Submitted:
01 October 2023
Posted:
02 October 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Information
2.2. Data Information
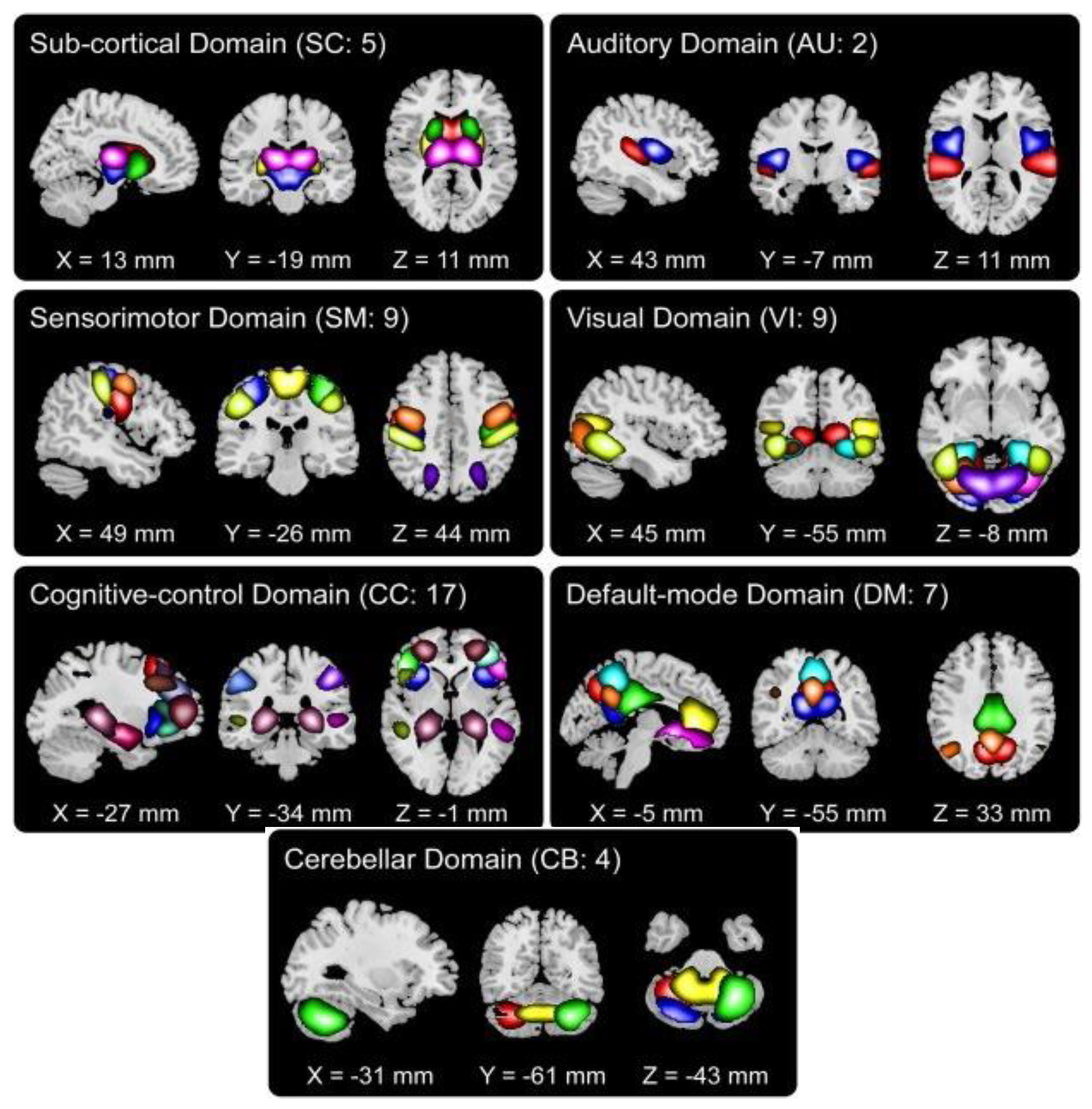
2.3. Data-Driven Node Estimation
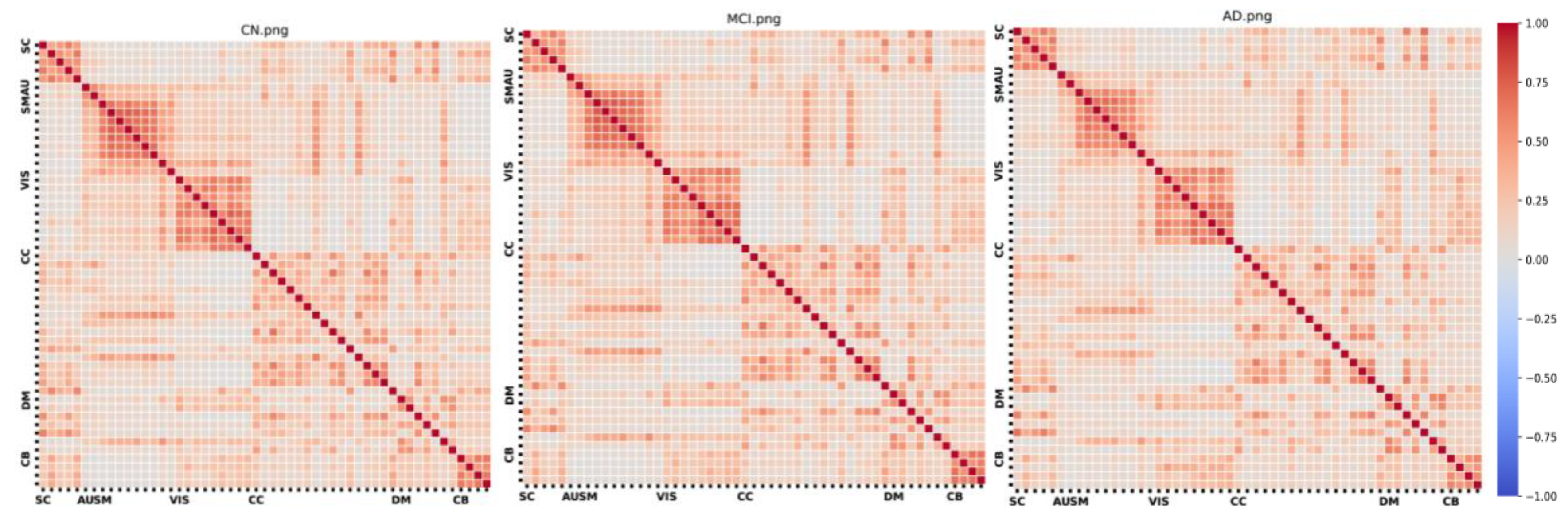
2.4. Estimation of Edge and Local/Global Metrics
2.5. Quantification of Node Size
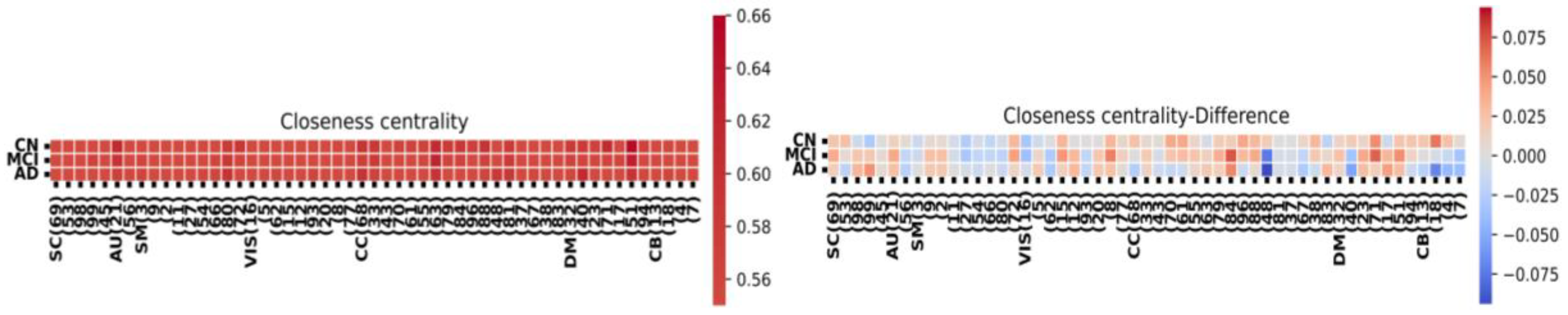
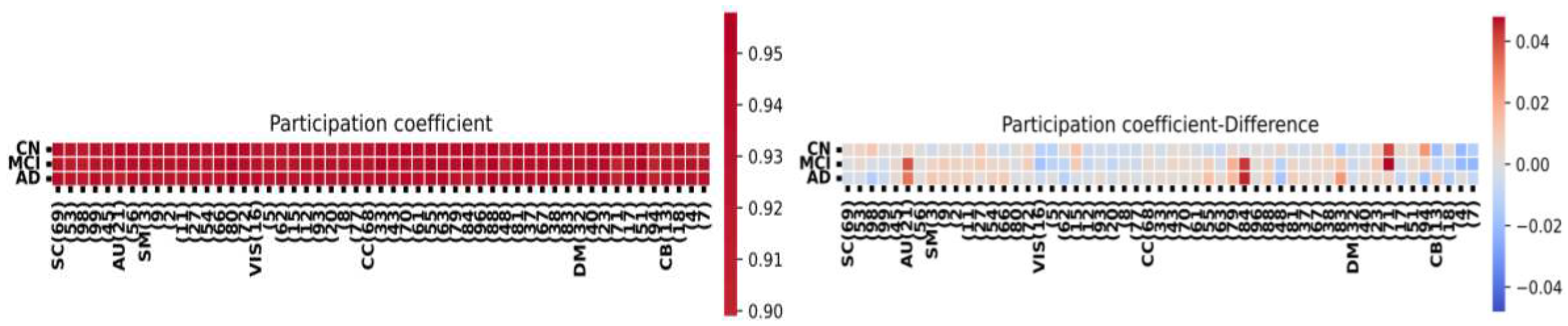
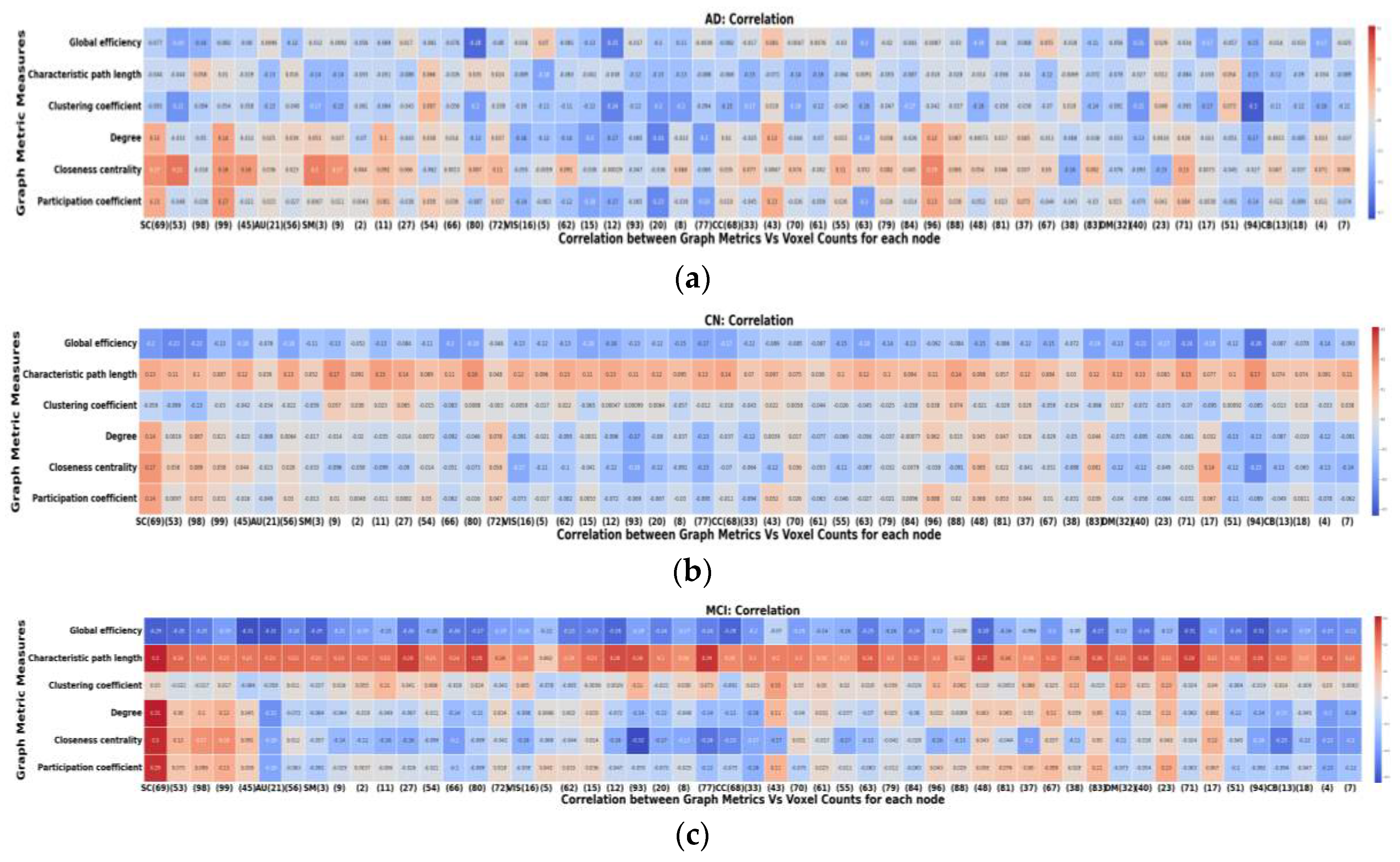
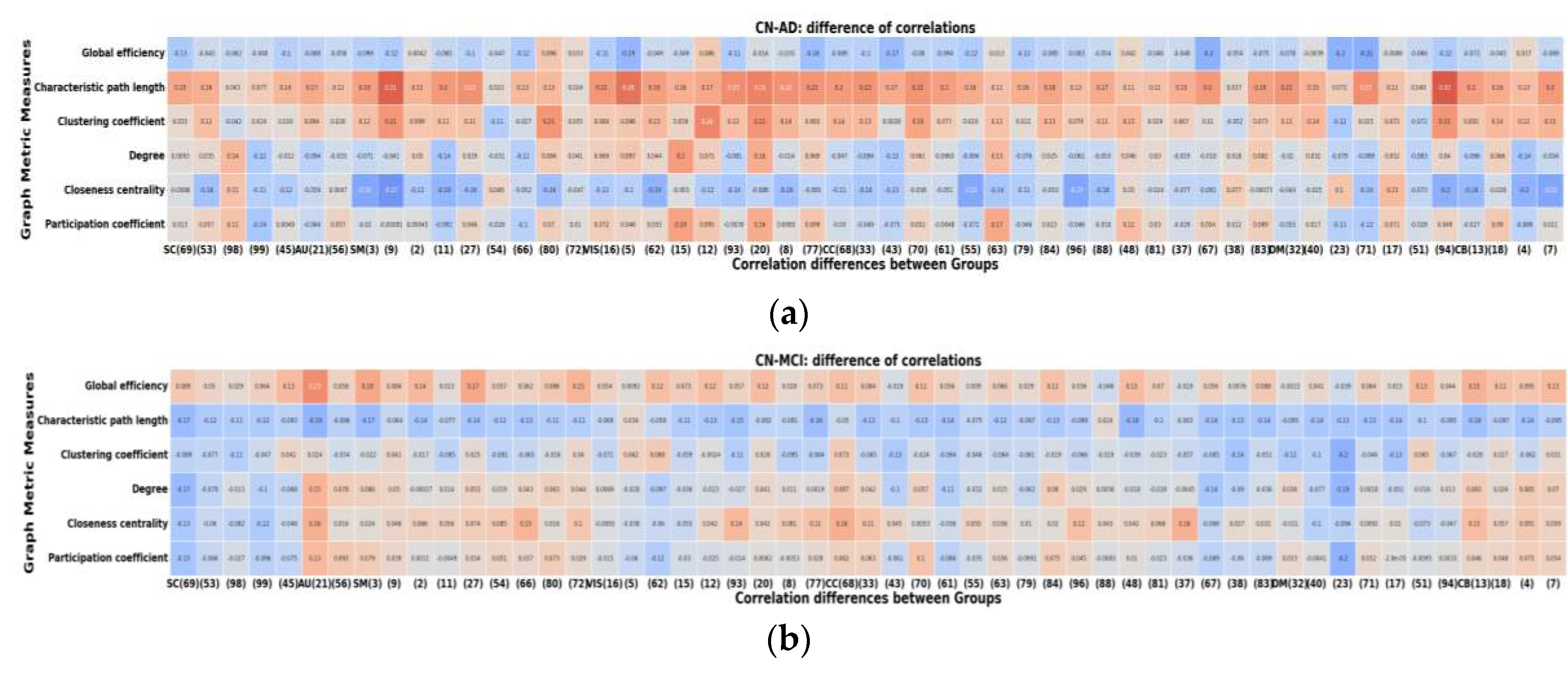
2.6. Node-Metric Coupling
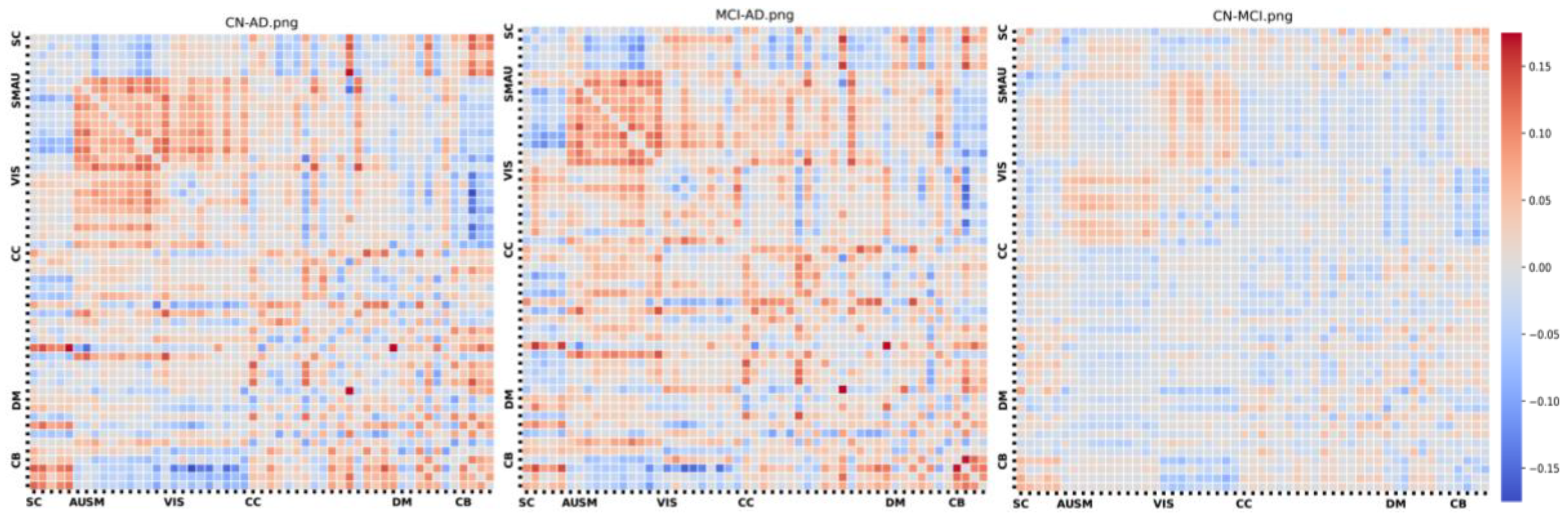
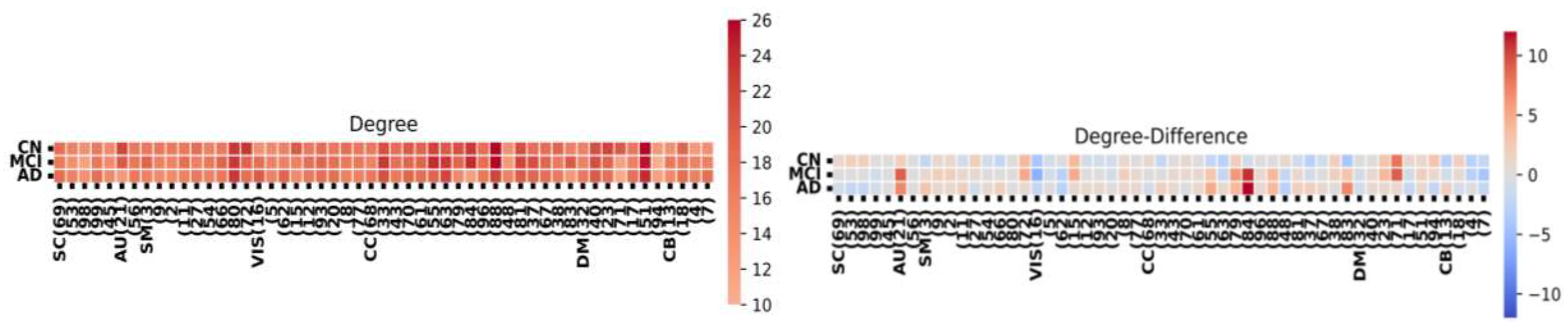
3. Results
4. Discussion
5. Conclusions
References
- Fornito, A. Graph theoretic analysis of human brain networks. In FMRI Techniques and Protocols; Springer, 2016; pp. 283–314. [Google Scholar]
- Telesford, Q.K.; Simpson, S.L.; Burdette, J.H.; Hayasaka, S.; Laurienti, P.J. The brain as a complex system: using network science as a tool for understanding the brain. Brain connectivity 2011, 1, 295–308. [Google Scholar] [CrossRef]
- Fornito, A.; Zalesky, A.; Bullmore, E. Fundamentals of brain network analysis; Academic Press, 2016. [Google Scholar]
- van den Heuvel, M.P.; Fornito, A. Brain networks in schizophrenia. Neuropsychology review 2014, 24, 32–48. [Google Scholar] [CrossRef]
- Falakshahi, H.; Vergara, V.M.; Liu, J.; Mathalon, D.H.; Ford, J.M.; Voyvodic, J.; Mueller, B.A.; Belger, A.; McEwen, S.; Potkin, S.G. ; and others, Meta-modal Information Flow: A Method for Capturing Multimodal Modular Disconnectivity in Schizophrenia. IEEE Transactions on Biomedical Engineering 2020. [Google Scholar] [CrossRef] [PubMed]
- Gordon, E.M.; Laumann, T.O.; Adeyemo, B.; Huckins, J.F.; Kelley, W.M.; Petersen, S.E. Generation and evaluation of a cortical area parcellation from resting-state correlations. Cerebral cortex 2016, 26, 288–303. [Google Scholar] [CrossRef] [PubMed]
- Glasser, M.F.; Coalson, T.S.; Robinson, E.C.; Hacker, C.D.; Harwell, J.A.Y.E.A.U.K.A.A.J.; Beckmann, C.F.; Jenkinson, M.; and others. A multi-modal parcellation of human cerebral cortex. Nature 2016, 536, 171–178. [Google Scholar] [CrossRef] [PubMed]
- Craddock, N.; Jones, I. Genetics of bipolar disorder. Journal of medical genetics 1999, 36, 585–594. [Google Scholar] [CrossRef] [PubMed]
- van den Heuvel, M.P.; de Lange, S.C.; Zalesky, A.; Seguin, C.; Yeo, B.T.; Schmidt, R. Proportional thresholding in resting-state fMRI functional connectivity networks and consequences for patient-control connectome studies: Issues and recommendations. Neuroimage 2017, 152, 437–449. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Fu, Z.; Sui, J.; Gao, S.; Xing, Y.; Lin, D.; Salman, M.; Abrol, A.; Rahaman, M.A.; Chen, J.; and others. NeuroMark: An automated and adaptive ICA based pipeline to identify reproducible fMRI markers of brain disorders. NeuroImage: Clinical 2020, 28, 102375. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Fan, Y. Group information guided ICA for fMRI data analysis. Neuroimage 2013, 69, 157–197. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Allen, E.A.; He, H.; Sui, J.; Wu, L.; Calhoun, V.D. Artifact removal in the context of group ICA: A comparison of single-subject and group approaches. Human brain mapping 2016, 37, 1005–1025. [Google Scholar] [CrossRef] [PubMed]
- M. Rubinov and O. Sporns, Complex network measures of brain connectivity: uses and interpretations. Neuroimage 2010, 52, 1059–1069. [Google Scholar] [CrossRef] [PubMed]
- Glasser, M.F.; Coalson, T.S.; Robinson, E.C.; Hacker, C.D.; Harwell, J.; Yacoub, E.; Ugurbil, K.; Andersson, J.; Beckmann, C.F.; Jenkinson, M. ; and others, A multi-modal parcellation of human cerebral cortex. Nature 2016, 536, 171–178. [Google Scholar] [CrossRef] [PubMed]
- Sporns, O. Graph theory methods: applications in brain networks. Dialogues in clinical neuroscience 2022. [Google Scholar] [CrossRef] [PubMed]
- Blevins, A.S.; Bassett, D.S. Reorderability of node-filtered order complexes. Physical Review E 2020, 101, 052311. [Google Scholar] [CrossRef] [PubMed]
- Anderson, R.J.; Long, C.M.; Calabrese, E.D.; Robertson, S.H.; Johnson, G.A.; Cofer, G.P.; R. J. O'Brien; Badea, A. Optimizing diffusion imaging protocols for structural connectomics in mouse models of neurological conditions. Frontiers in physics 2020, 8, 88. [Google Scholar] [CrossRef] [PubMed]
- Staiger, C.; Cadot, S.; Kooter, R.; Dittrich, M.; Muller, T.; Klau, G.W.; Wessels, L.F. A critical evaluation of network and pathway-based classifiers for outcome prediction in breast cancer. PloS one 2012, 7, e34796. [Google Scholar] [CrossRef] [PubMed]
- Yu, Q.; Du, Y.; Chen, J.; He, H.; Sui, J.; Pearlson, G.; Calhoun, V.D. Comparing brain graphs in which nodes are regions of interest or independent components: A simulation study. Journal of neuroscience methods 2017, 291, 61–68. [Google Scholar] [CrossRef] [PubMed]
| Group Categories | Number of Subjects | M/F count | Average Subject Age | |
|---|---|---|---|---|
| Male | Female | |||
| Healthy Controls (HC) | 947 | 384 | 563 | 74 |
| Mild Cognitive Impairment (MCI) | 368 | 199 | 169 | 73 |
| Alzheimer’s Disease (AD) | 213 | 115 | 98 | 75 |
| Graph Metrics | Formula |
|---|---|
| Node degree |
represents the connection weights. |
| Participation coefficient | |
| Closeness centrality | |
| Global efficiency |
shortest weighted path length between i and j |
| Characteristic path length | |
| Clustering coefficient |
Where the number of triangles is and represents the connection weights. |
| Graph Metric | HC | MCI | AD |
|---|---|---|---|
| Global Efficiency | 0.646 | 0.639 | 0.634 |
| Characteristic Path Length | 1.823 | 1.858 | 1.858 |
| Clustering Coefficient | 0.689 | 0.713 | 0.693 |
| Graph Metric | HC-MCI | HC-AD | MCI-AD |
|---|---|---|---|
| Global Efficiency | 4.410 | 2.667 | 6.361 |
| Characteristic Path Length | 1.505 | 1.6557 | 1.0 |
| Clustering Coefficient | 2.0458 | 0.4387 | 0.0005 |
| Graph Metric | HC-MCI | HC-AD | MCI-AD |
|---|---|---|---|
| Global Efficiency | 7.066(0.07) | *11.64(0.012) | *4.029(0.005) |
| Characteristic Path Length | *-9.304(0.035) | *-7.986(0.035) | 0.0(0.0) |
| Clustering Coefficient | *-5.668(0.024) | *-0.775(0.004) | *3.459(0.020) |
| correlation | |
|---|---|
| HC (significant value) | 0.12 |
| AD (significant value) | 0.26 |
| MCI (significant value) | 0.20 |
| MCI-CN | 0.11 |
| AD-CN | 0.12 |
| AD-MCI | 0.16 |
| Graph Metric | HC | AD | MCI | HC-MCI | AD-HC | AD-MCI |
|---|---|---|---|---|---|---|
| Global Efficiency | 0 | 3 | 0 | 24 | 4 | 0 |
| Characteristic Path Length | 53 | 4 | 53 | 0 | 48 | 53 |
| Clustering Coefficient | 5 | 3 | 17 | 5 | 30 | 46 |
| Degree | 8 | 10 | 16 | 10 | 9 | 17 |
| Closeness Centrality | 9 | 26 | 8 | 12 | 3 | 5 |
| Participation Coefficient | 14 | 8 | 14 | 10 | 15 | 17 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




