Submitted:
19 November 2023
Posted:
20 November 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
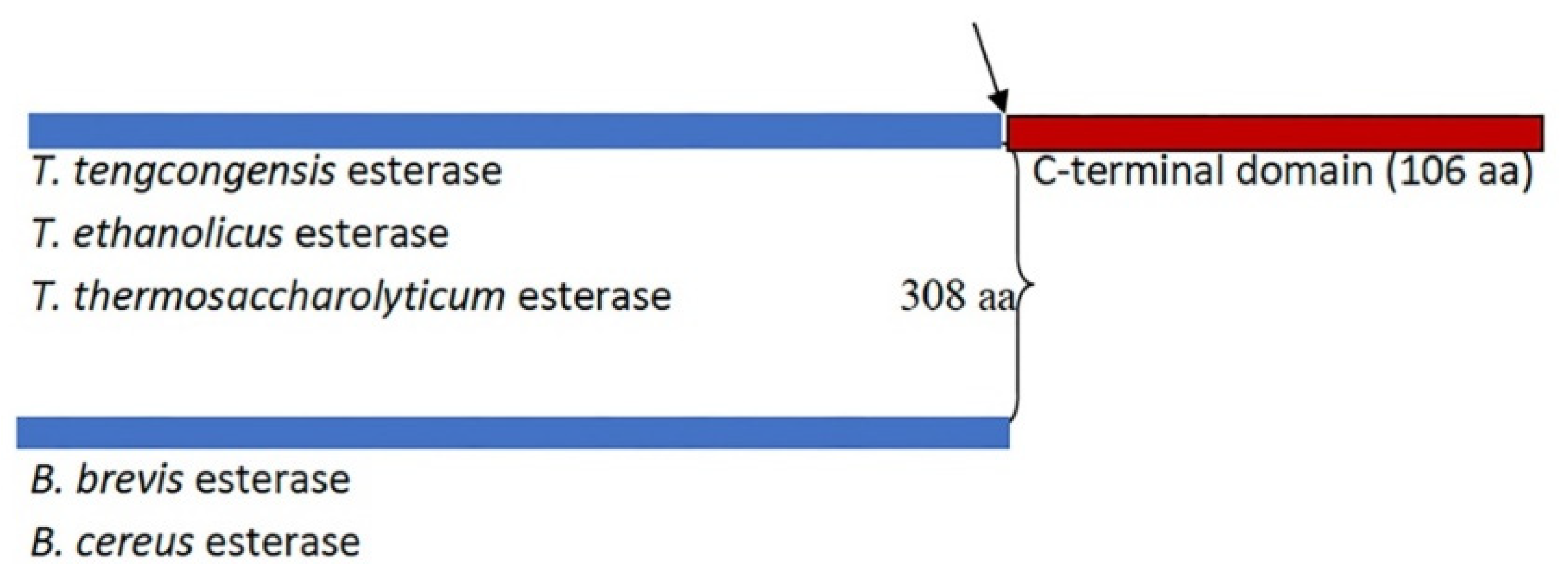
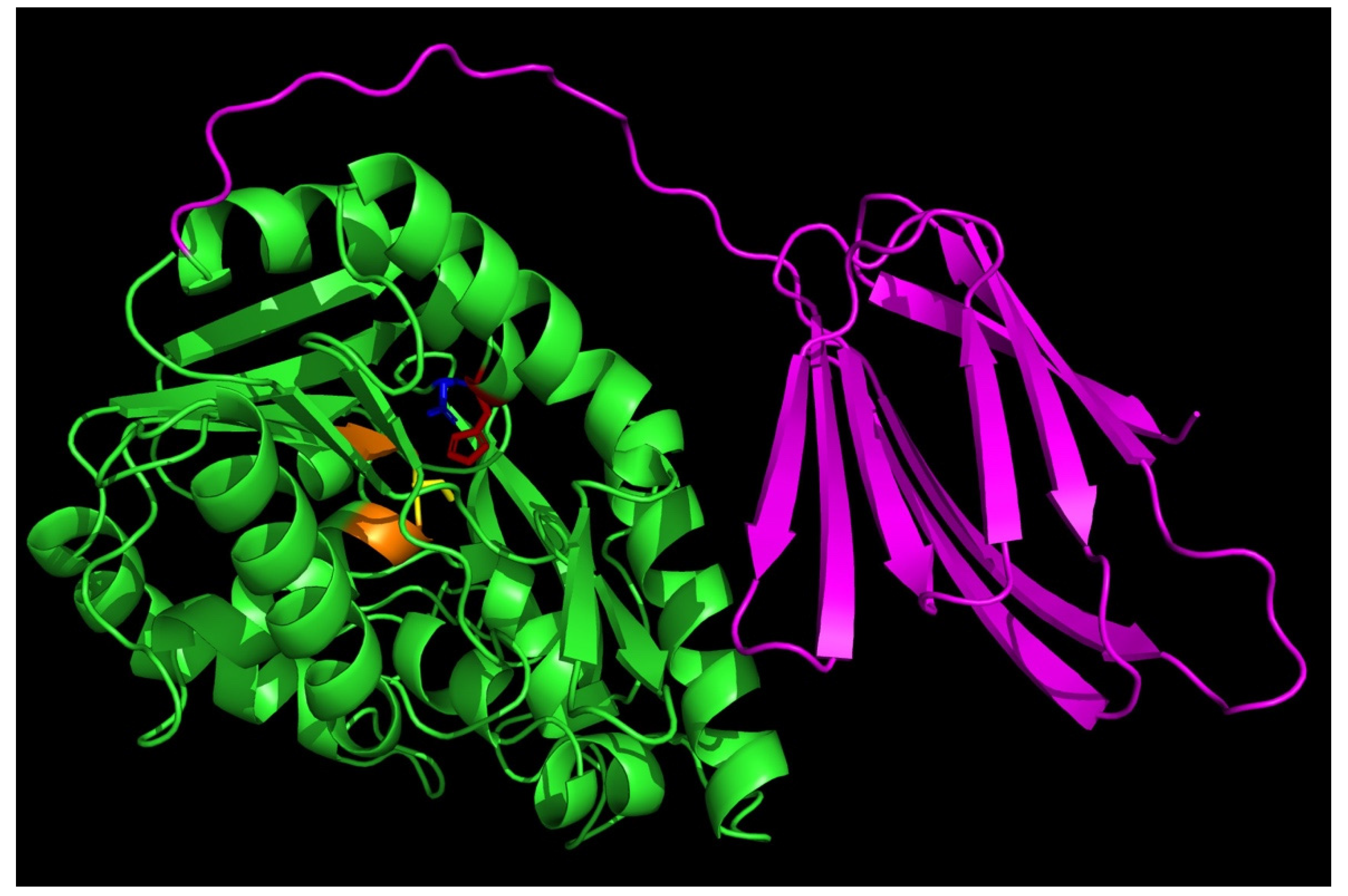
2.1. Alphafold-Predicted Structure of T. tengcongensis EstA3
2.2. Design, Cloning, Expression, and Purification of Esterases
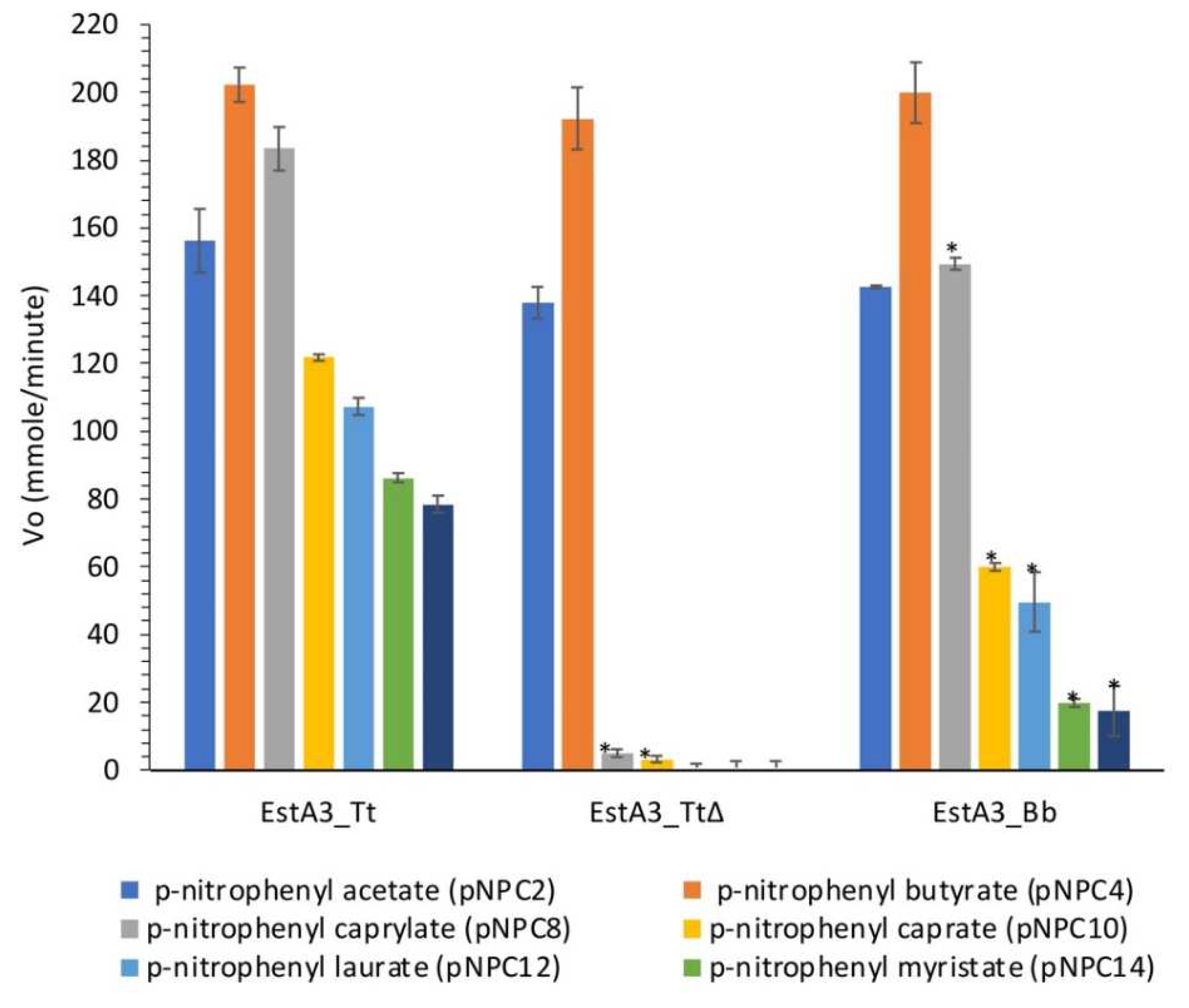
2.3. Hydrolytic Activities of Purified Esterases Reveal Different Substrate Preferences
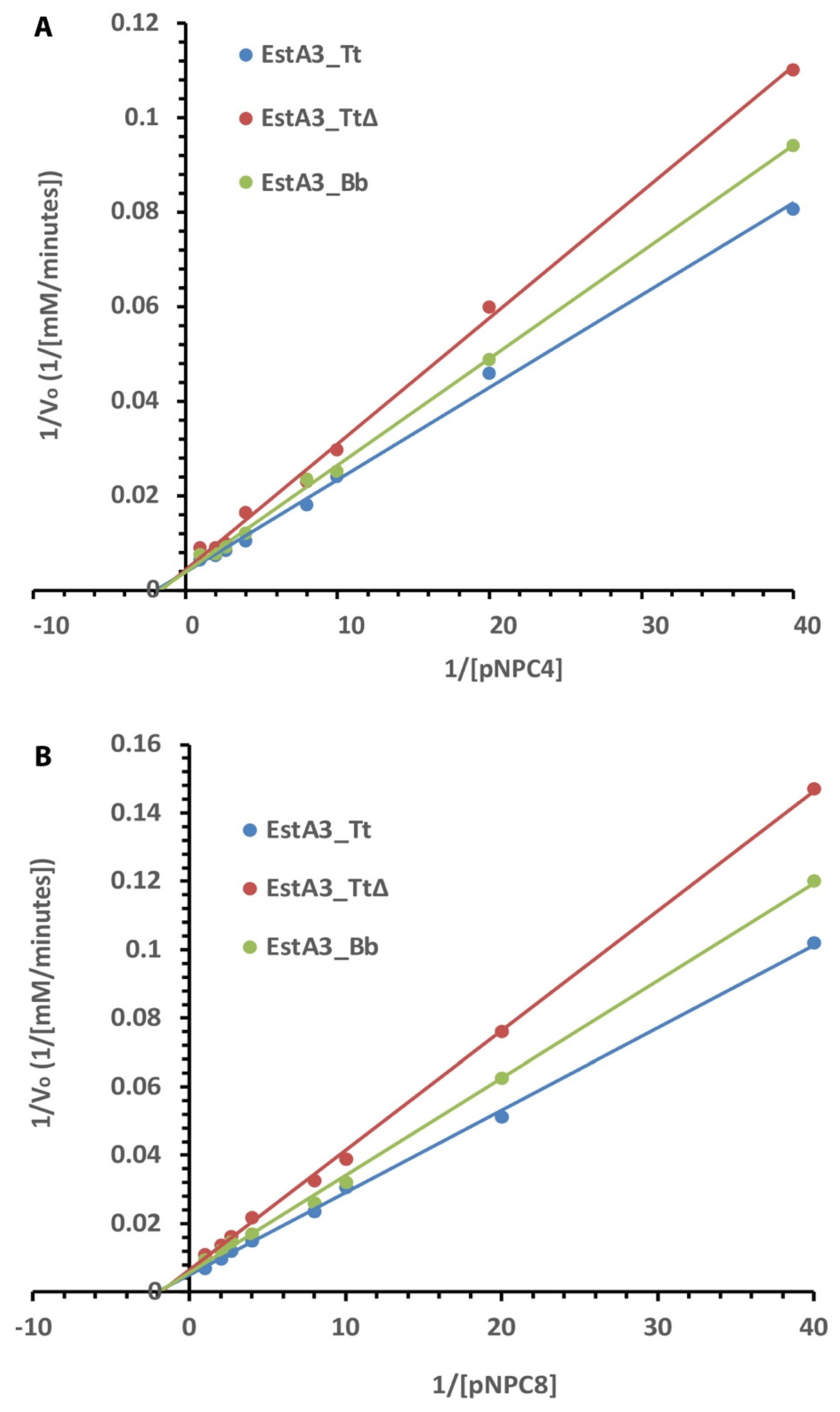
2.4. Kinetics of Activities of Purified Esterases on C4 and C8 Substrates
| Enzymes | Km (mM) | Vmax (mM/minutes) | Kcat x 105 (S-1) |
|---|---|---|---|
| pNPC4 | |||
| EstA3 | 0.53 | 263.17 | 2.94 |
| EstA3.bb | 0.59 | 256.41 | 1.51 |
| EstA3∆ | 0.64 | 238.10 | 1.50 |
| pNPC8 | |||
| EstA3 | 0.49 | 204.08 | 2.28 |
| EstA3.bb | 0.51 | 181.82 | 1.07 |
| EstA3∆ | 0.56 | 158.73 | 1.00 |
3. Discussion
3.1. Substrate Preference of EstA3_Tt, EstA3_Tt∆, and EstA3_Bb
3.2. The C-Terminal Tail Contributes to the Stability of EstA3
3.3. Role of the C-Terminal β Sandwich in Binding Long Chain Acyl Substrates
3.4. Structure-Guided Engineering of Esterases Could Generate New Catalytic Flexibility for Industrial Applications
4. Materials and Methods
4.1. Plasmids and Bacterial Strains
4.2. Chemicals
4.3. Protein Expression and Purification
4.4. Determination of Esterase Activity
4.5. Determination of Substrate Preference and Kinetic Parameters
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Andualema, B.; Gessesse, A. Microbial Lipases and Their Industrial Applications. Biotechnol. Faisalabad 2012, 11, 100–118. [Google Scholar] [CrossRef]
- Martínez-Martínez, M.; Coscolín, C.; Santiago, G.; Chow, J.; Stogios, P.J.; Bargiela, R.; Gertler, C.; Navarro-Fernández, J.; Bollinger, A.; Thies, S.; et al. Determinants and Prediction of Esterase Substrate Promiscuity Patterns. ACS Chem. Biol. 2018, 13, 225–234. [Google Scholar] [CrossRef]
- Holmquist, M. Alpha Beta-Hydrolase Fold Enzymes Structures, Functions and Mechanisms. Curr. Protein Pept. Sci. 2000, 1, 209–235. [Google Scholar] [CrossRef] [PubMed]
- Kohli, P.; Gupta, R. Medical Aspects of Esterases: A Mini Review. Int J Pharm Pharm Sci 2016, 8, 21–26. [Google Scholar]
- Panda, T.; Gowrishankar, B.S. Production and Applications of Esterases. Appl. Microbiol. Biotechnol. 2005, 67, 160–169. [Google Scholar] [CrossRef] [PubMed]
- Rao, L.; Xue, Y.; Zhou, C.; Tao, J.; Li, G.; Lu, J.R.; Ma, Y. A Thermostable Esterase from Thermoanaerobacter Tengcongensis Opening up a New Family of Bacterial Lipolytic Enzymes. Biochim. Biophys. Acta BBA-Proteins Proteomics 2011, 1814, 1695–1702. [Google Scholar] [CrossRef] [PubMed]
- Xue, Y.; Xu, Y.; Liu, Y.; Ma, Y.; Zhou, P. Thermoanaerobacter Tengcongensis Sp. Nov., a Novel Anaerobic, Saccharolytic, Thermophilic Bacterium Isolated from a Hot Spring in Tengcong, China. Int. J. Syst. Evol. Microbiol. 2001, 51, 1335–1341. [Google Scholar] [CrossRef] [PubMed]
- Steel, D.M.; Walker, J.M. Thermostable Proteins. Life Chem. Rep. Ser. 1991, 8, 49–96. [Google Scholar]
- Zhang, J.; Liu, J.; Zhou, J.; Ren, Y.; Dai, X.; Xiang, H. Thermostable Esterase from Thermoanaerobacter Tengcongensis: High-Level Expression, Purification and Characterization. Biotechnol. Lett. 2003, 25, 1463–1467. [Google Scholar] [CrossRef]
- Crowe, J.; Döbeli, H.; Gentz, R.; Hochuli, E.; Stüber, D.; Henco, K. 6xHis-Ni-NTA Chromatography as a Superior Technique in Recombinant Protein Expressiod/Purification. In Protocols for Gene Analysis; Humana Press: New Jersey, 1994; Vol. 31, pp. 371–388. [Google Scholar] [CrossRef]
- Rao, L.; Xue, Y.; Zheng, Y.; Lu, J.R.; Ma, Y. A Novel Alkaliphilic Bacillus Esterase Belongs to the 13th Bacterial Lipolytic Enzyme Family. PloS One 2013, 8, e60645. [Google Scholar] [CrossRef]
- Jensen, M.-B. V.; Horsfall, L.E.; Wardrope, C.; Togneri, P.D.; Marles-Wright, J.; Rosser, S.J. Characterisation of a New Family of Carboxyl Esterases with an OsmC Domain. Plos One 2016, 11, e0166128. [Google Scholar] [CrossRef]
- Latip, W.; Raja Abd Rahman, R.N.Z.; Leow, A.T.C.; Mohd Shariff, F.; Kamarudin, N.H.A.; Mohamad Ali, M.S. The Effect of N-Terminal Domain Removal towards the Biochemical and Structural Features of a Thermotolerant Lipase from an Antarctic Pseudomonas Sp. Strain AMS3. Int. J. Mol. Sci. 2018, 19, 560. [Google Scholar] [CrossRef]
- López-Fernández, J.; Barrero, J.J.; Benaiges, M.D.; Valero, F. Truncated Prosequence of Rhizopus Oryzae Lipase: Key Factor for Production Improvement and Biocatalyst Stability. Catalysts 2019, 9, 961. [Google Scholar] [CrossRef]
- Choi, Y.-J.; Lee, B. Culture Conditions for the Production of Esterase from Lactobacillus Casei CL96. Bioprocess Biosyst. Eng. 2001, 24, 59–63. [Google Scholar] [CrossRef]
- Samoylova, Y.V.; Sorokina, K.N.; Romanenko, M.V.; Parmon, V.N. Cloning, Expression and Characterization of the Esterase estUT1 from Ureibacillus Thermosphaericus Which Belongs to a New Lipase Family XVIII. Extremophiles 2018, 22, 271–285. [Google Scholar] [CrossRef] [PubMed]
- Fakruddin, M.; Mohammad Mazumdar, R.; Bin Mannan, K.S.; Chowdhury, A.; Hossain, M.N. Critical Factors Affecting the Success of Cloning, Expression, and Mass Production of Enzymes by Recombinant E. Coli. Int. Sch. Res. Not. 2013, 2013. [Google Scholar] [CrossRef]
- Olorunniji, F.J.; Buck, D.E.; Colloms, S.D.; McEwan, A.R.; Smith, M.C.M.; Stark, W.M.; Rosser, S.J. Gated Rotation Mechanism of Site-Specific Recombination by ϕC31 Integrase. Proc. Natl. Acad. Sci. 2012, 109, 19661–19666. [Google Scholar] [CrossRef]
- Bornscheuer, U.T.; Bessler, C.; Srinivas, R.; Krishna, S.H. Optimizing Lipases and Related Enzymes for Efficient Application. TRENDS Biotechnol. 2002, 20, 433–437. [Google Scholar] [CrossRef] [PubMed]
- Levisson, M.; Van Der Oost, J.; Kengen, S.W.M. Characterization and Structural Modeling of a New Type of Thermostable Esterase from Thermotoga Maritima. FEBS J. 2007, 274, 2832–2842. [Google Scholar] [CrossRef]
- López-López, O.; E Cerdan, M.; I Gonzalez Siso, M. New Extremophilic Lipases and Esterases from Metagenomics. Curr. Protein Pept. Sci. 2014, 15, 445–455. [Google Scholar] [CrossRef]
- Igunnu, A.; Joel, E.B.; Ezetulugo, L.N.; Malomo, S.O. Modulatory Effects of Mg2+ and Zn2+ Ions on Monoesterase Activity of Wild-Type and Mutant E. Coli Alkaline Phosphatases. Ilorin J. Sci. 2018, 5, 36–47. [Google Scholar] [CrossRef]
- Ferrer, M.; Bargiela, R.; Martínez-Martínez, M.; Mir, J.; Koch, R.; Golyshina, O.V.; Golyshin, P.N. Biodiversity for Biocatalysis: A Review of the α/β-Hydrolase Fold Superfamily of Esterases-Lipases Discovered in Metagenomes. Biocatal. Biotransformation 2015, 33, 235–249. [Google Scholar] [CrossRef]
- Arpigny, J.L.; JAEGER, K.-E. Bacterial Lipolytic Enzymes: Classification and Properties. Biochem. J. 1999, 343, 177–183. [Google Scholar] [CrossRef] [PubMed]
- Ejima, K.; Liu, J.; Oshima, Y.; Hirooka, K.; Shimanuki, S.; Yokota, Y.; Hemmi, H.; Nakayama, T.; Nishino, T. Molecular Cloning and Characterization of a Thermostable Carboxylesterase from an Archaeon, Sulfolobus Shibatae DSM5389: Non-Linear Kinetic Behavior of a Hormone-Sensitive Lipase Family Enzyme. J. Biosci. Bioeng. 2004, 98, 445–451. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, Y.; Miyamoto, K.; Ohta, H. A Novel Thermostable Esterase from the Thermoacidophilic Archaeon Sulfolobus Tokodaii Strain 7. FEMS Microbiol. Lett. 2004, 236, 97–102. [Google Scholar] [CrossRef]
- Kaiser, P.; Raina, C.; Parshad, R.; Johri, S.; Verma, V.; Andrabi, K.I.; Qazi, G.N. A Novel Esterase from Bacillus Subtilis (RRL 1789): Purification and Characterization of the Enzyme. Protein Expr. Purif. 2006, 45, 262–268. [Google Scholar] [CrossRef] [PubMed]
- Metin, K.; Burcu Bakir Ateslier, Z.; Basbulbul, G.; Halil Biyik, H. Characterization of Esterase Activity inGeobacillus Sp. HBB-4. J. Basic Microbiol. 2006, 46, 400–409. [Google Scholar] [CrossRef]
- Zhu, Y.; Li, J.; Cai, H.; Ni, H.; Xiao, A.; Hou, L. Characterization of a New and Thermostable Esterase from a Metagenomic Library. Microbiol. Res. 2013, 168, 589–597. [Google Scholar] [CrossRef]
- Zhang, Y.-J.; Chen, C.-S.; Liu, H.-T.; Chen, J.-L.; Xia, Y.; Wu, S.-J. Purification, Identification and Characterization of an Esterase with High Enantioselectivity to (S)-Ethyl Indoline-2-Carboxylate. Biotechnol. Lett. 2019, 41, 1223–1232. [Google Scholar] [CrossRef]
- Nakamura, A.M.; Godoy, A.S.; Kadowaki, M.A.S.; Polikarpov, I. The first structure of Bacillus licheniformis lipase BlEst2 in its propeptide and mature form revealing molecular details of inhibition by its C-terminal domains. (Department of Biotechnology and Biomedicine, Technical University of Denmark, Søltofts Plads 224, 2800 Kongens Lyngby, Denmark), (2021). (manuscript in preparation; to be submitted). [CrossRef]
- Sayari, A.; Frikha, F.; Miled, N.; Mtibaa, H.; Ali, Y.B.; Verger, R.; Gargouri, Y. N-Terminal Peptide of Rhizopus Oryzae Lipase Is Important for Its Catalytic Properties. FEBS Lett. 2005, 579, 976–982. [Google Scholar] [CrossRef]
- Huang, J.; Dai, S.; Chen, X.; Xu, L.; Yan, J.; Yang, M.; Yan, Y. Alteration of Chain-Length Selectivity and Thermostability of Rhizopus Oryzae Lipase via Virtual Saturation Mutagenesis Coupled with Disulfide Bond Design. Appl. Environ. Microbiol. 2023, 89, e01878–22. [Google Scholar] [CrossRef] [PubMed]
- Kovacic, F.; Babic, N.; Krauss, U.; Jaeger, K.-E. Classification of Lipolytic Enzymes from Bacteria. Aerob. Util. Hydrocarb. Oils Lipids 2019, 24, 255–289. [Google Scholar] [CrossRef]
- Lan, T.; Wang, X.-R.; Zeng, Q.-Y. Structural and Functional Evolution of Positively Selected Sites in Pine Glutathione S-Transferase Enzyme Family. J. Biol. Chem. 2013, 288, 24441–24451. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




