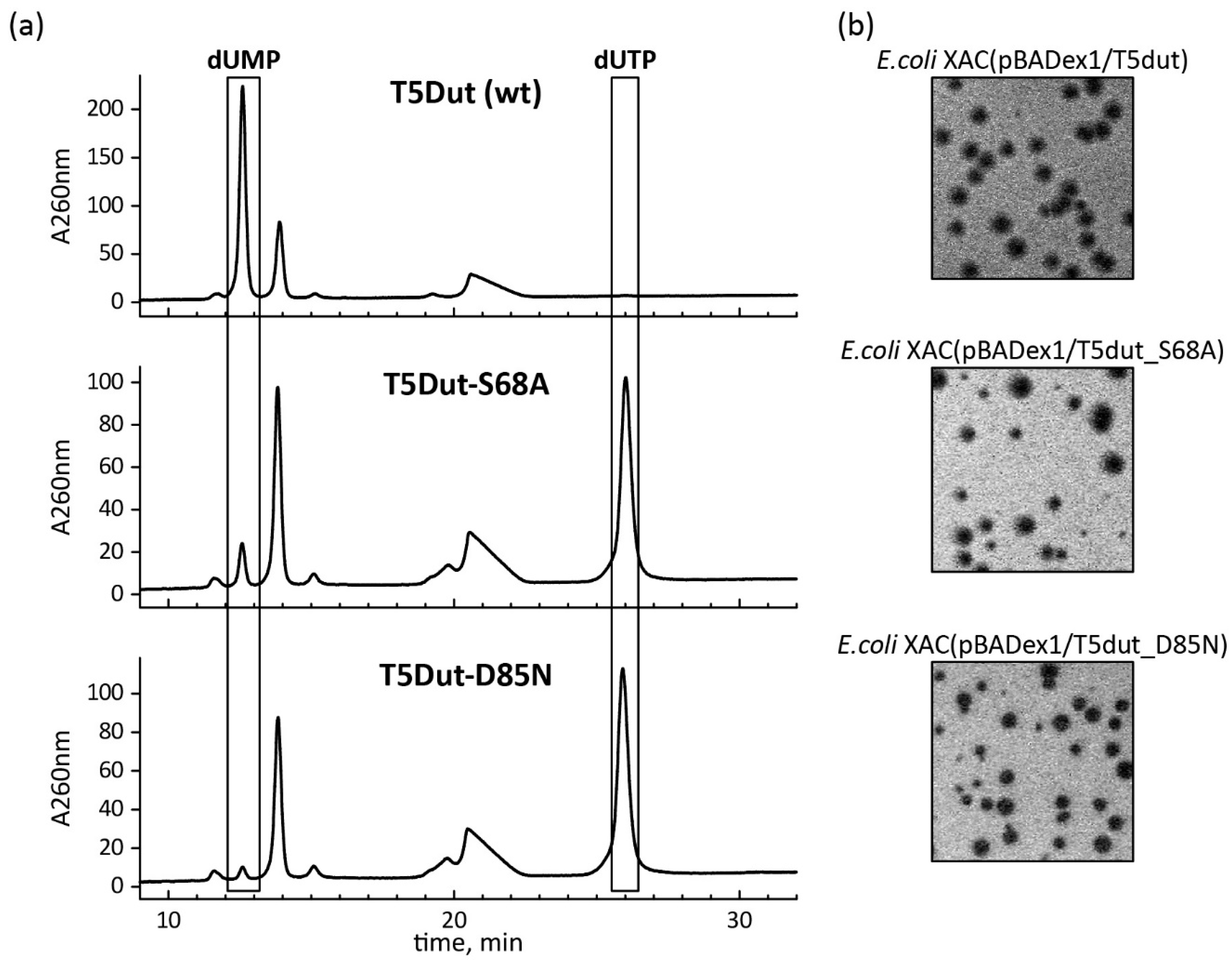
Figure 5). To terminate dUTPase activity the plasmids encoding mutant forms of T5 phage dUTPase containing single amino acid substitutions, T5Dut-S68A and T5Dut-D85N, were obtained. These mutations resulted in the complete loss of the enzymatic activity, however no impact on the execution of the additional function of phage dUTPase in the virus's development was noted (
Figure 6). Based on the results, we demonstrate that the enzymatic activity of the phage enzyme itself does not have a vital impact on the development of the phage. This indicates that the enzymatic activity and the newly detected function, combined within a single enzyme, are independent of each other.
3. Discussion
dUTPases participate in the regulation of the balance between dUTP and dTTP in cells and, as a result, have a significant impact on the DNA repair mechanism in living organisms. Despite the fact that mutant organisms encoding inactive enzymes remain viable, a complete knockout of the
dut gene have a lethal consequences [
3,
4]. Thus, it is believed that enzymes of this class performs an additional function in the cell's life processes that is independent of their enzymatic activity.
In this study, we demonstrate for the first time that the homotrimeric dUTPase of bacteriophage T5 performs an additional, non-enzymatic function in the virus's life cycle. Thus, the Dut protein of phage T5 could belong to the narrow group of dUTPases that have been confirmed to possess dual function. For example, the involvement of dUTPase from several staphylococcal bacteriophages (ϕ11, 80α, ϕNM1, etc.) in the process of derepression of pathogenicity islands (SaPI) during the infection of
Staphylococcus aureus cells by the phage has been demonstrated [
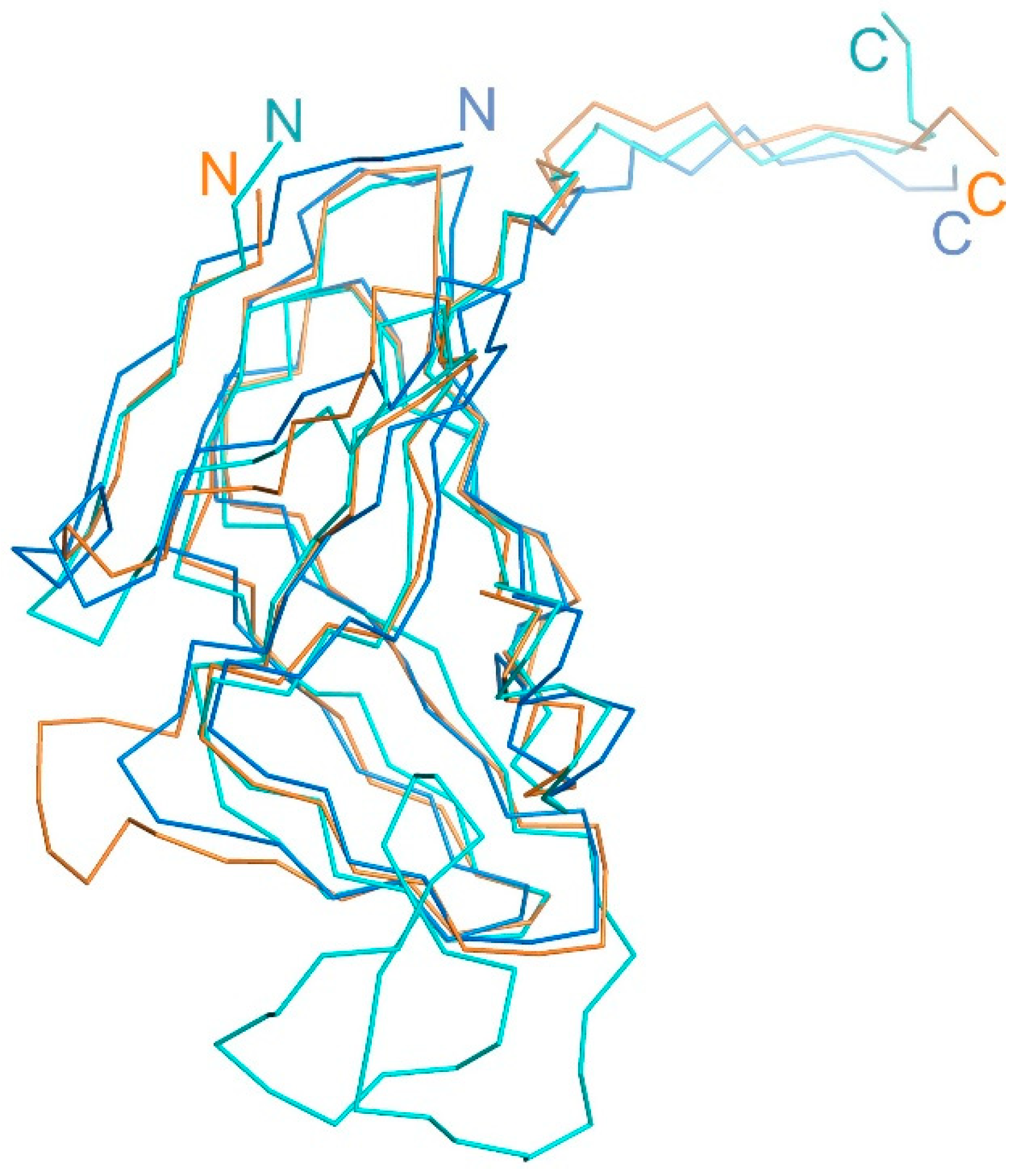
16]. The additional structural element on the surface of the trimeric (ϕ11, 80α) or dimeric (ϕNM1) forms of staphylococcal phage Dut plays a key role in carrying out this function. This element is formed by an extended polypeptide chain of up to 40 amino acid residues in length (
Figure 7).
Figure 7 shows the dUTPase of
M. smegmatis, which has a short specific loop in the C-terminus contributing to an additional protein function. Our experimental results clearly demonstrate that in the case of T5 Dut, the presence of an additional function is also attributed to the presence of an extra element in the enzyme structure (a short loop consisting of only 6 amino acids (30-GTNPAA-35)). While its reduction does not impact the overall spatial organization of the enzyme or its ability to cleave the substrate (dUTP), it has severe consequences for the development of the T5 bacteriophage itself. The fact, that the additional function of T5 Dut is likely realized at some stage of the phage's development, significantly distinguishes it from the dUTPases of bacteria and staphylococcal phages. However, similar to the mentioned examples of dUTPases in staphylococcal phages,
E.coli,
M. smegmatis, and potentially other organisms, the additional function of T5 Dut is independent of the enzymatic activity of the protein.
The details of the process in which this additional function of T5 Dut phage is realized are yet to be uncovered. However, based on the results of our additional experiments (under publication in a separate paper), it becomes evident that this enzyme plays a direct role in the phage particle morphogenesis process, and the new function is specifically realized at this stage of virus development in
E.coli cells. The fact that the Dut gene of T5 bacteriophage is necessary for the late stages of viral development partially explains the phenotypic manifestation of the mutation in the
dut gene, which results in an approximately 10-fold decrease in phage yield. It is possible that the phage has some "bypass pathways" to complete the formation of viral particles, despite such significant losses in yield. At the same time, we cannot dismiss the possibility that the absence of dUTPase in certain T5-like phages (
Figure S1) may lead to more dramatic consequences, despite their amino acid sequences being practically identical.
4. Materials and Methods
4.1. Construction of Plasmid
DNA cloning was performed using standard methods and techniques [
17]. Details of plasmid construction are described in the
Supplementary data.
The plasmid pZ/T5dut-am contained a fragment of the D14 and dut gene region of the T5 phage with an amber mutation in the dut gene. The plasmid pZ/T5dutΔloop contained a fragment of the D14 and dut gene region of the T5 phage with short deletions in the dut gene (88-93 and 96-104 bp). These plasmids were used to generate the corresponding mutants of the T5 phage.
The plasmid pBADex1/T5dut contains the wild-type dut gene under the control of an arabinose promoter. The plasmid pBADex1/T5dutΔloop encodes a mutant form of the T5 phage Dut, in which the amino acid sequence 28-FFGTNPAADL-37 is replaced with 28-FFNDL-32. Plasmids pBADex1/T5dut_S68A and pBADex1/T5dut_D85N encode mutant forms of the T5 phage dUTPase with the corresponding amino acid substitutions that result in loss of the enzymatic activity. These described plasmids were used for protein production and complementation tests.
4.2. Bacteriophages and Bacterial Strains
Strain E.coli TOP10 (F-, mcrA, Δ(mrr-hsdRMS-mcrBC), ϕ80lacZΔM15, ΔlacX74, deoR, recA1, araD139, Δ(ara-leu)7697, galU, galK, rpsL (strR), endA1, nupG) was used as the host for plasmids construction and propagation. Strain E.coli XA101 (ara, Δ(lac-pro), supD, gyrA, metB, argE-am, rpoB, thi) was used to cross of phage T5 and grow its amber mutant, T5dut-am. The non-permissive E.coli XAC strain (ara, Δ(lac-pro), gyrA, argE-am, rpoB, thi) was employed for the selection of T5 phage mutants carrying an amber mutation in the dut gene. The expression of both the wild-type T5 phage dut gene and its mutant variants was conducted in the E.coli XAC strain.
Bacteriophage T5
dut-am, carrying an amber mutation in the
dut gene, was generated through the recombination of phage T5 with the pZ/T5dut-am plasmid, as described in the
Supplementary data. Bacteriophage T5
dut(Δloop) encodes a mutant form of dUTPase, in which five amino acid residues (sequence 28-FFGTNPAADL-37) have been deleted and replaced with 28-FFNDL-32. It was obtained through the recombination of phage T5 with the pZ/T5dutΔloop plasmid.
4.3. Expression and Purification of Proteins
E.coli XAC strain transformed with the respective plasmids was used for the expression of the wild-type dut gene and its mutant form, dutΔloop. The cells were grown in LB medium supplemented with Amp (100 µg/mL) at +37°C until reaching an OD600 of 0.5. Gene expression was induced by adding L-(+)-Arabinose (Sigma-Aldrich, Taufkirchen, Germany) to a final concentration of 0.2% (w/v), and the cells were further incubated at +37°C for 4 hours. The cells were pelleted by centrifugation at 6,000g for 20 minutes. The cell pellet was then resuspended in lysis buffer (20 mM Tris HCl pH 8.0, 50 mM NaCl) and disrupted using a Gaulin Homogeniser (APV Homogeniser GmbH). The insoluble fraction was separated by centrifugation at 30,000g for 30 minutes. The soluble fraction was used for the isolation of the T5Dut and mutant form T5DutΔloop.
Cell extract was supplemented with streptamycin sulfate to a concentration of 2% (w/v), incubated for 1 hour at +4°C and centrifuged for 30 min at 20,000g. The supernatant was diluted 10 folds with buffer A (20 mM Tris HCl pH 8.5, 1 mM MgCl2), filtered through a 0.22 μm filter (Millipore, Billerica, MA, USA) and applied to a Mono Q 10/100 GL column (GE Healthcare, Chicago, IL, USA) equilibrated with buffer A. Elution was carried out at a flow rate 2 mL/min with a linear gradient of 0-0.3 M NaCl in buffer A for 50 min. Fractions containing T5Dut (T5DutΔloop) according to SDS PAGE were combined, diluted 3 folds with buffer B (20mM Na-Acetate, pH 5.0), supplemented with glacial acetic acid to a concentration of 6 mM and applied to a Mono S 5/50 GL column (GE Healthcare, Chicago, IL, USA) in buffer B. Elution carried out at a speed of 0.6 mL/min with a NaCl concentration gradient in buffer B: from 0 to 100 mM in 5 minutes and from 100 to 300 mM in 45 minutes. Fractions containing T5Dut (T5DutΔloop) according to SDS PAGE were pooled, concentrated by Amicon centrifugal concentrator MWCO 10 kDa (Millipore Ireland Ltd., Tullagreen, County Cork Carrigtwohill, Ireland) and applied to a Superdex 200 10/300 GL (GE Healthcare, Chicago, IL, USA) gel filtration column in buffer C (20mM Tris-HCl, pH 8.0, 150mM NaCl).
4.4. dUTPase Activity Test
The expression of the wild-type T5 dut gene and its mutant forms (dut-S68A, dut-D85N, and dut(Δloop) was conducted as mentioned above. A 0.5 mL suspension of cells with an OD600 of 0.5 was sonicated, and the soluble fraction was used to assess enzymatic activity. The E.coli XAC strain transformed with the pBADex1 plasmid was used as a negative control.
60 μL of buffer: 20 mM Tris-HCl, pH 8.0, 100 mM NaCl, 5 mM MgCl2 containing 200 mkM dUTP was supplemented with 0.25 μL of cell extract. The mixture was incubated for 30 min at +35°C, and next 180 μL of 20 mM ammonium acetate, pH 8.0, 10 mM EDTA was added, A 200 μL aliquot of the sample was immediately applied to a Mono Q 5/50 GL column (GE Healthcare, Chicago, IL, USA) equilibrated with 20 mM ammonium acetate, pH 8.0 buffer. Elution was carried out at flow rate was 0.6 mL/min for 30 minutes by linearly increasing the ammonium acetate concentration up to 1.2 M
Registration was done by absorption at 260 nm.
4.5. Crystallization and Crystallography
Screening for the initial crystallization conditions was performed using sitting-drop vapor diffusion method with commercial screen sets (Hampton Research, Aliso Viejo, CA, USA). 0.2 µL of protein (10-15 mg/mL) was combined with 0.2 µL of reservoir solution and incubated at +23°C. Crystals of T5Dut were obtained by drop equilibrating against 0.075 M Tris-HCl, pH 8.5, 1.5 M ammonium sulfate, 25% (v/v) glycerol (A4 from Crystal Screen Cryo, Hampton Research, Aliso Viejo, CA, USA), appeared overnight and reached their full size within 3 days. T5DutΔloop crystals were obtained with 0.085 M Hepes sodium, pH 7.5, 1.7% (v/v) polyethylene glycol 400, 1.7 M ammonium sulfate, 15% (v/v) glycerol (D3 Crystal Screen Cryo, Hampton Research, Aliso Viejo, CA, USA) in a same manner. Crystals were harvested and flash-frozen.
Diffraction data from T5Dut crystal was collected on beamline P11 at PETRA III, DESY, Hamburg, Germany. Reflection data set were processed using XDS [
18]. Diffraction data of T5DutΔloop were collected using a home source Rigaku XtaLAB Synergy-S laboratory system (The Woodlands, TX, USA) [
19]. Reflection data were processed and merged using CrysAlis software 42.89a [
20]. The structures were determined by molecular replacement with Phaser [
21], using the structure of dUTPase from
E.coli, determined at 1.9 Å resolution (pdb id 1DUP), as a search model. Water molecules and metal ions were removed from the model. The initial model was subjected to crystallographic refinement with REFMAC5 [
22,
23]. Manual rebuilding of the model was carried out in Coot [
24]. The final cycle with an occupancy refinement was performed in Phenix [
25]. Data and refinement statistics are summarized in
Table 2. The atom coordinates and structure factors were deposited in the Protein Data Bank.
Figures were prepared using PyMOL [
26].
5. Conclusions
We have identified a structural element within T5 Dut, a small loop 30-GTNPAA-35 that is only 6 amino acids long, which is responsible for the implementation of a novel function. Reducing the size of this loop has dramatic consequences for the development of the phage T5i n E.coli cells, similar to the complete removal of Dut. Additionally, we have demonstrated that the enzymatic activity is implemented separately and does not influence the novel function. We speculate that the novel function is realized during the morphogenesis stage of the phage particle.
Supplementary Materials
The following supporting information can be downloaded at the website of this paper posted on Preprints.org, Figure S1: Sequence alignment of trimeric dUTPases.
Author Contributions
Conceptualization, A.S.G., U.D. and A.G.G.; methodology, A.S.G., V.M., U.D. and A.G.G.; software, A.G.G.; validation, A.S.G., U.D. and A.G.G.; formal analysis, A.S.G., V.M., U.D. and A.G.G.; investigation, A.S.G., U.D. and A.G.G.; resources, A.S.G., V.M., U.D. and A.G.G.; data curation, A.S.G., V.M., U.D. and A.G.G.; writing—original draft preparation, A.S.G., U.D. and A.G.G.; writing—review and editing, A.S.G., V.M., U.D., G.S. and A.G.G.; visualization, A.G.G.; supervision, A.G.G; project administration, A.G.G.; funding acquisition, A.G.G. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by Russian Science Foundation, grant number 23-24-00510.
Institutional Review Board Statement
Not applicable.
Data Availability Statement
MDPI Research Data Policies.
Acknowledgments
We thank MDPI for the opportunity to publish the results of our research free of charge.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Kerepesi, C.; Szabó, J.E.; Papp-Kádár, V.; Dobay, O.; Szabó, D.; Grolmusz, V.; Vértessy, B.G. Life without dUTPase. Frontiers in Microbiology 2016, 7, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Vértessy, B.G.; Tóth, J. Keeping Uracil Out of DNA: Physiological Role, Structure and Catalytic Mechanism of dUTPases. Acc. Chem. Res. 2009, 42, 97–106. [Google Scholar] [CrossRef] [PubMed]
- el-Hajj, H.H.; Zhang, H.; Weiss, B. Lethality of a Dut (Deoxyuridine Triphosphatase) Mutation in Escherichia Coli. Journal of bacteriology 1988, 170, 1069–1075. [Google Scholar] [CrossRef] [PubMed]
- Pecsi, I.; Hirmondo, R.; Brown, A.C.; Lopata, A.; Parish, T.; Vertessy, B.G.; Tóth, J. The Dutpase Enzyme Is Essential in Mycobacterium Smegmatis. PLoS ONE 2012, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Gadsden, M.H.; McIntosh, E.M.; Game, J.C.; Wilson, P.J.; Haynes, R.H. dUTP Pyrophosphatase Is an Essential Enzyme in Saccharomyces Cerevisiae. EMBO Journal 1993, 12, 4425–4431. [Google Scholar] [CrossRef] [PubMed]
- Pálinkás, H.L.; Rácz, G.A.; Gál, Z.; Hoffmann, O.I.; Tihanyi, G.; Róna, G.; Gócza, E.; Hiripi, L.; Vértessy, B.G. Crispr/Cas9-Mediated Knock-out of Dutpase in Mice Leads to Early Embryonic Lethality. Biomolecules 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Tarbouriech, N.; Buisson, M.; Seigneurin, J.-M.; Cusack, S.; Burmeister, W.P. The Monomeric dUTPase from Epstein-Barr Virus Mimics Trimeric dUTPases. Structure 2005, 13, 1299–1310. [Google Scholar] [CrossRef] [PubMed]
- Harkiolaki, M.; Dodson, E.J.; Bernier-Villamor, V.; Turkenburg, J.P.; González-Pacanowska, D.; Wilson, K.S. The Crystal Structure of Trypanosoma Cruzi dUTPase Reveals a Novel dUTP/dUDP Binding Fold. Structure 2004, 12, 41–53. [Google Scholar] [CrossRef] [PubMed]
- Cedergren-Zeppezauer, E.S.; Larsson, G.; Olof Nyman, P.; Dauter, Z.; Wilson, K.S. Crystal Structure of a dUTPase. Nature 1992, 355, 740–743. [Google Scholar] [CrossRef]
- Ariza, M.E.; Cox, B.; Martinez, B.; Mena-Palomo, I.; Zarate, G.J.; Williams, M.V. Viral dUTPases: Modulators of Innate Immunity. Biomolecules 2022, 12, 227. [Google Scholar] [CrossRef]
- Tormo-Más, M.Á.; Mir, I.; Shrestha, A.; Tallent, S.M.; Campoy, S.; Lasa, Í.; Barbé, J.; Novick, R.P.; Christie, G.E.; Penadés, J.R. Moonlighting Bacteriophage Proteins Derepress Staphylococcal Pathogenicity Islands. Nature 2010, 465, 779–782. [Google Scholar] [CrossRef] [PubMed]
- Tormo-Más, M.Á.; Donderis, J.; García-Caballer, M.; Alt, A.; Mir-Sanchis, I.; Marina, A.; Penadés, J.R. Phage dUTPases Control Transfer of Virulence Genes by a Proto-Oncogenic G Protein-like Mechanism. Molecular Cell 2013, 49, 947–958. [Google Scholar] [CrossRef] [PubMed]
- Maiques, E.; Quiles-Puchalt, N.; Donderis, J.; Ciges-Tomas, J.R.; Alite, C.; Bowring, J.Z.; Humphrey, S.; Penadés, J.R.; Marina, A. Another Look at the Mechanism Involving Trimeric dUTPases in Staphylococcus Aureus Pathogenicity Island Induction Involves Novel Players in the Party. Nucleic Acids Res 2016, 44, 5457–5469. [Google Scholar] [CrossRef] [PubMed]
- Warner, H.R.; Thompson, R.B.; Mozer, T.J.; Duncan, B.K. The Properties of a Bacteriophage T5 Mutant Unable to Induce Deoxyuridine 5’-Triphosphate Nucleotidohydrolase. Synthesis of Uracil-Containing T5 Deoxyribonucleic Acid. Journal of Biological Chemistry 1979, 254, 7534–7539. [Google Scholar] [CrossRef] [PubMed]
- Penadés, J.R.; Donderis, J.; García-Caballer, M.; Tormo-Más, M.Á.; Marina, A. dUTPases, the Unexplored Family of Signalling Molecules. Current Opinion in Microbiology 2013, 16, 163–170. [Google Scholar] [CrossRef] [PubMed]
- Nyíri, K.; Harris, M.J.; Matejka, J.; Ozohanics, O.; Vékey, K.; Borysik, A.J.; Vértessy, B.G. HDX and Native Mass Spectrometry Reveals the Different Structural Basis for Interaction of the Staphylococcal Pathogenicity Island Repressor Stl with Dimeric and Trimeric Phage dUTPases. Biomolecules 2019, 9, 488. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T.; Russell, D.W.; Green, M.R. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, 1989; ISBN 978-0-87969-309-1. [Google Scholar]
- Kabsch, W. Integration, Scaling, Space-Group Assignment and Post-Refinement. Acta Crystallographica Section D: Biological Crystallography 2010, 66, 133–144. [Google Scholar] [CrossRef] [PubMed]
- Rigaku XtaLAB Synergy-S Available online: https://www.rigaku.com/products/crystallography/synergys.
- Rigaku; Oxford; Diffraction CrysAlisPro 2019, Rigaku Oxford Diffraction, Yarnton, UK.
- McCoy, A.J.; Grosse-Kunstleve, R.W.; Adams, P.D.; Winn, M.D.; Storoni, L.C.; Read, R.J. Phaser Crystallographic Software. Journal of Applied Crystallography 2007, 40, 658–674. [Google Scholar] [CrossRef]
- Murshudov, G.N.; Skubák, P.; Lebedev, A. a.; Pannu, N.S.; Steiner, R. a.; Nicholls, R. a.; Winn, M.D.; Long, F.; Vagin, A. a. REFMAC5 for the Refinement of Macromolecular Crystal Structures. Acta Crystallographica Section D: Biological Crystallography 2011, 67, 355–367. [Google Scholar] [CrossRef]
- Kovalevskiy, O.; Nicholls, R.A.; Long, F.; Carlon, A.; Murshudov, G.N. Overview of Refinement Procedures within REFMAC 5: Utilizing Data from Different Sources. Acta Crystallographica Section D: Structural Biology 2018, 74, 215–227. [Google Scholar] [CrossRef]
- Emsley, P.; Lohkamp, B.; Scott, W.G.; Cowtan, K. Features and Development of Coot. Acta Crystallographica Section D: Biological Crystallography 2010, 66, 486–501. [Google Scholar] [CrossRef] [PubMed]
- Liebschner, D.; Afonine, P. V.; Baker, M.L.; Bunkoczi, G.; Chen, V.B.; Croll, T.I.; Hintze, B.; Hung, L.W.; Jain, S.; McCoy, A.J.; et al. Macromolecular Structure Determination Using X-Rays, Neutrons and Electrons: Recent Developments in Phenix. Acta Crystallographica Section D: Structural Biology 2019, 75, 861–877. [Google Scholar] [CrossRef] [PubMed]
- DeLano, W.L. The PyMOL Molecular Graphics System Available online: http://www.pymol.org/.
Figure 1.
Phenotypic manifestation of an amber mutation in the T5 phage dut gene. The mutant phage, T5dut-am, was plated with the same dilution onto a lawn of nonsuppressing (XAC) and suppressing (XA101) E.coli strains.
Figure 1.
Phenotypic manifestation of an amber mutation in the T5 phage dut gene. The mutant phage, T5dut-am, was plated with the same dilution onto a lawn of nonsuppressing (XAC) and suppressing (XA101) E.coli strains.
Figure 2.
(а) Sequence alignment of trimeric dUTPases (identical residues are shown in blue, homologous residues are in green); extra loop shown by magenta, mutated residues of the enzymatic active center (Ser68 and Asp85) are boxed; (b) schematic ribbon representation of the monomer wild-type T5Dut structure. (α-helices in red and β-strands in blue) phage-specific insert highlighted in magenta; (c) ribbon model of the monomer of T5Dut (orange), superimposed on the E.coli dUTPases monomer (blue, pdb id 1DUD); (d) trimer of T5Dut structure. Extra loop shown by magenta.
Figure 2.
(а) Sequence alignment of trimeric dUTPases (identical residues are shown in blue, homologous residues are in green); extra loop shown by magenta, mutated residues of the enzymatic active center (Ser68 and Asp85) are boxed; (b) schematic ribbon representation of the monomer wild-type T5Dut structure. (α-helices in red and β-strands in blue) phage-specific insert highlighted in magenta; (c) ribbon model of the monomer of T5Dut (orange), superimposed on the E.coli dUTPases monomer (blue, pdb id 1DUD); (d) trimer of T5Dut structure. Extra loop shown by magenta.
Figure 3.
Superimposed structural view of the T5Dut (orange, pdb id 8QKY) and mutant form T5DutΔloop (grey, pdb id 8QLD); (a) Loop region close-up; (b) monomers superposition.
Figure 3.
Superimposed structural view of the T5Dut (orange, pdb id 8QKY) and mutant form T5DutΔloop (grey, pdb id 8QLD); (a) Loop region close-up; (b) monomers superposition.
Figure 4.
(а) Phenotypic manifestation of the loop reduction in the dut gene of phage T5. The mutant phage T5dut(Δloop) was plated at the equal titre onto a lawn of host E.coli strains XAC and the same cells carrying the plasmid for complementation pBADex1/T5dut; (b) Enzymatic activity T5Dut and T5Dut(Δloop).
Figure 4.
(а) Phenotypic manifestation of the loop reduction in the dut gene of phage T5. The mutant phage T5dut(Δloop) was plated at the equal titre onto a lawn of host E.coli strains XAC and the same cells carrying the plasmid for complementation pBADex1/T5dut; (b) Enzymatic activity T5Dut and T5Dut(Δloop).
Figure 5.
(a) Superimposed structural view of the dUTPases from E.coli in complex with substrate analogue deoxyuridine-5’-diphosphate (monomers are colored shades of blue, pdb id 1DUD) and T5 (monomers are colored shades of orange, pdb id 8QKY); (b) Active-site close-up with the catalytically important residues, mutated residues of active center (Ser68 and Asp85) are boxed.
Figure 5.
(a) Superimposed structural view of the dUTPases from E.coli in complex with substrate analogue deoxyuridine-5’-diphosphate (monomers are colored shades of blue, pdb id 1DUD) and T5 (monomers are colored shades of orange, pdb id 8QKY); (b) Active-site close-up with the catalytically important residues, mutated residues of active center (Ser68 and Asp85) are boxed.
Figure 6.
(a) Analysis of the enzymatic activity of T5 Dut mutant forms T5Dut-S68A and T5Dut-D85N (b) and their gene's ability to complement amber mutation in the T5 phage dut gene.
Figure 6.
(a) Analysis of the enzymatic activity of T5 Dut mutant forms T5Dut-S68A and T5Dut-D85N (b) and their gene's ability to complement amber mutation in the T5 phage dut gene.
Figure 7.
Superimposed structural view of the T5Dut (orange, pdb id 8QKY), E.coli (blue, pdb id 1DUD) and S.aureus (cyan, pdb id 4GV8) dUTPases monomers.
Figure 7.
Superimposed structural view of the T5Dut (orange, pdb id 8QKY), E.coli (blue, pdb id 1DUD) and S.aureus (cyan, pdb id 4GV8) dUTPases monomers.
Table 1.
Efficiency of plating of T5wt and T5dut-am phages on various E.coli hosts.
Table 1.
Efficiency of plating of T5wt and T5dut-am phages on various E.coli hosts.
| E.coli |
Efficiency of plating compared to E.coli XA101 |
| strain |
T5wt |
T5dut-am |
XA101 (Su+)
XAC (Su0) |
1
0.97 |
1
0.33 |
Table 2.
Crystallographic data collection and refinement statistics.
Table 2.
Crystallographic data collection and refinement statistics.
| Data Collection |
|---|
| |
T5Dut |
T5DutΔloop |
| Wavelength (Å) |
1.0332 |
1.5418 |
| Resolution range (Å) |
50.00 - 2.00
(2.05 - 2.00)a
|
23.00 – 2.10
(2.20 – 2.10) |
| Space group |
P43 |
P65 |
Cell parameters
a,b,c (Å)
α, β, γ ( ◦ ) |
78.43, 78.43, 86.94
90.0, 90.0, 90.0 |
88.69, 88.74, 100.01
90.0, 90.0, 120.0 |
| Collection temperature (K) |
100 |
120 |
| Total reflections |
452,788 (20,225) |
233,256 (30,793) |
| Unique reflections |
35,488 (2,637) |
25,912 (3,239) |
| Rmerge (%) |
7.3 (101.5) |
9.4 (51.4) |
| Multiplicity |
12.76 (6.67) |
9.00 (9.51) |
| Completeness (%) |
99.9 (99.9) |
99.3 (100.0) |
| Mean I/sigma(I) |
22.91 (1.86) |
28.14 (4.22) |
| Wilson B-factor (Å2 ) |
35.1 |
23.8 |
| CC1/2 |
0.99 (0.72) |
0.99 (0.95) |
| Refinement |
| Resolution range |
26.76 - 2.00
(2.06 - 2.00) |
20.00 – 2.10
(2.15 – 2.10) |
| Reflections used in refinement |
35,476 (2,598) |
23,632 (1,746) |
| Reflections used for R-free |
1,775 (137) |
1,160 (76) |
| R-work, % |
17.76 (27.97) |
22.05 (31.7) |
| R-free, % |
21.09 (33.25) |
26.41 (40.6) |
| RMSD bond lengths (Å) |
0.007 |
0.007 |
| RMSD bond angles (◦) |
0.843 |
1.342 |
| Ramachandran favored (%) |
98.51 |
97.12 |
| Ramachandran allowed (%) |
1.49 |
2.88 |
| Ramachandran outliers (%) |
0.00 |
0.00 |
| Average B-factor (Å2) |
40.21 |
33.46 |
| macromolecules |
40.24 |
33.48 |
| ligands |
43.65 |
42.32 |
| solvent |
39.63 |
30.51 |
| PDB ID |
8QKY |
8QLD |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).