Submitted:
12 September 2023
Posted:
13 September 2023
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sources of recombinant RBD203-N1 proteins
2.2. Production of the five variant RBD antigens in Pichia pastoris X-33
2.3. Vaccine formulations and preclinical study design
2.4. Serological antibody measurements by ELISA
2.5. Pseudovirus assay for determination of neutralizing antibodies
3. Results
3.1. Expression, purification, and characterization of recombinant proteins of different variants of SARS-CoV-2 RBD203
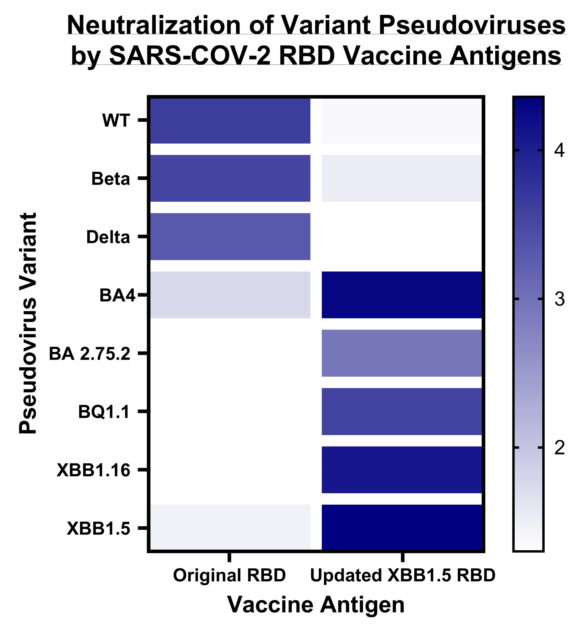
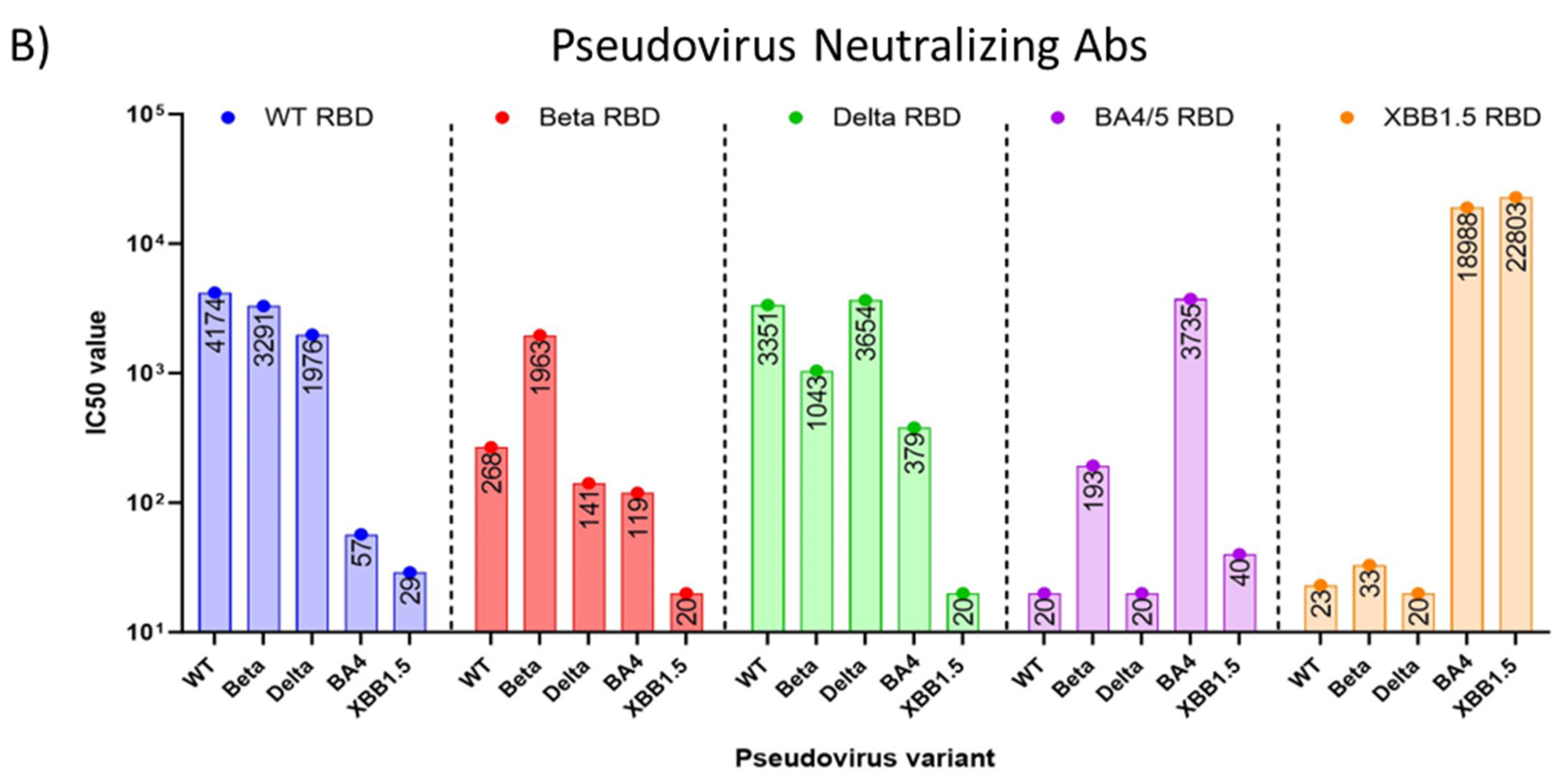
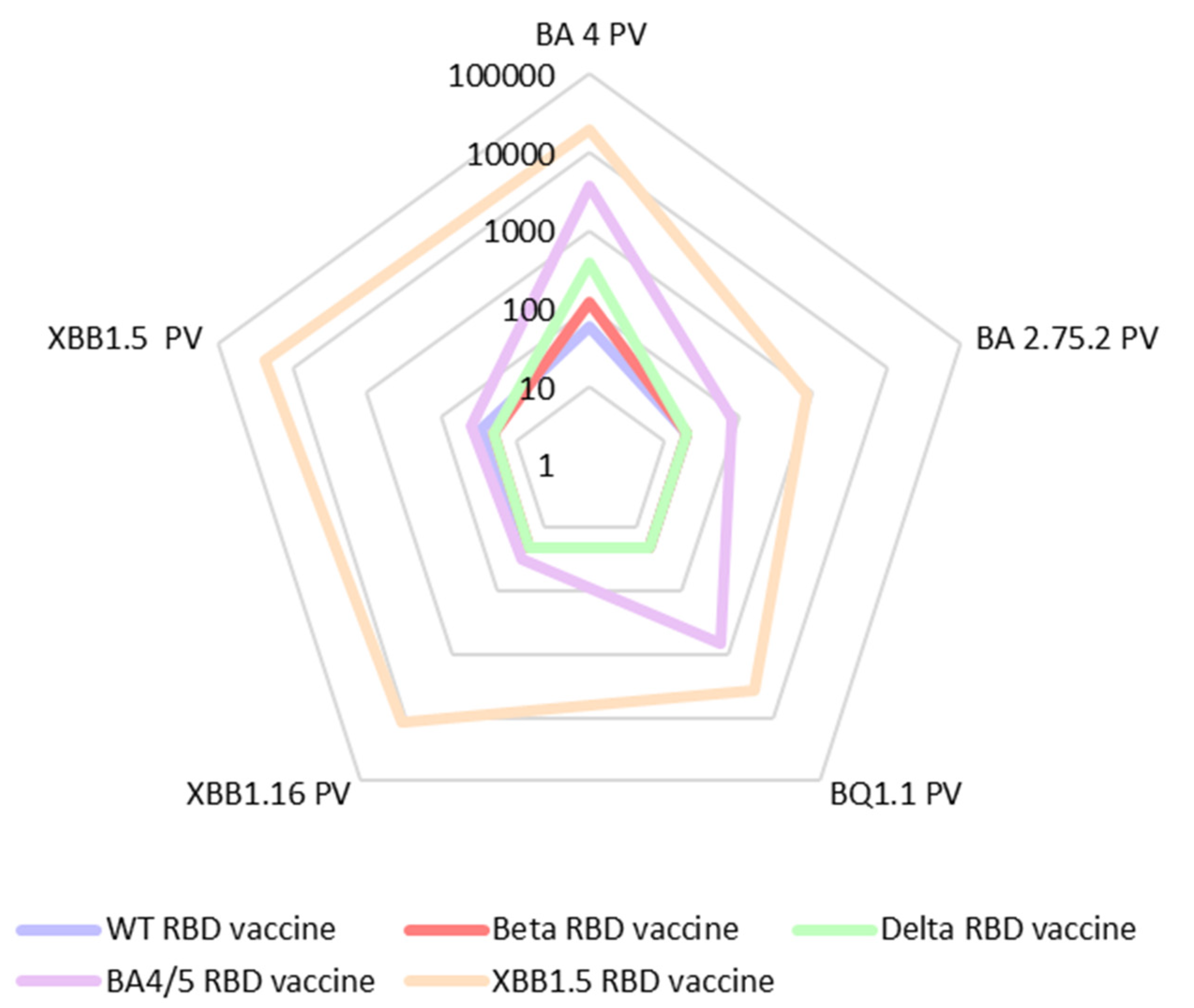
3.2. Immune response induced by adjuvanted RBD vaccines in mice
4. Discussion
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- M. Carabelli et al., “SARS-CoV-2 variant biology: immune escape, transmission and fitness,” Nat Rev Microbiol, vol. 21, no. 3, Art. no. 3, Mar. 2023. [CrossRef]
- N. K. Channabasappa, A. K. N. K. Channabasappa, A. K. Niranjan, and T. B. Emran, “SARS-CoV-2 variant omicron XBB.1.5: challenges and prospects – correspondence,” Int J Surg, vol. 109, no. 4, pp. 1054–1055, Mar. 2023. [Google Scholar] [CrossRef]
- V. Gerardi, M. A. V. Gerardi, M. A. Rohaim, R. F. E. Naggar, M. O. Atasoy, and M. Munir, “Deep Structural Analysis of Myriads of Omicron Sub-Variants Revealed Hotspot for Vaccine Escape Immunity,” Vaccines (Basel), vol. 11, no. 3, p. 668, Mar. 2023. [Google Scholar] [CrossRef]
- P. Qu et al., “Enhanced evasion of neutralizing antibody response by Omicron XBB.1.5, CH.1.1, and CA.3.1 variants,” Cell Rep, vol. 42, no. 5, p. 20 May 1124; 43. [CrossRef]
- et al. , “A recombinant spike-XBB.1.5 protein vaccine induces broad-spectrum immune responses against XBB.1.5-included Omicron variants of SARS-CoV-2,” MedComm, vol. 4, no. 3, p. 2023. [Google Scholar] [CrossRef]
- et al. , “Neutralization of BQ.1, BQ.1.1, and XBB with RBD-Dimer Vaccines,” N Engl J Med, vol. 388, no. 12, pp. 1142–1145, Mar. 2023. [Google Scholar] [CrossRef]
- M. Imai et al., “Efficacy of Antiviral Agents against Omicron Subvariants BQ.1.1 and XBB,” N Engl J Med, vol. 388, no. 1, pp. 89–91, Jan. 2023. [CrossRef]
- J. Pollet et al., “Receptor-binding domain recombinant protein on alum-CpG induces broad protection against SARS-CoV-2 variants of concern,” Vaccine, vol. 40, no. 26, pp. 3655–3663, Jun. 2022. [CrossRef]
- J. Pollet et al., “SARS-CoV-2 RBD219-N1C1: A yeast-expressed SARS-CoV-2 recombinant receptor-binding domain candidate vaccine stimulates virus neutralizing antibodies and T-cell immunity in mice,” Hum Vaccin Immunother, vol. 17, no. 8, pp. 2356–2366, Aug. 2021. [CrossRef]
- M. Pino et al., “A yeast-expressed RBD-based SARS-CoV-2 vaccine formulated with 3M-052-alum adjuvant promotes protective efficacy in non-human primates,” Sci Immunol, vol. 6, no. 61, p. 3634. [CrossRef]
- P. J. Hotez et al., “From concept to delivery: a yeast-expressed recombinant protein-based COVID-19 vaccine technology suitable for global access,” Expert Rev Vaccines, vol. 22, no. 1, pp. 2023. [CrossRef]
- “Statement on the antigen composition of COVID-19 vaccines.” https://www.who.int/news/item/18-05-2023-statement-on-the-antigen-composition-of-covid-19-vaccines (accessed Aug. 18, 2023).
- J. Pollet et al., “SARS-CoV-2 RBD219-N1C1: A yeast-expressed SARS-CoV-2 recombinant receptor-binding domain candidate vaccine stimulates virus neutralizing antibodies and T-cell immunity in mice,” Human Vaccines & Immunotherapeutics, vol. 17, no. 8, pp. 2356–2366, Aug. 2021. [CrossRef]
- J. Lee et al., “Process development and scale-up optimization of the SARS-CoV-2 receptor binding domain–based vaccine candidate, RBD219-N1C1,” Appl Microbiol Biotechnol, vol. 105, no. 10, pp. 4153– 4165, 2021. [CrossRef]
- W.-H. Chen et al., “Yeast-expressed recombinant SARS-CoV-2 receptor binding domain RBD203-N1 as a COVID-19 protein vaccine candidate,” Protein Expr Purif, vol. 190, p. 106003, Feb. 2022. [CrossRef]
- J. Nie et al., “Quantification of SARS-CoV-2 neutralizing antibody by a pseudotyped virus-based assay,” Nat Protoc, vol. 15, no. 11, Art. no. 11, Nov. 2020. [CrossRef]
- et al. , “SARS-CoV-2 Omicron variant: Antibody evasion and cryo-EM structure of spike protein-ACE2 complex,” Science, vol. 375, no. 6582, pp. 760–764, Feb. 2022. [Google Scholar] [CrossRef]
- R. Uraki et al., “Humoral immune evasion of the omicron subvariants BQ.1.1 and XBB,” The Lancet Infectious Diseases, vol. 23, no. 1, pp. 30–32, Jan. 2023. [CrossRef]
- J. Yang et al., “Low levels of neutralizing antibodies against XBB Omicron subvariants after BA.5 infection,” Sig Transduct Target Ther, vol. 8, no. 1, Art. no. 1, Jun. 2023. [CrossRef]
- W. Wang et al., “Antigenic cartography of well-characterized human sera shows SARS-CoV-2 neutralization differences based on infection and vaccination history,” Cell Host Microbe, vol. 30, no. 12, pp. 1745-1758.e7, Dec. 2022. [CrossRef]

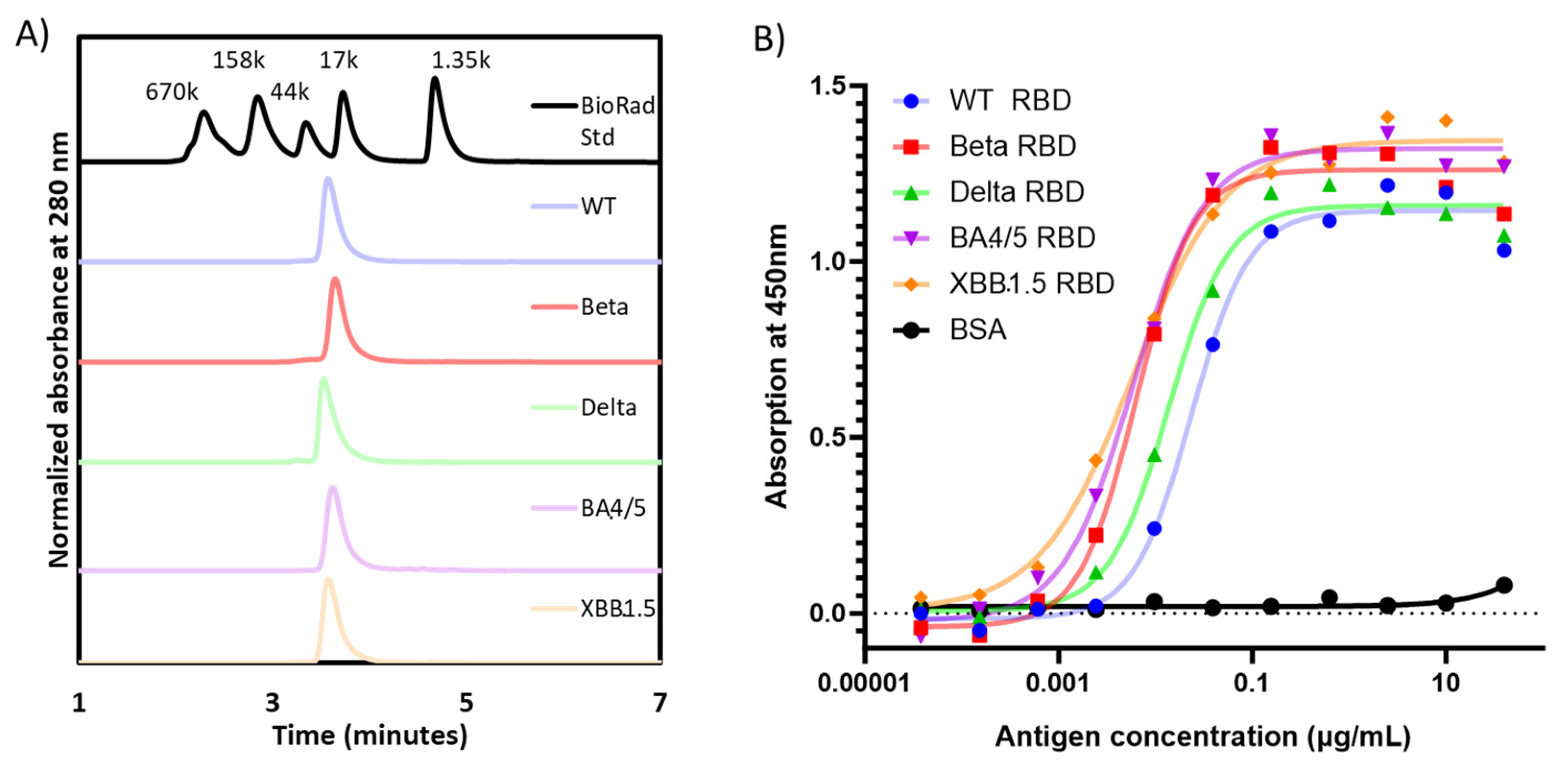
| RBD Variant | Purification Yield [RBD (mg) / fermentation supernatant (L)] |
SDS-PAGE with Coomassie Blue staining (non-reduced) | Homogeneity by SE-HPLC (%) | Radius by dynamic light scattering (nm) |
|
|---|---|---|---|---|---|
| MW (kDa) | Purity (%) | ||||
| WT-RBD | 270.5 | 25.3 | 94.1 | 99.7 | 2.57±0.01 |
| Beta-RBD | 204.3 | 23.8 | 93.2 | 99.7 | 2.63±0.01 |
| Delta-RBD | 294.5 | 25.1 | 93.9 | 98.4 | 2.73±0.02 |
| BA.4/5-RBD | 46.2 | 24.7 | 94.2 | 96.4 | 2.58±0.02 |
| XBB.1.5-RBD | 78.0 | 25.1 | 93.2 | 99.4 | 2.71±0.03 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




