1. Introduction
The primary pathogen of maize (Zea mays L.) in sub-Saharan Africa, the Islands of Mauritius and Reunion, and other parts of the world is the maize streak virus (MSV), which causes maize streak disease (Shepherd et al., 2010; Emeraghi et al., 2021). According to Shepherd et al. (2010), MSV is a species and type of member of the genus virus in the family Geminiviridae. The latter also includes the genera Curtovirus and Begomovirus. Virus species contain distinctive gemininate particles and a single-component, single-stranded, circular DNA genome that is around 2.7 kb long. The MSV genome is transcribed in both directions, as is the case for every member of the Geminiviridae family (Willment et al., 2007). Transcripts are independently regulated and begin in the noncoding long intergenic region (LIR). According to Palmer and Rybicki (1998) and Rojas et al. (2018), the noncoding short intergenic region (SIR) is where complementary-sense DNA replication, polyadenylation, and transcription termination all begin. While C1 and C2 are expressed from the complementary strand, open reading frames V1 and V2 are translated from the virion sense strand. From these ORFs, four useful proteins are encoded. These proteins include the replication-initiator protein Rep (41.3 kDa), the replication-associated protein RepA (31.5-31.6 kDa), translated from an unspliced RNA of the C1 ORF, the movement protein MP (10.9 kDa), and the coat protein CP (26.9-27.0 kDa), according to Palmer and Rybicki (1998). Different species of leafhoppers are natural transmitters of viruses. MSV is spread by a number of Cicadulina species (Shepherd et al., 2010; Emeraghi et al., 2021).et alet alet alet alet alet alet alMultiple isolates of Maize streak virus (MSV) have been identified, exhibiting variations in symptom severity on maize plants and their potential to infect resistant maize varieties (Monjane et al., 2011; Sime et al., 2021). Plants that are afflicted by severe isolates exhibit pronounced chlorosis throughout the leaf lamina, resulting in necrosis, stunted growth, and the production of subpar cobs. Conversely, mild isolates only induce limited streaks or spots, minimal stunting, and no significant effects on cob development (Bosque-Pérez, 2000; Shepherd et al., 2010; Emeraghi et al., 2021). According to a study conducted by Willment et al. (2007), it was shown that the presence of chlorotic streaks on leaves can be attributed to the degradation of chloroplasts.et alet alet alet alet al. The literature has documented the existence of infectious clones derived from many parts of Africa for isolates of Maize streak virus (MSV) (Martin & Shepherd, 2009; Shepherd et al., 2010). The analysis of sequences acquired from several clones exhibiting varied symptoms revealed a high degree of similarity (>95%). This finding indicates that only a few number of variations in the viral genomes are accountable for the variations in the intensity of symptoms (Varsani et al., 2008). The genomic region encompassing the 5’ terminus of the C1 ORF (Rep/RepA protein) and the LIR of the Nigerian clone of MSV-N (Boulton et al., 1991) was shown to be associated with determinants related to host range, onset of symptoms, and symptom type, namely the severity of chlorosis and streak length. The determination of streak width was based on the genomic segment corresponding to the virion-sense. According to the findings of Boulton et al. (1991), a specific nucleotide change at position 2473 (where the severe clone had a "A" and the mild clone had a "G") within the LIR region had an impact on both the severity of symptoms and the spectrum of hosts affected by the MSV-N clone. Previous research has indicated that the localization of the factors influencing the ability of a virus to cause disease may differ among various isolates of Maize streak virus (MSV) and may be distributed throughout many regions of the MSV genome (Martin & Rybicki, 2002). Previous research has employed chimeric geminivirus clones to ascertain the factors responsible for different biological characteristics (Boulton et al., 1991; Martin and Rybicki, 2002). The researchers in this study employed chimeric constructions of MSV in order to discern the factors that contribute to the pathogenicity of closely related mild and severe symptom clones from different African isolates (Martin and Rybicki, 2002). In a study conducted by Unseld et al. (2000), pseudo-recombinants and recombinants were generated by combining several begomoviruses to investigate the factors responsible for symptom development.et alet al. In the current effort to ascertain viral genes implicated in virulence and potential interactions, chimeric clones were generated, including segments of the genomes from a moderate clone (pMSV-KL) and a severe clone (pMSV-Km), both originating from isolates obtained in Kenya. The initial samples were procured from maize-infected plants located in the Kenyan Lake Victoria region, designated as MSV-KL, as well as from the wild grass Setaria verticillata, designated as MSV-Km. After the process of cloning, it was observed that pMSV-Kl induced narrow and discontinuous moderate chlorotic streaks in infected plants, along with mild stooping. On the other hand, pMSV-Km resulted in the development of long and broad severe chlorotic streaks, characterized by a yellowish white color, followed by considerable stunting of the infected plants. The virulence of these chimeric pMSV genomes, which were obtained from pMSV-KL and pMSV-Km, was assessed on a susceptible maize hybrid in a controlled and replicable environmental setting.
2. Materials and Methods
Construction of dimeric clones
In this study, the process of isolating whole genomic DNA from maize leaves infected with Maize Streak Virus (MSV) was conducted. To separate the viral DNA from the host nucleic acids, agarose gel electrophoresis was employed using a gel concentration of 0.8%. The double-stranded replicative form of the MSV DNA was extracted from the agarose gel and subjected to purification using Geneclean II, following the guidelines provided by the manufacturer (Bio101, Inc., La Jolla, CA). Two tandem dimeric copies were generated by inserting DNA fragments into the bacterial transformation vector pUC19 following digestion with the restriction enzyme BamH1. The recombinant plasmids were stored in two different conditions: E. Coli strain DH5a at a temperature of -80 °C, and as viral DNA in a buffer solution containing 10 mM Tris-HCl (pH 8.0) and 1 mM EDTA, kept at a temperature of 4 °C. The infectious clones utilized in the current investigation, namely pMSV-KL and pMSV-Km, served as the origin of the chimeras.
Characterization of virulence using vascular puncture inoculations (VPI)
In order to validate the previously documented virulences of pMSV-Km and pMSV-KL (Ngwira, 1997), dimeric clones were employed to infect the Pioneer hybrid P3379, which is known to be susceptible. The viral particle inoculation (VPI) method, as previously published by Redinbaugh et al. (2001), was utilized for infecting the kernels. The plants that were infected were kept in containers within a designated greenhouse or growth chambers located at the Ohio Agricultural Research Center (OARDC) in Wooster. VPI infected a total of fifty seeds for each clone of MSV. For the subsequent evaluation of the illness, a random selection of five plants displaying symptoms was made from each clone following inoculation. The visual assessment of chlorotic patches on leaf area was conducted to establish the proportion of coverage, using a rating scale ranging from 0 to 5. A rating of 0 indicated the absence of symptoms, while a rating of 5 indicated the presence of highly severe symptoms (Shepherd et al., 2005). Furthermore, the measurements of the heights of the five infected plants were taken by determining the distance from the soil’s stem collar to the apex of the fourth leaf. The data collected in this study were obtained from three separate replications conducted using a totally randomized methodology. Measurements were taken at 7-day intervals, starting from 14 days and continuing up to 28 days following the initiation of VPI.
Construction of recombinant chimeric clones
The construction of recombinant chimeric clones involved the utilization of conserved restriction endonuclease sites present in the viral genomes and multiple cloning sites (MCSs) of pUC19. This allowed for the interchange of DNA fragments between pMSV-Km(Severe) and pMSV-KL(Mild) using established cloning procedures. In a study conducted by Redinbaugh
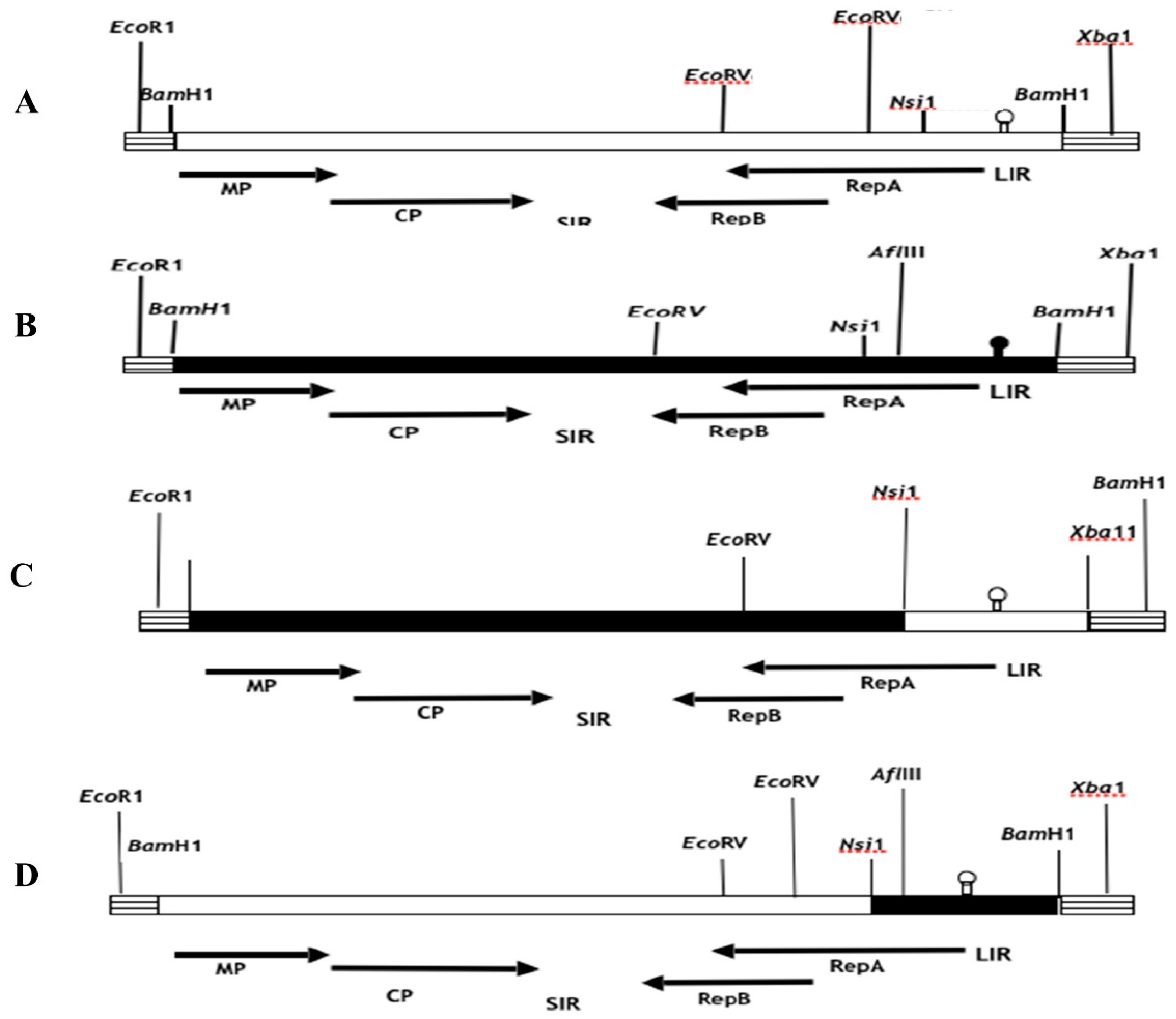
et al. (2001), the genomic MSV inserts in pUC19 were analyzed for their orientation using a series of strategically positioned restriction enzymes in both the MSV and pUC19 genomes. The selection process for constructing recombinant clones involved choosing clones with identical orientations, so guaranteeing the exchange of fragments in the same direction. The fragments necessary for the assembly of recombinant clones were derived from the monomeric versions of the parental clones contained within the pUC19 vector, which has a size of around 5.3 kilobases. In addition, pieces were directly removed from the ancestral dimeric clones, which were likewise present in pUC19. In order to acquire parental DNA monomers from the dimeric clones, the infectious clones of the latter were initially subjected to digestion employing specific viral restriction enzyme sites, namely Nsi1, Xho1, and BglII, as seen in
Figure 1. The parental monomeric clones, with a resultant size of 5.3 kb, were retrieved using the process of gel electrophoresis and afterwards religated at a temperature of 14 °C for an overnight duration. The resulting ligation mixes were then employed for the transformation of
E. Coli cells, namely DH5alpha or XL1Blue strains. Putative recombinant clones were tested in colonies by the utilization of restriction enzyme analysis. The DNA template used for this analysis was produced using the Wizard Miniprep kit (Promega, Madison, WI) in accordance with the instructions provided by the manufacturer.
et alE. Coli
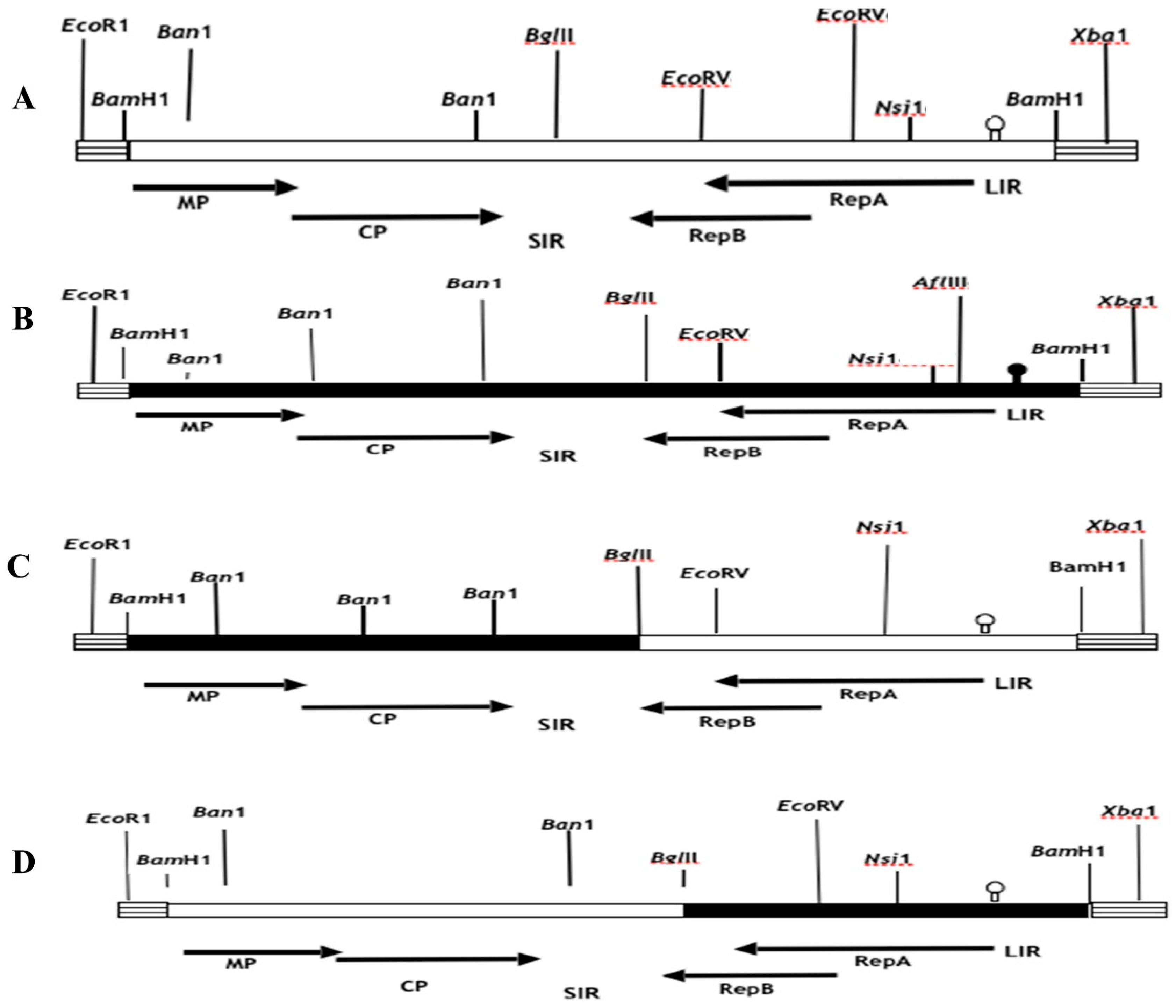
In order to generate recombinant chimeric clones pMSV-Km (KLLIR) and pMSV-KL(KmLIR), a reciprocal exchange of a 0.5 kilobase DNA fragment was performed subsequent to Nsi1 and Xba1 restriction enzyme digestion, as seen in
Figure 1 and
Figure 2. The Nsi1 restriction site, positioned at 2187, is situated at the 5’-end of the C1 open reading frame (ORF). Conversely, the Xba1 restriction site is found within the multiple cloning site (MCS) of the pUC19 vector. The interchange of viral sequences between nucleotides 2187-2687, which includes the 5’ end of the C1 open reading frame (ORF) and the whole long intergenic region (LIR) of Maize streak virus (MSV), was facilitated by the digestion of Nsi1 and Xba1 enzymes. In a similar manner, chimeric clones pMSV-Km (KLMP: CP) and pMSV-KL (KmMP:CP) were constructed, including the MP and CP ORFs together with the SIR through the exchange of EcoR1-BglII DNA pieces (refer to
Figure 2). The recombinants pMSV-Km (KLRep) and pMSV-KL(KmRep) were generated by including an EcoR1-BglII fragment from the parental pMSV-Km(Severe) and pMSV-KL(Mild), respectively, into pMSV-KL(KmLIR) and pMSV-Km(KLLIR) (
Figure 2). Therefore, the pMSV-Km (KL Rep) and pMSV-KL (Km Rep) constructs demonstrate reciprocal exchanges, wherein the C2 open reading frame (ORF) and a majority of the C1 ORF (627 out of 816 nucleotides) were inherited from one parental clone, while the remaining portion of the genome originated from the other parent (
Figure 2).
The use of clone-specific restriction enzyme sites facilitated the validation of recombinant genomes following each cloning or subcloning operation in
E.coli, employing the suitable endonucleases (
Figure 1 and
Figure 2). The chimeric clones’ infectious dimers were generated by incorporating BamH1 inserts into the pUC19 vector. The DNA of MSV, which had been digested with BamH1, was eluted subsequent to agarose gel electrophoresis. In order to acquire tandem dimeric repeats in the pUC19 plasmid, the BamH1-digested MSV DNA was ligated with the suitably digested and dephosphorylated pUC19 vector at a ratio of 1:6, and the ligation reaction was carried out overnight at a temperature of 14 °C. The transformation of competent cells of the HB101 strain of
E. coli (Gibco), commercially available, was carried out using ligation mixes as per the directions provided by the manufacturer. The colonies that included potential dimeric chimerical clones were subjected to screening using miniprep. The process of preparing large quantities of plasmids containing dimeric chimerical DNAs was carried out using silica-based anion-exchanger columns known as "Wizard Maxiprep" from Promega (Madison, WI), following the instructions provided by the manufacturer. The plasmid DNA was diluted in a solution containing 10 mM Tris, pH 8.0, and 1 mM EDTA to a concentration of 0.2 g l-1. This diluted DNA was then employed as an inoculum to infect the maize hybrid P3379 using the Viral Particle Infection (VPI) method. The evaluation of disease severity caused by each clone was conducted on a sample of five randomly chosen infected plants, as previously reported, and the timing of initial symptom manifestation was documented. Furthermore, the percentage of leaf area occupied by chlorotic spots in plants displaying symptoms was determined for leaf three. This was accomplished by employing a digital picture camera (Sony Digital Mavica, MVC-FD91) and quantifying the data using picture Pro Plus software (version 3.0 from Media Cybernetic, L.P.) as described by Martin and Rybicki (1998). The symptoms shown by plants that were infected with recombinant MSV DNA constructs were compared to those of control plants that were infected with parental mild and severe clones, as well as to mock-inoculated and uninfected plants, respectively. Three leaves were subjected to measurement, and afterwards, the obtained values were averaged.
Analysis of MSV genome sequences and coat proteins in infected plants
The extraction of total plant DNA was conducted on plants that were infected with both parental and recombinant DNA clones. This extraction procedure followed a modified version of the CTAB extraction approach. The PCR technique was employed to amplify the specific MSV genome area, and the resulting degenerated DNA fragments were subjected to purification using the Wizard PCR Preps DNA purification equipment manufactured by Promega (Madison, WI), following the instructions provided by the manufacturer. The constructions were validated by the use of the enzymes EcoRV, AflIII, and Ban1, resulting in characteristic digestions (data not shown). In order to validate the anticipated swapped pieces, the sequences of recombinant viral DNAs were ascertained and afterwards compared to the sequences of the initial cloned MSV DNA. The quantification of viral accumulation was conducted using a F(ab)2-Protein A enzyme-linked immunosorbent assay (ELISA), following the methodology outlined in a previous study by Todd et al. (2010). A sample of leaf tissue weighing 0.5 grams was collected from the sixth leaf of plants that were infected with parental and recombinant viral clones. This collection took place four weeks after viral pathogen inoculation (VPI). The enzyme-linked immunosorbent assay (ELISA) technique was employed to ascertain the relative quantities of viral coat protein (CP) in the samples. Control samples were obtained by using leaves from plants that were mock-inoculated. The spectrophotometer was utilized to measure the responses at a wavelength of 405 nm. The mean absorbencies obtained from the different treatments were compared using SPSS (SPSS Inc. Version 6.0).
Data analysis
The data pertaining to plant height, disease severity, and the percentage of chlorotic leaf area were analyzed using a repeated measures analysis of variance. Measurements were made at 1, 2, and 3 weeks following the first planting. The analysis was conducted using SAS PROC MIXED PC version 8.01, developed by SAS Institute Inc. in Cary, NC. The Tukey method, a technique for adjusting means in the context of multiple comparisons, was employed in this study to determine significant differences between means at a significance level of p = 0.05.
3. Results
Symptom phenotypes incited by recombinant clones
The severity of symptoms and stunting incited on inoculated P3379 hybrid maize plants by infectious dimeric clones, pMSV-Km and pMSV-KL, differentiated these clones into severe and mild symptomatic clones (
Table 1). Specifically, clone pMSV-Km incited long, wide and severe chlorotic streaks (yellowish white) accompanied by significant stunting of infected plants. In contrast, plants infected with clone pMSV-KL developed narrow, discontinuous, mildly chlorotic (light green) streaks accompanied by mild stunting statistically indistinguishable from both mock and uninoculated plants (
Figure 3 and
Figure 4). More severe symptoms were observed on thirteen different maize inbred lines inoculated with MSV-Km, as measured by AUDPC, and disease incidence was lower for MSV-Kl (
Table 1).
The LIR and 5’ end of the Rep A ORF
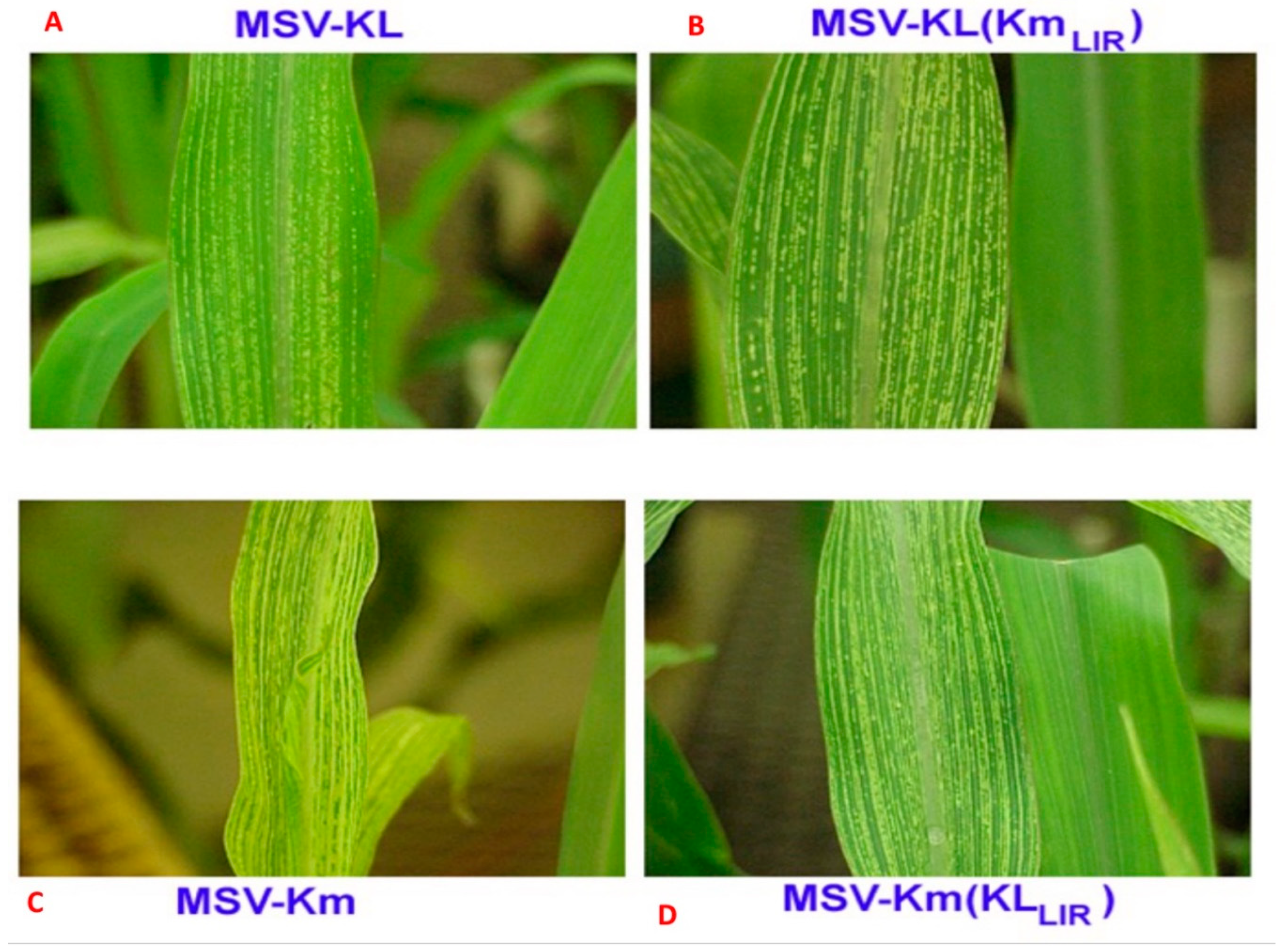
To test the contribution of the LIR plus the 5’ end of the RepA ORF to virulence, VPI inoculation of recombinant clones [pMSV-Km(KL
LIR] resulted in intermediate symptom expression in infected plants. The pMSV-Km(KL
LIR) exhibited symptoms similar to those of the severe parental clone pMSV-Km(Severe) (
Figure 3;
Table 1). Plants infected with this chimeric clone expressed long, wide and sometimes discontinuous moderately severe chlorotic streaks accompanied by intermediate stunting (
Figure 3;
Table 1). The latent period for symptom expression of this clone was indistinguishable from that of pMSV-Km(Severe), with symptoms appearing commonly 40 days after planting (
Table 1). In contrast, plants infected with pMSV-KL(Km
LIR) had a shorter latent period and exhibited slightly longer and wider streaks than those of the mild parental pMSV-KL(Mild) (
Figure 3).
The SIR, complementary and virion-sense sequences
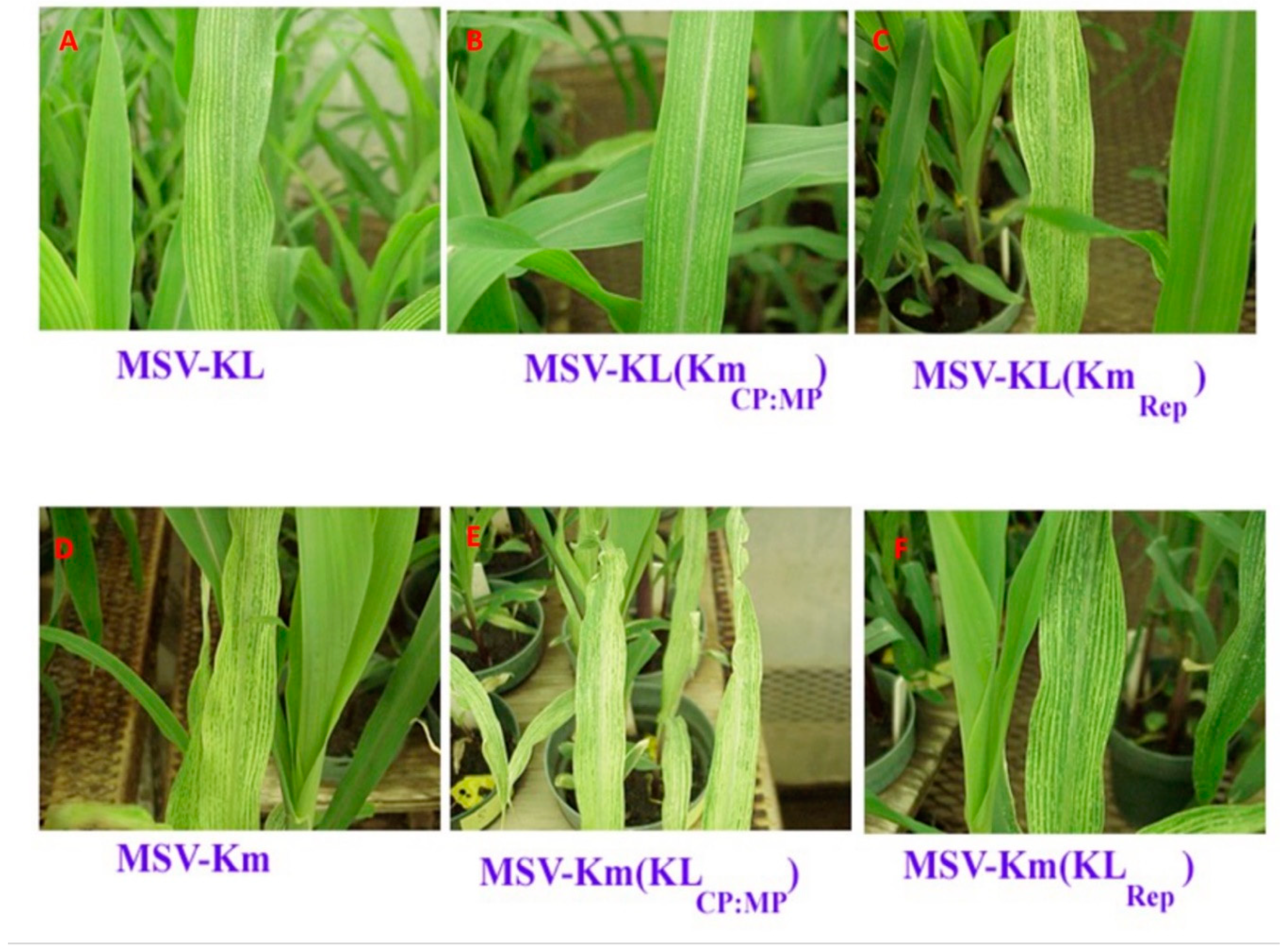
In order to investigate the impact of the SIR and complementary- and virion-sense sequences on the pathogenicity of Maize streak virus (MSV), four chimeric clones were generated, namely pMSV-Km (KL Rep), pMSV-KL (Km Rep), pMSVKm (KLMP:CP), and pMSV-KL(KmMP:CP) (
Figure 4). Plants that were infected with pMSV-Km (KL Rep) exhibited much less severe chlorotic streaks and stunting compared to plants infected with pMSV-Km (Severe), as seen in
Figure 4 and summarized in
Table 2. Nevertheless, there was no significant difference in the latent time for symptom manifestation between this clone and pMSV-Km(Severe) (
Table 2). Moreover, the symptoms elicited by pMSV-Km (KL Rep) were found to be of intermediate severity and exhibited substantial differences when compared to those caused by pMSV-KL (Mild), as shown in
Table 2. On the other hand, it was shown that pMSV-KL(KmRep) elicited symptoms that were comparable to those induced by pMSV-Km(Severe) (
Table 2). Similar to the cases of pMSV-Km (KL RepA) and pMSV-Km (Severe), there were no significant differences seen in the latent periods for symptom manifestation between pMSV-KL (KmRep) and pMSV-KL (Mild).
The introduction of chimeric clones containing the MP and CP ORFs, along with the SIR, resulted in significant alterations in the degree of chlorosis and stunting seen. The chimera pMSV-Km(KLMP:CP) elicited leaf symptoms of greater severity compared to the parental pMSV-Km(Severe) (
Figure 3;
Table 2). Nevertheless, the latent periods for both of these clones had an identical duration of 4.5 days. On the other hand, there was a notable decrease in chlorosis and stunting in plants that were infected with pMSV-KL(KmMP:CP) in comparison to the original pMSV-KL(Mild) clone. The onset of illness symptoms in this clone was seen to occur 1.5 days subsequent to the manifestation of symptoms in the pMSV-KL(Mild) variant.
Accumulation of virions of parental and recombinant clones
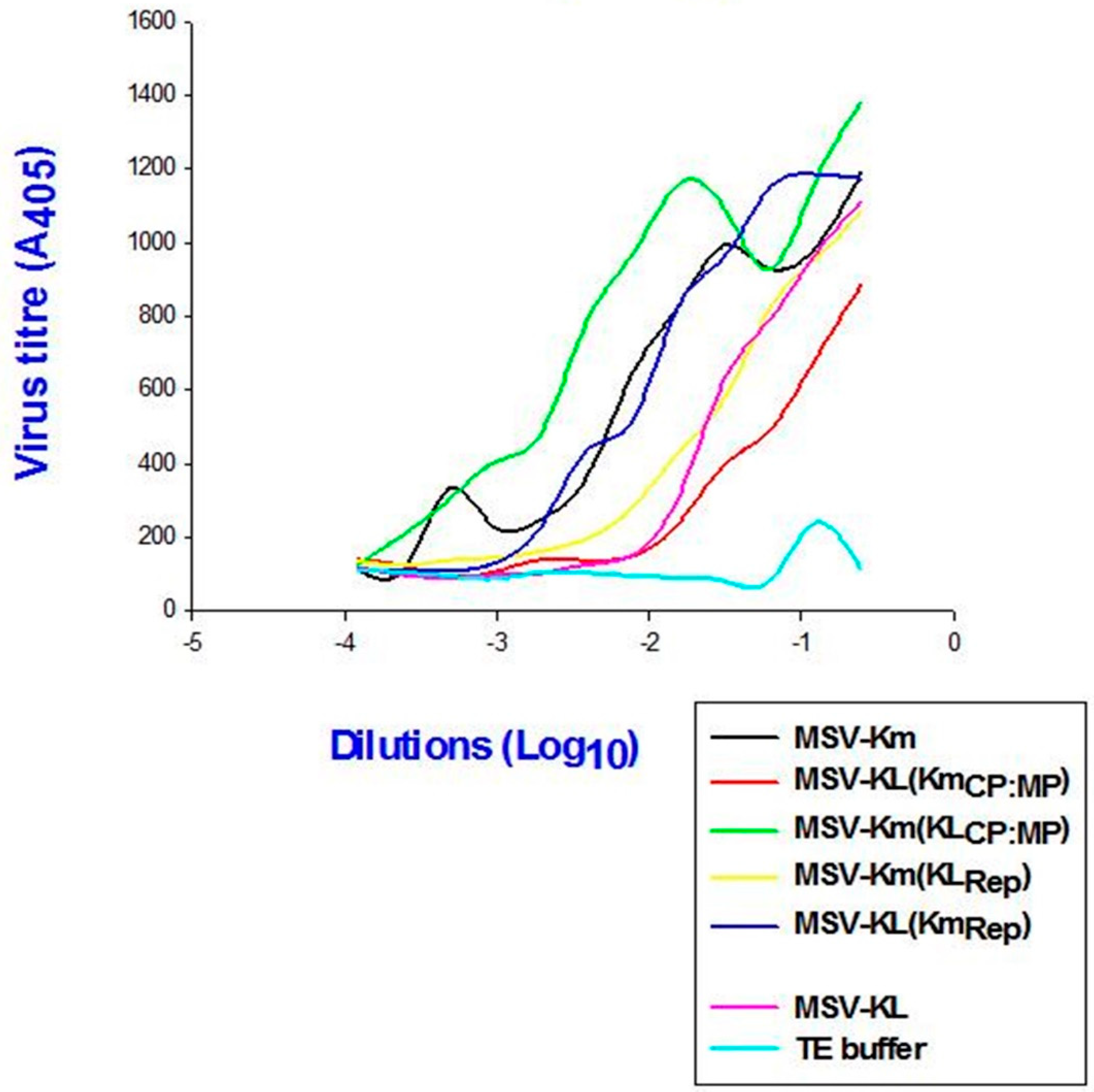
In order to evaluate the impact of the modified virulence on the viral load within the plants, the titer was measured. The findings indicated a positive correlation between viral accumulation in plants infected with P3379 and disease severity, as determined by the quantity of CP antigen (
Figure 5). The plants infected with pMSV-Km(KLMP:CP) exhibited the highest level of viral accumulation among all clones, accompanied with the most severe symptoms. Conversely, the lowest level of virus accumulation was found in plants infected with pMSV-KL(KmMP:CP), which displayed the mildest symptoms.
4. Discussion
The investigation focused on the genetic underpinnings of symptom severity in MSV. This was achieved by creating recombinant clones that included specific segments of genomic regions from two separate clones, namely pMSV-Km(Severe) and pMSV-KL(Mild). The Nsi1-BamH1 segment, which includes the LIR and the 5’ terminus of the Rep/RepA gene (specifically, the first 189 nucleotides), was found to be associated with the severity of symptoms. In a previous study conducted by Boulton et al. (1991), it was demonstrated that a mutation occurred in the promoter region of the Rep/RepA genes, specifically changing the TATA sequence to TGTA. This mutation was found to have several effects, including a reduction in the severity of chlorosis and length of streaks, an increase in latency, and a restriction in the host range of clone pMSV-Ns. There was speculation on the impact of the mutation in the promoter region on the transcription process of the complementary sense genes, leading to a reduction in viral replication. Nevertheless, in a separate investigation, the alignment of the LIR sequences of pMSV-KL(Mild) and pMSV-Km (Severe) revealed the presence of a Rep/RepA promoter harboring a TATA sequence (Edema, 2001). In order to investigate whether variations in the LIR and 5’ end of the Rep/RepA ORFs were responsible for the observed differences in MSV symptom severity between the two clones, the present study conducted VPIs on maize hybrid P3379 kernels. Chimeric clones pMSV-Km (KLLIR) and pMSV-KL (KmLIR) were utilized, wherein the LIR and the 5’-terminus of the Rep A/Rep ORF were exchanged between the severe and mild parental clones. The symptoms exhibited by the two chimeric clones were found to be intermediate in severity compared to those of the recipient parental clones. The observed intermediate phenotype suggests that the influence of LIR alone on symptom severity was only limited. Therefore, our findings align with prior research. However, several research conducted thus far have linked the LIR to the severity of MSV symptoms, albeit to various extents (Schnippenkoetter et al., 2001). The results of these investigations indicate that there are variations in the determinants found in the LIR across MSV clones exhibiting different levels of symptom severity.
The BglII-Nsi1 segment, encompassing the complete C2 open reading frame (ORF) and a significant portion of the RepA ORF, was found to be a predictor of symptom intensity. Clones harboring this fragment exhibited exacerbated symptoms and elevated levels of viral CP antigen in leaf tissues. The severity of symptoms shown by the pMSV-Km (KL RepA) variant was comparable to that of the donor clone pMSV-KL (Mild), whereas the pMSV-KL (Km RepA) variant had a symptom severity equal to that of the donor pMSV-Km (Severe). The observed link between the severity of symptoms and the accumulation of CP antigen implies a potential association between the development of symptoms and the accumulation of the virus. The aforementioned view aligns with the role of RepA and C2 gene products, specifically RepA and Rep, in the replication of MSV, as described by Palmer and Rybicki in 1998. The activation of transcription of the virion MP and CP genes has been observed in studies using RepA and/or Rep (Roja et al., 2018). The mild phenotype seen in pMSV-KL(Mild) may be attributed to reduced replicative activity of the Rep or RepA proteins. Additional experimentation is required to conduct a detailed mapping of the determinants present within the Rep/RepA open reading frames (ORFs) that contribute to the aforementioned activities and the severity of associated symptoms.
A noteworthy discovery was made about the chimeric clone pMSV-Km (KLMP), whereby it was shown that the sequences derived from pMSV-KL (Mild) were responsible for inducing the most severe symptoms. In contrast, pMSV-KL (KmMP:CP) elicited the least severe symptoms and had relatively modest levels of accumulation. The observed effects can be attributed to variations in either the CP or the SIR, or both, as the MP sequences of pMSV-Km (severe) and pMSV-KL (mild) were found to be similar. The multifunctionality of the CP is evident in its involvement in intracellular processes, as well as its collaboration with the MP in intercellular movement and genome encapsidation (Zhang et al., 2001). On the other hand, the SIR serves a dual purpose as the origin of replication for complementary-sense DNA and as a site for polyadenylation and termination of transcription (Willment et al., 2007).
The present findings demonstrate a correlation between the exacerbation of symptom severity and an elevation in viral CNS protein antigen levels. The potential impact of viral infiltration into host tissues may have played a role in the observed rise in concentration, as it might have strengthened the contact between the CP and the MP, so promoting intercellular movement. Moreover, it is possible that the effects of CP on the conversion of double-stranded DNA to single-stranded DNA may have been expedited. Previous research by Palmer and Rybicki (2001) proposed that mutations occurring at the polyadenylation signal of the SIR may potentially diminish the processing efficiency and/or stability of viral messenger ribonucleic acids (mRNAs), hence influencing the virulence of MSV. The findings provided in this study support the concept that the genomes of naturally occurring strains of MSV include numerous nucleotides present in both coding and noncoding regions, which have a role in determining the severity of symptoms associated with MSV. In contrast, Boulton et al. (1991) reached the conclusion that the intensity of symptoms is governed by a solitary nucleotide present in the LIR.
The inclusion of the MP genome fragment from pMSV-KL(Mild) in the chimera resulted in an exacerbation of symptom severity observed in pMSV-Km(Severe). Conversely, the MP genome fragment derived from pMSV-Km(Severe) attenuated the mild symptoms associated with pMSV-KL(Mild). Consequently, the substitution of the fragment resulted in contrasting effects in both the severe and mild clones. The observed impact is somewhat unexpected, as one could have anticipated that the fragment derived from the severe clone would amplify symptom manifestation in the mild clone, and conversely, the fragment from the moderate clone would have a similar effect on the severe clone. Considering the established functions of the genome components within the fragment, namely cell-to-cell movement (MP), nucleus ingress and egress, encapsidation (CP), and genome replication (SIR), it is reasonable to anticipate that these components derived from the severe clone would exhibit superior performance in executing these functions compared to those obtained from the mild clone. Given the available information, it appears that this impact did not occur, suggesting the potential involvement of additional effects related to the components. The available information suggests that the symptom result is determined by the compatibility of the interaction between the components of the fragment and the recipient genome, as well as host variables. Therefore, in the case of the mild clone fragment, the functions were more effectively achieved compared to its individual components, however in the case of the severe clone fragment, the achievement was less effective than that of its individual components. Put simply, the impact of these elements is contingent upon their interaction with the other components inside the genome. It may be premature to engage in speculation on the nature of these interactions, as there is currently a lack of data supporting the notion that these interactions are directly influencing the intensity of symptoms.
An alternative hypothesis is that these pieces have varying impacts on the host plant’s reaction to the clone. One or more of the components of the fragment may impact the plant’s capacity to inhibit the replication of viral genomes via a process known as posttranscriptional gene silencing (PTGS). Therefore, it may be inferred that the impact of the severe clone fragment on this mechanism would be comparatively lesser when compared to the mild clone. Bisaro (2006) shown that post-transcriptional gene silencing (PTGS) is facilitated by small interfering RNAs (siRNAs) and microRNAs (miRNAs). Nevertheless, it is important to note that there is currently no available information regarding any such demonstration conducted on MSV or mastrevirus proteins. The experiment yielded consistent results in terms of the varying severities of symptoms observed in the chimeras. These results were replicated both within and across trials, indicating a high level of reproducibility. The latter observation suggested that the observed effects were heritable features encoded within the genomes of the chimeras. The assessment of symptom severity did not include an examination of the influence of host plant genotype and environmental factors. Nevertheless, a previous investigation conducted by Redinbaugh et al. (2001) yielded contrasting results, as no disparities were observed in the relative severities of symptoms among MSV isolates obtained from Kenya. These isolates were collected from various maize genotypes, ranging from highly susceptible to moderately resistant to resistant. This finding suggests that symptom severity is likely governed by viral genetic factors rather than being primarily influenced by the genotype of the host plant. Furthermore, a notable observation was made in one instance during the current study. Maize plants that were inoculated with these chimeras or their parental clones and subjected to temporary exposure to elevated light and temperature conditions exhibited enhanced plant growth. Consequently, the severity of symptoms was correspondingly alleviated for all clones. This finding suggests that the severity of symptoms may be influenced by growth conditions, but not the relative severity among different clones.
The heightened severity of symptoms seen in pMSV-Km (KLMP:CP) indicates a potential mechanism for the exacerbation of severity in naturally occurring MSV populations. The presence of naturally occurring mixed infections has been documented in previous studies (Owor et al., 2007). In addition, the occurrence of a recombination event, together with the subsequent effective establishment and proliferation of the recombinant genome within the population, is necessary for the manifestation of its impact on the population. The postulated events have been substantiated by existing evidence (Owor et al., 2007). There is existing evidence, as reported by Martin et al. (2015) and van der Walt et al. (2008), that supports the presence of variability in genome sequences among wild populations of MSV.
The current findings presented in this study further illustrate the existence of sequences that have the capacity to cause increased severity through recombination. Nevertheless, the absence of evidence supporting the coexistence of these sequences within a natural MSV population is apparent. Previous investigations (Owor et al., 2007) have shown evidence of recombination occurring between MSV and mastrevirus genomes. However, it should be noted that none of the identified recombinant sequences in previous studies were as extensive as the ones discussed in the current paper. Furthermore, it was shown that none of the recombinants had a higher propensity to induce more severe symptoms on maize compared to closely comparable nonrecombinant viruses (Martin et al., 2002). The potential impact of recombination on symptom severity was not documented for either the recombinant genomes or the recombinant MSV genomes within mixed wild populations.
Although there have been no reports of increased disease severity resulting from recombination among sequences of MSV genomes in field-collected isolates, it is worth noting that recombination is observed to be prevalent among certain naturally occurring begomoviruses (Shi et al., 2014). This phenomenon has been suggested to be a contributing factor in the emergence of various severe begomovirus diseases (Varsani et al., 2008). The recombination event between the begomovirus species East African cassava mosaic virus (EACMV) and African cassava mosaic virus (ACMV), and their subsequent infection of cassava (Manihot esculenta Crantz) in East Africa (Uganda) and West Africa (Cameroon) is of particular significance for the African continent (Pita et al., 2001). The recombination events encompassed both pseudo-recombinants, which occurred between two strains of East African cassava mosaic virus (EACMV), namely EACMV-UG2 (DNA A) and EACMV-UG3 (DNA B), and interspecific recombination, which took place between EACMV-UG2 DNA-A and DNA-A of African cassava mosaic virus (ACMV), specifically inside the CP ORF (coding region) of the former genome. This interspecific recombination involved a segment of 459 nucleotides. Nevertheless, it was improbable that this recombinant strain was accountable for the heightened severity of the sickness (Munoz et al., 1997). The latter phenomenon arose as a result of the concurrent infections of East African cassava mosaic virus (EACMV) and African cassava mosaic virus (ACMV), which elicited a synergistic reaction leading to an exacerbation in symptom severity. Moreover, the presence of the recombinant was detected in both moderate and severe strains of EACMV-UG2, as reported by Pita et al. (2001). This finding provides additional evidence that the recombination event did not significantly influence the degree of symptoms observed in the synergistic interaction with ACMV.
However, it is worth noting that there are potential for recombination. The presence of genetically varied yet closely related viral genomes has been seen in natural isolates of MSV (van der Walt et al., 2008). Additionally, co-infections involving various mastreviruses infecting the same host have been documented (Willment et al., 2007). The findings of the aforementioned study by Martin and Shepherd (2009) indicate that the genetic makeup of MSV-Tas and MSV-VM may consist of genetic material derived from many viral species. These results provide credence to the aforementioned hypotheses. Moreover, the present study’s results pertaining to the dimeric constructions derived from recombinant clones demonstrate the potential existence of a naturally occurring recombinant variant that exhibits either heightened or diminished symptomatology.
5. Conclusion
The mode of transmission for the sequences responsible for exacerbating the severity of MSV symptoms remains uncertain, as it is unclear whether they are transferred in trans or through recombinational processes involving hypothetical mild and severe parental genomes. Previous studies have demonstrated that infection with MSV can be accomplished by the supplementation of defective MSV movement (Palmer and Rybicki, 2001) or replication functions in trans (Palmer and Rybicki, 2001). These results suggest that the exacerbation of symptom severity caused by factors may have a transitory effect. Nevertheless, the current investigation did not attempt to replicate the observed intensification or reduction in symptom severity among trans individuals by employing infectious clones of pMSV-KL(Mild) and pMSV-Km (Severe) in combination infections. Additional research will be required in order to have a more comprehensive understanding of the viral population dynamics in combination infections involving heterologous MSV genomes (Owor et al., 2007).
Although this study aimed to delineate the genetic factors contributing to symptom severity, it is important to acknowledge that the existence of a genetic basis for symptom severity remains uncertain. In this study, the genotypes examined exhibit diverse roles in essential viral life cycle processes, including genome replication and intra- and intercellular migration. Consequently, these genetic factors have an impact on the severity of symptoms. Nevertheless, it is conceivable that these genetic factors have a direct impact on the host plant, leading to variations in the severity of symptoms, therefore enabling the aforementioned essential activities. In other terms, these genetic components have the potential to establish the host plant’s environment, which is evident in the intensity of symptoms, therefore facilitating the replication of the virus and its systemic infection within the plant
Author Contributions
Conceptualization, R.E and D.T.G; Writing-original draft preparation; R.E; Analysis, R.E Original writing, R.E; Writing-review and editing, R.E and D.T.G All authors have read and agreed to the published version of the manuscript.
Funding
Funding was part of the first author’s student fellowship from the Rockefeller Foundation.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Complete genome sequence data for clones MSV-Km has been deposited in GenBank/EMBL/DDBJ under accession numbers AF395891.GI: 18996461.
Acknowledgments
We thank Margret Redingburg, Dave Fulton, Kristen Willie and John Abt for invaluable technical support and Prof. N. H. Gordon (Case Western Reserve University, School of Medicine, Cleveland, Ohio 44106) for advice on statistical analyses. We thank Emmanuel Amphonsah for review of various versions of the Manuscript. The first author was on student fellowship from the Rockefeller Foundation. MSV was held and manipulated under APHIS and MUA permits. 43266 and 99R0033, respectively.
Competing Interests
The authors declare no conflict of interest.
References
- Bisaro, D.M (2006). Silencing suppression by geminivirus proteins. Virology, 344(1), 158–168. [CrossRef]
- Bosque-Pérez, NA (2000). Eight decades of maize streak virus research. Virus Research, 71(1–2), 107–121. [CrossRef]
- Boulton, MI, King, DI, Donson, J, & Davies, JW (1991). Point substitutions in a promoter-like region and the V1 gene affect the host range and symptoms of maize streak virus. Virology, 183, 114–121.
- Boulton, MI, King, DI, Markham, PG, Pinner, MS, & Davies, JW (1991). Host range and symptoms are determined by specific domains of the maize streak virus genome. Virology, 181(1), 312–318. [CrossRef]
- Edema, R (2001). The genetics of virulence of the maize streak virus (MSV). Dissertation of The Ohio State University.
- Emeraghi, M, Achigan-Dako, EG, Nwaoguala, CNC & Oselebe, H (2021). Maize streak virus research in Africa: an end or a crossroad. Theor Appl Genet 134, 3785–3803. [CrossRef]
- Martin, DP, & Rybicki, E. P. (2002). Investigation of Maize streak virus Pathogenicity Determinants Using Chimaeric Genomes. Virology, 300(2), 180–188. [CrossRef]
- Martin, D. P., & Shepherd, D. N. (2009). The epidemiology, economic impact and control of maize streak disease. Food Security, 1, 305–315.
- Munoz, C., Zhou, X., Otim-Nape, G. W., Liu, Y., Robinson, D. J., Calvert, L., & Harrison, B. D. (1997). Evidence that DNA-A of a geminivirus associated with severe cassava mosaic disease in Uganda has arisen by interspecific recombination. Journal of General Virology, 78(8), 2101–2111. [CrossRef]
- Owor, B. E., Martin, D. P., Shepherd, D. N., Edema, R., Monjane, A. L. and Rybicki, E. P.(2007). Genetic analysis of maize streak virus isolates from Uganda reveals widespread distribution of a recombinant variant. Journal of General Virology, 88 (11), 3154–3165.
- Palmer, K. E., & Rybicki, E. P. (1998). The Molecular Biology of Mastreviruses (pp. 183–234). [CrossRef]
- Palmer, K. E., & Rybicki, EP (2001). Investigation of the potential of Maize streak virus to act as an infectious gene vector in maize plants. Archives of Virology, 146(6), 1089–1104. [CrossRef]
- Pita, JS, Fondong, VN, Sangaree, A, Otim-Nape, GW, Ogwal, S, & Fauquet, CM (2001). Recombination, pseudo recombination and synergism of geminiviruses are determinant keys to the epidemic of severe cassava mosaic disease in Uganda. Journal of General Virology, 82, 655–665.
- Redinbaugh, MG, Louie, R, Ngwira, P, Edema, R, Gordon, DT, & Bisaro, DM (2001). Transmission of viral RNA and DNA to maize kernels by vascular puncture inoculation. Journal of Virological Methods, 98(2), 135–143. [CrossRef]
- Schnippenkoetter, WH, Martin, DP, Hughes, FL, Fyvie, M, Rojas, MA, Macedo, MA, Maliano, MR, Soto-Aguilar, M, Souza, JO, Briddon, RW, Kenyon, L, Rivera Bustamante, RF, Murilo Zerbini, F, Adkins, S, Legg, JP, Kvarnheden, A, Wintermantel, WA, Sudarshana, MR, Peterschmitt, M, Lapidot, M, Martin, DR, Moriones, E, Inoue-Nagata, AK and Gilbertson, RL (2018). World Management of Geminiviruses. Annual Review of Phytopathology 56:1, 637-677.
- Shen, WH, & Hohn, B (1991). Mutational analysis of the small intergenic region of maize streak virus. Virology, 183(2), 721–730. [CrossRef]
- Shepherd, DN, Martin, DP, van der Walt, E, Dent, K, Varsani, A, & Rybicki, EP (2010). Maize streak virus: an old and complex ‘emerging’’ pathogen.’ Molecular Plant Pathology, 11, 1–12.
- Sime SS, Menkir A, Adetimirin VO, Gedil M andKumar PL. Validation of Diagnostic Markers for Streak Virus Disease Resistance in Maize. Agriculture. (2021) 11(2):130. [CrossRef]
- Todd, JC, Ammar, ED, Redinbaugh, MG, Hoy, C, & Hogenhout, SA. (2010). Plant host range and leafhopper transmission of Maize fine streak virus. Phytopathology, 100, 1138–1145.
- Unseld, S, Ringel, M, Höfer, P, Höhnle, M, Jeske, H, Bedford, ID, Markham, PG, & Frischmuth, T (2000). Host range and symptom variation of pseudorecombinant virus produced by two distinct bipartite geminiviruses. Archives of Virology, 145(7), 1449–1454. [CrossRef]
- van der Walt, E, Martin, DP, Varsani, A, Polston, JE, & Rybicki, EP (2008). Experimental observations of rapid Maize streak virus evolution reveal a strand-specific nucleotide substitution bias. Virology Journal, 5(1), 104. [CrossRef]
- Willment, JA, James, D, von Wechmar, MB, & Rybicki, EP (2001). The relative infectivities and genomic characterisation of three distinct mastreviruses from South Africa. Archives of Virology, 146(6), 1075–1088. [CrossRef]
- Willment, JA, Martin, DP, Palmer, KE, Schnippenkoetter, WH, Shepherd, DN, & Rybicki, EP (2007). Identification of long intergenic region sequences involved in maize streak virus replication. Journal of General Virology, 88(6), 1831–1841. [CrossRef]
- Zhang, W, Olson, NH, Baker,T S, Faulkner, L, Agbandje-McKenna, M, Boulton, MI, Davies, JWand McKenna, R (2001). Structure of the Maize streak virus geminate particle.Virology 279(2):471-7. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).