Submitted:
12 October 2023
Posted:
12 October 2023
You are already at the latest version
Abstract
Keywords:
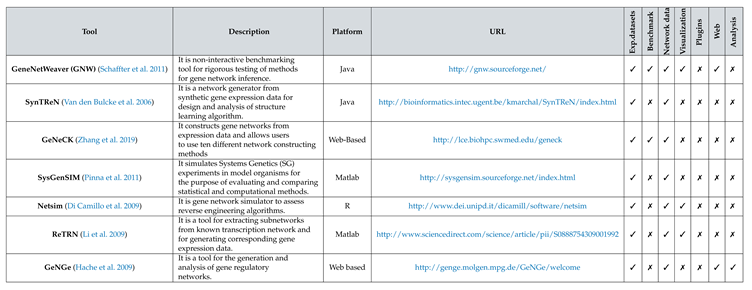
1. Introduction
- A lightweight tool built upon NextJS, Python, R and Flask that provides the user to infer, visualize, and network analysis of large networks with ease.
- Offered many complex network analysis facilities applicable to any network.
- Server-side rendering is used for interactive visualization of large networks to reduce client burden.
- A parallel inference engine has been integrated that can make any inference method handle large networks efficiently.
- A benchmarking facility has been integrated to assess the prediction quality of inference methods on user-given networks. Our tool can also assess and compare any newly developed inference method with 11 candidate methods.
- We evaluate the performance of eleven (11) candidate methods using our tool and compare their qualitative performance and the characteristics of the networks produced.
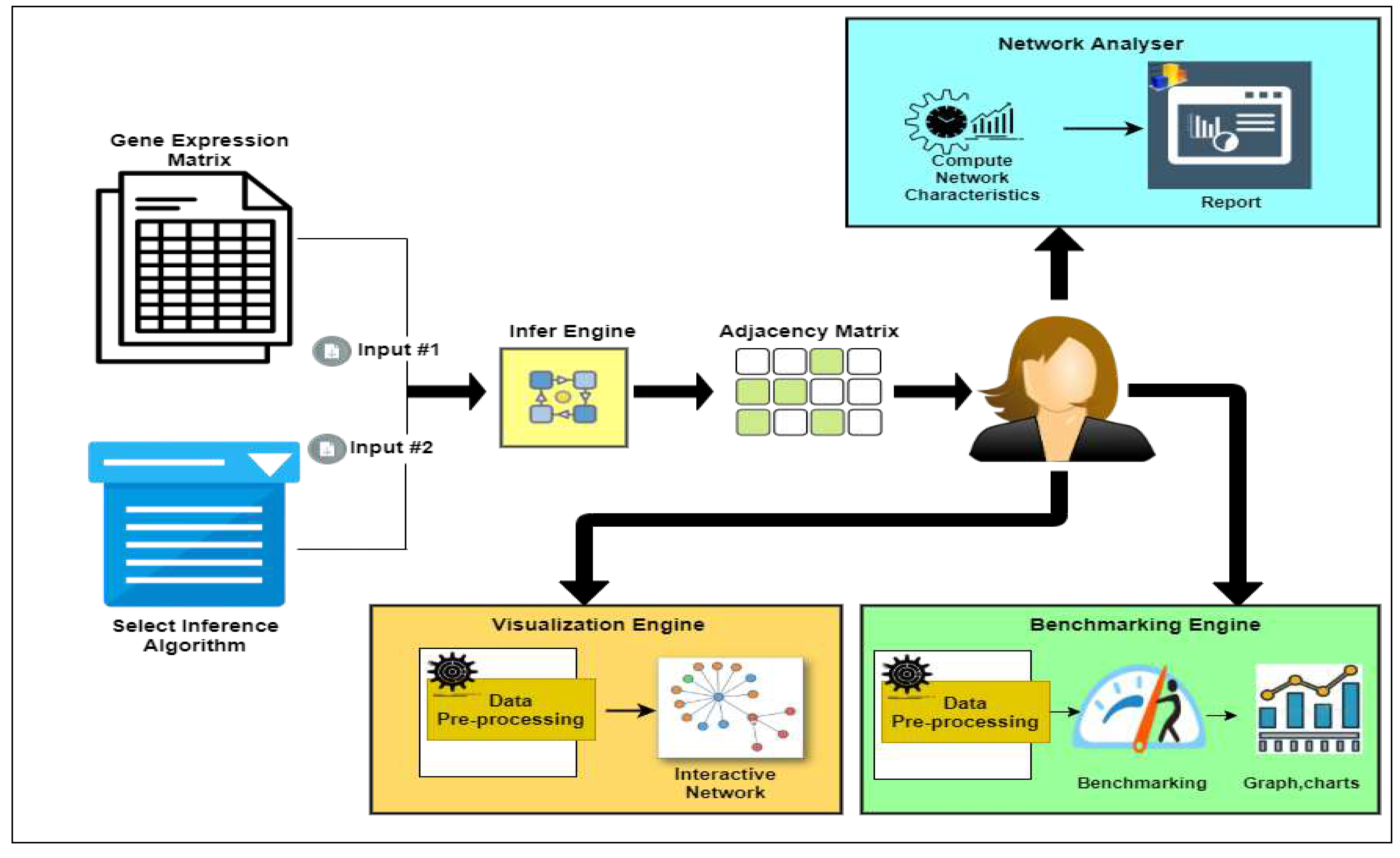
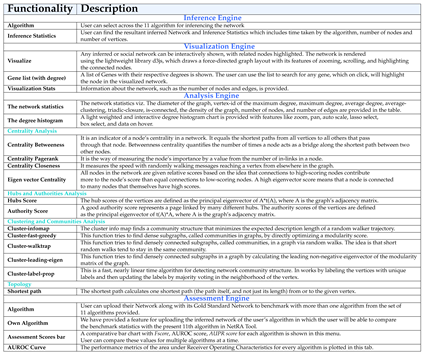
2. The Workflow of NetRA Tool
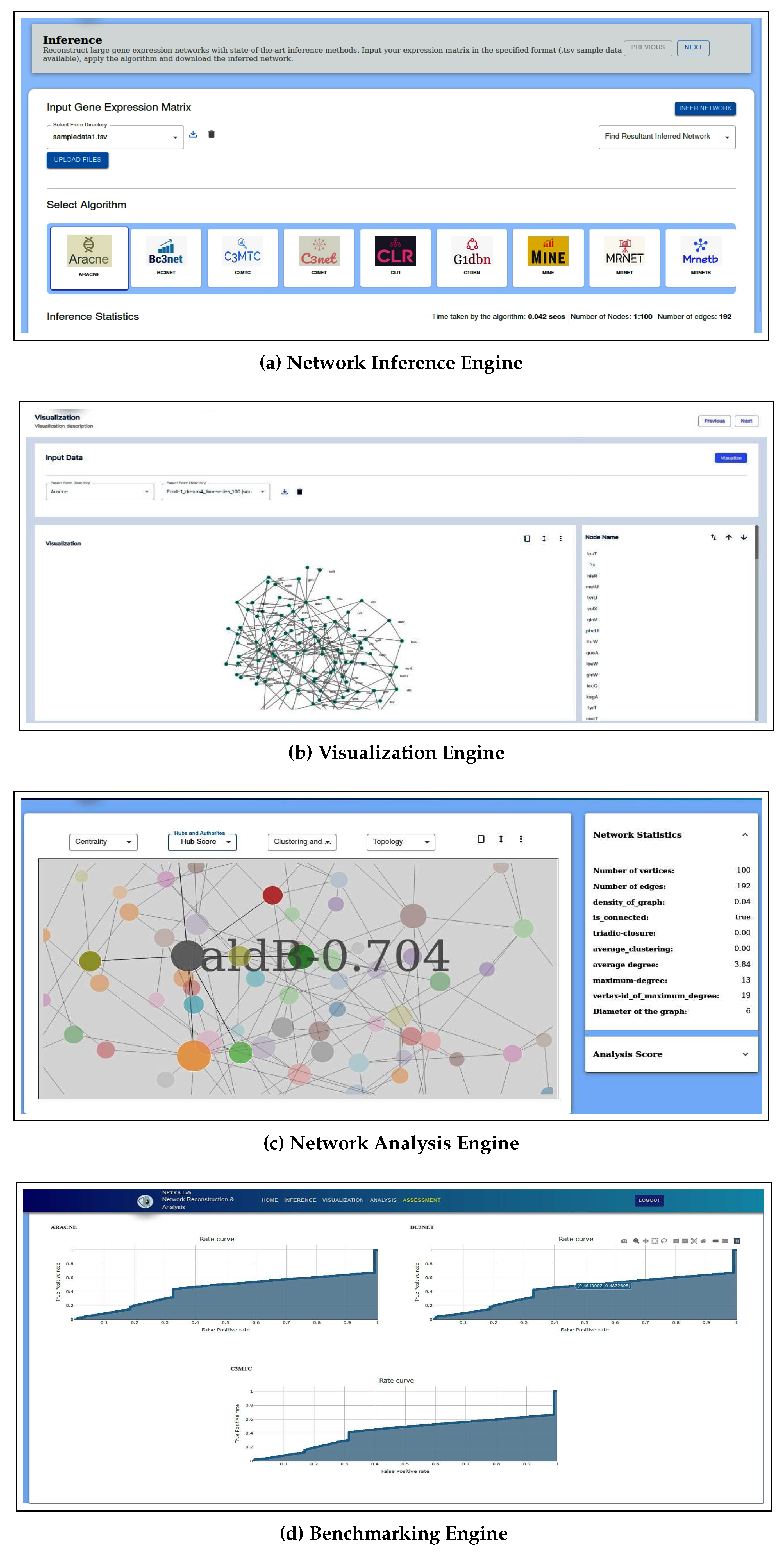
2.1. Inference Engine
2.1.1. Parallel Framework for Large -Scale Inference
2.2. Visualization Engine
2.3. Analysis Engine
2.4. Assessment Engine
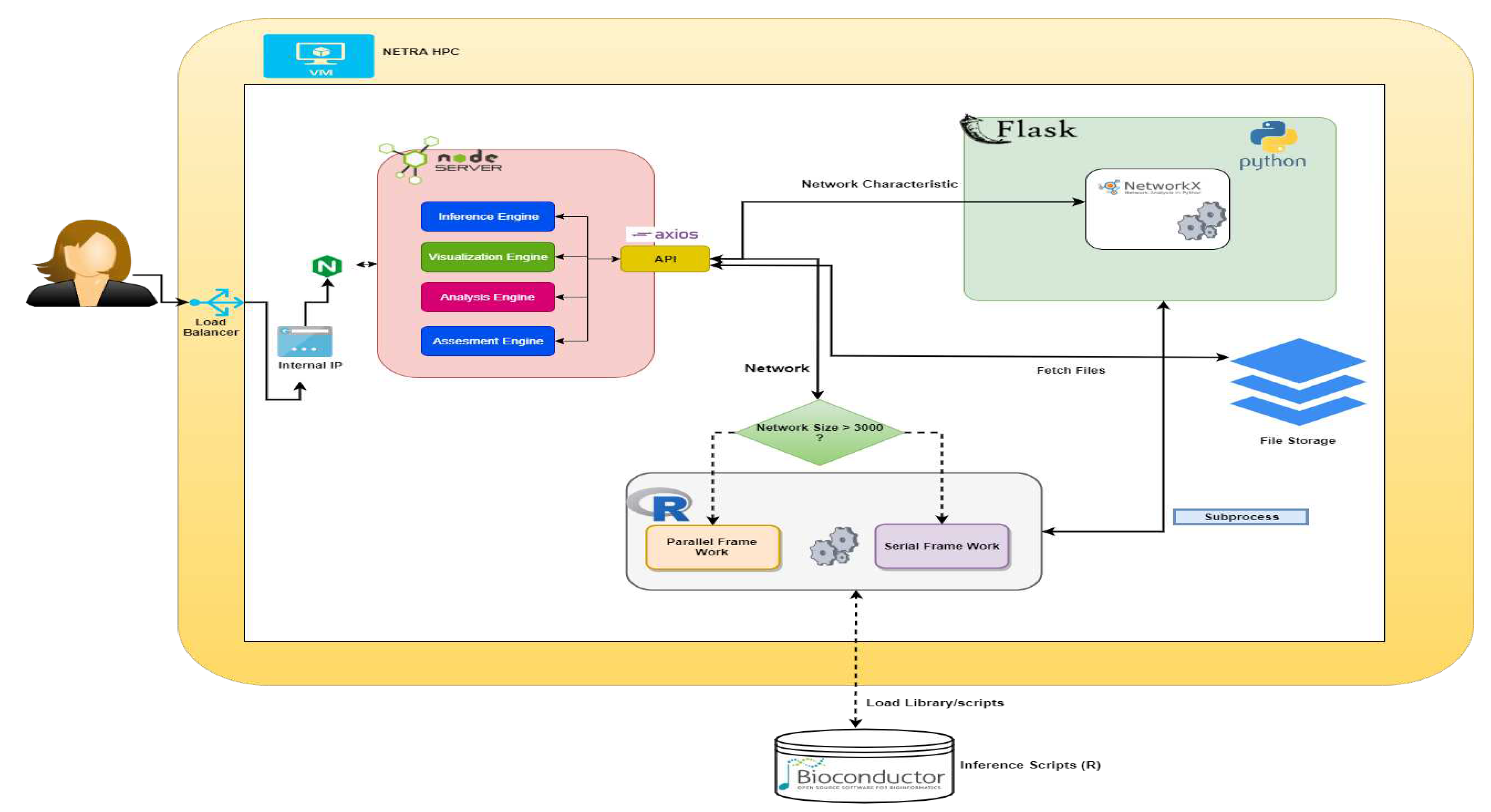
3. Inner Engineering of NetRA Tool
4. Qualitative Assessment of Inference Methods
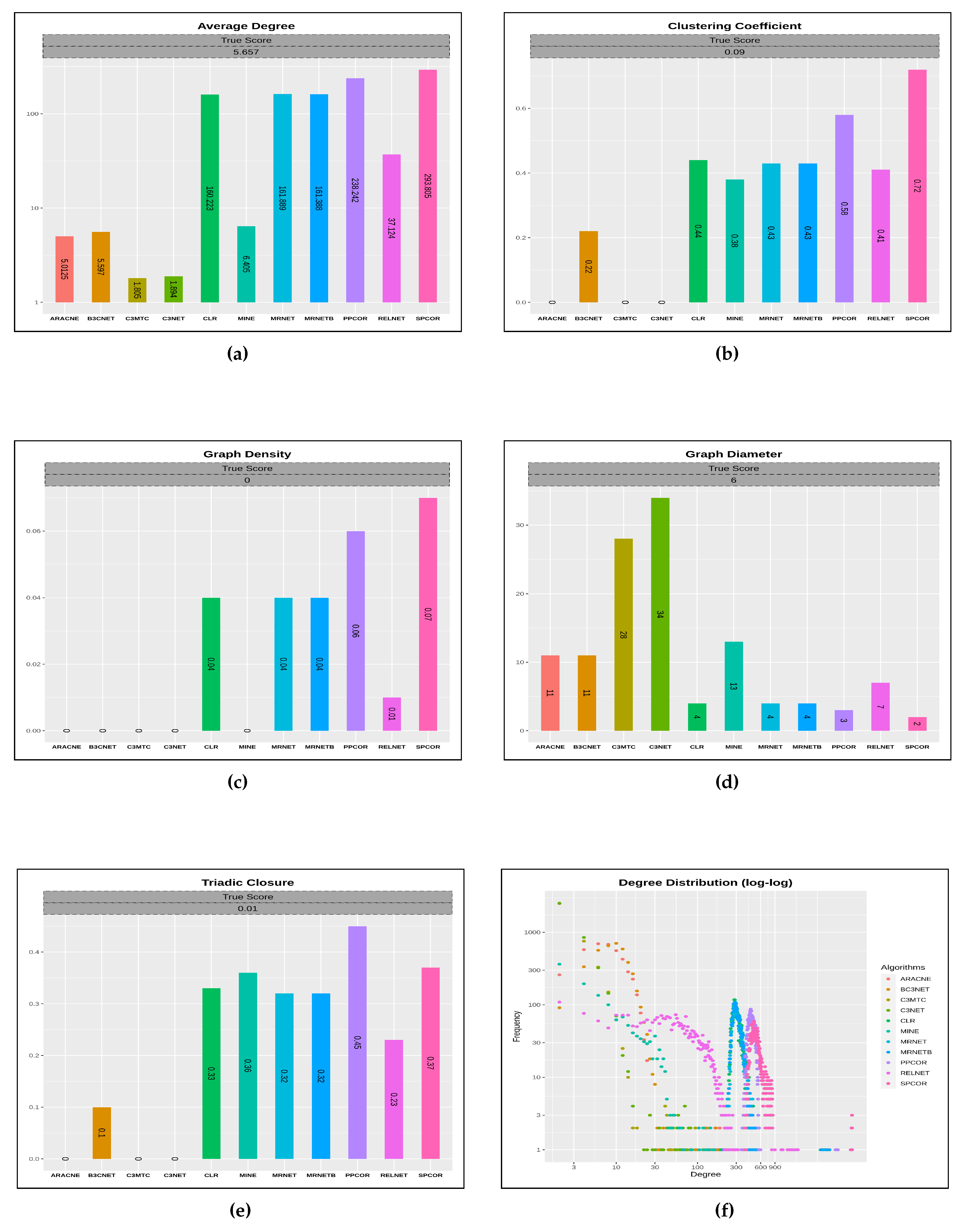
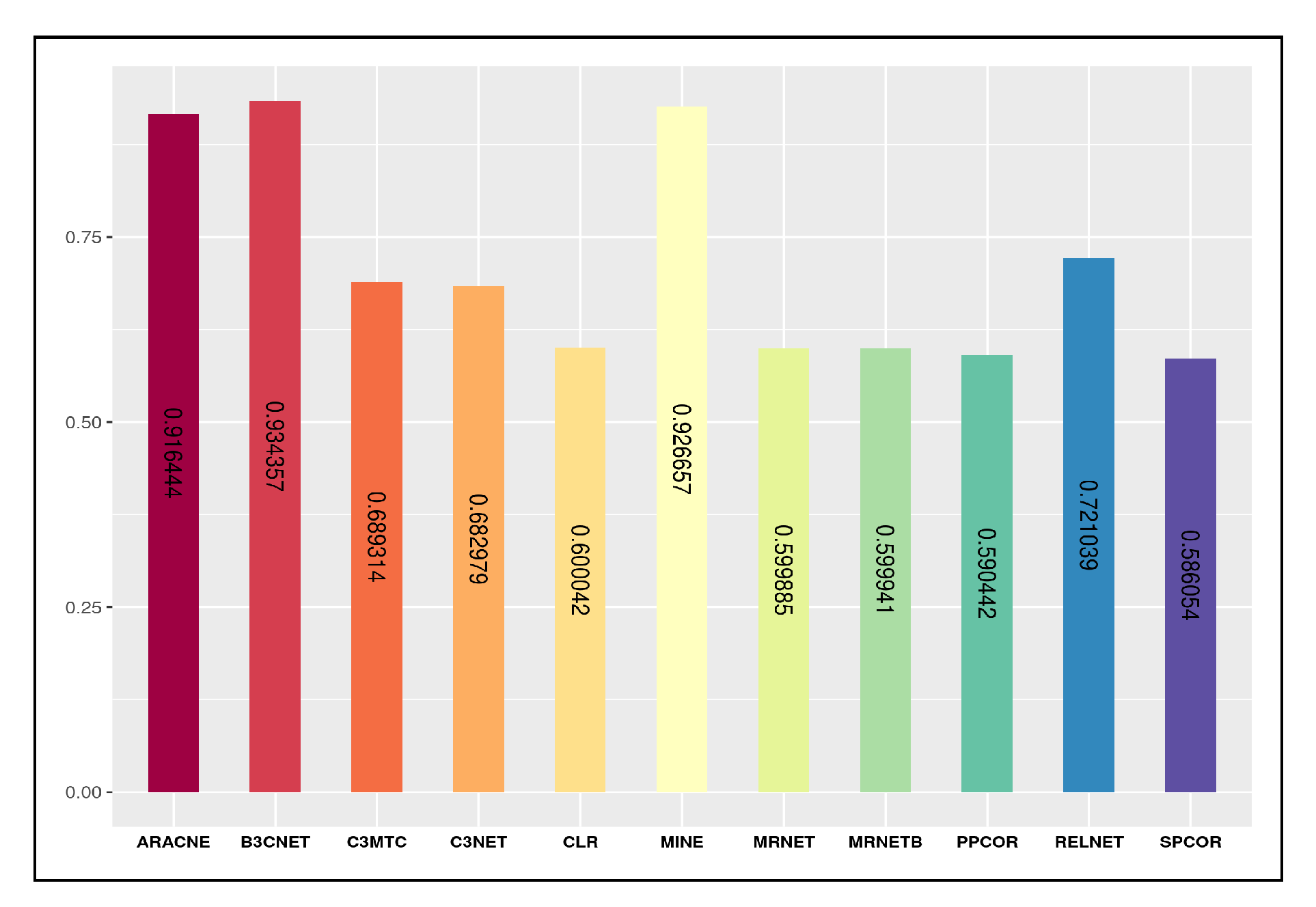
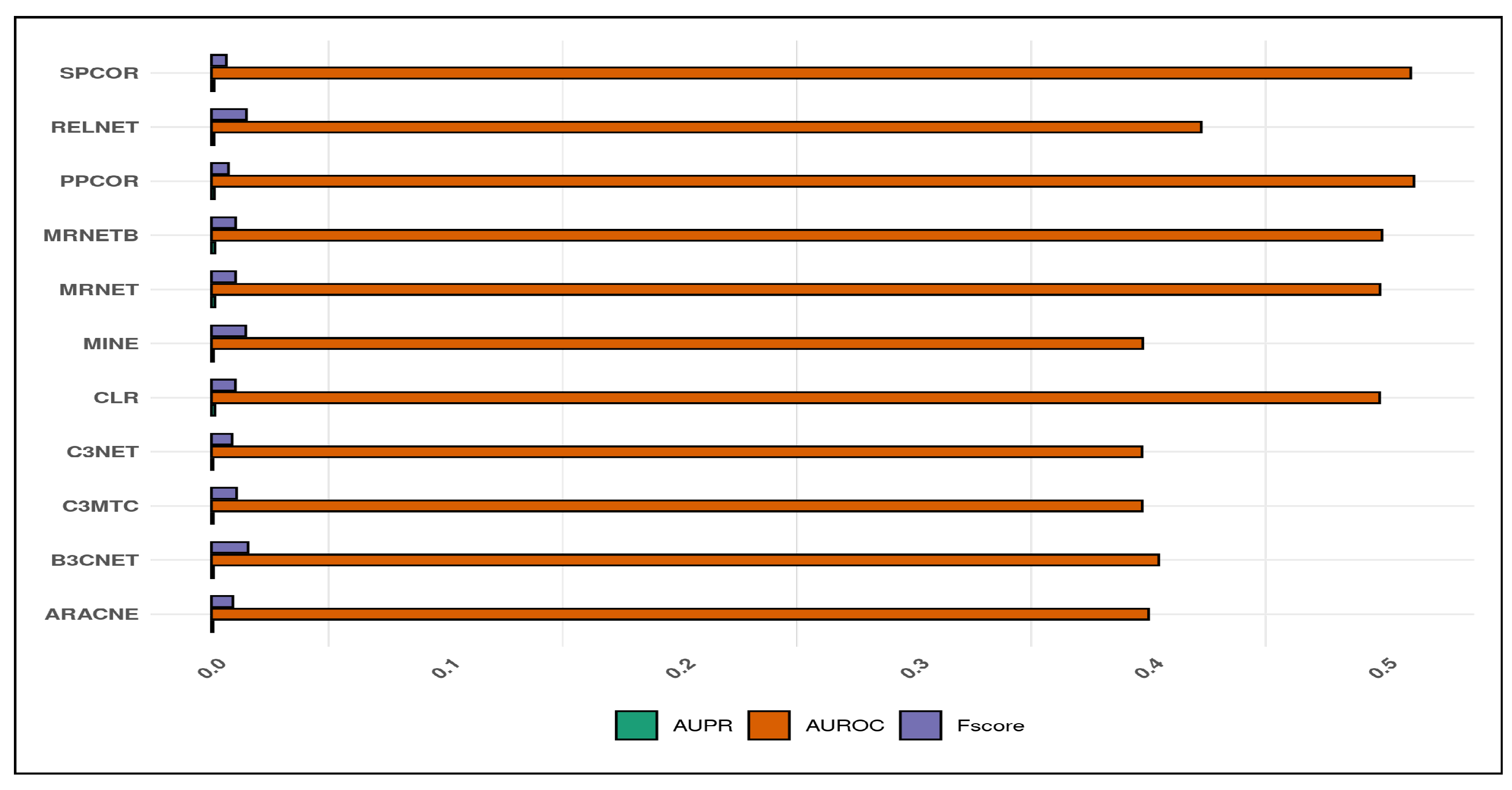
4.1. Characteristics of Predicted Networks
4.2. Comparative Analysis of Inference Quality
5. Conclusion
Data Availability Statement
Acknowledgments
References
- Pietro Hiram Guzzi and Swarup Roy. Biological Network Analysis: Trends, Approaches, Graph Theory, and Algorithms. Elsevier, 2020.
- Alberto de la Fuente. What are gene regulatory networks? In Handbook of research on computational methodologies in gene regulatory networks, pages 1–27. IGI Global, 2010.
- Binon Teji, Jayanta K Das, Swarup Roy, and Dinabandhu Bhandari. Predicting missing links in gene regulatory networks using network embeddings: A qualitative assessment of selective embedding techniques. In Intelligent Systems, pages 143–154. Springer, 2022.
- Binon Teji and Swarup Roy. Missing link identification from node embeddings using graph auto encoders and its variants. In 2022 OITS International Conference on Information Technology (OCIT), pages 1–6. IEEE, 2022.
- Ardeshir Bayat. Science, medicine, and the future-bioinformatics. BMJ-BRITISH MEDICAL JOURNAL, 324(7344):1018–1022, 2002.
- Thomas Schaffter, Daniel Marbach, and Dario Floreano. Genenetweaver: in silico benchmark generation and performance profiling of network inference methods. Bioinformatics, 27(16):2263–2270, 2011.
- Tim Van den Bulcke, Koenraad Van Leemput, Bart Naudts, Piet van Remortel, Hongwu Ma, Alain Verschoren, Bart De Moor, and Kathleen Marchal. Syntren: a generator of synthetic gene expression data for design and analysis of structure learning algorithms. BMC bioinformatics, 7(1):1–12, 2006.
- Minzhe Zhang, Qiwei Li, Donghyeon Yu, Bo Yao, Wei Guo, Yang Xie, and Guanghua Xiao. Geneck: a web server for gene network construction and visualization. BMC bioinformatics, 20(1):1–7, 2019.
- Andrea Pinna, Nicola Soranzo, Ina Hoeschele, and Alberto de la Fuente. Simulating systems genetics data with sysgensim. Bioinformatics, 27(17):2459–2462, 2011.
- Barbara Di Camillo, Gianna Toffolo, and Claudio Cobelli. A gene network simulator to assess reverse engineering algorithms. Annals of the New York Academy of Sciences, 1158(1):125–142, 2009.
- Yong Li, Yanming Zhu, Xi Bai, Hua Cai, Wei Ji, and Dianjing Guo. Retrn: A retriever of real transcriptional regulatory network and expression data for evaluating structure learning algorithm. Genomics, 94(5):349–354, 2009.
- Hendrik Hache, Christoph Wierling, Hans Lehrach, and Ralf Herwig. Genge: systematic generation of gene regulatory networks. Bioinformatics, 25(9):1205–1207, 2009.
- Robert C Gentleman, Vincent J Carey, Douglas M Bates, Ben Bolstad, Marcel Dettling, Sandrine Dudoit, Byron Ellis, Laurent Gautier, Yongchao Ge, Jeff Gentry, et al. Bioconductor: open software development for computational biology and bioinformatics. Genome biology, 5(10):1–16, 2004.
- Jeremiah J Faith, Boris Hayete, Joshua T Thaden, Ilaria Mogno, Jamey Wierzbowski, Guillaume Cottarel, Simon Kasif, James J Collins, and Timothy S Gardner. Large-scale mapping and validation of escherichia coli transcriptional regulation from a compendium of expression profiles. PLoS biology, 5(1):e8, 2007.
- Adam A Margolin, Ilya Nemenman, Katia Basso, Chris Wiggins, Gustavo Stolovitzky, Riccardo Dalla Favera, and Andrea Califano. Aracne: an algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. In BMC bioinformatics, volume 7, pages 1–15. BioMed Central, 2006.
- Patrick E Meyer, Kevin Kontos, Frederic Lafitte, and Gianluca Bontempi. Information-theoretic inference of large transcriptional regulatory networks. EURASIP journal on bioinformatics and systems biology, 2007:1–9, 2007.
- Ricardo de Matos Simoes and Frank Emmert-Streib. Bagging statistical network inference from large-scale gene expression data. PloS one, 7(3):e33624, 2012.
- Ricardo de Matos Simoes and Frank Emmert-Streib. Influence of statistical estimators of mutual information and data heterogeneity on the inference of gene regulatory networks. PLoS One, 6(12):e29279, 2011.
- Gökmen Altay and Frank Emmert-Streib. Inferring the conservative causal core of gene regulatory networks. BMC systems biology, 4(1):1–13, 2010.
- Patrick Meyer, Daniel Marbach, Sushmita Roy, and Manolis Kellis. Information-theoretic inference of gene networks using backward elimination. In BioComp, pages 700–705, 2010.
- Seongho Kim. ppcor: an r package for a fast calculation to semi-partial correlation coefficients. Communications for statistical applications and methods, 22(6):665, 2015.
- Atul J Butte. Mutual information relevance networks: functional genomic networks built from pair-wise entropy measurements. PhD thesis, Massachusetts Institute Of Technology, 2002.
- Softya Sebastian, Swarup Roy, and Jugal Kalita. A generic parallel framework for inferring large-scale gene regulatory networks from expression profiles: Application to alzheimer’s disease network. Briefings in Bioinformatics, 2022.
- Softya Sebastian, Swarup Roy, and Jugal Kalita. Parallel framework for inferring genome scale gene regulatory networks. bioRxiv, 2021.
- Sumit Dutta and Swarup Roy. Complex network visualisation using javascript: A review. Intelligent Systems, pages 45–53, 2022.
- Gabor Csardi, Tamas Nepusz, et al. The igraph software package for complex network research. InterJournal, complex systems, 1695(5):1–9, 2006.
- Patrick E Meyer, Frederic Lafitte, and Gianluca Bontempi. minet: Ar/bioconductor package for inferring large transcriptional networks using mutual information. BMC bioinformatics, 9(1):1–10, 2008.
| 1 | |
| 2 | |
| 3 | |
| 4 | |
| 5 |
 |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




