Submitted:
10 August 2023
Posted:
11 August 2023
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Material and Methods
2.1. Oligo Design and Vector Construction
2.2. Cell Culture
2.3. Virus Production
2.4. Determination of Infection Conditions
2.5. Single Cell Clone Selection, Genomic DNA Preparation and Sequencing
2.6. DNA and Protein Sequence Analysis
2.7. Immunoblot Analysis
2.8. Transcriptome Analysis
2.8.1. RNA-seq
2.8.2. DEG (Differentially Expressed Gene) Analysis
2.8.3. Gene Set Enrichment Analyses (GSEA)
2.8.4. Gene Ontology (GO) Enrichment and STRING Analysis
2.8.5. Statistical Analysis
3. Results
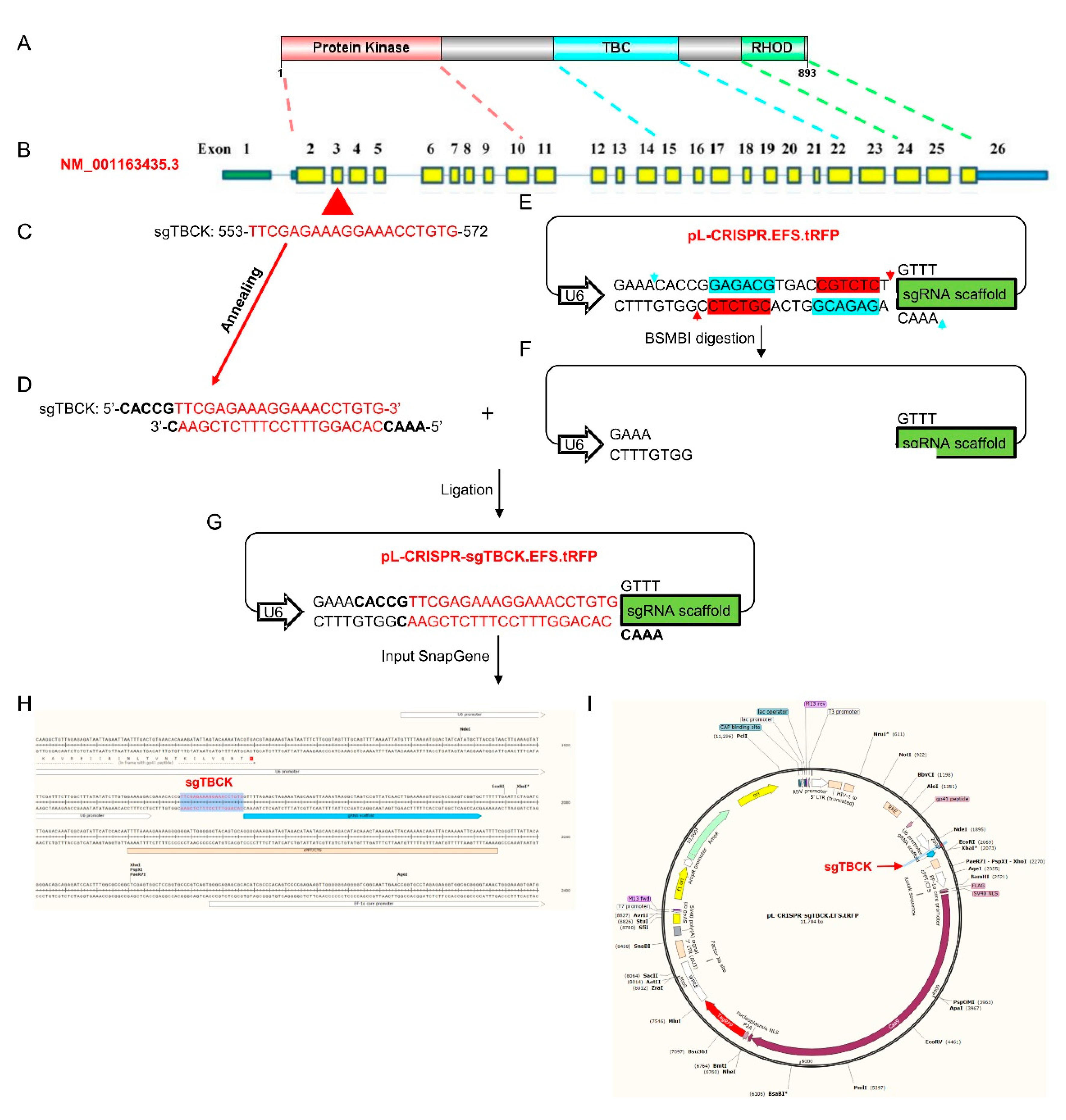
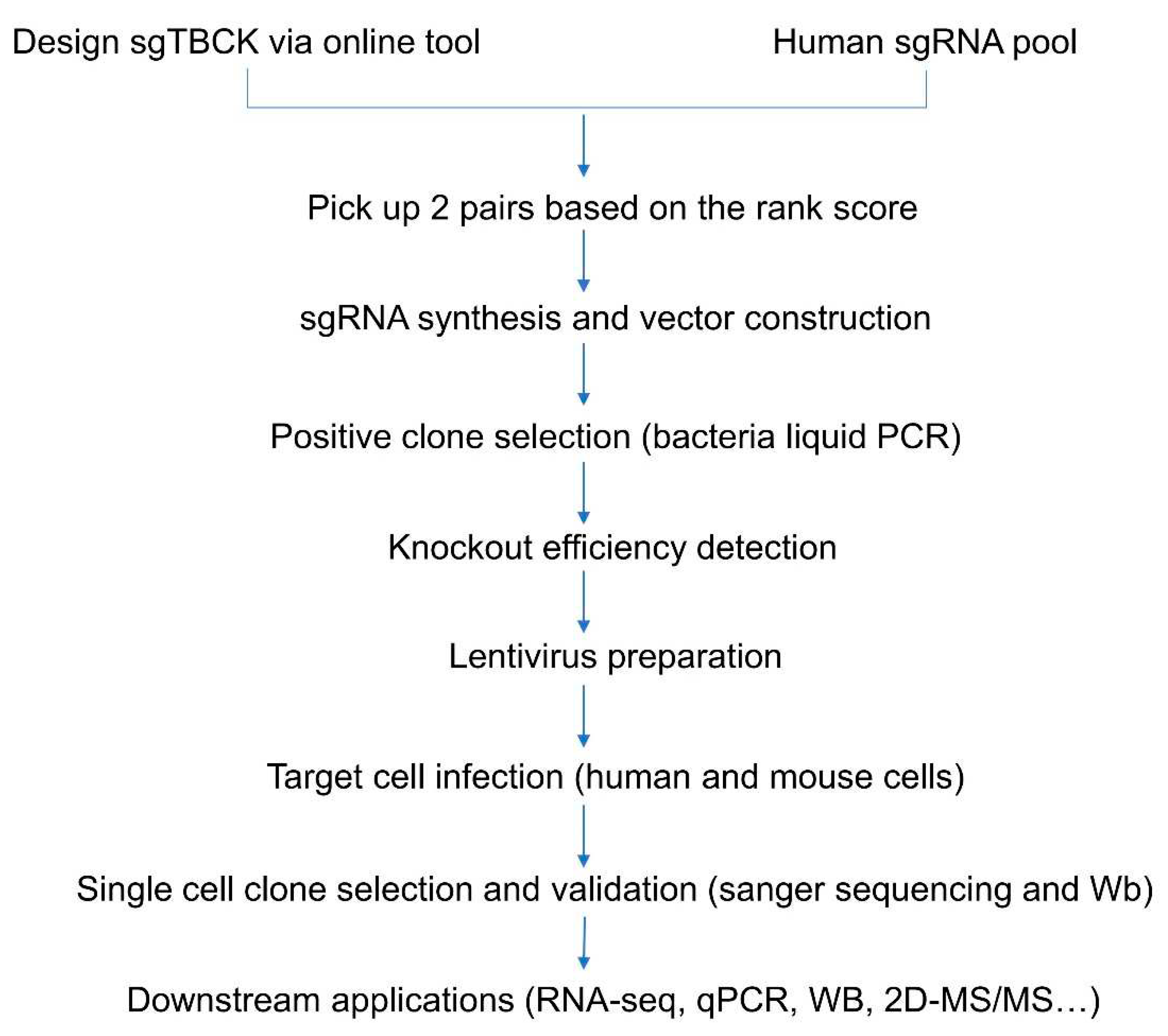
3.1. Design of sgRNAs Targeted to the TBCK Gene
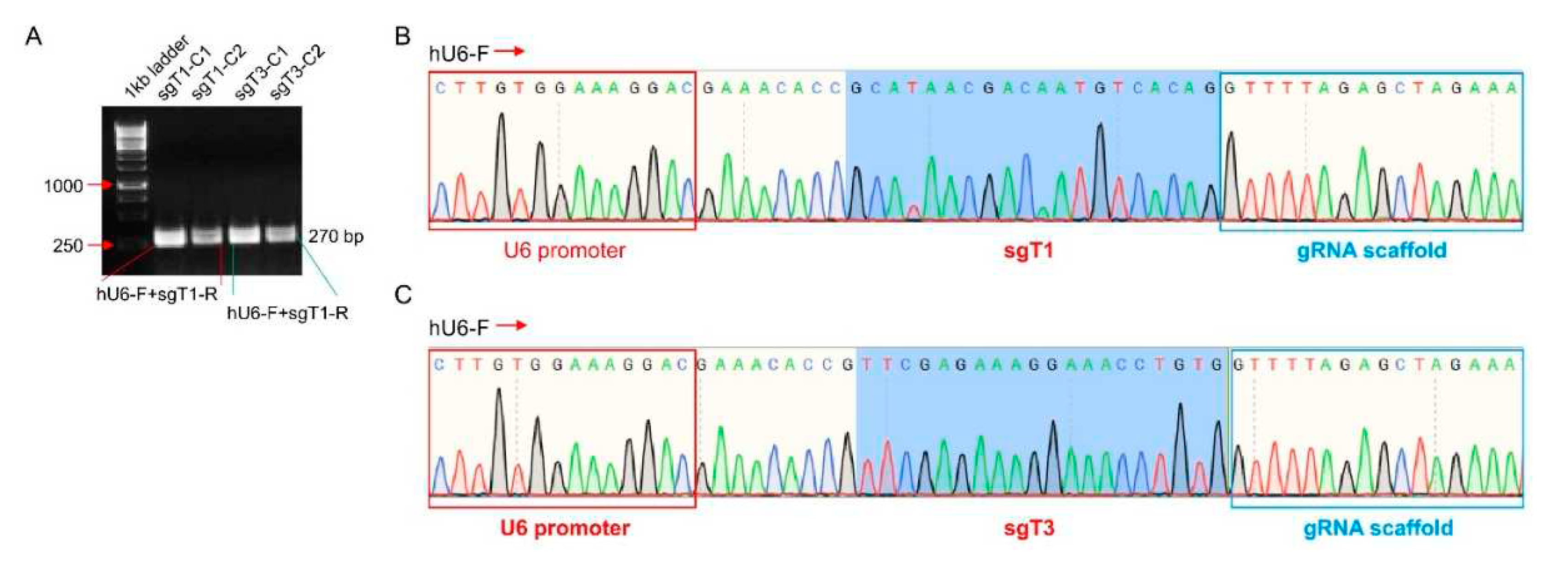
3.2. TBCK Knockout Vector Construction
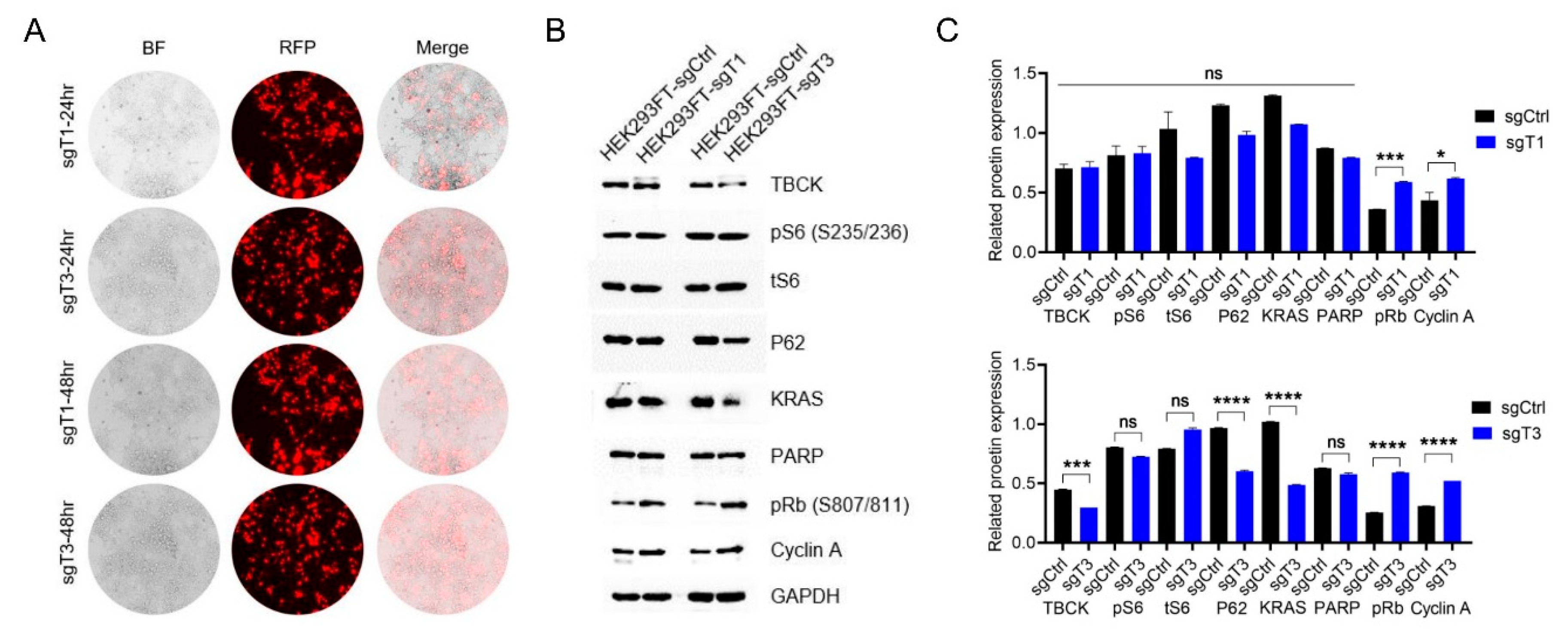
3.3. Knockout Efficiency of TBCK in HEK293FT Cells
3.4. Lentivirus Preparation and Target Cell Infection

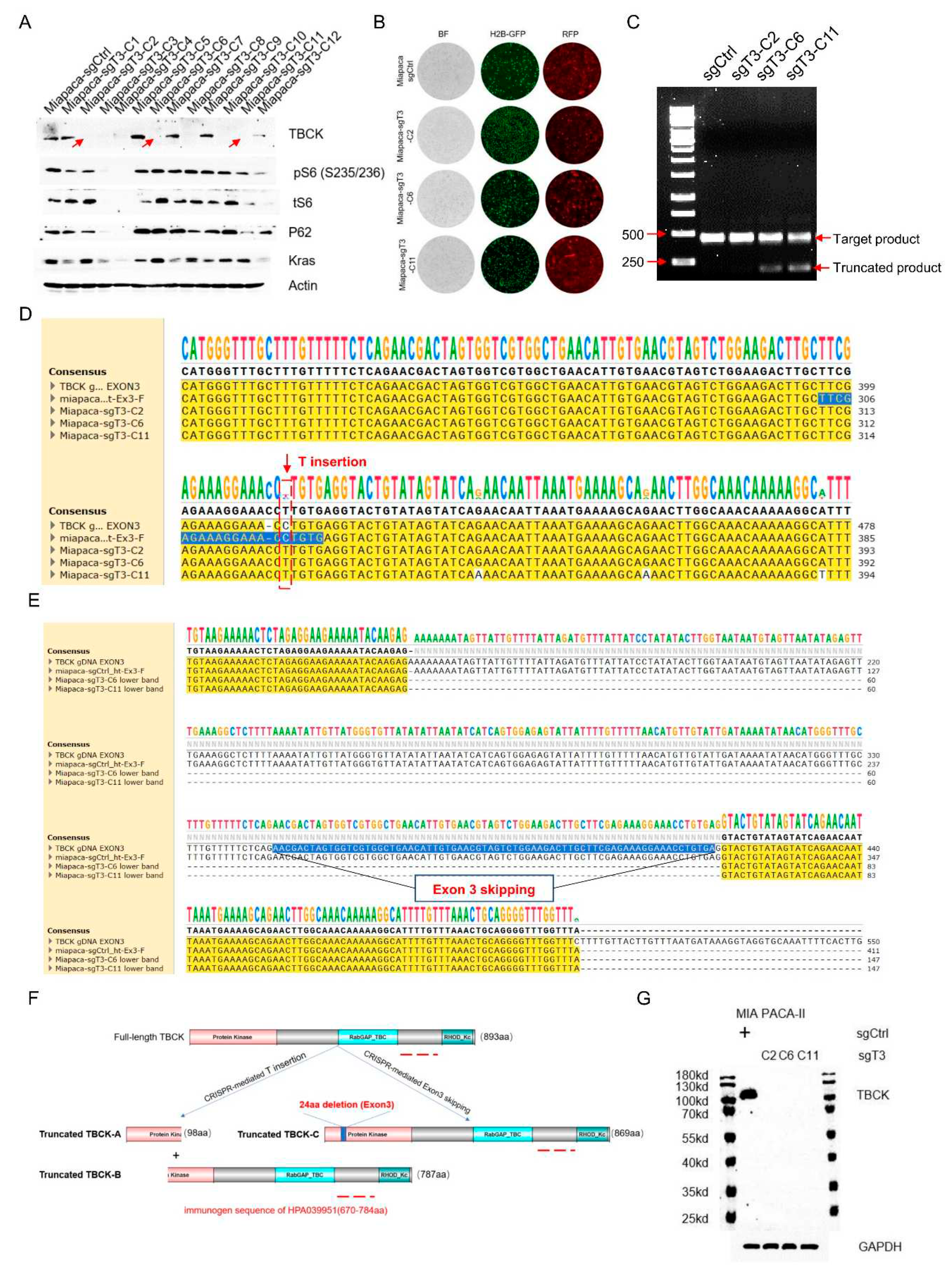
3.5. Single Cell Clone Selection and Validation in MIA PaCa-2 Cell Model
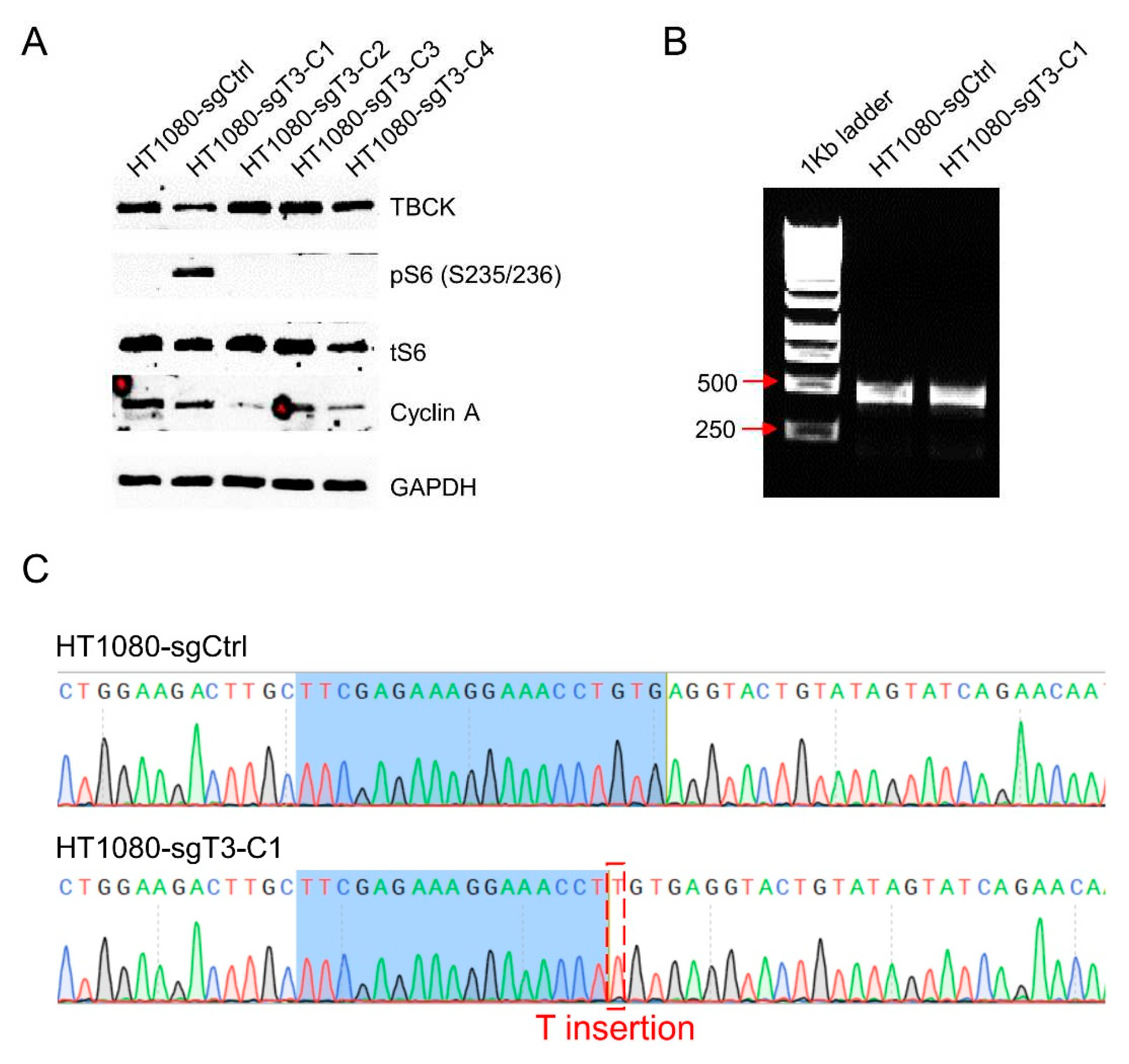
3.6. The Knockout Vector Can Be Applied in Fibrosarcoma HT1080 Cell Model
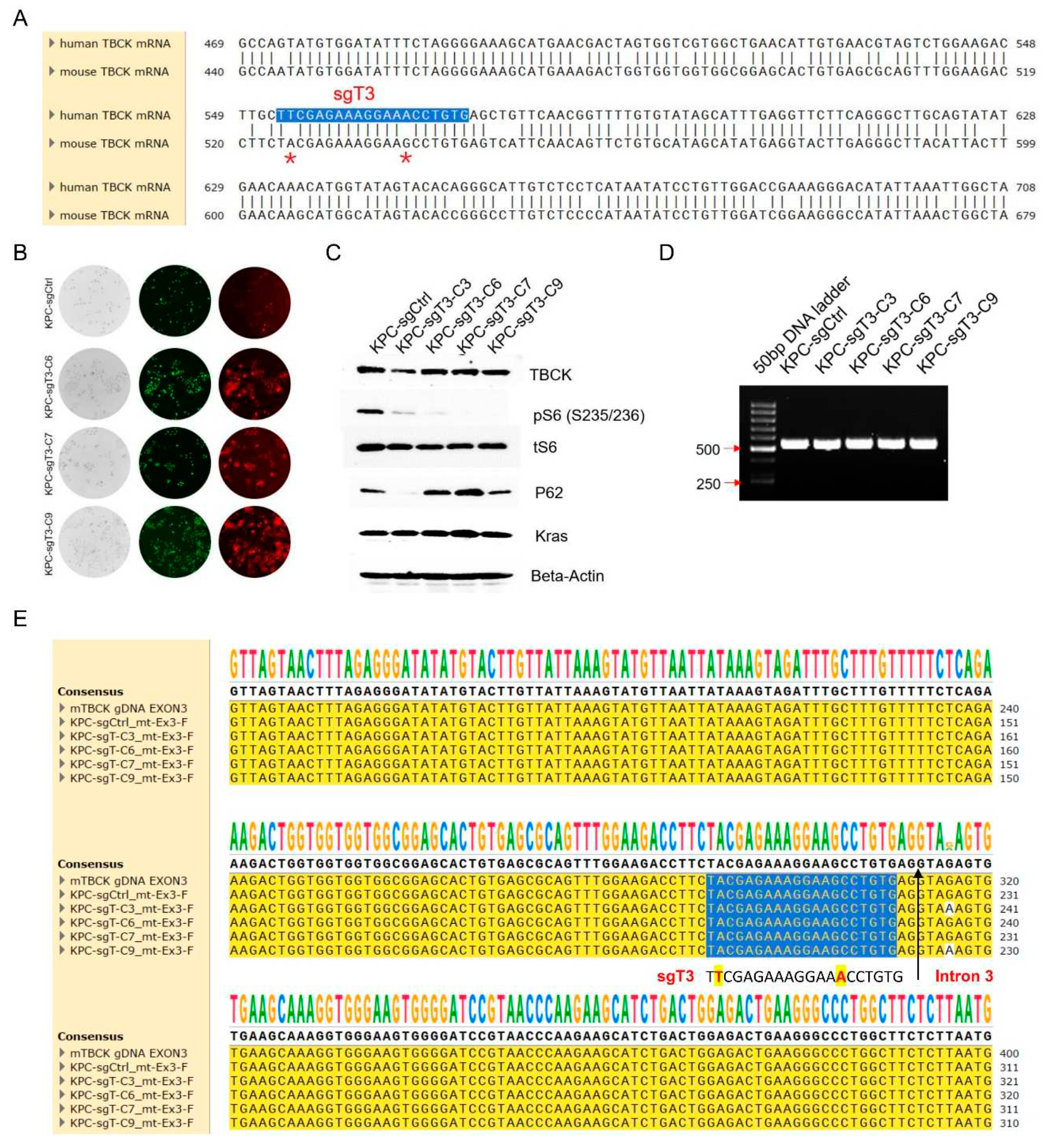
3.7. The Human Specific sgRNA against TBCK Showed Little Effects on Mouse Cells
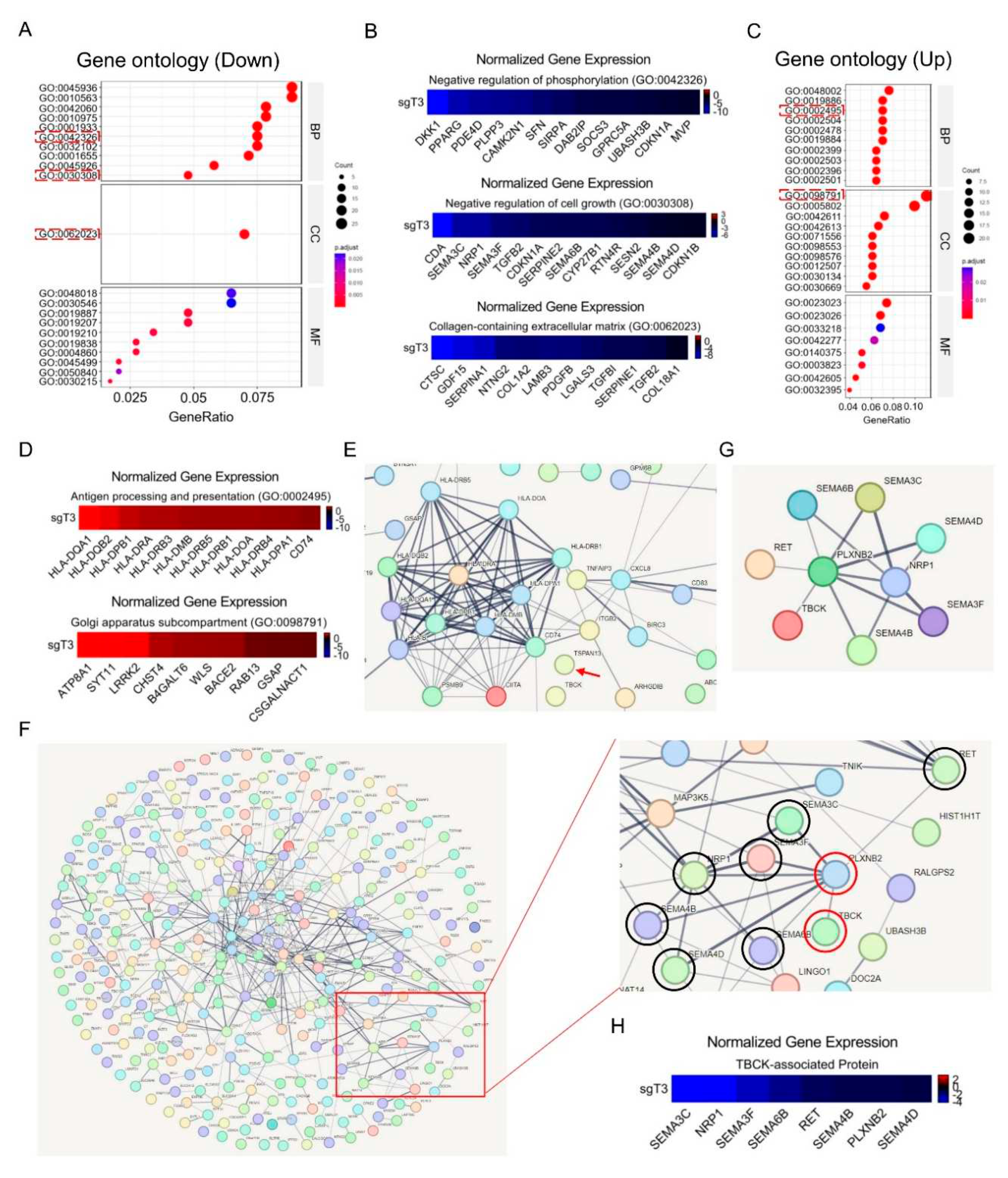
3.8. RNA-seq Application for TBCK Knockout Single Clone in PDAC Model
4. Discussion
5. Conclusions
Abbreviations
- CRISPR Clustered Regularly Interspaced Short Palindromic Repeats
- Cas9 CRISPR-associated 9
- TBCK TBC1 domain containing kinase
- HCC Hepatocellular carcinoma
- sgRNA Single guide RNA
- siRNA Small interfering RNA
- sgCtrl Single guide control RNA
- sgT1 Single guide TBCK RNA (oligo 1)
- sgT3 Single guide TBCK RNA (oligo 3)
- RNA-seq RNA sequencing
- DEG Differentially Expressed Genes
- GSEA Gene set enrichment analyses
- GO Gene Ontology
- PCR Polymerase Chain Reaction
- RFP Red Fluorescent Protein
- GFP Green Fluorescent Protein
- pRB Phosphorylated Rb1
- gDNA Genomic deoxyribonucleic acid
- KPC3 C57/BL6 genetic background mouse cell line with Kras and TP53 mutations
- PDAC Pancreatic ductal adenocarcinoma
- Figure S Supplementary Figure
- Table S Supplementary Table
References
- J. Wu and G. Lu Multiple functions of TBCK protein in neurodevelopment disorders and tumors Oncol Lett 21 (2021) 17. [CrossRef]
- J. X. Chong, V. Caputo, I. G. Phelps, L. Stella, L. Worgan, J. C. Dempsey, A. Nguyen, V. Leuzzi, R. Webster, A. Pizzuti, C. T. Marvin, G. E. Ishak, S. Ardern-Holmes, Z. Richmond, G. University of Washington Center for Mendelian, M. J. Bamshad, X. R. Ortiz-Gonzalez, M. Tartaglia, M. Chopra and D. Doherty Recessive Inactivating Mutations in TBCK, Encoding a Rab GTPase-Activating Protein, Cause Severe Infantile Syndromic Encephalopathy Am J Hum Genet 98 (2016) 772-781. [CrossRef]
- E. J. Bhoj, D. Li, M. Harr, S. Edvardson, O. Elpeleg, E. Chisholm, J. Juusola, G. Douglas, M. J. Guillen Sacoto, K. Siquier-Pernet, A. Saadi, C. Bole-Feysot, P. Nitschke, A. Narravula, M. Walke, M. B. Horner, D. L. Day-Salvatore, P. Jayakar, S. A. Vergano, M. A. Tarnopolsky, M. Hegde, L. Colleaux, P. Crino and H. Hakonarson Mutations in TBCK, Encoding TBC1-Domain-Containing Kinase, Lead to a Recognizable Syndrome of Intellectual Disability and Hypotonia Am J Hum Genet 98 (2016) 782-788. [CrossRef]
- X. R. Ortiz-Gonzalez, J. A. Tintos-Hernandez, K. Keller, X. Li, A. R. Foley, D. X. Bharucha-Goebel, S. K. Kessler, S. W. Yum, P. B. Crino, M. He, D. C. Wallace and C. G. Bonnemann Homozygous boricua TBCK mutation causes neurodegeneration and aberrant autophagy Ann Neurol 83 (2018) 153-165. [CrossRef]
- J. A. Tintos-Hernandez, A. Santana, K. N. Keller and X. R. Ortiz-Gonzalez Lysosomal dysfunction impairs mitochondrial quality control and is associated with neurodegeneration in TBCK encephaloneuronopathy Brain Commun 3 (2021) fcab215. [CrossRef]
- D. Nair, A. Diaz-Rosado, E. Varella-Branco, I. Ramos, A. Black, R. Angireddy, J. Park, S. Murali, A. Yoon, B. Ciesielski, W. T. O'Brien, M. R. Passos-Bueno and E. Bhoj Heterozygous variants in TBCK cause a mild neurologic syndrome in humans and mice Am J Med Genet A (2023). [CrossRef]
- K. Komurov, D. Padron, T. Cheng, M. Roth, K. P. Rosenblatt and M. A. White Comprehensive mapping of the human kinome to epidermal growth factor receptor signaling J Biol Chem 285 (2010) 21134-21142. [CrossRef]
- Y. Liu, X. Yan and T. Zhou TBCK influences cell proliferation, cell size and mTOR signaling pathway PLoS One 8 (2013) e71349. [CrossRef]
- J. Wu, Q. Li, Y. Li, J. Lin, D. Yang, G. Zhu, L. Wang, D. He, G. Lu and C. Zeng A long type of TBCK is a novel cytoplasmic and mitotic apparatus-associated protein likely suppressing cell proliferation J Genet Genomics 41 (2014) 69-72. [CrossRef]
- I. Panagopoulos, L. Gorunova, T. Viset and S. Heim Gene fusions AHRR-NCOA2, NCOA2-ETV4, ETV4-AHRR, P4HA2-TBCK, and TBCK-P4HA2 resulting from the translocations t(5;8;17)(p15;q13;q21) and t(4;5)(q24;q31) in a soft tissue angiofibroma Oncol Rep 36 (2016) 2455-2462. [CrossRef]
- E. A. Kim, J. H. Jang, E. G. Sung, I. H. Song, J. Y. Kim and T. J. Lee MiR-1208 Increases the Sensitivity to Cisplatin by Targeting TBCK in Renal Cancer Cells Int J Mol Sci 20 (2019). [CrossRef]
- J. Gao, L. Xi, R. Yu, H. Xu, M. Wu and H. Huang Differential mutation detection capability through capture-based targeted sequencing in plasma samples in hepatocellular carcinoma Frontiers in Oncology 11 (2021) 596789.
- J. Wu, G. Lu, J. Wu, H. Yang, Z. Yu, S. Mu and H. Zhang Application of fusion PCR to the amplification of full-length ORF sequences of different splicing variants of NuMA1 from HeLa cells Acta Biochim Biophys Sin (Shanghai) 49 (2017) 962-965. [CrossRef]
- M. W. Davis and E. M. Jorgensen ApE, A Plasmid Editor: A Freely Available DNA Manipulation and Visualization Program Front Bioinform 2 (2022) 818619. [CrossRef]
- J. Ren, L. Wen, X. Gao, C. Jin, Y. Xue and X. Yao DOG 1.0: illustrator of protein domain structures Cell Res 19 (2009) 271-273. [CrossRef]
- J. Chang, H. J. Chang, H. Wu, J. Wu, M. Liu, W. Zhang, Y. Hu, X. Zhang, J. Xu, L. Li, P. Yu and J. Zhu Constructing a novel mitochondrial-related gene signature for evaluating the tumor immune microenvironment and predicting survival in stomach adenocarcinoma J Transl Med 21 (2023) 191. [CrossRef]
- J. Zhu, W. Zhang, J. Chang, J. Wu, H. Wu, X. Zhang, Z. Ou, T. Tang, L. Li, M. Liu and Y. Xin Identification and Validation of a Mitochondria Calcium Uptake-Related Gene Signature for Predicting Prognosis in COAD J Cancer 14 (2023) 741-758. [CrossRef]
- E. Y. Chen, C. M. Tan, Y. Kou, Q. Duan, Z. Wang, G. V. Meirelles, N. R. Clark and A. Ma'ayan Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool BMC Bioinformatics 14 (2013) 128. [CrossRef]
- M. V. Kuleshov, M. R. Jones, A. D. Rouillard, N. F. Fernandez, Q. Duan, Z. Wang, S. Koplev, S. L. Jenkins, K. M. Jagodnik, A. Lachmann, M. G. McDermott, C. D. Monteiro, G. W. Gundersen and A. Ma'ayan Enrichr: a comprehensive gene set enrichment analysis web server 2016 update Nucleic Acids Res 44 (2016) W90-97. [CrossRef]
- Z. Xie, A. Bailey, M. V. Kuleshov, D. J. B. Clarke, J. E. Evangelista, S. L. Jenkins, A. Lachmann, M. L. Wojciechowicz, E. Kropiwnicki, K. M. Jagodnik, M. Jeon and A. Ma'ayan Gene Set Knowledge Discovery with Enrichr Curr Protoc 1 (2021) e90. [CrossRef]
- A. Liberzon, C. Birger, H. Thorvaldsdottir, M. Ghandi, J. P. Mesirov and P. Tamayo The Molecular Signatures Database (MSigDB) hallmark gene set collection Cell Syst 1 (2015) 417-425. [CrossRef]
- A. Subramanian, P. Tamayo, V. K. Mootha, S. Mukherjee, B. L. Ebert, M. A. Gillette, A. Paulovich, S. L. Pomeroy, T. R. Golub, E. S. Lander and J. P. Mesirov Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles Proc Natl Acad Sci U S A 102 (2005) 15545-15550. [CrossRef]
- V. K. Mootha, C. M. Lindgren, K. F. Eriksson, A. Subramanian, S. Sihag, J. Lehar, P. Puigserver, E. Carlsson, M. Ridderstrale, E. Laurila, N. Houstis, M. J. Daly, N. Patterson, J. P. Mesirov, T. R. Golub, P. Tamayo, B. Spiegelman, E. S. Lander, J. N. Hirschhorn, D. Altshuler and L. C. Groop PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes Nat Genet 34 (2003) 267-273. [CrossRef]
- D. Szklarczyk, A. Franceschini, S. Wyder, K. Forslund, D. Heller, J. Huerta-Cepas, M. Simonovic, A. Roth, A. Santos, K. P. Tsafou, M. Kuhn, P. Bork, L. J. Jensen and C. von Mering STRING v10: protein-protein interaction networks, integrated over the tree of life Nucleic Acids Res 43 (2015) D447-452. [CrossRef]
- Y. Cui, J. Xu, M. Cheng, X. Liao and S. Peng Review of CRISPR/Cas9 sgRNA Design Tools Interdiscip Sci 10 (2018) 455-465. [CrossRef]
- V. Pattanayak, S. Lin, J. P. Guilinger, E. Ma, J. A. Doudna and D. R. Liu High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity Nat Biotechnol 31 (2013) 839-843. [CrossRef]
- J. G. Doench, N. Fusi, M. Sullender, M. Hegde, E. W. Vaimberg, K. F. Donovan, I. Smith, Z. Tothova, C. Wilen, R. Orchard, H. W. Virgin, J. Listgarten and D. E. Root Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9 Nat Biotechnol 34 (2016) 184-191. [CrossRef]
- J. G. Doench, E. Hartenian, D. B. Graham, Z. Tothova, M. Hegde, I. Smith, M. Sullender, B. L. Ebert, R. J. Xavier and D. E. Root Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation Nat Biotechnol 32 (2014) 1262-1267. [CrossRef]
- N. E. Sanjana, O. Shalem and F. Zhang Improved vectors and genome-wide libraries for CRISPR screening Nat Methods 11 (2014) 783-784. [CrossRef]
- A. Holmgaard, A. L. Askou, J. N. E. Benckendorff, E. A. Thomsen, Y. Cai, T. Bek, J. G. Mikkelsen and T. J. Corydon In Vivo Knockout of the Vegfa Gene by Lentiviral Delivery of CRISPR/Cas9 in Mouse Retinal Pigment Epithelium Cells Mol Ther Nucleic Acids 9 (2017) 89-99. [CrossRef]
- B. O'Boyle, S. Shrestha, K. Kochut, P. A. Eyers and N. Kannan Computational tools and resources for pseudokinase research Methods Enzymol 667 (2022) 403-426. [CrossRef]
- K. S. Metz, E. M. Deoudes, M. E. Berginski, I. Jimenez-Ruiz, B. A. Aksoy, J. Hammerbacher, S. M. Gomez and D. H. Phanstiel Coral: Clear and Customizable Visualization of Human Kinome Data Cell Syst 7 (2018) 347-350 e341. [CrossRef]
- V. Perrot, J. Vazquez-Prado and J. S. Gutkind Plexin B regulates Rho through the guanine nucleotide exchange factors leukemia-associated Rho GEF (LARG) and PDZ-RhoGEF Journal of Biological Chemistry 277 (2002) 43115-43120.
- P. Conrotto, S. Corso, S. Gamberini, P. M. Comoglio and S. Giordano Interplay between scatter factor receptors and B plexins controls invasive growth Oncogene 23 (2004) 5131-5137.
- D. Sumathipala, P. Stromme, C. Gilissen, J. Corominas, E. Frengen and D. Misceo TBCK Encephaloneuropathy With Abnormal Lysosomal Storage: Use of a Structural Variant Bioinformatics Pipeline on Whole-Genome Sequencing Data Unravels a 20-Year-Old Clinical Mystery Pediatr Neurol 96 (2019) 74-75. [CrossRef]
- M. Jinek, K. Chylinski, I. Fonfara, M. Hauer, J. A. Doudna and E. Charpentier A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity Science 337 (2012) 816-821. [CrossRef]
- P. Mali, L. Yang, K. M. Esvelt, J. Aach, M. Guell, J. E. DiCarlo, J. E. Norville and G. M. Church RNA-guided human genome engineering via Cas9 Science 339 (2013) 823-826. [CrossRef]
- L. Cong, F. A. Ran, D. Cox, S. Lin, R. Barretto, N. Habib, P. D. Hsu, X. Wu, W. Jiang, L. A. Marraffini and F. Zhang Multiplex genome engineering using CRISPR/Cas systems Science 339 (2013) 819-823. [CrossRef]
- M. Jinek, A. East, A. Cheng, S. Lin, E. Ma and J. Doudna RNA-programmed genome editing in human cells Elife 2 (2013) e00471. [CrossRef]
- Y. Fu, J. A. Foden, C. Khayter, M. L. Maeder, D. Reyon, J. K. Joung and J. D. Sander High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells Nat Biotechnol 31 (2013) 822-826. [CrossRef]
- A. Veres, B. S. Gosis, Q. Ding, R. Collins, A. Ragavendran, H. Brand, S. Erdin, C. A. Cowan, M. E. Talkowski and K. Musunuru Low incidence of off-target mutations in individual CRISPR-Cas9 and TALEN targeted human stem cell clones detected by whole-genome sequencing Cell Stem Cell 15 (2014) 27-30. [CrossRef]
- F. A. Ran, P. D. Hsu, C. Y. Lin, J. S. Gootenberg, S. Konermann, A. E. Trevino, D. A. Scott, A. Inoue, S. Matoba, Y. Zhang and F. Zhang Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity Cell 154 (2013) 1380-1389. [CrossRef]
- J. P. Guilinger, D. B. Thompson and D. R. Liu Fusion of catalytically inactive Cas9 to FokI nuclease improves the specificity of genome modification Nat Biotechnol 32 (2014) 577-582. [CrossRef]
- P. D. Hsu, D. A. Scott, J. A. Weinstein, F. A. Ran, S. Konermann, V. Agarwala, Y. Li, E. J. Fine, X. Wu, O. Shalem, T. J. Cradick, L. A. Marraffini, G. Bao and F. Zhang DNA targeting specificity of RNA-guided Cas9 nucleases Nat Biotechnol 31 (2013) 827-832. [CrossRef]
- A. L. Kolodkin, D. J. Matthes and C. S. Goodman The semaphorin genes encode a family of transmembrane and secreted growth cone guidance molecules Cell 75 (1993) 1389-1399. [CrossRef]
- S. Hu and L. Zhu Semaphorins and their receptors: from axonal guidance to atherosclerosis Frontiers in Physiology 9 (2018) 1236.
- G. M. Stella, A. Corino, G. Berzero, S. Kolling, A. R. Filippi and S. Benvenuti Brain metastases from lung cancer: Is MET an actionable target? Cancers 11 (2019) 271.
- W. Mu, Z. Wang and M. Zoller Ping-Pong-Tumor and Host in Pancreatic Cancer Progression Front Oncol 9 (2019) 1359. [CrossRef]
- Z. He and M. Tessier-Lavigne Neuropilin is a receptor for the axonal chemorepellent Semaphorin III Cell 90 (1997) 739-751.
- S. Soker, S. Takashima, H. Q. Miao, G. Neufeld and M. Klagsbrun Neuropilin-1 is expressed by endothelial and tumor cells as an isoform-specific receptor for vascular endothelial growth factor Cell 92 (1998) 735-745.
- M. L. Gagnon, D. R. Bielenberg, Z. e. Gechtman, H.-Q. Miao, S. Takashima, S. Soker and M. Klagsbrun Identification of a natural soluble neuropilin-1 that binds vascular endothelial growth factor: In vivo expression and antitumor activity Proceedings of the National Academy of Sciences 97 (2000) 2573-2578.

| Experiment | Gene | Forward Primer, 5' → 3′ | Reverse Primer, 5' → 3′ | Accession Number |
|---|---|---|---|---|
| sgCtrl | Control | CACCGACGGAGGCTAAGCGTCGCAA | AAACTTGCGACGCTTAGCCTCCGTC | NA |
| sgT1 | hTBCK | CACCGCATAACGACAATGTCACAG | AAACCTGTGACATTGTCGTTATGC | NM_001163435.3 |
| sgT3 | hTBCK | CACCGTTCGAGAAAGGAAACCTGTG | AAACCACAGGTTTCCTTTCTCGAAC | NM_001163435.3 |
| RT-PCR | U6 | GAGGGCCTATTTCCCATGATT | NA | |
| hTBCK | GTGTGTCAGAAGAAGGGTGAGT | AAACCAAACCCCTGCAGTTTA | NG_034057.3 | |
| mTBCK | GGTGGATGGGGTGCTTACAT | CTCCGGGCTAGGGGAATAAG | NC_000069.7 |
| Source | Gene_id | UID | seq | PAM | Exon Number | On-Target Efficacy Score | Rank for sgTBCK via Online Tool |
|---|---|---|---|---|---|---|---|
| human_geckov2_library_a | TBCK | HGLibA_48598 | TGAACATTGTGAACGTAGTC | TGG | 3 | 0.4328 | 182 |
| TBCK | HGLibA_48599 | CTCCCATTTCAGCGTCCTTC | GGG | 2 | 0.2272 | 258 | |
| TBCK | HGLibA_48600 | AGCCGAGGCAAAGAAGGTAA | AGG | 2 | 0.5571 | 86 | |
| human_geckov2_library_b | TBCK | HGLibB_48539 | TTCGAGAAAGGAAACCTGTG | AGG | 3 | 0.7099 | 8 |
| TBCK | HGLibB_48540 | AAGAAAATTATTTCAGAGCT | TGG | 7 | 0.3958 | 201 | |
| TBCK | HGLibB_48541 | TTGCTTCCACAAACATCATG | TGG | 2 | 0.6266 | 36 | |
| Online tool sgRNA designer | TBCK | NA | GCATAACGACAATGTCACAG | TGG | 12 | 0.7724 | 3 |
| Cell Line Name | Species | Tumor Type | Genetic Information | TBCK Expression |
|---|---|---|---|---|
| MiaPaca-2 | Homo Sapiens | PDAC | Kras (G12C); c-Myc (WT); TP53(R248W); RB1(WT) | High |
| HT1080 | Homo Sapiens | Fibrosarcoma | Kras (WT); c-Myc (WT); TP53(WT); RB1(WT) | High |
| KPC3 mouse cell line | Mus musculus | Mouse pancreatic neoplasm | Kras (G12D); c-Myc (WT); TP53(R270H); RB1(WT) | High |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




