Submitted:
09 November 2023
Posted:
10 November 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Ethical Approval
2.2. Experimental Animals and Testis Tissue Sample Collection
2.3. Determination of Hormonal Concentration
2.4. Histological Assessment
2.5. RNA Extraction and Qualification
2.6. Quality Control of Sequence Reads
2.7. Reads Mapping and Sequence Analysis
2.8. Predicting miRNAs
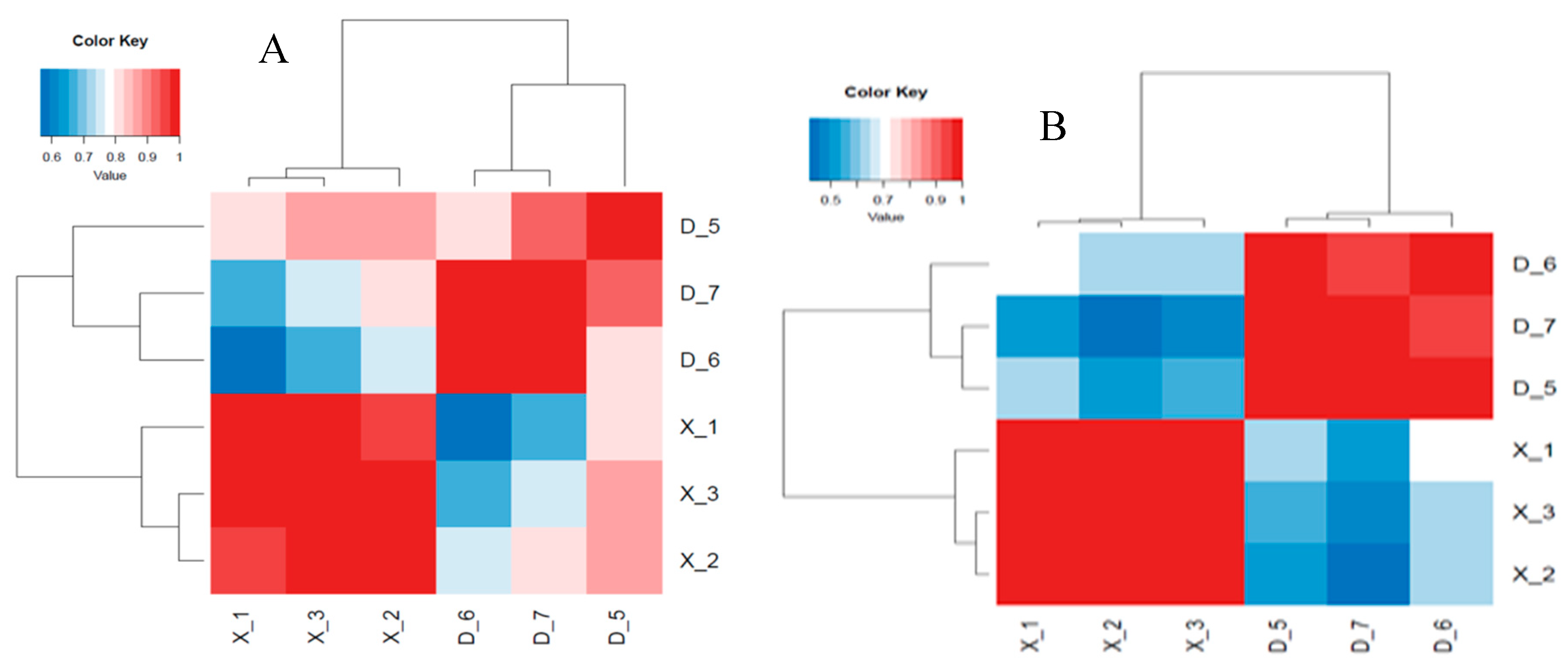
2.9. Expression Normalization and Correlation Analysis of Samples
2.10. Differential Expression Analysis
2.11. Target Prediction and Function Annotation
2.12. Network Analysis of Differentially Expressed miRNA Target Genes
2.13. RT-qPCR Validation and Statistical Analysis
| Gene name | Sequence of primers (5’- 3’) | Product length (bp) | Tm (◦C) |
|---|---|---|---|
| THAP3 | F: TCTGGGGAGAGAAAGGTCCTC R: AGGGCATAGCTGTGATCGGA |
227 | 60.27 |
| FOSB | F: GAGAAGAGAAGGGTTCGCCG R: CTAGCTGATCTGTCTCCGCC |
106 | 60.46 |
| SLC2A5 | F: CTAGCTGATCTGTCTCCGCC R: GTCGACGGTGGAAACTCCTT |
203 | 60.03 |
| CCNT2 | F: ACCCAGTTAGTAAGAGCAAGCA R: CAGGGATCTCCCAATTGGACC |
160 | 59.36 |
| CMC1 | F: ACCCCTCAGAGCAGCAT R: TGGGAAGCTTCTGTAGCCTTT |
300 | 57.33 |
| GAPDH | F: CTTCGGCATTGTGGAGGG R: GGAGGCAGGGATGATGTTCT |
130 | 61.3 |
| MiRNA ID (5′-3′) | Stem-loop RT primer (5′-3′) | MiRNA specific forward primer |
|---|---|---|
| bta-miR-11971 | 5’--GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACTTCTCA --3’ | 5’--CGCGTGAGGGGCAGAGAG --3’ |
| bta-miR-2284c | 5’-- GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACAAAACC --3’ | 5’--GCGCGAAAAAGTTCGTTTT --3’ |
| bta-miR-3956 | 5’--GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACCCTCTG --3’ | 5’--CGACGTGGATGCTGAAGGT --3’ |
| bta-miR-135b | 5’--GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACTCACAT --3’ | 5’--CGCGTATGGCTTTTCATTCCT --3’ |
| bta-miR-299-2 | 5’--GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACATGGTT --3’ | 5’--CGCGGTATGTGGGACGGTA --3’ |
| bta-miR-433 | 5’--GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACACACCG --3’ | 5’--GCGATCATGATGGGCTCCT --3’ |
| bta-miR-191 | 5’--GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACCAGCTG --3’ | 5’--CGCAACGGAATCCCAAAAG --3’ |
| bta-miR-20a | 5’--GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACCTACCT --3’ | 5’--GCGCGTAAAGTGCTTATAGTGC --3’ |
| 4_37054-5p(cli-miR-9632-3p) | 5’--GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACAGTCCG --3’ | 5’--GCGAACTTTTGCCCCTAGTAA --3’ |
| 7_42582-5p | 5’--GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACTGTTAG --3’ | 5’--GCGCGTGAGAGGTCTGTAATAA --3’ |
| 6_40791-3p | 5’--GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACGGTCCC --3’ | 5’--GCGTGAGTGTGCATCCACG --3’ |
| 21_23590-5p(mmu-miR-665-3p) | 5’--GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACGCTCAT --3’ | 5’-- CGCCCAGGAGAATGAGGG --3’ |
3.2. Results
3.2.1. Hormonal Concentration
3.2.2. Testicular Tissue Morphologic Characteristics of Six Samples
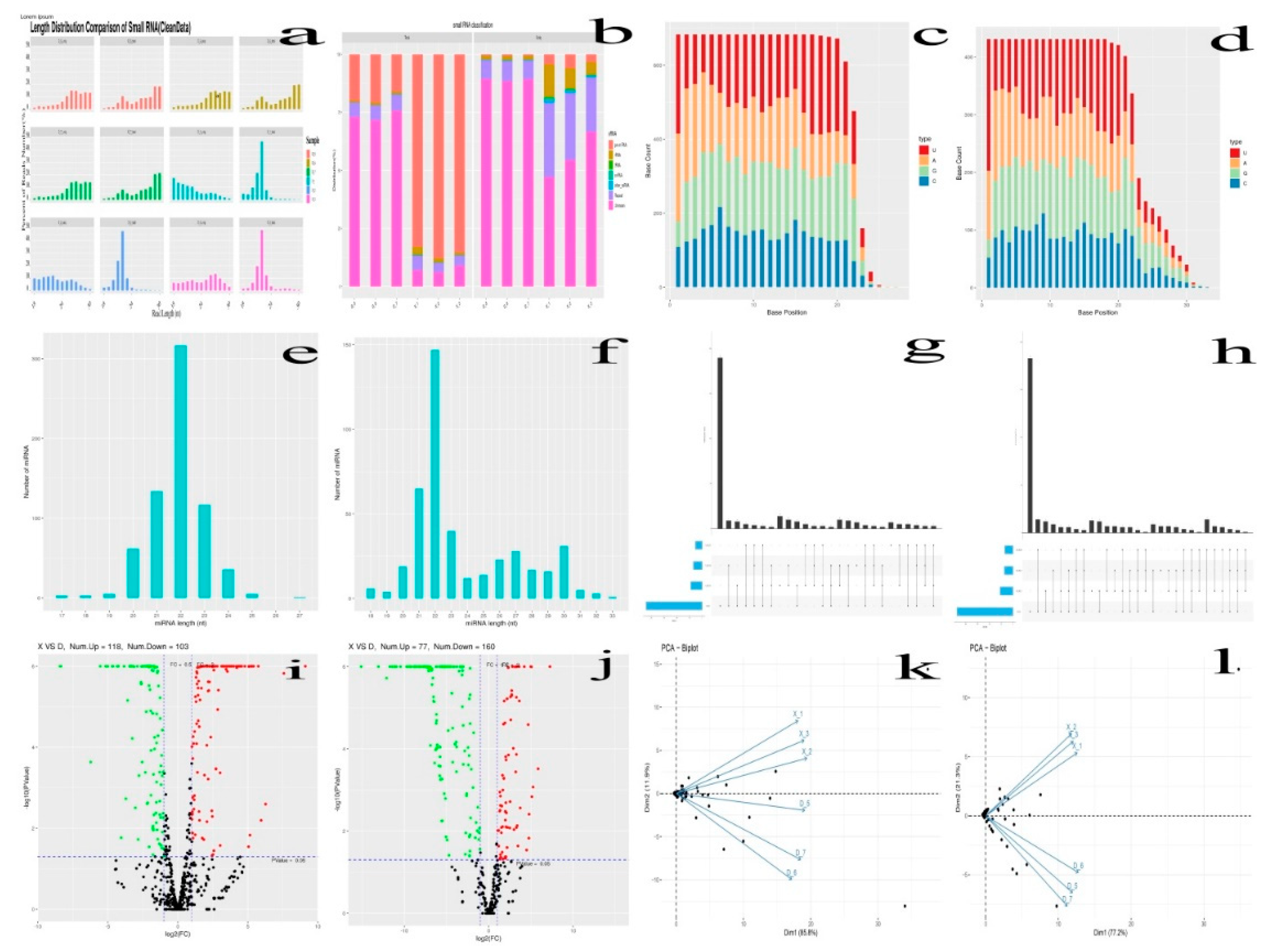
3.3.2. Quality Control
3.3.3. Reads Mapping and Filtration
3.3.5. The miRNAs Family and Expression Level Analysis
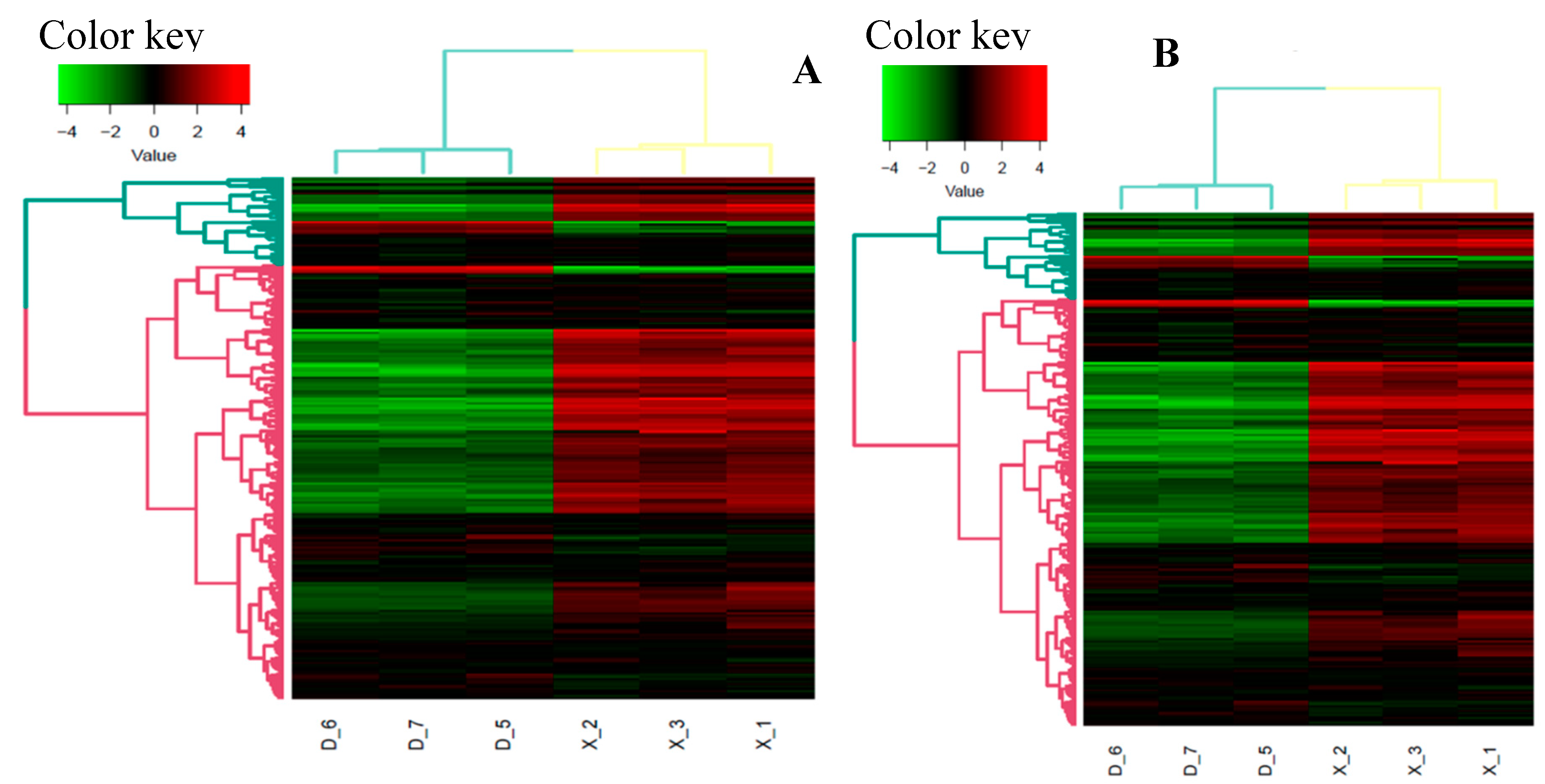
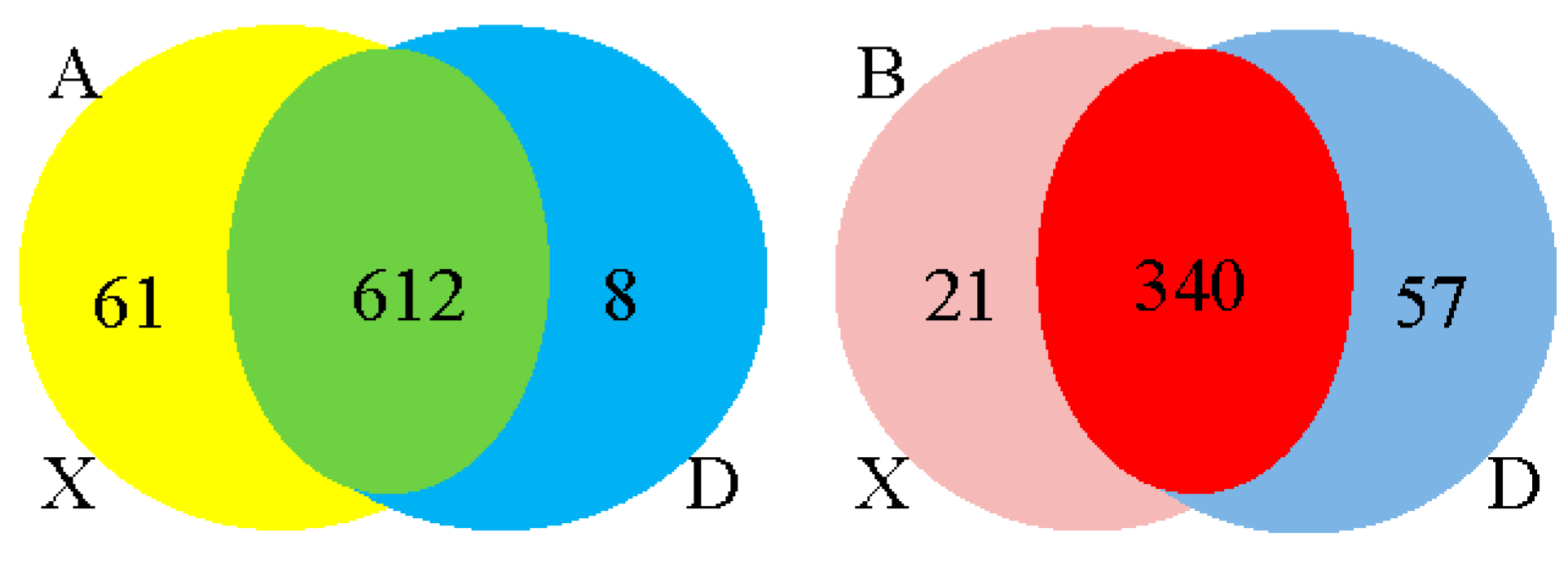
3.3.6. Differential Expression Analysis of miRNAs and Target Genes
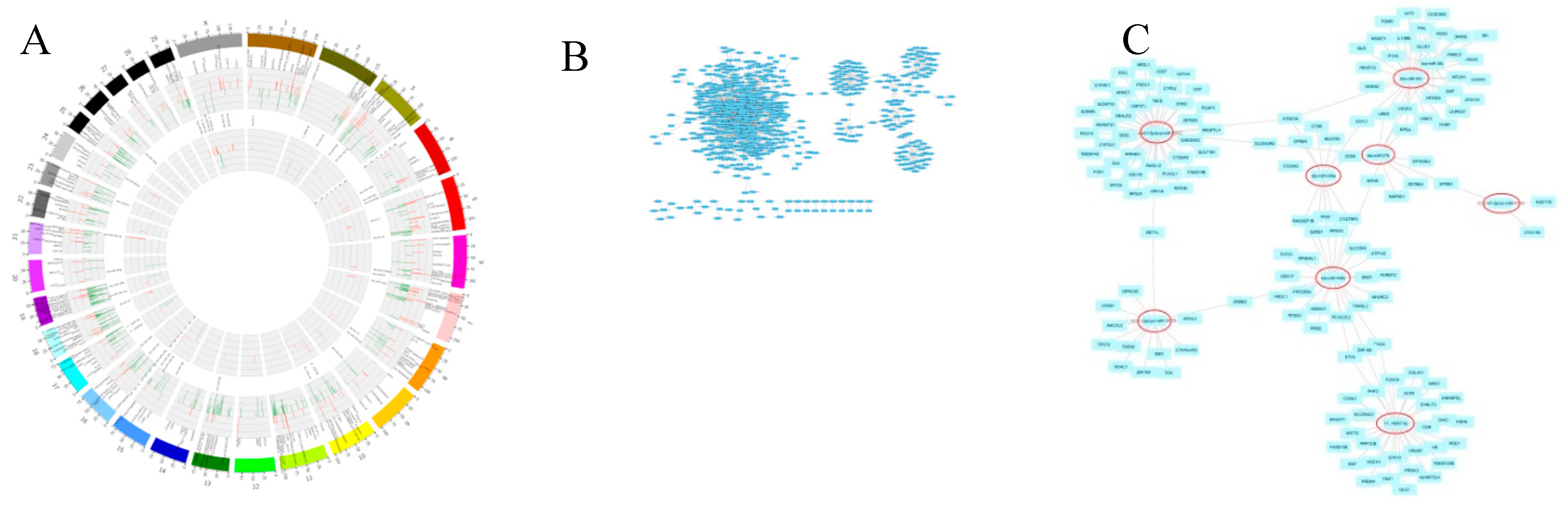
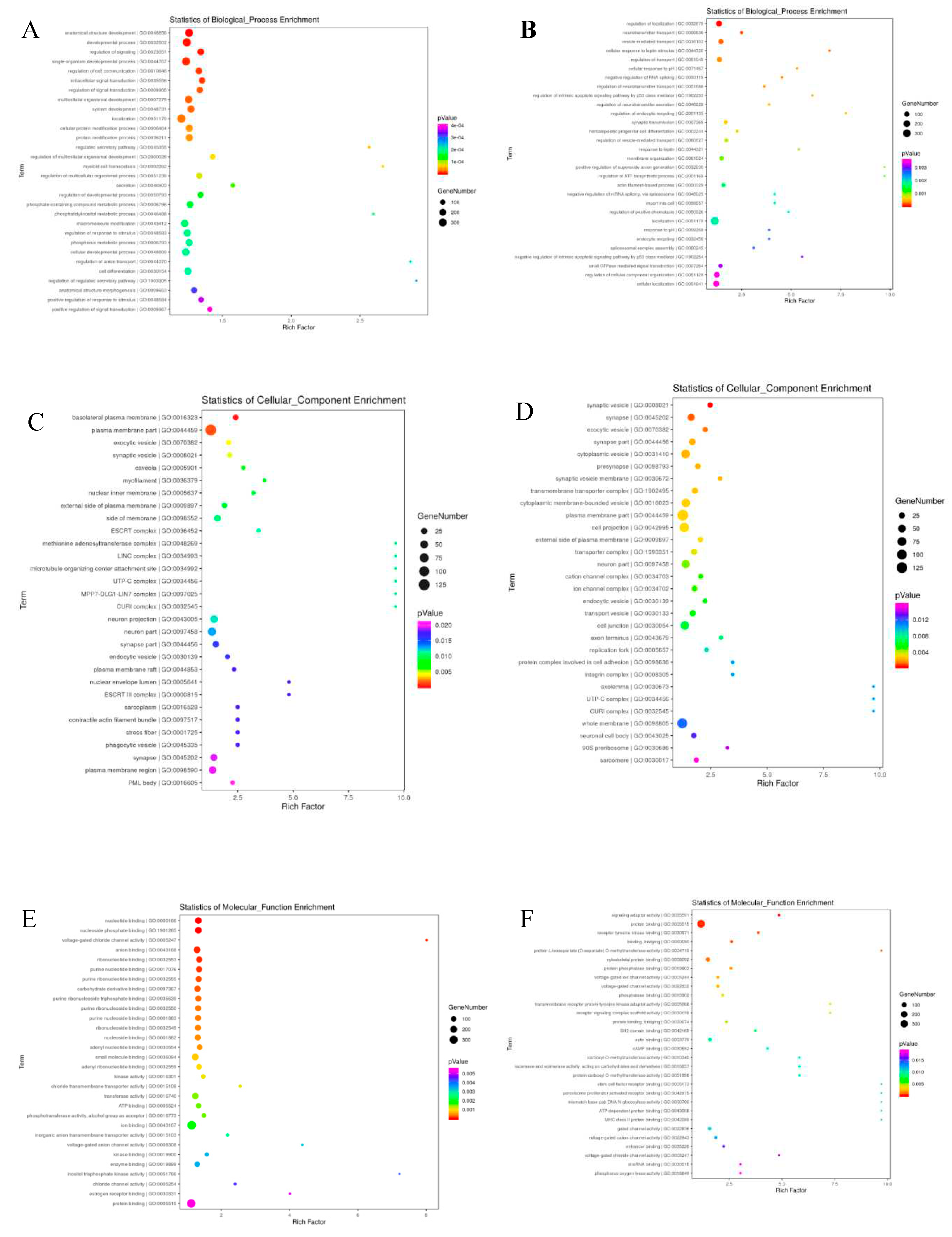
3.3.7. Target Prediction, Functional Enrichment, and Network Analysis
3.3.8. Functional Genes of Differentially Expressed (DE) in the GO Terms
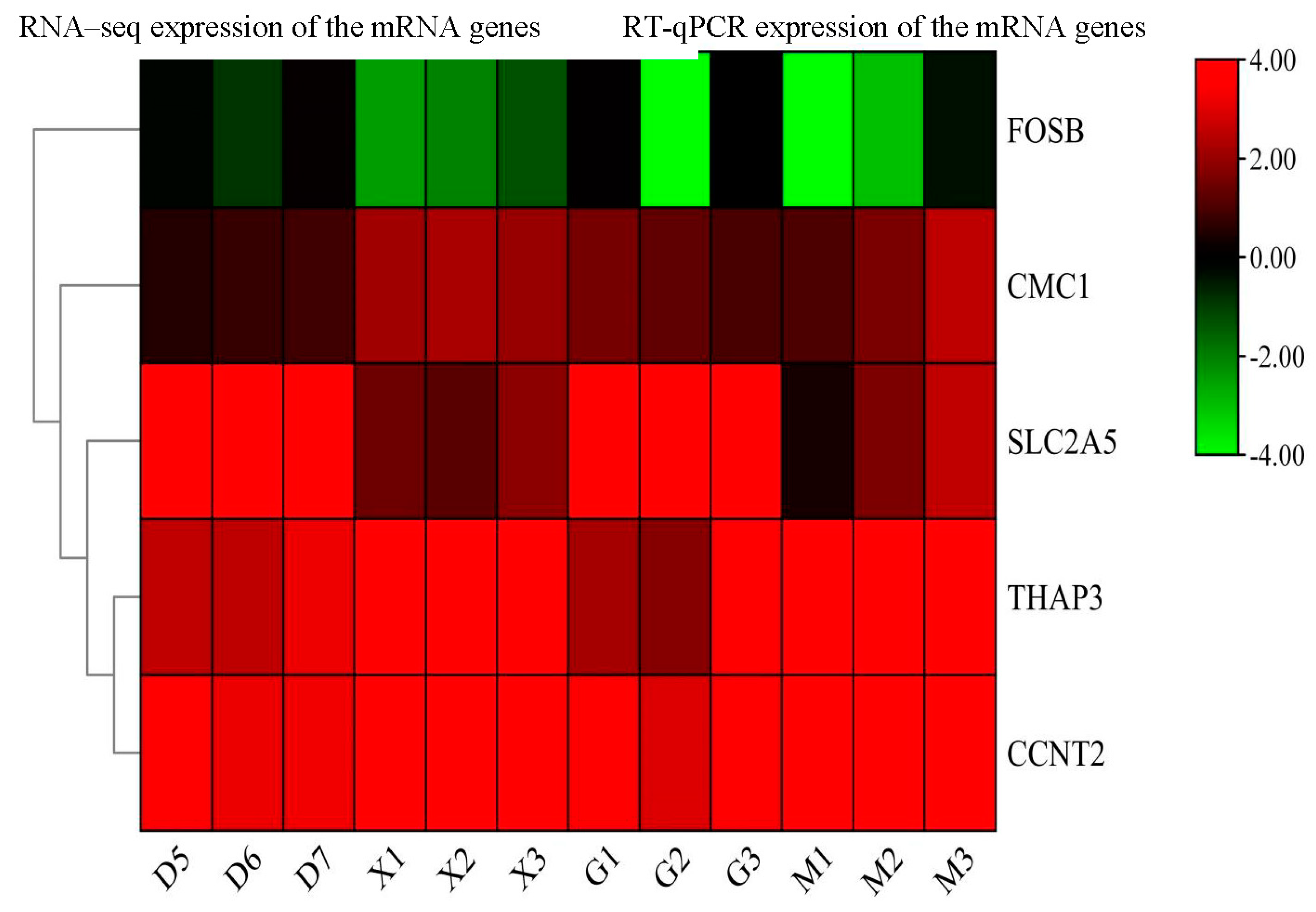
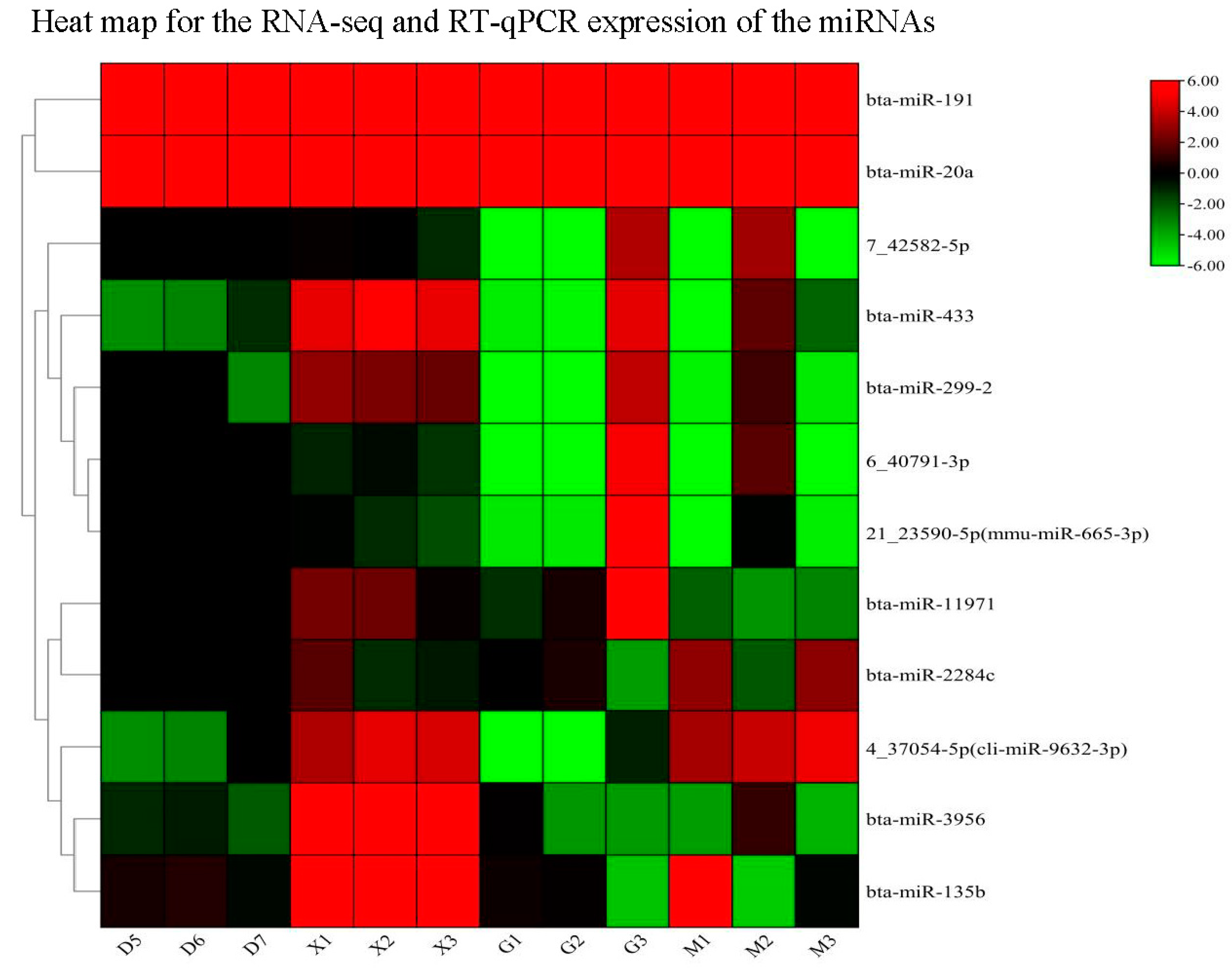
3.3.9. RT-qPCR Validation
3.4. Discussion
3.5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ala, U. (2020). Competing Endogenous RNAs, Non-Coding RNAs and Diseases: An Intertwined Story. In Cells (Vol. 9, Issue 7). [CrossRef]
- Ansari-Lari, M., Kafi, M., Sokhtanlo, M., & Ahmadi, H. N. (2010). Reproductive performance of Holstein dairy cows in Iran. Tropical Animal Health and Production, 42(6), 1277–1283. [CrossRef]
- Bai, M., Sun, L., Jia, C., Li, J., Han, Y., Liu, H., Chen, Y., & Jiang, H. (2019). Integrated analysis of miRNA and mRNA expression profiles reveals functional miRNA-targets in development testes of small tail han sheep. G3: Genes, Genomes, Genetics, 9(2), 523–533. [CrossRef]
- Baker, P. J., Pakarinen, P., Huhtaniemi, I. T., Abel, M. H., Charlton, H. M., Kumar, T. R., & O’Shaughnessy, P. J. (2003). Failure of normal Leydig cell development in follicle-stimulating hormone (FSH) receptor-deficient mice, but not FSHbeta-deficient mice: role for constitutive FSH receptor activity. Endocrinology, 144(1), 138–145. [CrossRef]
- Baohong, Qinglian, Wang, A., & Pan, X. (2007). MicroRNAs and Their Regulatory Roles in Animals and Plants. Journal Cellular Physiology, 210(May), 279–289. [CrossRef]
- Buchold, G. M., Coarfa, C., Kim, J., Milosavljevic, A., Gunaratne, P. H., & Matzuk, M. M. (2010). Analysis of microRNA expression in the prepubertal testis. PloS One, 5(12), e15317. [CrossRef]
- Carreau, S., Bouraima-Lelong, H., & Delalande, C. (2012). Role of estrogens in spermatogenesis. Frontiers in Bioscience (Elite Edition), 4(1), 1–11. [CrossRef]
- Cheng, C.-Y., Hwang, C.-I., Corney, D. C., Flesken-Nikitin, A., Jiang, L., Öner, G. M., Munroe, R. J., Schimenti, J. C., Hermeking, H., & Nikitin, A. Y. (2014). miR-34 cooperates with p53 in suppression of prostate cancer by joint regulation of stem cell compartment. Cell Reports, 6(6), 1000–1007. [CrossRef]
- Ding, H., Luo, Y., Liu, M., Huang, J., & Xu, D. (2016). Histological and transcriptome analyses of testes from Duroc and Meishan boars. Scientific Reports, 6(January), 1–13. [CrossRef]
- Fabian, M. R., Sonenberg, N., & Filipowicz, W. (2010). Regulation of mRNA translation and stability by microRNAs. Annual Review of Biochemistry, 79, 351–379. [CrossRef]
- Fang, X., Qin, L., Yu, H., Jiang, P., Xia, L., Gao, Z., Yang, R., Zhao, Y., Yu, X., & Zhao, Z. (2021). Comprehensive analysis of miRNAs and target mRNAS between immature and mature testis tissue in Chinese red steppes cattle. Animals, 11(11). [CrossRef]
- Friedländer, M. R., Mackowiak, S. D., Li, N., Chen, W., & Rajewsky, N. (2012). miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Research, 40(1), 37–52. [CrossRef]
- Gao, Y., Wu, F., Ren, Y., Zhou, Z., Chen, N., Huang, Y., Lei, C., Chen, H., & Dang, R. (2020). MiRNAs Expression Profiling of Bovine (Bos taurus) Testes and Effect of bta-miR-146b on Proliferation and Apoptosis in Bovine Male Germline Stem Cells. International Journal of Molecular Sciences, 21(11). [CrossRef]
- Ge, R.-S., Li, X., & Wang, Y. (2021). Leydig Cell and Spermatogenesis. Advances in Experimental Medicine and Biology, 1288, 111–129. [CrossRef]
- Gong, W., Pan, L., Lin, Q., Zhou, Y., Xin, C., Yu, X., Cui, P., Hu, S., & Yu, J. (2013). Transcriptome profiling of the developing postnatal mouse testis using next-generation sequencing. Science China. Life Sciences, 56(1), 1–12. [CrossRef]
- Grande, G., Barrachina, F., Soler-Ventura, A., Jodar, M., Mancini, F., Marana, R., Chiloiro, S., Pontecorvi, A., Oliva, R., & Milardi, D. (2022). The Role of Testosterone in Spermatogenesis: Lessons From Proteome Profiling of Human Spermatozoa in Testosterone Deficiency. Frontiers in Endocrinology, 13, 852661. [CrossRef]
- Han, Y., & Peñagaricano, F. (2016). Unravelling the genomic architecture of bull fertility in Holstein cattle. BMC Genetics, 17(1), 143. [CrossRef]
- Hayashi, K., Chuva de Sousa Lopes, S. M., Kaneda, M., Tang, F., Hajkova, P., Lao, K., O’Carroll, D., Das, P. P., Tarakhovsky, A., Miska, E. A., & Surani, M. A. (2008). MicroRNA biogenesis is required for mouse primordial germ cell development and spermatogenesis. PLoS ONE, 3(3), 1–9. [CrossRef]
- Huang, J., Ju, Z., Li, Q., Hou, Q., Wang, C., Li, J., Li, R., Wang, L., Sun, T., Hang, S., Gao, Y., Hou, M., & Zhong, J. (2011). Solexa sequencing of novel and differentially expressed microRNAs in testicular and ovarian tissues in Holstein cattle. International Journal of Biological Sciences, 7(7), 1016–1026. [CrossRef]
- Johnston, H., Baker, P. J., Abel, M., Charlton, H. M., Jackson, G., Fleming, L., Kumar, T. R., & O’Shaughnessy, P. J. (2004). Regulation of Sertoli cell number and activity by follicle-stimulating hormone and androgen during postnatal development in the mouse. Endocrinology, 145(1), 318–329. [CrossRef]
- Kastelic, J. P., & Thundathil, J. C. (2008). Breeding Soundness Evaluation and Semen Analysis for Predicting Bull Fertility. Reproduction in Domestic Animals, 43(SUPPL.2), 368–373. [CrossRef]
- Kim, D., Langmead, B., & Salzberg, S. L. (2015). HISAT: a fast spliced aligner with low memory requirements. Nature Methods, 12(4), 357–360. [CrossRef]
- Kim, D., Yeong, Sung, & Jong-Hyuk. (2017). Regulatory role of microRNAs in the proliferation and differentiation of adipose-derived stem cells. Histology and Histopathology, 32(1), 1–10. [CrossRef]
- La, Y., Ma, F., Ma, X., Bao, P., Chu, M., Liang, C., Guo, X., Yin, M., Li, J., & Yan, P. (2022). Different expression of LHR, PRLR, GH and IGF1 during testicular development of yak. Reproduction in Domestic Animals = Zuchthygiene, 57(2), 221–227. [CrossRef]
- La, Y., Ma, X., Bao, P., Chu, M., Guo, X., Liang, C., & Yan, P. (2023). Identification and profiling of microRNAs during yak’s testicular development. BMC Veterinary Research, 19(1), 53. [CrossRef]
- Li, Yao, Li, J., Fang, C., Shi, L., Tan, J., Xiong, Y., Fan, B., & Li, C. (2016). Genome-wide differential expression of genes and small RNAs in testis of two different porcine breeds and at two different ages. Scientific Reports, 6, 26852. [CrossRef]
- Li, Z., Lan, X., Guo, W., Sun, J., Huang, Y., Wang, J., Huang, T., Lei, C., Fang, X., & Chen, H. (2012). Comparative Transcriptome Profiling of Dairy Goat MicroRNAs from Dry Period and Peak Lactation Mammary Gland Tissues. PLoS ONE, 7(12). [CrossRef]
- Liang, X., Zhou, D., Wei, C., Luo, H., Liu, J., Fu, R., & Cui, S. (2012). MicroRNA-34c enhances murine male germ cell apoptosis through targeting ATF1. PloS One, 7(3), e33861. [CrossRef]
- Liu, H., Khan, I. M., Liu, Y., Khan, N. M., Ji, K., Yin, H., Wang, W., Zhou, X., & Zhang, Y. (2022). A Comprehensive Sequencing Analysis of Testis-Born miRNAs in Immature and Mature Indigenous Wandong Cattle (Bos taurus). Genes, 13(12). [CrossRef]
- Lu, Q., Gore, M., Zhang, Q., Camenisch, T., Boast, S., Casagranda, F., Lai, C., Skinner, M. K., Klein, R., Matsushima, G. K., Earp, H. S., Goff, S. P., & Lemke, G. (1999). Tyro-3 family receptors are essential regulators of mammalian spermatogenesis. Nature, 398(6729), 723–728. [CrossRef]
- Luo, G., Hu, S., Lai, T., Wang, J., Wang, L., & Lai, S. (2020). MiR-9-5p promotes rabbit preadipocyte differentiation by suppressing leptin gene expression. Lipids in Health and Disease, 19(1), 126. [CrossRef]
- Mäkelä, J.-A., Koskenniemi, J. J., Virtanen, H. E., & Toppari, J. (2019). Testis Development. Endocrine Reviews, 40(4), 857–905. [CrossRef]
- Navarro, F., & Lieberman, J. (2015). miR-34 and p53: New Insights into a Complex Functional Relationship. PloS One, 10(7), e0132767. [CrossRef]
- Niu, Z., Goodyear, S. M., Rao, S., Wu, X., Tobias, J. W., Avarbock, M. R., & Brinster, R. L. (2011). MicroRNA-21 regulates the self-renewal of mouse spermatogonial stem cells. Proceedings of the National Academy of Sciences of the United States of America, 108(31), 12740–12745. [CrossRef]
- Sahlu, B. W., Zhao, S., Wang, X., Umer, S., Zou, H., & Huang, J. (2020). Long noncoding RNAs: new insights in modulating mammalian spermatogenesis. 8, 1–12. [CrossRef]
- Salzman, D. W., Nakamura, K., Nallur, S., Dookwah, M. T., Metheetrairut, C., Slack, F. J., & Weidhaas, J. B. (2016). miR-34 activity is modulated through 5’-end phosphorylation in response to DNA damage. Nature Communications, 7, 10954. [CrossRef]
- Schulster, M., Bernie, A. M., & Ramasamy, R. (2016). The role of estradiol in male reproductive function. Asian Journal of Andrology, 18(3), 435–440. [CrossRef]
- Smith, L. B., & Walker, W. H. (2014). The regulation of spermatogenesis by androgens. Seminars in Cell & Developmental Biology, 30, 2–13. [CrossRef]
- Staub, C., & Johnson, L. (2018). Review: Spermatogenesis in the bull. 1–9. [CrossRef]
- Sun, H. Z., Chen, Y., & Guan, L. L. (2019). MicroRNA expression profiles across blood and different tissues in cattle. Scientific Data, 6(1), 1–8. [CrossRef]
- Vigneault, F., Ter-Ovanesyan, D., Alon, S., Eminaga, S., Christodoulou, D. C., Seidman, J. G., Eisenberg, E., & Church, G. M. (2012). High-throughput multiplex sequencing of miRNA. Current Protocols in Human Genetics, SUPPL.73. [CrossRef]
- Wang, H., Zhong, J., Chai, Z., Zhu, J., & Xin, J. (2018). Comparative expression profile of microRNAs and piRNAs in three ruminant species testes using next-generation sequencing. Reproduction in Domestic Animals, 53(4), 963–970. [CrossRef]
- Waqas, M. S., Ciccarelli, M., Oatley, M. J., Kaucher, A. V., Tibary, A., & Oatley, J. M. (2019). Enhanced sperm production in bulls following transient induction of hypothyroidism during prepubertal development. Journal of Animal Science, 97(4), 1468–1477. [CrossRef]
- Wu, J., Zhu, H., Song, W., Li, M., Liu, C., Li, N., Tang, F., Mu, H., Liao, M., Li, X., Guan, W., Li, X., & Hua, J. (2014). Identification of conservative MicroRNAs in saanen dairy goat testis through deep sequencing. Reproduction in Domestic Animals, 49(1), 32–40. [CrossRef]
- Xu, C., Wu, S., Zhao, W., Mipam, T., Liu, J., Liu, W., Yi, C., Shah, M. A., Yu, S., & Cai, X. (2018). Differentially expressed microRNAs between cattleyak and yak testis. Scientific Reports, 8(1), 1–12. [CrossRef]
- Yao, C., Liu, Y., Sun, M., Niu, M., Yuan, Q., Hai, Y., Guo, Y., Chen, Z., Hou, J., Liu, Y., & He, Z. (2015). MicroRNAs and DNA methylation as epigenetic regulators of mitosis, meiosis and spermiogenesis. Reproduction (Cambridge, England), 150(1), R25-34. [CrossRef]
- Yao, Qian, Chen, Y., & Zhou, X. (2019). The roles of microRNAs in epigenetic regulation. Current Opinion in Chemical Biology, 51, 11–17. [CrossRef]
- Yin, Z., Xu, X., Tan, Y., Cao, H., Zhou, W., Dong, X., & Mao, H. (2021). Expression analysis of microRNAs and their target mRNAs of testes with high and low sperm motility in domestic pigeons (Columba livia). Genomics, 113(1), 257–264. [CrossRef]
- Zhang, B., Yan, Z., Gao, Y., Li, J., Wang, Z., Wang, P., Yang, Q., Huang, X., & Gun, S. (2022). Integrated analysis of miRNA and mRNA expression profiles in testes of Landrace and Hezuo boars. Frontiers in Veterinary Science, 9. [CrossRef]
- Zhao, S., Heng, N., Sahlu, B. W., Wang, H., & Zhu, H. (2021). Long noncoding rnas: Recent insights into their role in male infertility and their potential as biomarkers and therapeutic targets. International Journal of Molecular Sciences, 22(24). [CrossRef]
- Zhao, S., Sun, W., Chen, S.-Y., Li, Y., Wang, J., Lai, S., & Jia, X. (2022). The exploration of miRNAs and mRNA profiles revealed the molecular mechanisms of cattle-yak male infertility. Frontiers in Veterinary Science, 9(5). [CrossRef]
- Zhao, S., Wang, H., Hu, Z., Sahlu, B. W., Heng, N., Gong, J., Wang, H., & Zhu, H. (2022). Identification of spermatogenesis-related lncRNA in Holstein bull testis after sexual maturity based on transcriptome analysis. Animal Reproduction Science, 247(October), 107146. [CrossRef]



| Sample | T (Testosterone) (ng/mL) | Average | Standard Deviation | Estrogen (ng/mL) |
Average | Standard Deviation | Progesterone (ng/mL) |
Average | Standard Deviation |
|---|---|---|---|---|---|---|---|---|---|
| X1 | 0.002 | 0.003 | 0.001 | 65.05 | 62.12 | 7.07 | 0.12 | 0.12 | 0.01 |
| X2 | 0.004 | 54.05 | 0.11 | ||||||
| X3 | 0.004 | 67.25 | 0.13 | ||||||
| D5 | 0.199 | 0.166 | 0.078 | 73.03 | 57.22 | 21.41 | 0.14 | 0.16 | 0.04 |
| D6 | 0.222 | 32.85 | 0.14 | ||||||
| D7 | 0.076 | 65.78 | 0.2 |
| Sample | Raw reads number | Raw bases | Clean reads number | Clean bases | Clean rate (%) | Q20 (%) | Q30 (%) |
|---|---|---|---|---|---|---|---|
| D_5 | 86639342 | 12995901300 | 85382674 | 12773620656 | 98.29 | 98.72 | 95.89 |
| X_1 | 84967166 | 12745074900 | 83607760 | 12511803713 | 98.17 | 98.6 | 95.59 |
| D_6 | 89752696 | 13462904400 | 88281454 | 13216123838 | 98.17 | 97.98 | 93.77 |
| X_3 | 88444394 | 13266659100 | 87142022 | 13044924344 | 98.33 | 98.58 | 95.55 |
| X_2 | 88046786 | 13207017900 | 86695100 | 12978591207 | 98.27 | 98.56 | 95.42 |
| D_7 | 88614166 | 13292124900 | 87268088 | 13060990754 | 98.26 | 98.71 | 95.86 |
| Sample | Num. of Raw Reads | Clean Reads% | Remove Adapter% | Insert Null% | N% | too_short% | Poly-A% | too_long% | low_quality% |
|---|---|---|---|---|---|---|---|---|---|
| D_5 | 15259349 | 89.05% | 0.02% | 0.00% | 0.10% | 1.54% | 0.17% | 9.05% | 0.08% |
| D_6 | 13463407 | 87.95% | 0.03% | 0.01% | 0.10% | 2.72% | 0.13% | 8.97% | 0.09% |
| D_7 | 14244562 | 85.08% | 0.02% | 0.01% | 0.16% | 2.48% | 0.17% | 11.98% | 0.10% |
| X_1 | 13956441 | 70.09% | 0.02% | 0.04% | 0.11% | 28.52% | 0.05% | 1.06% | 0.11% |
| X_2 | 15869310 | 85.48% | 0.01% | 0.01% | 0.09% | 13.35% | 0.04% | 0.94% | 0.08% |
| X_3 | 19881140 | 91.83% | 0.01% | 0.01% | 0.10% | 6.72% | 0.05% | 1.18% | 0.10% |
| ID | Type of reads | Sum | Hairpin | rRNA | tRNA | snRNA | other_ncRNA | Repeat | Unknown |
|---|---|---|---|---|---|---|---|---|---|
|
D_5 |
Uniq | 2493381(100%) | 16835(0.68%) | 36744(1.47%) | 444(0.02%) | 3092(0.12%) | 3524(0.14%) | 202394(8.12%) | 2230348(89.45%) |
| Total | 13588224(100%) | 2690486(19.8%) | 115201(0.85%) | 802(0.01%) | 6017(0.04) | 7400(0.05%) | 847312(6.24%) | 9921006(73.01%) | |
|
D_6 |
Uniq | 2129032(100%) | 15962(0.75%) | 36429(1.71%) | 493(0.02%) | 3261(0.15%) | 3686(0.17%) | 186561(8.76%) | 1882640(88.43%) |
| Total | 11841624(100%) | 2457259(20.75%) | 128723(1.09%) | 946(0.01) | 6199(0.05%) | 8011(0.07%) | 756444(6.39%) | 8484042(71.65%) | |
|
D_7 |
Uniq | 2197903(100%) | 14767(0.67%) | 35337(1.61%) | 445(0.02%) | 3103(0.14%) | 3418(0.16%) | 179476(8.17%) | 1961357(89.24%) |
| Total | 12119198(100%) | 1939019(16%) | 130467(1.08) | 875(0.01%) | 5963(0.05%) | 8218(0.07%) | 856157(7.06%) | 9178499(75.14%) | |
|
X_1 |
Uniq | 588558(100%) | 25673(4.36%) | 82808(14.07%) | 1274(0.22%) | 6312(1.07%) | 6934(1.18%) | 188052(31.95%) | 277505(47.15%) |
| Total | 9781701(100%) | 8104780(82.86%) | 345098(3.53%) | 2382(0.02%) | 10984(0.11%) | 17844(0.18%) | 617283(6.31%) | 683330(6.99%) | |
|
X_2 |
Uniq | 487860(100%) | 29003(5.94%) | 43555(8.93%) | 795(0.16%) | 3818(0.78%) | 4272(0.88%) | 140384(28.78%) | 266033(54.53%) |
| Total | 13564919(100%) | 11914763(87.84%) | 242554(1.79%) | 1652(0.01%) | 6957(0.05%) | 11732(0.09%) | 548659(4.04%) | 838602(6.18%) | |
|
X_3 |
Uniq | 949314(100%) | 31531(3.32%) | 52596(5.54%) | 781(0.08%) | 4423(0.47%) | 5030(0.53%) | 222152(23.4%) | 632801(66.66%) |
| Total | 18256548(100%) | 15570998(85.29%) | 214214(1.17%) | 1541(0.01%) | 7468(0.04%) | 11602(0.06%) | 810797(4.44%) | 1639928(8.98%) |
| Known miRNA | P value | FC | Target genes | Up/Down |
|---|---|---|---|---|
| bta-miR-11971 | 0.00 | 190.67 | SUPT6H, PCSK7, ALKBH5, CRELD1, CMC1, DAPK3, UNC5B, FAM78A | Up |
| bta-miR-2284c | 0.00 | 77.16 | SUPT7L, CXCL12 | Up |
| bta-miR-3956 | 0.00 | 53.70 | TM9SF4, TNK2, TRIM2, AFAP1, ABL1, TRIOBP, RNF122, GLRB | Up |
| bta-miR-135b | 0.00 | 42.48 | KCTD10, GPATCH11, NCKIPSD, ALDH2, TTC14, DUSP27, UBOX5, HMG20A, BCL2L14 | Up |
| bta-miR-299-2 | 0.00 | 39.50 | BET1,,NRN1,RAB11FIP2,CD1E,GLCE,ADARB2,ATM,SYAP1, TTC9C | Up |
| bta-miR-433 | 0.00 | 37.56 | LRRTM3,MAPRE2,PAK4,SLC35D1,KCNJ14,MXI1 | Up |
| bta-miR-191 | 0.0000 | 0.4188 | AJAP1, SLC39A13, KCTD17, PRICKLE1, USP22, IER2, DSG1 | Down |
| bta-miR-20a | 0.0000 | 0.4153 | PLEKHM1, SAAL1, CFL2, ZNFX1, APP, RAB29, CAMK1G, METAP1, KCTD7 | Down |
| Novel miRNAs | ||||
| 4_37054-5p(cli-miR-9632-3p) | 0.00000 | 153.87736 | GATAD2B,CD47, HEATR5A,USP43, NANOG, CHSY1, SLC38A3, ALX3, EFR3B | Up |
| 7_42582-5p | 0.00030 | 58.00028 | FCRL5, EMP1, TTPAL, CAMK2A, ESD, NMNAT2, CYP4V2, AKAP6 | Up |
| 6_40791-3p | 0.00085 | 38.35687 | BTBD7, FBXW11, KCNJ2, C18H19orf12, CMC1, PLD3, RACK1, SLC39A14 | Up |
| 21_23590-5p(mmu-miR-665-3p) | 0.00146 | 34.79795 | RHBDD1,FGF2,C1QTNF5,PLA2G3,BCO2,HDAC5,PRELP,TRMT12 | Up |
| Groups | Terms | DEGs | GO_Accession | Some of the Genes ID associated with the GO terms |
|---|---|---|---|---|
|
X vs D |
Sexual reproduction | 279 | GO:0019953 | NOS3, CLIC4, BCL2L11, UBE3A, PLCD4,APP, TYRO3, UBE2Q1, TBPL1, ACVR2A, AGFG1, TEKT3, CD46, HMGA2, HSPA1L, ADAMTS1, DDX25, PRSS37, ALKBH5, PAIP2, PRDM1, B4GALT1, STRBP, LGR4, MCM8, PDE5A, TRIM36, RGS2,BCL6, FOSL1, STAT3, LEP, SPIN1, NPPC, PLA2G3, CIB1, NR2F2, FKBP6, OVOL1, RAD23B, DAZL, ALKBH5, CDC25B, FNDC3A, VIPAS39, RNF2 |
| Male gonad development | 39 | GO:0008584 | NKX3-1, KITLG, LRRC6, ACVR2A, PDGFRA, HMGA2, BCL2L11,FNDC3A | |
| Germ cell development | 103 | GO:0007281 | PDE5A, TBPL1, AGFG1, PRDM1, STRBP, DDX25, DAZL, PLA2G3, CIB1, RNF2, CDC25B, FNDC3A, NPPC | |
| Germ cell migration | 2 | GO:0008354 | TGFBR1,CXADR | |
| Spermatid development | 58 | GO:0007286 | TBPL1, AGFG1, STRBP, DDX25, PLA2G3, CIB1, FNDC3A | |
| Fusion of sperm to egg | 3 | GO:0007342 | CD9,SPESP1,SPAM1 | |
| Sperm part | 62 | GO:0097223 | TRIM36, CABYR, CAV1, TEKT3, CD46, POMT1, AK1, FNDC3A, CD46 | |
| Lateral plasma membrane | 24 | GO:0016328 | NSG1, AXIN1, BVES, CLDN12, ARL2 | |
| Negative regulation of androgen secretion | 1 | GO:2000835 | GH1 | |
| Androgen secretion | 3 | GO:0035935 | NKX3-1,CSN1S1, GH1 | |
| Sertoli cell development | 3 | GO:0060009 | FNDC3A,FNDC3A,ARID4A | |
| Male genitalia development | 9 | GO:0030539 | BMP5, CTNNB1, DHCR24, ASB1, FGF8, SRD5A2, AR, PDGFRA, HSD17B3 | |
| Reproductive process | 498 | GO:0022414 | STAT5A, PAIP2, FZR1, PRDM1, B4GALT1, STRBP, LGR4, FEM1B, MAP2K1, ESPL1, STAT5B, RGS2, ASCL2, PDGFRA, KITLG, DHCR24, KRT19, BCL2L11, DRD2, UBE2A, TYRO3, APP, TBPL1, ACVR2A, TEKT3, CD46, ALKBH5, ERCC4, MED1, MAPK14, MCM8, TTPA, PDE5A, WNT2, CASP4, IL1A, TRIM36, CRHBP, CYP27B1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




