Submitted:
18 October 2023
Posted:
19 October 2023
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Description of Data
2.1.1. Soil Data
- MM_CA: Calcium content in a particular plot.
- MM_CEC: Cation capacity exchange in a particular plot.
- MM_K: Potasium content in a particular plot.
- MM_MG: Magnesium content in a particular plot.
- MM_OM: Organic matter content in a particular plot.
- MM_P1: Phosporous content in a particular plot.
- MM_PH: PH level in a particular plot.
- MM_CLAY: Clay content in a particular plot.
- MM_SAND: Sand content in a particular plot.
- MM_SILT: Silt content in a particular plot.
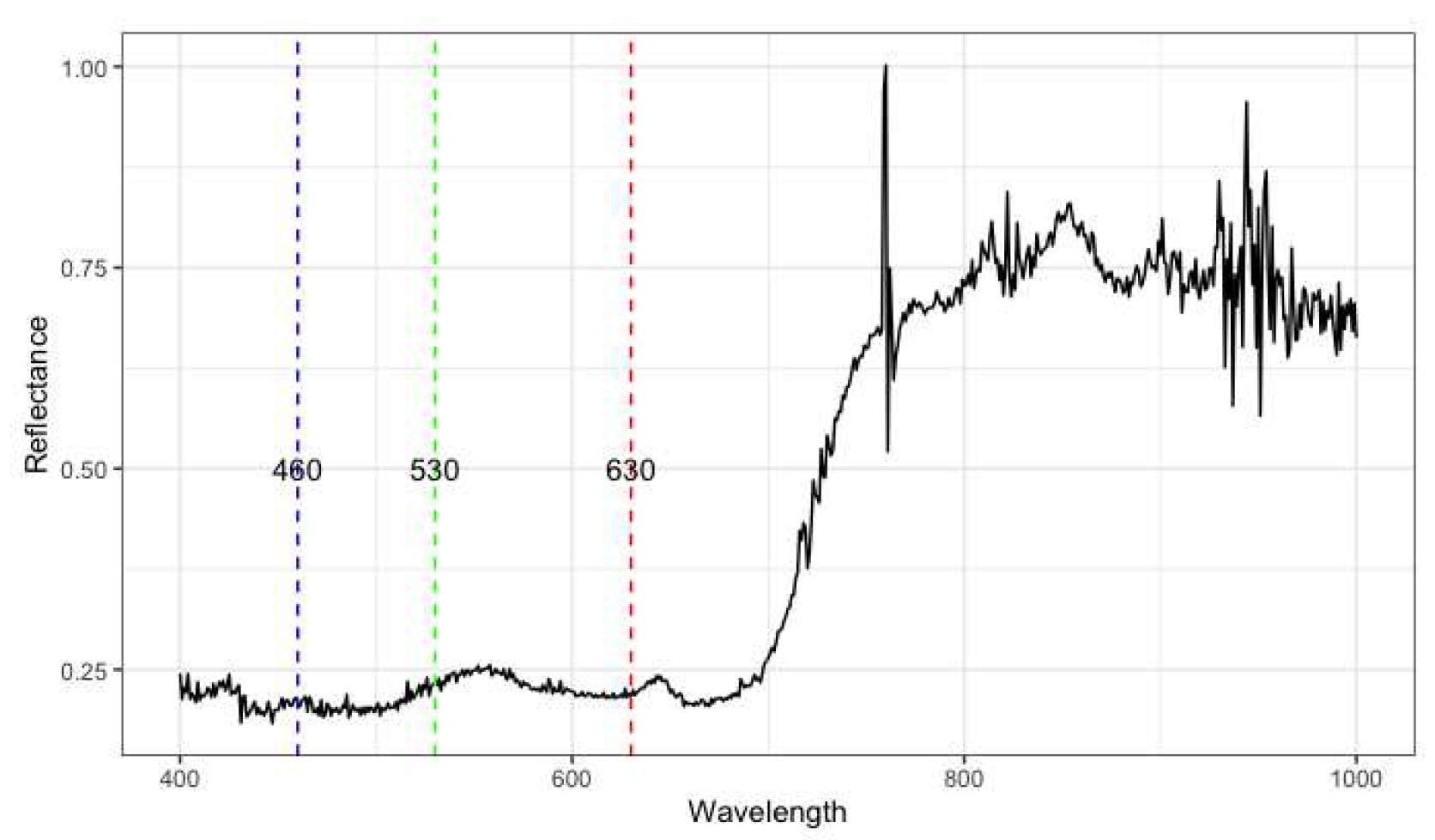
2.1.2. Hyperspectral Reflectance Data
2.1.3. Yield Data
- All observations for band values less than 400 nm were removed. This was done since we noticed many anomalies in the readings in those bands.
- All datapoints which had negative hyperspectral values in any of the bands ranging from 400 nm to 1000 nm were removed.
- All datapoints which had negative seed yield values were removed.
3. Methodology
3.1. Yield Prediction Using Vegetation Indices
3.2. Ensemble Models Using Vegetation Indices
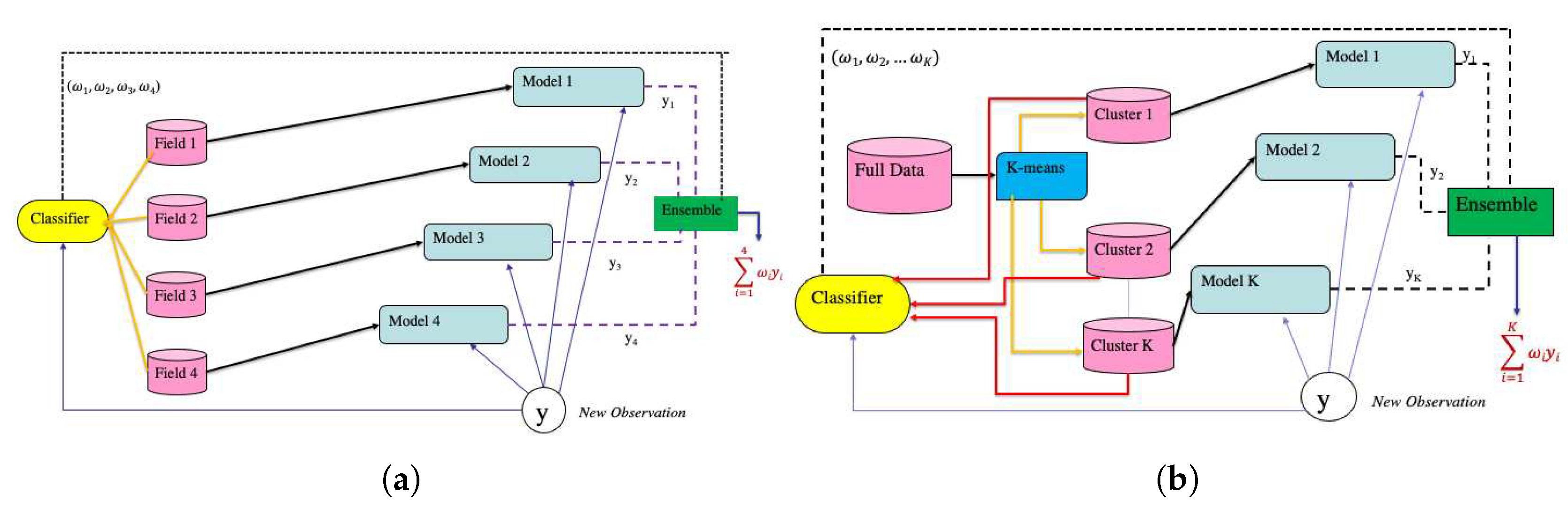
3.2.1. Field Weighted Ensemble Model
- Let denote the dataset for the kth field (dataset for kth task), where denotes the predictors (vegetation indices) and the response (seed yield), . Divide into and , where and denote the training and test set for respectively.
- Fit a random forest regression model [22] to every .
- Let , where for every data point in denote a new dataset. Here denotes the field membership of . Combine row wise to create a combined dataset .
- Fit a multinomial logistic regression classifier [23] on . This classifer provides the ensemble weights for new observations.
3.2.2. Cluster Ensemble Model
- Divide into K homogeneous groups using k-means [24] and elbow method. Let be a variable denoting the cluster memberships of .
- Based on divide into K groups , . is hence the dataset for the kth task.
- Fit a random forest regression model to every , .
- Fit a logistic regression classifier to to determine the ensemble weights.
3.3. Yield Prediction Using Soil Data
4. Results and Discussion
4.1. Hyperspectral Reflectance Data
4.1.1. Analysis Using Derived Vegetation Indices
4.1.2. Ensemble Models
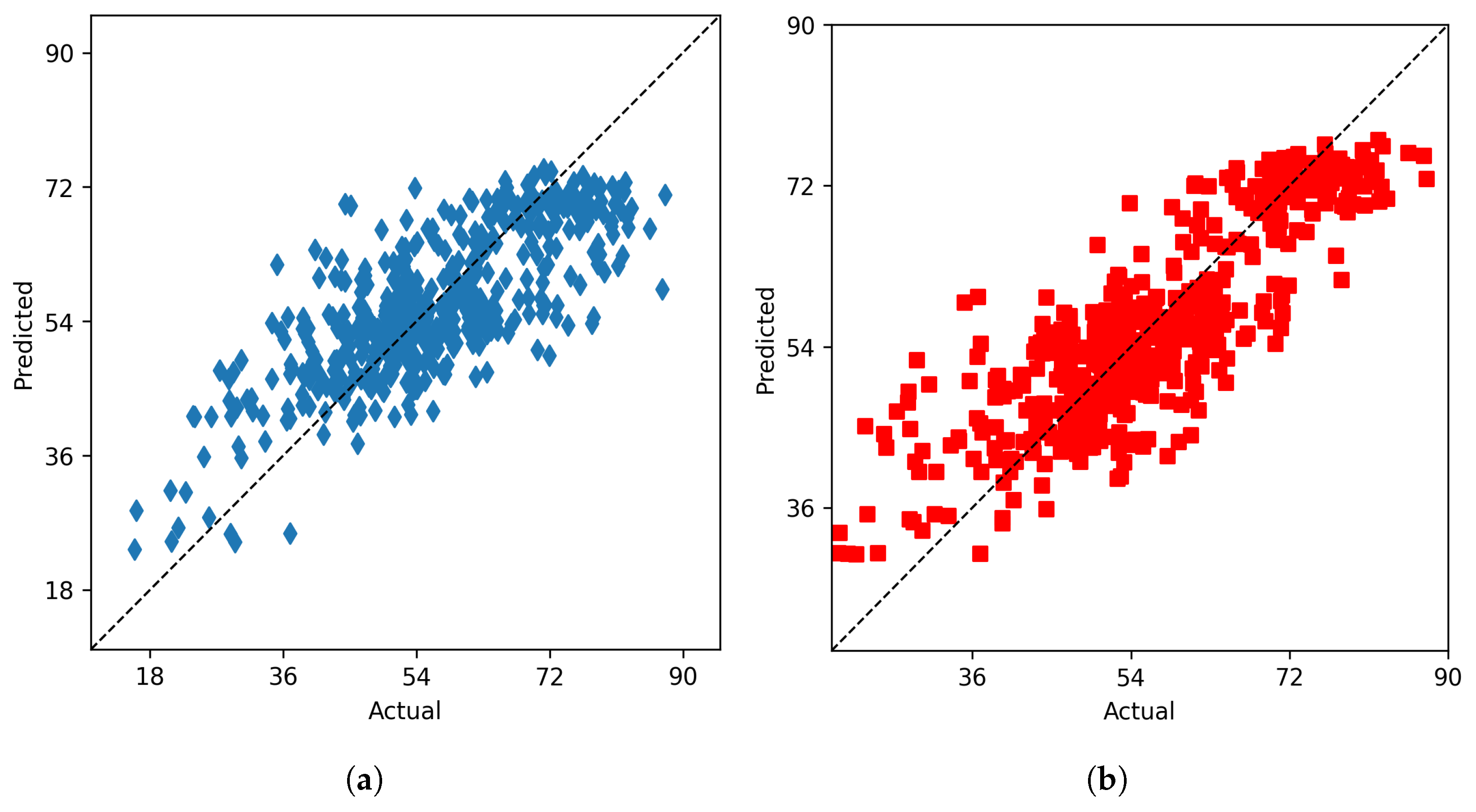
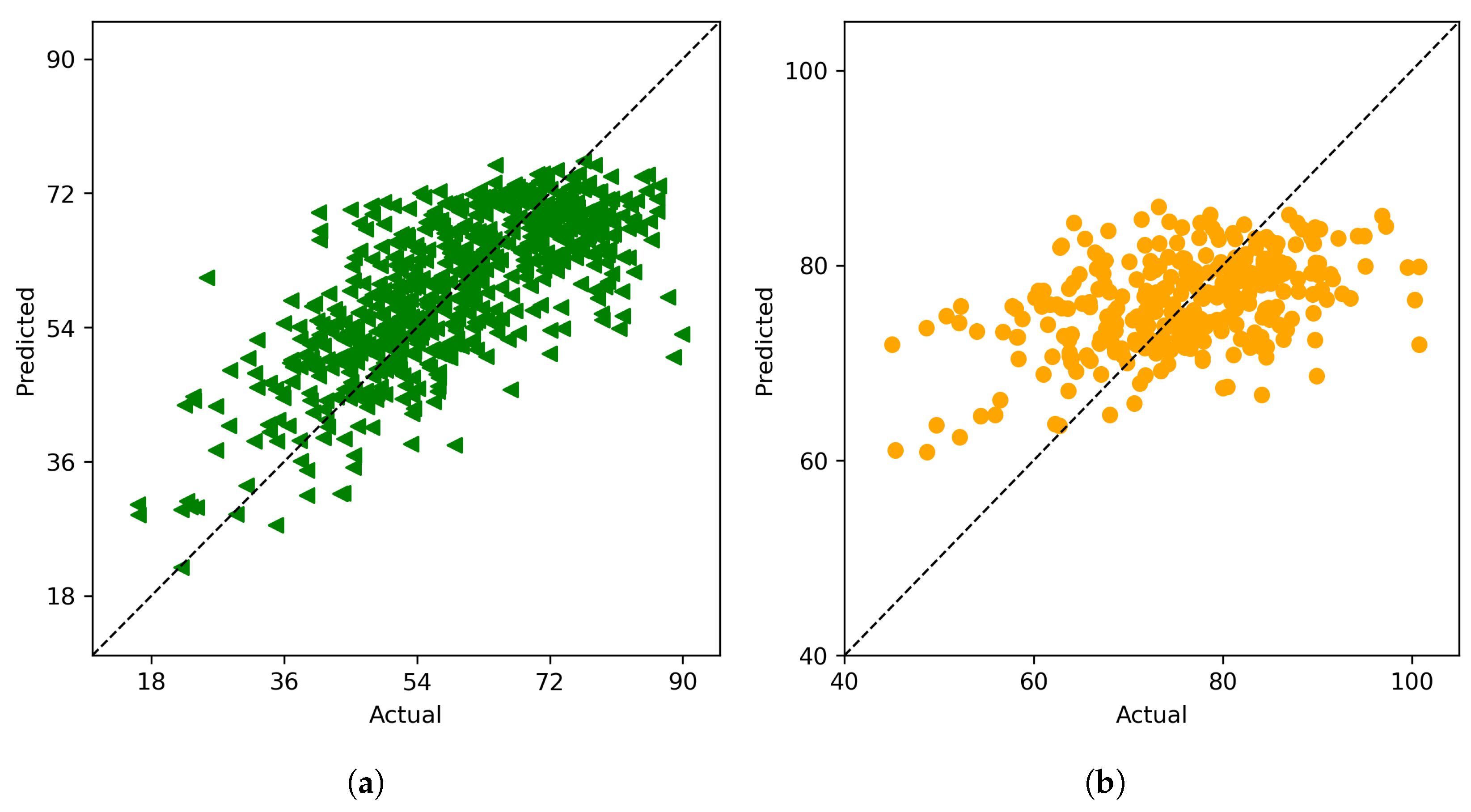
4.1.3. Field Weighted Ensemble Model
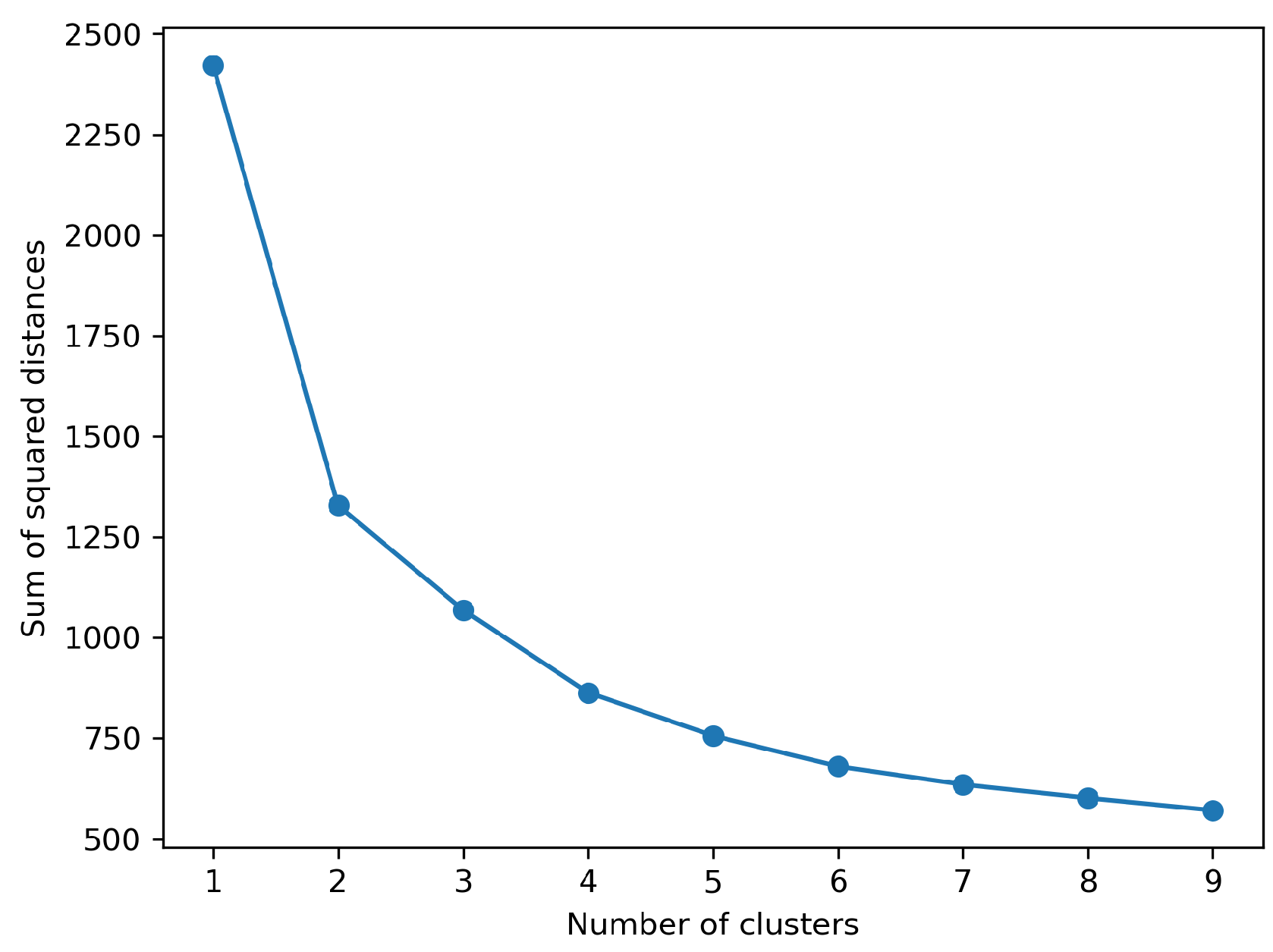
4.1.4. Cluster Ensemble Model
| Cluster No. | Number of observations |
| 1 | 2100 |
| 2 | 428 |
4.2. Analysis Using Soil Data
| Hyperparameter | Values |
|---|---|
| bootstrap | True, False |
| max_depth | 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, None |
| max_features | auto, sqrt |
| min_samples_leaf | 1, 2, 4 |
| min_samples_split | 2, 5, 10 |
| n_estimators | 200, 400, 600, 800, 1000, 1200, 1400, 1600, 1800, 2000 |
| Hyperparameter | Value |
|---|---|
| n_estimators | 400 |
| min_samples_split | 2 |
| min_samples_leaf | 1 |
| max_features | sqrt |
| max_depth | None |
| bootstrap | False |
4.3. Feature Importance
| Variable | Value |
|---|---|
| Population | 0.318527 |
| DDI_1 | 0.088559 |
| Clre_1 | 0.070222 |
| MM_P1 | 0.068096 |
| Clre_3 | 0.036757 |
| MM_CLAY | 0.035482 |
| MM_PH | 0.033846 |
| DPI_1 | 0.033274 |
| GNDV_3 | 0.031830 |
| Clre_2 | 0.029058 |
| MM_SILT | 0.027715 |
| MM_K | 0.023197 |
| MM_MG | 0.020231 |
| GNDV_1 | 0.017468 |
| MM_SAND | 0.016942 |
| Cl_1 | 0.015149 |
| Cl_3 | 0.014004 |
| DDI_2 | 0.013723 |
| DDI_3 | 0.013481 |
| MM_OM | 0.013148 |
| Model | Accuracy |
| Linear Regression | 0.69 |
| Ridge Regression | 0.68 |
| LASSO | 0.68 |
| Random Forest | 0.73 |
| Number of features | Accuracy |
| 5 | 0.74 |
| 3 | 0.70 |
| 2 | 0.68 |
5. Conclusions
Author Contributions
Acknowledgments
Appendix A Table of Vegetation Indices
| Full form | Spectral Index/Ratio | Formula |
|---|---|---|
| Curvature index | Cl | |
| Chlorophyll Index red-edge | Clre | |
| Datt1 | ||
| Datt4 | ||
| Datt6 | ||
| Double difference index | DDI | |
| Double peak index | DPI | |
| Gitelson2 | ||
| Green normalized difference vegetation index | GNDVI | |
| Modified chlorphyll absorption ratio index | MCARI | |
| MCARI3 | ||
| Modified normalized difference | MND1 | |
| MND2 | ||
| Modified simple ratio | mSR | |
| Modified simple ratio 2 | mSR2 | |
| MERIS terrestrail cholrophyll index | MTCI | |
| Modified traingular vegetation index 1 | MTVI1 | |
| Normalized difference 550/531 | ND1 | |
| Normalized difference 682/553 | ND2 | |
| Normalized difference chlorophyll | NDchl | |
| Normalized difference red edge | NDRE | |
| Normalized difference vegetation index | NDVI1 | |
| NDVI2 | ||
| NDVI3 | ||
| Normalized pigment cholrophyll index | NPCL | |
| Normalized difference pigment index | NPQI | |
| Optimized soil-adjusted vegetation index | OSAVI | |
| Plant biochemical index | PBI | |
| Plant pigment ratio | PPR | |
| Physiological reference index | PRI | |
| Pigment-specific normalized difference | PSNDb1 | |
| PSNDc1 | ||
| PSNDc2 | ||
| Plant senescence reflectance index | PSRI | |
| Pigment-specific simple ratio | PSSRc1 | |
| PSSRc2 | ||
| Photosynthetic vigor ratio | PVR | |
| Plant water index | PWI | |
| Renormalized difference vegetation index | RDVI | |
| Red-edge stress vegatation index | RVSI | |
| Soil-adjusted vegatation index | SAVI | |
| Structure intensive pigment index | SIPI | |
| Simple ratio | SR1 | |
| SR2 | ||
| SR3 | ||
| SR4 | ||
| Disease -water stress index 4 | DSWI-4 | |
| Simple ratio pigment index | SRPI | |
| Transformed chlorophyll absorption ratio | TCARI | |
| Traingular cholrophyll index | TCI | |
| Triangular vegetation index | TVI | |
| Water band index | WBI |
References
- Singh, D.P.; Singh, A.K.; Singh, A. Plant breeding and cultivar development; Academic Press, 2021.
- Li, Z.; Chen, Z.; Cheng, Q.; Duan, F.; Sui, R.; Huang, X.; Xu, H. UAV-Based Hyperspectral and Ensemble Machine Learning for Predicting Yield in Winter Wheat. Agronomy 2022, 12. [Google Scholar] [CrossRef]
- Yoosefzadeh-Najafabadi, M.; Torabi, S.; Tulpan, D.; Rajcan, I.; Eskandari, M. Genome-wide association studies of soybean yield-related hyperspectral reflectance bands using machine learning-mediated data integration methods. Frontiers in plant science 2021, p. 2555. [CrossRef]
- Chiozza, M.V.; Parmley, K.A.; Higgins, R.H.; Singh, A.K.; Miguez, F.E. Comparative prediction accuracy of hyperspectral bands for different soybean crop variables: From leaf area to seed composition. Field Crops Research 2021, 271, 108260. [Google Scholar] [CrossRef]
- Shook, J.; Gangopadhyay, T.; Wu, L.; Ganapathysubramanian, B.; Sarkar, S.; Singh, A.K. Crop yield prediction integrating genotype and weather variables using deep learning. PLOS ONE 2021, 16, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Riera, L.G.; Carroll, M.E.; Zhang, Z.; Shook, J.M.; Ghosal, S.; Gao, T.; Singh, A.; Bhattacharya, S.; Ganapathysubramanian, B.; Singh, A.K.; Sarkar, S. Deep Multiview Image Fusion for Soybean Yield Estimation in Breeding Applications. Plant Phenomics 2021, 2021, 9846470. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Carroll, M.E.; Singh, A.; Swetnam, T.L.; Merchant, N.; Sarkar, S.; Singh, A.K.; Ganapathysubramanian, B. UAS-Based Plant Phenotyping for Research and Breeding Applications. Plant Phenomics 2021, 2021, 9840192. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.K.; Singh, A.; Sarkar, S.; Ganapathysubramanian, B.; Schapaugh, W.; Miguez, F.E.; Carley, C.N.; Carroll, M.E.; Chiozza, M.V.; Chiteri, K.O.; Falk, K.G.; Jones, S.E.; Jubery, T.Z.; Mirnezami, S.V.; Nagasubramanian, K.; Parmley, K.A.; Rairdin, A.M.; Shook, J.M.; Van der Laan, L.; Young, T.J.; Zhang, J., High-Throughput Phenotyping in Soybean. In High-Throughput Crop Phenotyping; Zhou, J.; Nguyen, H.T., Eds.; Springer International Publishing: Cham, 2021; pp. 129–163 [CrossRef]
- Nagasubramanian, K.; Jones, S.; Sarkar, S.; Singh, A.K.; Singh, A.; Ganapathysubramanian, B. Hyperspectral band selection using genetic algorithm and support vector machines for early identification of charcoal rot disease in soybean stems. Plant methods 2018, 14, 86. [Google Scholar] [CrossRef] [PubMed]
- Parmley, K.A.; Higgins, R.H.; Ganapathysubramanian, B.; Sarkar, S.; Singh, A.K. Machine Learning Approach for Prescriptive Plant Breeding. Scientific Reports 2019, 9, 17132. [Google Scholar] [CrossRef] [PubMed]
- Parmley, K.; Nagasubramanian, K.; Sarkar, S.; Ganapathysubramanian, B.; Singh, A.K. Development of Optimized Phenomic Predictors for Efficient Plant Breeding Decisions Using Phenomic-Assisted Selection in Soybean. Plant Phenomics 2019, 2019, 5809404. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.; Sharma, B.; Chingtham, P. Forecast of Earthquake Magnitude for North-West (NW) Indian Region Using Machine Learning Techniques 2023. [CrossRef]
- Sarkar, S.; Ganapathysubramanian, B.; Singh, A.; Fotouhi, F.; Kar, S.; Nagasubramanian, K.; Chowdhary, G.; Das, S.K.; Kantor, G.; Krishnamurthy, A.; et al. . Cyber-agricultural systems for crop breeding and sustainable production. Trends in Plant Science 2023. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.K.; Singh, A.; Sarkar, S.; Ganapathysubramanian, B.; Schapaugh, W.; Miguez, F.E.; Carley, C.N.; Carroll, M.E.; Chiozza, M.V.; Chiteri, K.O. High-throughput phenotyping in soybean. High-throughput crop phenotyping 2021, pp. 129–163. [CrossRef]
- Herr, A.W.; Adak, A.; Carroll, M.E.; Elango, D.; Kar, S.; Li, C.; Jones, S.E.; Carter, A.H.; Murray, S.C.; Paterson, A.; others. Unoccupied aerial systems imagery for phenotyping in cotton, maize, soybean, and wheat breeding. Crop Science 2023, 63, 1722–1749. [Google Scholar] [CrossRef]
- Young, T.J.; Jubery, T.Z.; Carley, C.N.; Carroll, M.; Sarkar, S.; Singh, A.K.; Singh, A.; Ganapathysubramanian, B. “Canopy fingerprints” for characterizing three-dimensional point cloud data of soybean canopies. Frontiers in Plant Science 2023, 14, 1141153. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.; Tayal, V.K. Analysis of Twitter Sentiment to Predict Financial Trends. 2023 International Conference on Artificial Intelligence and Smart Communication (AISC). IEEE, 2023, pp. 1027–1031. [CrossRef]
- Huang, F.; Xie, G.; Xiao, R. Research on Ensemble Learning. 2009 International Conference on Artificial Intelligence and Computational Intelligence, 2009, Vol. 3, pp. 249–252. [CrossRef]
- Khaledian, Y.; Miller, B.A. Selecting appropriate machine learning methods for digital soil mapping. Applied Mathematical Modelling 2020, 81, 401–418. [Google Scholar] [CrossRef]
- Bai, G.; Ge, Y.; Hussain, W.; Baenziger, P.S.; Graef, G. A multi-sensor system for high throughput field phenotyping in soybean and wheat breeding. Computers and Electronics in Agriculture 2016, 128, 181–192. [Google Scholar] [CrossRef]
- Raschka, S. MLxtend: Providing machine learning and data science utilities and extensions to Python’s scientific computing stack. The Journal of Open Source Software 2018, 3. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests. Machine Learning 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Abdillah, A.; Sutisna, A.; Tarjiah, I.; Fitria, D.; Widiyarto, T. Application of Multinomial Logistic Regression to analyze learning difficulties in statistics courses. Journal of Physics: Conference Series 2020, 1490, 012012. [Google Scholar] [CrossRef]
- MacQueen, J. Some methods for classification and analysis of multivariate observations. Proceedings of the Fifth Berkeley Symposium on Mathematical Statistics and Probability, Volume 1: Statistics; University of California Press: Berkeley, California, 1967; pp. 281–297. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; Vanderplas, J.; Passos, A.; Cournapeau, D.; Brucher, M.; Perrot, M.; Duchesnay, E. Scikit-learn: Machine Learning in Python. Journal of Machine Learning Research 2011, 12, 2825–2830. [Google Scholar]
| Field No. | Number of observations |
| 1 | 770 |
| 2 | 912 |
| 3 | 800 |
| 4 | 679 |
| Model | Accuracy |
| Linear Regression | 0.46 |
| Ridge Regression | 0.45 |
| LASSO | 0.41 |
| Random Forest | 0.55 |
| Number of features | Accuracy |
| 40 | 0.55 |
| 20 | 0.53 |
| 10 | 0.50 |
| 5 | 0.47 |
| Field No. | Accuracy |
| 1 | 0.09 |
| 2 | -0.17 |
| 3 | -0.19 |
| 4 | -0.005 |
| Field ensemble | 0.21 |
| Cluster No. | Accuracy |
| 1 | 0.39 |
| 2 | 0.28 |
| Cluster ensemble | 0.51 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




