Submitted:
25 October 2023
Posted:
26 October 2023
You are already at the latest version
Abstract
Keywords:
Introduction
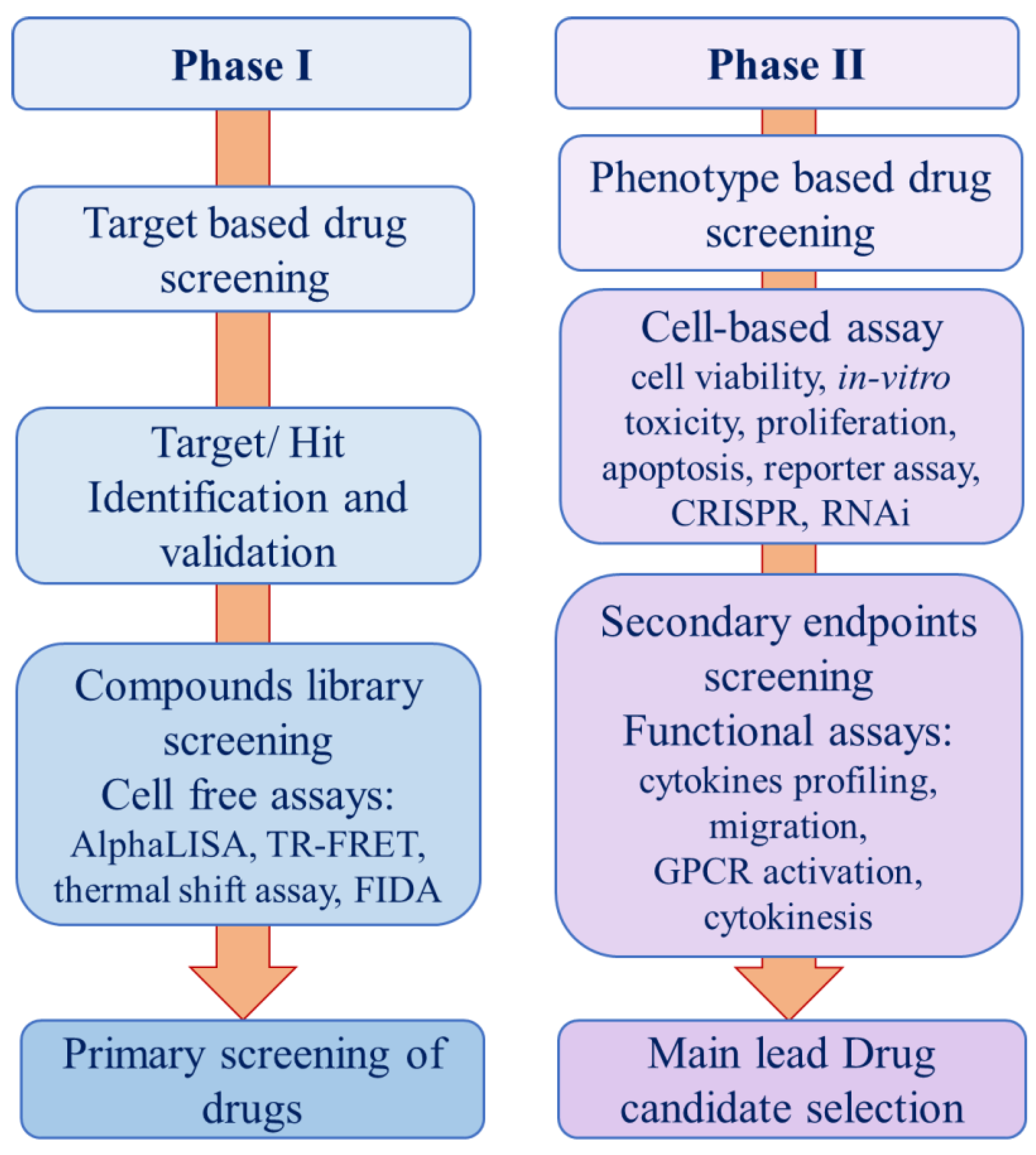
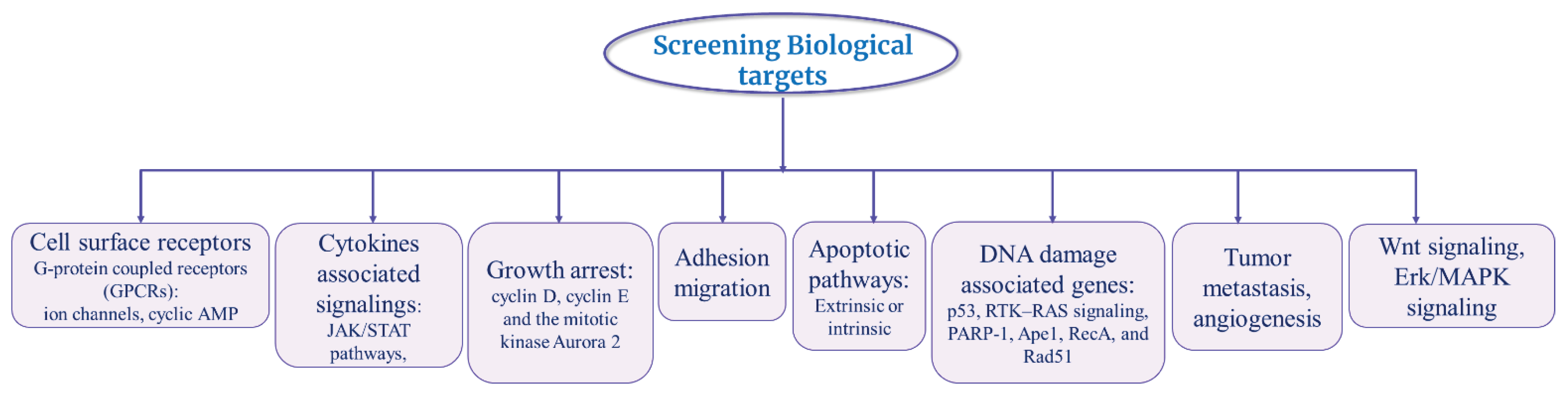
Target-Based Screening
Phenotype Based Screening
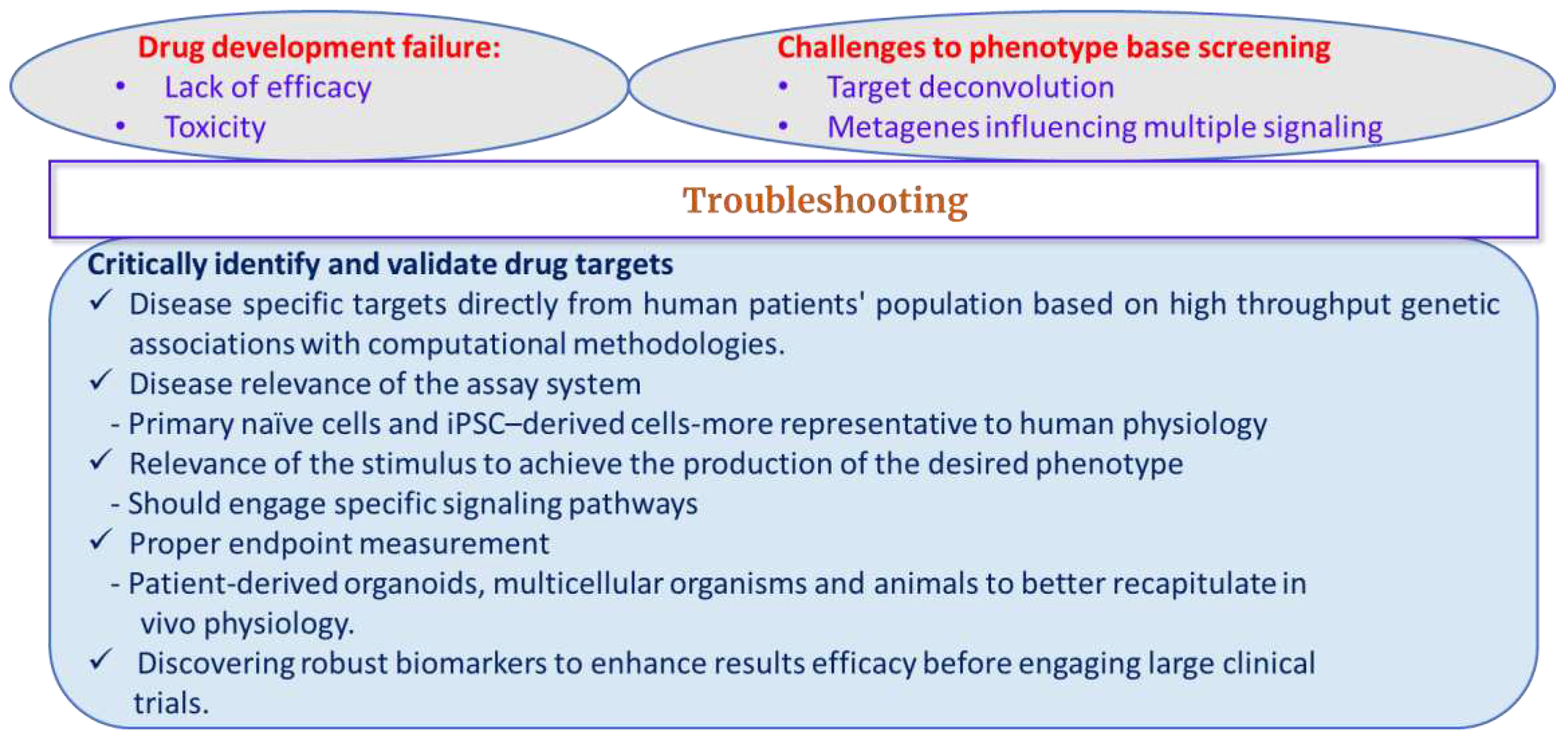
Challenges to Phenotypic Screening
Screening for Genotoxic Compounds
Application in Tumor Chemoresistance and Cancer Treatment
Conclusion
| Druggable target | Compounds | Ref. |
|---|---|---|
| P-glycoprotein | LY335979, R101933 and XR9576 | [114-116] |
| Mometasone furoate, NSC23925, NSC77037, pimozide, acacetin and loxapine | [119-122] | |
| β-catenin (Wnt signaling) | CCT031374, CCT036477, and CCT7070535 | [135, 136] |
| MEK1 | Class 1 and class 2 substituted 4-anilino-3-quinolinecarbonitriles | [137] |
| HDAC | Trichostatin A (TSA), suberoylanilide hydroxamic acid (SAHA), LBH589, and PXD101 | [139] |
| KRAS | Demethoxyviridin, mithramycin, triphenyl tetrazolium (TPT), sulfinyl cytidine derivative (SC-D) | [138] |
| PI3K/PTEN | N’-[(1-benzyl-1H-indol-3-yl)methylene]benzenesulfonohydrazide (CID1340132) | [140] |
| JAK/STAT | RUS0910-G009 | [141] |
| p53 | CP-31398, PRIMA1 and Nutlins | [156] |
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Macarron, R. , et al., Impact of high-throughput screening in biomedical research. Nat Rev Drug Discov 2011, 10, 188–195. [Google Scholar] [CrossRef] [PubMed]
- Wigglesworth, M.J. , et al., Increasing the delivery of next generation therapeutics from high throughput screening libraries. Curr Opin Chem Biol 2015, 26, 104–110. [Google Scholar] [CrossRef] [PubMed]
- Pereira, D.A. and J.A. Williams, Origin and evolution of high throughput screening. Br J Pharmacol 2007, 152, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Shun, T.Y. , et al., Identifying actives from HTS data sets: practical approaches for the selection of an appropriate HTS data-processing method and quality control review. J Biomol Screen 2011, 16, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Dreiman, G.H.S. , et al., Changing the HTS Paradigm: AI-Driven Iterative Screening for Hit Finding. SLAS Discov 2021, 26, 257–262. [Google Scholar] [CrossRef] [PubMed]
- Tsoli, M. , et al., Integration of genomics, high throughput drug screening, and personalized xenograft models as a novel precision medicine paradigm for high risk pediatric cancer. Cancer Biol Ther 2018, 19, 1078–1087. [Google Scholar] [CrossRef] [PubMed]
- Mayr, L.M. and D. Bojanic, Novel trends in high-throughput screening. Curr Opin Pharmacol 2009, 9, 580–588. [Google Scholar] [CrossRef] [PubMed]
- Szymański, P., M. Markowicz, and E. Mikiciuk-Olasik, Adaptation of high-throughput screening in drug discovery-toxicological screening tests. Int J Mol Sci 2012, 13, 427–452. [Google Scholar] [PubMed]
- Milinkovic, A. and E. Martínez, Nevirapine in the treatment of HIV. Expert Rev Anti Infect Ther 2004, 2, 367–373. [Google Scholar] [CrossRef]
- FDA approved drugs generated from high throughput screening efforts I - 20190719. 2019.
- Margineanu, D.G. , Systems biology, complexity, and the impact on antiepileptic drug discovery. Epilepsy Behav 2014, 38, 131–42. [Google Scholar] [CrossRef]
- Kell, D.B. , Finding novel pharmaceuticals in the systems biology era using multiple effective drug targets, phenotypic screening and knowledge of transporters: where drug discovery went wrong and how to fix it. Febs j 2013, 280, 5957–5980. [Google Scholar] [CrossRef] [PubMed]
- Ericson, E. , et al., Off-target effects of psychoactive drugs revealed by genome-wide assays in yeast. PLoS Genet 2008, 4, e1000151. [Google Scholar] [CrossRef] [PubMed]
- Lim, H. , et al., Large-Scale Off-Target Identification Using Fast and Accurate Dual Regularized One-Class Collaborative Filtering and Its Application to Drug Repurposing. PLoS Comput Biol 2016, 12, e1005135. [Google Scholar] [CrossRef] [PubMed]
- Lin, A. , et al., Off-target toxicity is a common mechanism of action of cancer drugs undergoing clinical trials. Sci Transl Med 2019. 11(509).
- Koeberle, A. and O. Werz, Multi-target approach for natural products in inflammation. Drug Discov Today 2014, 19, 1871–1882. [Google Scholar] [CrossRef] [PubMed]
- Konecny, G.E. , et al., Activity of the dual kinase inhibitor lapatinib (GW572016) against HER-2-overexpressing and trastuzumab-treated breast cancer cells. Cancer Res 2006, 66, 1630–1639. [Google Scholar] [CrossRef] [PubMed]
- Flinn, I.W. , et al., Duvelisib, a novel oral dual inhibitor of PI3K-δ,γ, is clinically active in advanced hematologic malignancies. Blood 2018, 131, 877–887. [Google Scholar] [CrossRef] [PubMed]
- Murai, R. , et al., A novel screen using the Reck tumor suppressor gene promoter detects both conventional and metastasis-suppressing anticancer drugs. Oncotarget 2010, 1, 252. [Google Scholar] [CrossRef]
- An, W.F. and N. Tolliday, Cell-based assays for high-throughput screening. Mol Biotechnol 2010, 45, 180–186. [Google Scholar] [CrossRef]
- Takenaka, T. , Classical vs reverse pharmacology in drug discovery. BJU Int 2001. 88 Suppl 2: p. 7-10; discussion 49-50.
- Lage, O.M. , et al., Current Screening Methodologies in Drug Discovery for Selected Human Diseases. Mar Drugs 2018, 16. [Google Scholar]
- Sriram, K. and P.A. Insel, G Protein-Coupled Receptors as Targets for Approved Drugs: How Many Targets and How Many Drugs? Mol Pharmacol 2018, 93, 251–258. [Google Scholar] [CrossRef]
- Kenny, C.H. , et al., Development of a fluorescence polarization assay to screen for inhibitors of the FtsZ/ZipA interaction. Analytical biochemistry 2003, 323, 224–233. [Google Scholar] [CrossRef]
- Burns, S. , et al., Identification of Small-Molecule Inhibitors of Protein Kinase B (PKB/AKT) in an AlphaScreen™ High-Throughput Screen. SLAS Discovery 2006, 11, 822–827. [Google Scholar] [CrossRef] [PubMed]
- Sudo, K. , et al., High-throughput screening of low molecular weight NS3-NS4A protease inhibitors using a fluorescence resonance energy transfer substrate. Antiviral Chemistry and Chemotherapy 2005, 16, 385–392. [Google Scholar] [CrossRef]
- Swaney, S. , et al., Characterization of a high-throughput screening assay for inhibitors of elongation factor P and ribosomal peptidyl transferase activity. Journal of biomolecular screening 2006, 11, 736–742. [Google Scholar] [CrossRef] [PubMed]
- Allen, M., J. Reeves, and G. Mellor, High throughput fluorescence polarization: a homogeneous alternative to radioligand binding for cell surface receptors. Journal of biomolecular screening 2000, 5, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Xu, J. , et al., Ion-channel assay technologies: quo vadis? Drug Discovery Today 2001, 6, 1278–1287. [Google Scholar] [CrossRef]
- Parker, G.J. , et al., Development of high throughput screening assays using fluorescence polarization: nuclear receptor-ligand–binding and kinase/phosphatase assays. SLAS Discovery 2000, 5, 77–88. [Google Scholar] [CrossRef]
- Bagal, S.K. , et al. , Ion channels as therapeutic targets: a drug discovery perspective. J Med Chem 2013, 56, 593–624. [Google Scholar]
- Morachis, J.M., R. Huang, and B.M. Emerson, Identification of kinase inhibitors that target transcription initiation by RNA polymerase II. Oncotarget 2011, 2, 18–28. [Google Scholar] [CrossRef]
- Glickman, J.F. , Assay Development for Protein Kinase Enzymes, in Assay Guidance Manual, S. Markossian, et al., Editors. 2004, Eli Lilly & Company and the National Center for Advancing Translational Sciences: Bethesda (MD).
- Harner, M.J., A. O. Frank, and S.W. Fesik, Fragment-based drug discovery using NMR spectroscopy. J Biomol NMR 2013, 56, 65–75. [Google Scholar] [CrossRef]
- Bollag, G. , et al., Clinical efficacy of a RAF inhibitor needs broad target blockade in BRAF-mutant melanoma. Nature 2010, 467, 596–599. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, I.H. , Drug discovery for neglected diseases: molecular target-based and phenotypic approaches. J Med Chem 2013, 56, 7719–7726. [Google Scholar] [CrossRef]
- Chen, G.Q. , et al., Phenotype and target-based chemical biology investigations in cancers. Natl Sci Rev 2019, 6, 1111–1127. [Google Scholar] [CrossRef] [PubMed]
- Iqbal, N. and N. Iqbal, Imatinib: a breakthrough of targeted therapy in cancer. Chemother Res Pract 2014, 2014, 357027. [Google Scholar] [PubMed]
- Rossari, F., F. Minutolo, and E. Orciuolo, Past, present, and future of Bcr-Abl inhibitors: from chemical development to clinical efficacy. J Hematol Oncol 2018, 11, 84. [Google Scholar] [PubMed]
- Dong, B. , et al., Cryptotanshinone suppresses key onco-proliferative and drug-resistant pathways of chronic myeloid leukemia by targeting STAT5 and STAT3 phosphorylation. Sci China Life Sci 2018, 61, 999–1009. [Google Scholar] [CrossRef]
- Liu, C. , et al., Antitumor Effects of Blocking Protein Neddylation in T315I-BCR-ABL Leukemia Cells and Leukemia Stem Cells. Cancer Res 2018, 78, 1522–1536. [Google Scholar] [CrossRef]
- Misra, S. , et al., Both BRCA1-wild type and -mutant triple-negative breast cancers show sensitivity to the NAE inhibitor MLN4924 which is enhanced upon MLN4924 and cisplatin combination treatment. Oncotarget 2020, 11, 784–800. [Google Scholar] [CrossRef]
- Krejci, P., K. Pejchalova, and W.R. Wilcox, Simple, mammalian cell-based assay for identification of inhibitors of the Erk MAP kinase pathway. Investigational New Drugs 2007, 25, 391–395. [Google Scholar] [CrossRef]
- Chambers, C. , et al., Measuring intracellular calcium fluxes in high throughput mode. Combinatorial chemistry & high throughput screening 2003, 6, 355–362. [Google Scholar]
- Kariv, I. , et al., High throughput quantitation of cAMP production mediated by activation of seven transmembrane domain receptors. Journal of biomolecular screening 1999, 4, 27–32. [Google Scholar] [CrossRef]
- Eggert, U.S. , et al. , Parallel chemical genetic and genome-wide RNAi screens identify cytokinesis inhibitors and targets. PLoS biology 2004, 2, e379. [Google Scholar]
- Beck, V., A. Pfitscher, and A. Jungbauer, GFP-reporter for a high throughput assay to monitor estrogenic compounds. Journal of biochemical and biophysical methods 2005, 64, 19–37. [Google Scholar] [CrossRef]
- Yarrow, J.C. , et al., Screening for cell migration inhibitors via automated microscopy reveals a Rho-kinase inhibitor. Chemistry & biology 2005, 12, 385–395. [Google Scholar]
- Vincent, F. , et al., Developing predictive assays: the phenotypic screening "rule of 3". Sci Transl Med 2015, 7, 293ps15. [Google Scholar] [CrossRef]
- Engle, S.J. and F. Vincent, Small molecule screening in human induced pluripotent stem cell-derived terminal cell types. J Biol Chem 2014, 289, 4562–4570. [Google Scholar] [CrossRef]
- McNeish, J. , et al., High-throughput Screening in Embryonic Stem Cell-derived Neurons Identifies Potentiators of α-Amino-3-hydroxyl-5-methyl-4-isoxazolepropionate-type Glutamate Receptors. Journal of Biological Chemistry 2010, 285, 17209–17217. [Google Scholar] [CrossRef]
- Ashlock, M.A. and E.R. Olson, Therapeutics development for cystic fibrosis: a successful model for a multisystem genetic disease. Annu Rev Med 2011, 62, 107–25. [Google Scholar] [CrossRef]
- Van Goor, F. , et al., Correction of the F508del-CFTR protein processing defect in vitro by the investigational drug VX-809. Proc Natl Acad Sci U S A 2011, 108, 18843–18848. [Google Scholar] [CrossRef]
- Lee, G. , et al., Large-scale screening using familial dysautonomia induced pluripotent stem cells identifies compounds that rescue IKBKAP expression. Nat Biotechnol 2012, 30, 1244–1248. [Google Scholar] [CrossRef]
- Barberis, A. , et al., Yeast as a screening tool. Drug Discovery Today: Technologies 2005, 2, 187–192. [Google Scholar] [CrossRef]
- Balgi, A.D. and M. Roberge, Screening for chemical inhibitors of heterologous proteins expressed in yeast using a simple growth-restoration assay, in Cell-Based Assays for High-Throughput Screening. 2009, Springer. p. 125-137.
- Puri, A.W. and M. Bogyo, Using small molecules to dissect mechanisms of microbial pathogenesis. ACS chemical biology 2009, 4, 603–616. [Google Scholar] [CrossRef] [PubMed]
- Zlitni, S., J. E. Blanchard, and E.D. Brown, High-throughput screening of model bacteria, in Cell-Based Assays for High-Throughput Screening. 2009, Springer. p. 13-27.
- Hong, C.C. , Large-scale small-molecule screen using zebrafish embryos, in Cell-Based Assays for High-Throughput Screening. 2009, Springer. p. 43-55.
- Zon, L.I. and R.T. Peterson, In vivo drug discovery in the zebrafish. Nature reviews Drug discovery 2005, 4, 35–44. [Google Scholar] [CrossRef] [PubMed]
- O’Rourke, E.J., A. L. Conery, and T.I. Moy, Whole-animal high-throughput screens: the C. elegans model, in Cell-Based Assays for High-Throughput Screening. 2009, Springer. p. 57-75.
- Agee, A. and D. Carter, Whole-organism screening: plants, in Cell-Based Assays for High-Throughput Screening. 2009, Springer. p. 77-95.
- Norambuena, L., N. V. Raikhel, and G.R. Hicks, Chemical genomics approaches in plant biology. Plant Systems Biology, 345–354.
- Stern, H.M. , et al., Small molecules that delay S phase suppress a zebrafish bmyb mutant. Nat Chem Biol 2005, 1, 366–370. [Google Scholar] [CrossRef]
- Hong, C.C. , et al., Artery/Vein Specification Is Governed by Opposing Phosphatidylinositol-3 Kinase and MAP Kinase/ERK Signaling. Current Biology 2006, 16, 1366–1372. [Google Scholar] [CrossRef]
- Swinney, D.C. , The value of translational biomarkers to phenotypic assays. Front Pharmacol 2014, 5, 171. [Google Scholar] [CrossRef] [PubMed]
- Ioannidis, J.P. , Biomarker failures. Clin Chem 2013, 59, 202–204. [Google Scholar] [CrossRef]
- Chau, C.H. , et al., Validation of analytic methods for biomarkers used in drug development. Clin Cancer Res 2008, 14, 5967–5976. [Google Scholar] [CrossRef]
- Spreafico, R. , et al., Advances in Genomics for Drug Development. Genes (Basel) 2020, 11. [Google Scholar] [CrossRef]
- Guo, J.B. and X.J. Li, [Impacts of modern biology on drug discovery]. Sheng Li Ke Xue Jin Zhan 2007, 38, 25–31. [Google Scholar]
- Hong, A.L. , et al., Integrated genetic and pharmacologic interrogation of rare cancers. Nat Commun 2016, 7: p. 11987.
- Lin, X. , et al., Life beyond kinases: structure-based discovery of sorafenib as nanomolar antagonist of 5-HT receptors. Journal of medicinal chemistry 2012, 55, 5749–5759. [Google Scholar] [CrossRef] [PubMed]
- Schneidewind, T. , et al., Morphological profiling identifies a common mode of action for small molecules with different targets. ChemBioChem 2020, 21, 3197–3207. [Google Scholar] [CrossRef] [PubMed]
- Han, X. , et al., Discovery of RG7834: the first-in-class selective and orally available small molecule hepatitis B virus expression inhibitor with novel mechanism of action. Journal of Medicinal Chemistry 2018, 61, 10619–10634. [Google Scholar] [CrossRef] [PubMed]
- Moffat, J.G. , et al., Opportunities and challenges in phenotypic drug discovery: an industry perspective. Nature reviews Drug discovery 2017, 16, 531–543. [Google Scholar] [CrossRef] [PubMed]
- Vincent, F. , et al., Hit triage and validation in phenotypic screening: considerations and strategies. Cell Chemical Biology 2020, 27, 1332–1346. [Google Scholar] [CrossRef] [PubMed]
- Warchal, S.J., A. Unciti-Broceta, and N.O. Carragher, Next-generation phenotypic screening. Future medicinal chemistry 2016, 8, 1331–1347. [Google Scholar] [CrossRef] [PubMed]
- Comess, K.M. , et al., Emerging approaches for the identification of protein targets of small molecules-a practitioners’ perspective. Journal of medicinal chemistry 2018, 61, 8504–8535. [Google Scholar] [CrossRef] [PubMed]
- Lee, J. and M. Bogyo, Target deconvolution techniques in modern phenotypic profiling. Curr Opin Chem Biol 2013, 17, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.-J. , et al., Multi-target drugs: the trend of drug research and development. PloS one 2012, 7, e40262. [Google Scholar]
- Rena, G., D. G. Hardie, and E.R. Pearson, The mechanisms of action of metformin. Diabetologia 2017, 60, 1577–1585. [Google Scholar] [CrossRef]
- Keller, M.P. and A.D. Attie, Physiological insights gained from gene expression analysis in obesity and diabetes. Annual review of nutrition 2010, 30, 341–364. [Google Scholar] [CrossRef] [PubMed]
- Van’t Veer, L.J. and R. Bernards, Enabling personalized cancer medicine through analysis of gene-expression patterns. Nature 2008, 452, 564–570. [Google Scholar] [CrossRef] [PubMed]
- Bertos, N.R. and M. Park, Breast cancer—one term, many entities? The Journal of clinical investigation 2011, 121, 3789–3796. [Google Scholar] [CrossRef] [PubMed]
- Huang, E. , et al., Gene expression phenotypic models that predict the activity of oncogenic pathways. Nature genetics 2003, 34, 226–230. [Google Scholar] [CrossRef] [PubMed]
- Loboda, A. , et al., A gene expression signature of RAS pathway dependence predicts response to PI3K and RAS pathway inhibitors and expands the population of RAS pathway activated tumors. BMC medical genomics 2010, 3, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Loboda, A. , et al., EMT is the dominant program in human colon cancer. BMC medical genomics 2011, 4, 1–10. [Google Scholar] [CrossRef]
- Méndez-Lucio O, N.J. , Vite-Caritino H, Prieto-Martínez FD, Medina-Franco JL., One Drug for Multiple Targets: A Computational Perspective. J Mex Chem Soc. 2016, 60, 168–181. [Google Scholar]
- Hsieh, J.H. , et al., Identifying Compounds with Genotoxicity Potential Using Tox21 High-Throughput Screening Assays. Chem Res Toxicol 2019, 32, 1384–1401. [Google Scholar] [CrossRef]
- Michod, D. and C. Widmann, DNA-damage sensitizers: potential new therapeutical tools to improve chemotherapy. Critical reviews in oncology/hematology 2007, 63, 160–171. [Google Scholar] [CrossRef]
- Evans, T.J. , et al., Mutant cells defective in DNA repair pathways provide a sensitive high-throughput assay for genotoxicity. DNA repair 2010, 9, 1292–1298. [Google Scholar] [CrossRef]
- Ji, K. , et al., A Novel Approach Using DNA-Repair–Deficient Chicken DT40 Cell Lines for Screening and Characterizing the Genotoxicity of Environmental Contaminants. Environmental health perspectives 2009, 117, 1737–1744. [Google Scholar] [CrossRef] [PubMed]
- Buerstedde, J.-M. and S. Takeda, Increased ratio of targeted to random integration after transfection of chicken B cell lines. Cell 1991, 67, 179–188. [Google Scholar] [CrossRef] [PubMed]
- Yamazoe, M. , et al., Reverse genetic studies of the DNA damage response in the chicken B lymphocyte line DT40. DNA repair 2004, 3, 1175–1185. [Google Scholar] [CrossRef] [PubMed]
- Walmsley, R.M. and M. Tate, The GADD45a-GFP GreenScreen HC assay. Genetic Toxicology: Principles and Methods, 231–250.
- Knight, A.W. , et al., Evaluation of high-throughput genotoxicity assays used in profiling the US EPA ToxCast™ chemicals. Regulatory Toxicology and Pharmacology 2009, 55, 188–199. [Google Scholar] [CrossRef] [PubMed]
- Hastwell, P.W. , et al., High-specificity and high-sensitivity genotoxicity assessment in a human cell line: Validation of the GreenScreen HC GADD45a-GFP genotoxicity assay. Mutation Research/Genetic Toxicology and Environmental Mutagenesis 2006, 607, 160–175. [Google Scholar] [CrossRef] [PubMed]
- Knight, A.W. , et al., Evaluation of high-throughput genotoxicity assays used in profiling the US EPA ToxCast chemicals. Regul Toxicol Pharmacol 2009, 55, 188–199. [Google Scholar] [CrossRef]
- Walmsley, R.M. and M. Tate, The GADD45a-GFP GreenScreen HC assay. Methods Mol Biol 2012, 817, 231–250. [Google Scholar] [PubMed]
- Giuliano, K.A. , et al., Early safety assessment using cellular systems biology yields insights into mechanisms of action. J Biomol Screen 2010, 15, 783–797. [Google Scholar] [CrossRef] [PubMed]
- Brattain, M.G. , et al., Heterogeneity of malignant cells from a human colonic carcinoma. Cancer Res 1981, 41, 1751–1756. [Google Scholar]
- Xing, J.Z. , et al., Microelectronic cell sensor assay for detection of cytotoxicity and prediction of acute toxicity. Toxicology in Vitro 2006, 20, 995–1004. [Google Scholar] [CrossRef]
- Fox, J.T. , et al., High-throughput genotoxicity assay identifies antioxidants as inducers of DNA damage response and cell death. Proceedings of the National Academy of Sciences 2012, 109, 5423–5428. [Google Scholar] [CrossRef] [PubMed]
- Sikdar, N. , et al., DNA damage responses by human ELG1 in S phase are important to maintain genomic integrity. Cell Cycle 2009, 8, 3199–3207. [Google Scholar] [CrossRef] [PubMed]
- Scholes, D.T. , et al., Multiple regulators of Ty1 transposition in Saccharomyces cerevisiae have conserved roles in genome maintenance. Genetics 2001, 159, 1449–1465. [Google Scholar] [CrossRef] [PubMed]
- Sykora, P. , et al., Next generation high throughput DNA damage detection platform for genotoxic compound screening. Sci Rep 2018, 8, 2771. [Google Scholar] [CrossRef] [PubMed]
- Gatti, L. and F. Zunino, Overview of tumor cell chemoresistance mechanisms. Methods Mol Med 2005, 111, 127–48. [Google Scholar]
- Gottesman, M.M., T. Fojo, and S.E. Bates, Multidrug resistance in cancer: role of ATP-dependent transporters. Nat Rev Cancer 2002, 2, 48–58. [Google Scholar] [CrossRef] [PubMed]
- Gottesman, M.M. and V. Ling, The molecular basis of multidrug resistance in cancer: the early years of P-glycoprotein research. FEBS Lett 2006, 580, 998–1009. [Google Scholar] [CrossRef] [PubMed]
- Sharom, F.J. , ABC multidrug transporters: structure, function and role in chemoresistance. Pharmacogenomics 2008, 9, 105–127. [Google Scholar] [CrossRef] [PubMed]
- Sparreboom, A. , et al., Limited oral bioavailability and active epithelial excretion of paclitaxel (Taxol) caused by P-glycoprotein in the intestine. Proc Natl Acad Sci U S A 1997, 94, 2031–2035. [Google Scholar] [CrossRef]
- Kartner, N., J. R. Riordan, and V. Ling, Cell surface P-glycoprotein associated with multidrug resistance in mammalian cell lines. Science 1983, 221, 1285–1288. [Google Scholar] [CrossRef]
- Shen, D.W. , et al., Multiple drug-resistant human KB carcinoma cells independently selected for high-level resistance to colchicine, adriamycin, or vinblastine show changes in expression of specific proteins. J Biol Chem 1986, 261, 7762–7770. [Google Scholar] [CrossRef] [PubMed]
- Roe, M. , et al., Reversal of P-glycoprotein mediated multidrug resistance by novel anthranilamide derivatives. Bioorg Med Chem Lett 1999, 9, 595–600. [Google Scholar] [CrossRef] [PubMed]
- Dantzig, A.H. , et al., Selectivity of the multidrug resistance modulator, LY335979, for P-glycoprotein and effect on cytochrome P-450 activities. J Pharmacol Exp Ther 1999, 290, 854–862. [Google Scholar] [PubMed]
- Mistry, P. , et al., In vitro and in vivo reversal of P-glycoprotein-mediated multidrug resistance by a novel potent modulator, XR9576. Cancer Res 2001, 61, 749–758. [Google Scholar] [PubMed]
- Bates, S.F. , et al., Reversal of multidrug resistance: lessons from clinical oncology. Novartis Found Symp 2002, 243, 83–96. [Google Scholar] [PubMed]
- Dalton, W. , Detection of multidrug resistance gene expression in multiple myeloma. Leukemia 1997, 11, 1166–1169. [Google Scholar] [CrossRef] [PubMed]
- Susa, M. , et al., Multidrug resistance reversal agent, NSC77037, identified with a cell-based screening assay. J Biomol Screen 2010, 15, 287–296. [Google Scholar] [CrossRef] [PubMed]
- Winter, S.S. , et al., High-throughput screening for daunorubicin-mediated drug resistance identifies mometasone furoate as a novel ABCB1-reversal agent. J Biomol Screen 2008, 13, 185–193. [Google Scholar] [CrossRef]
- Duan, Z., E. Choy, and F.J. Hornicek, NSC23925, identified in a high-throughput cell-based screen, reverses multidrug resistance. PLoS One 2009, 4, e7415. [Google Scholar] [CrossRef]
- Ivnitski-Steele, I. , et al., High-throughput flow cytometry to detect selective inhibitors of ABCB1, ABCC1, and ABCG2 transporters. Assay Drug Dev Technol 2008, 6, 263–276. [Google Scholar] [CrossRef]
- Gurdon, J.B., T. R. Elsdale, and M. Fischberg, Sexually mature individuals of Xenopus laevis from the transplantation of single somatic nuclei. Nature 1958, 182, 64–65. [Google Scholar] [CrossRef] [PubMed]
- Hass, R., J. von der Ohe, and H. Ungefroren, The Intimate Relationship among EMT, MET and TME: AT (ransdifferentiation) E (nhancing) M (ix) to Be Exploited for Therapeutic Purposes. Cancers, 3674. [Google Scholar]
- Saito, S. , et al., Potential application of cell reprogramming techniques for cancer research. Cellular and Molecular Life Sciences.
- Welch, D.R. , Tumor heterogeneity—a ‘contemporary concept’founded on historical insights and predictions. Cancer research 2016, 76, 4–6. [Google Scholar] [CrossRef] [PubMed]
- Denny, S.K. , et al., Nfib Promotes Metastasis through a Widespread Increase in Chromatin Accessibility. Cell 2016, 166, 328–342. [Google Scholar] [CrossRef] [PubMed]
- Teng, S. , et al., Tissue-specific transcription reprogramming promotes liver metastasis of colorectal cancer. Cell Res 2020, 30, 34–49. [Google Scholar] [CrossRef] [PubMed]
- Van Keymeulen, A. , et al., Reactivation of multipotency by oncogenic PIK3CA induces breast tumour heterogeneity. Nature 2015, 525, 119–123. [Google Scholar] [CrossRef] [PubMed]
- Katoh, M. , Canonical and non-canonical WNT signaling in cancer stem cells and their niches: Cellular heterogeneity, omics reprogramming, targeted therapy and tumor plasticity (Review). Int J Oncol 2017, 51, 1357–1369. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.W., Y. Z. Nie, and H. Taniguchi, Cellular reprogramming and hepatocellular carcinoma development. World J Gastroenterol 2013, 19, 8850–8860. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.A. and P.W. Laird, Cancer epigenetics comes of age. Nat Genet 1999, 21, 163–167. [Google Scholar] [CrossRef] [PubMed]
- Wang, D. , et al., Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature 2011, 474, 390–394. [Google Scholar] [CrossRef]
- Qin, L. , et al., WNT5A promotes stemness characteristics in nasopharyngeal carcinoma cells leading to metastasis and tumorigenesis. Oncotarget 2015, 6, 10239–10252. [Google Scholar] [CrossRef]
- Ewan, K. , et al., A useful approach to identify novel small-molecule inhibitors of Wnt-dependent transcription. Cancer Res 2010, 70, 5963–5973. [Google Scholar] [CrossRef]
- Bialkowska, A.B. and V.W. Yang, High-throughput screening strategies for targeted identification of therapeutic compounds in colorectal cancer. Future Oncol 2012, 8, 259–272. [Google Scholar] [CrossRef]
- Mallon, R. , et al., Identification of 4-anilino-3-quinolinecarbonitrile inhibitors of mitogen-activated protein/extracellular signal-regulated kinase 1 kinase. Mol Cancer Ther 2004, 3, 755–762. [Google Scholar] [CrossRef]
- Torrance, C.J. , et al. , Use of isogenic human cancer cells for high-throughput screening and drug discovery. Nat Biotechnol 2001, 19, 940–945. [Google Scholar]
- Liu, S. , et al., Targeting enhancer reprogramming to mitigate MEK inhibitor resistance in preclinical models of advanced ovarian cancer. J Clin Invest 2021, 131. [Google Scholar]
- Li, H.F. , et al., A high-throughput screen with isogenic PTEN+/+ and PTEN-/- cells identifies CID1340132 as a novel compound that induces apoptosis in PTEN and PIK3CA mutant human cancer cells. J Biomol Screen 2011, 16, 383–393. [Google Scholar] [CrossRef]
- Yuliantie, E. , et al., High-throughput screening for small molecule inhibitors of the type-I interferon signaling pathway. Acta Pharm Sin B 2018, 8, 889–899. [Google Scholar] [CrossRef]
- Dillon, K.J., G. C. Smith, and N.M. Martin, A FlashPlate assay for the identification of PARP-1 inhibitors. J Biomol Screen 2003, 8, 347–352. [Google Scholar] [CrossRef]
- Bapat, A. , et al., Novel small-molecule inhibitor of apurinic/apyrimidinic endonuclease 1 blocks proliferation and reduces viability of glioblastoma cells. J Pharmacol Exp Ther 2010, 334, 988–998. [Google Scholar] [CrossRef]
- Peterson, E.J. , et al., High-throughput screening for RecA inhibitors using a transcreener adenosine 5’-O-diphosphate assay. Assay Drug Dev Technol 2012, 10, 260–268. [Google Scholar] [CrossRef]
- Sexton, J.Z. , et al. , Novel Inhibitors of E. coli RecA ATPase Activity. Curr Chem Genomics 2010, 4, 34–42. [Google Scholar]
- Huang, F. , et al., Identification of specific inhibitors of human RAD51 recombinase using high-throughput screening. ACS Chem Biol 2011, 6, 628–635. [Google Scholar] [CrossRef]
- Huang, L. , et al., High-Throughput Strategies for the Discovery of Anticancer Drugs by Targeting Transcriptional Reprogramming. Front Oncol 2021, 11, 762023. [Google Scholar] [CrossRef]
- Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 2008, 455, 1061–1068. [CrossRef]
- Ding, L. , et al., Somatic mutations affect key pathways in lung adenocarcinoma. Nature 2008, 455, 1069–1075. [Google Scholar] [CrossRef]
- Jones, S. , et al., Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 2008, 321, 1801–1806. [Google Scholar] [CrossRef]
- Steeg, P.S. and Q. Zhou, Cyclins and breast cancer. Breast Cancer Res Treat 1998, 52, 17–28. [Google Scholar] [CrossRef]
- Miyoshi, Y. , et al. , Association of centrosomal kinase STK15/BTAK mRNA expression with chromosomal instability in human breast cancers. Int J Cancer 2001, 92, 370–373. [Google Scholar]
- Dhillon, N.K. and M. Mudryj, Cyclin E overexpression enhances cytokine-mediated apoptosis in MCF7 breast cancer cells. Genes Immun 2003, 4, 336–342. [Google Scholar] [CrossRef]
- Niu, D., G. Wang, and X. Wang, Up-regulation of cyclin E in breast cancer via estrogen receptor pathway. Int J Clin Exp Med 2015, 8, 910–915. [Google Scholar]
- Warner, S.L. , et al., Comparing Aurora A and Aurora B as molecular targets for growth inhibition of pancreatic cancer cells. Mol Cancer Ther 2006, 5, 2450–2458. [Google Scholar] [CrossRef]
- Wang, W., S. H. Kim, and W.S. El-Deiry, Small-molecule modulators of p53 family signaling and antitumor effects in p53-deficient human colon tumor xenografts. Proc Natl Acad Sci U S A 2006, 103, 11003–11008. [Google Scholar] [CrossRef]
- Maloney, A. and P. Workman, HSP90 as a new therapeutic target for cancer therapy: the story unfolds. Expert Opin Biol Ther 2002, 2, 3–24. [Google Scholar] [CrossRef]
- Magwenyane, A.M. , et al., Heat Shock Protein 90 (HSP90) Inhibitors as Anticancer Medicines: A Review on the Computer-Aided Drug Discovery Approaches over the Past Five Years. Comput Math Methods Med 2022, 2022, 2147763. [Google Scholar] [CrossRef]
- Bos, R. , et al., Levels of hypoxia-inducible factor-1 alpha during breast carcinogenesis. J Natl Cancer Inst 2001, 93, 309–314. [Google Scholar] [CrossRef]
- Semenza, G.L. , HIF-1 and tumor progression: pathophysiology and therapeutics. Trends Mol Med 2002, 8 (Suppl. S4), S62–S67. [Google Scholar] [CrossRef]
- Masoud, G.N. and W. Li, HIF-1α pathway: role, regulation and intervention for cancer therapy. Acta Pharm Sin B 2015, 5, 378–389. [Google Scholar] [CrossRef]
- Workman, P. , Scoring a bull’s-eye against cancer genome targets. Curr Opin Pharmacol 2001, 1, 342–352. [Google Scholar] [CrossRef]
- Mahlknecht, U. and D. Hoelzer, Histone acetylation modifiers in the pathogenesis of malignant disease. Mol Med 2000, 6, 623–644. [Google Scholar] [CrossRef]
- Turlais, F. , et al., High-throughput screening for identification of small molecule inhibitors of histone acetyltransferases using scintillating microplates (FlashPlate). Anal Biochem 2001, 298, 62–68. [Google Scholar] [CrossRef]
- Dekker, F.J. and H.J. Haisma, Histone acetyl transferases as emerging drug targets. Drug Discov Today 2009, 14, 942–948. [Google Scholar] [CrossRef]
- Miles, D.W. , Update on HER-2 as a target for cancer therapy: herceptin in the clinical setting. Breast Cancer Res 2001, 3, 380–384. [Google Scholar] [CrossRef]
- Kelland, L.R. , et al., Preclinical antitumor activity and pharmacodynamic studies with the farnesyl protein transferase inhibitor R115777 in human breast cancer. Clin Cancer Res 2001, 7, 3544–3550. [Google Scholar]
- Johnston, S.R. , Farnesyl transferase inhibitors: a novel targeted tnerapy for cancer. Lancet Oncol 2001, 2, 18–26. [Google Scholar] [CrossRef]
- Nicholls, H. , Aromatase inhibitors continue their ATAC on tamoxifen. Trends Mol Med 2002, 8 (Suppl. S4), S12–3. [Google Scholar] [CrossRef]
- Buzdar, A.U. , Anastrozole (Arimidex)--an aromatase inhibitor for the adjuvant setting? Br J Cancer 2001, 85 (Suppl. S2), 6–10. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




