Submitted:
28 July 2023
Posted:
31 July 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Computer-Aided Drug Design
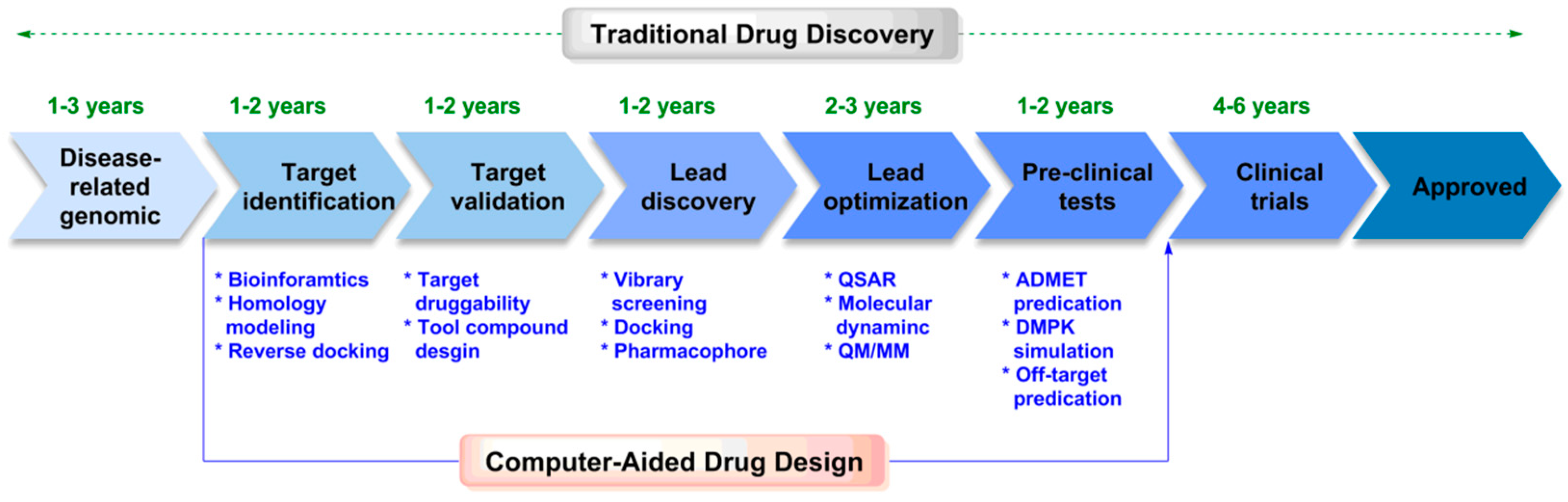
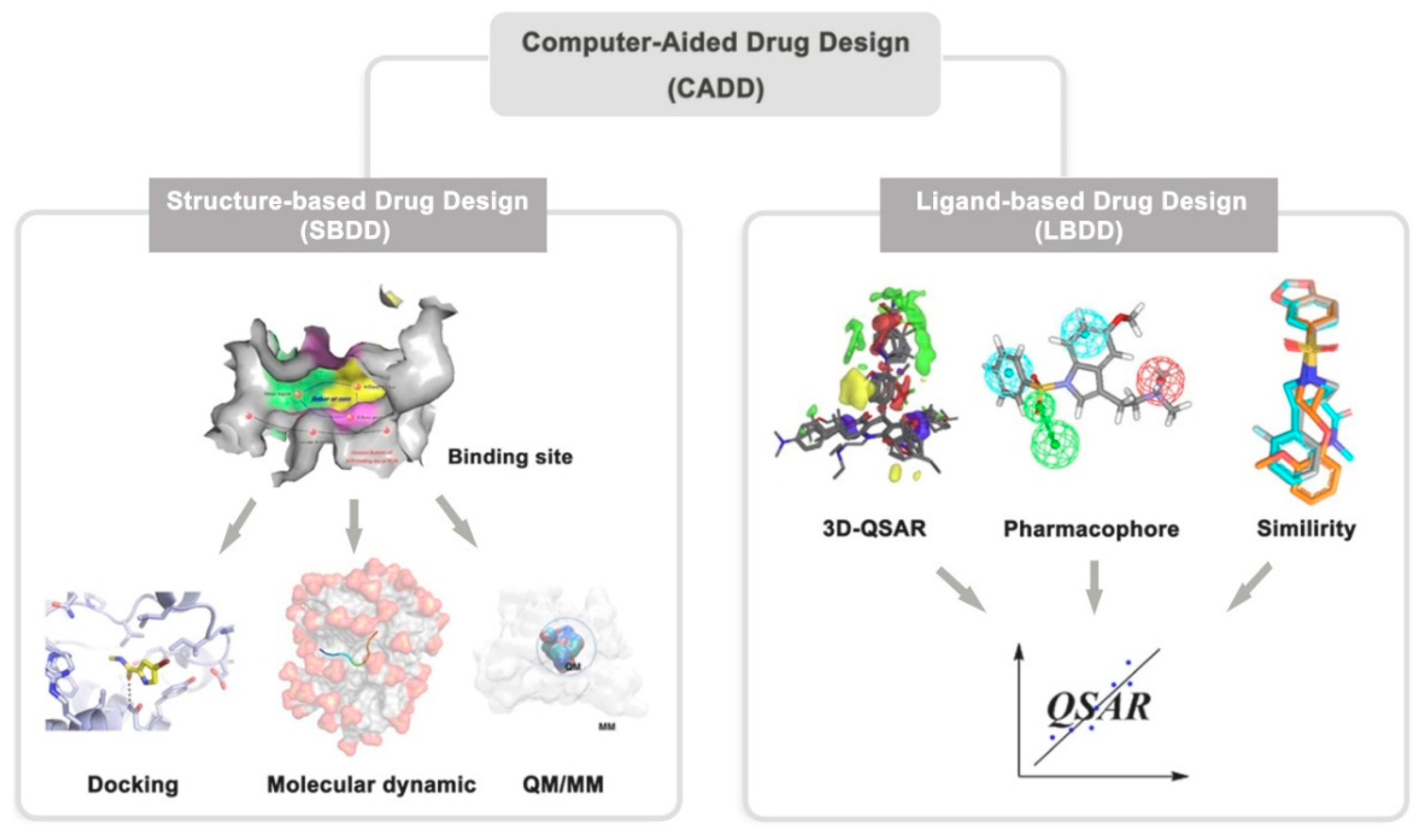
2.1. Structure-Based Drug Design
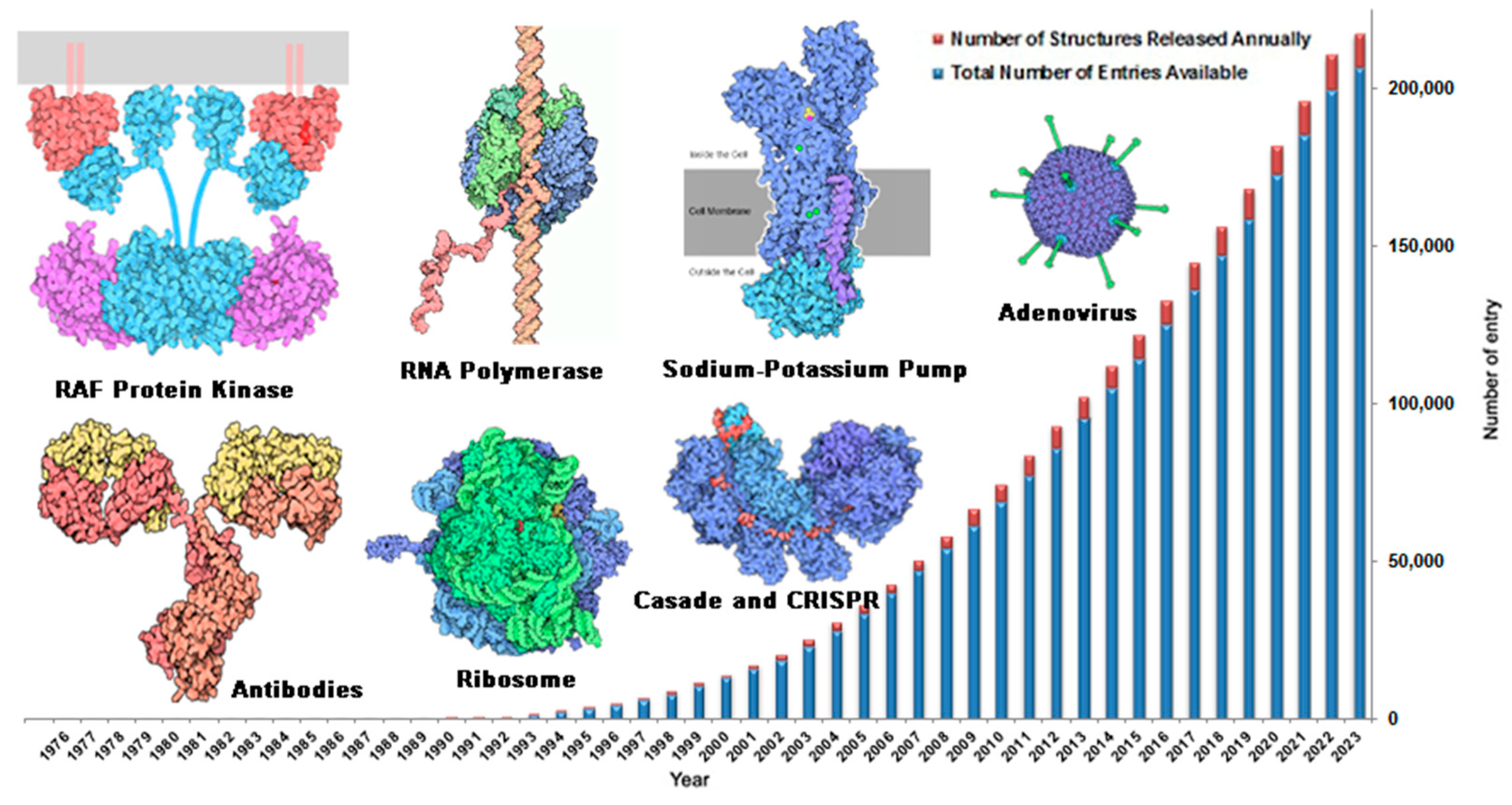
2.1.1. Preparation of Target Structures
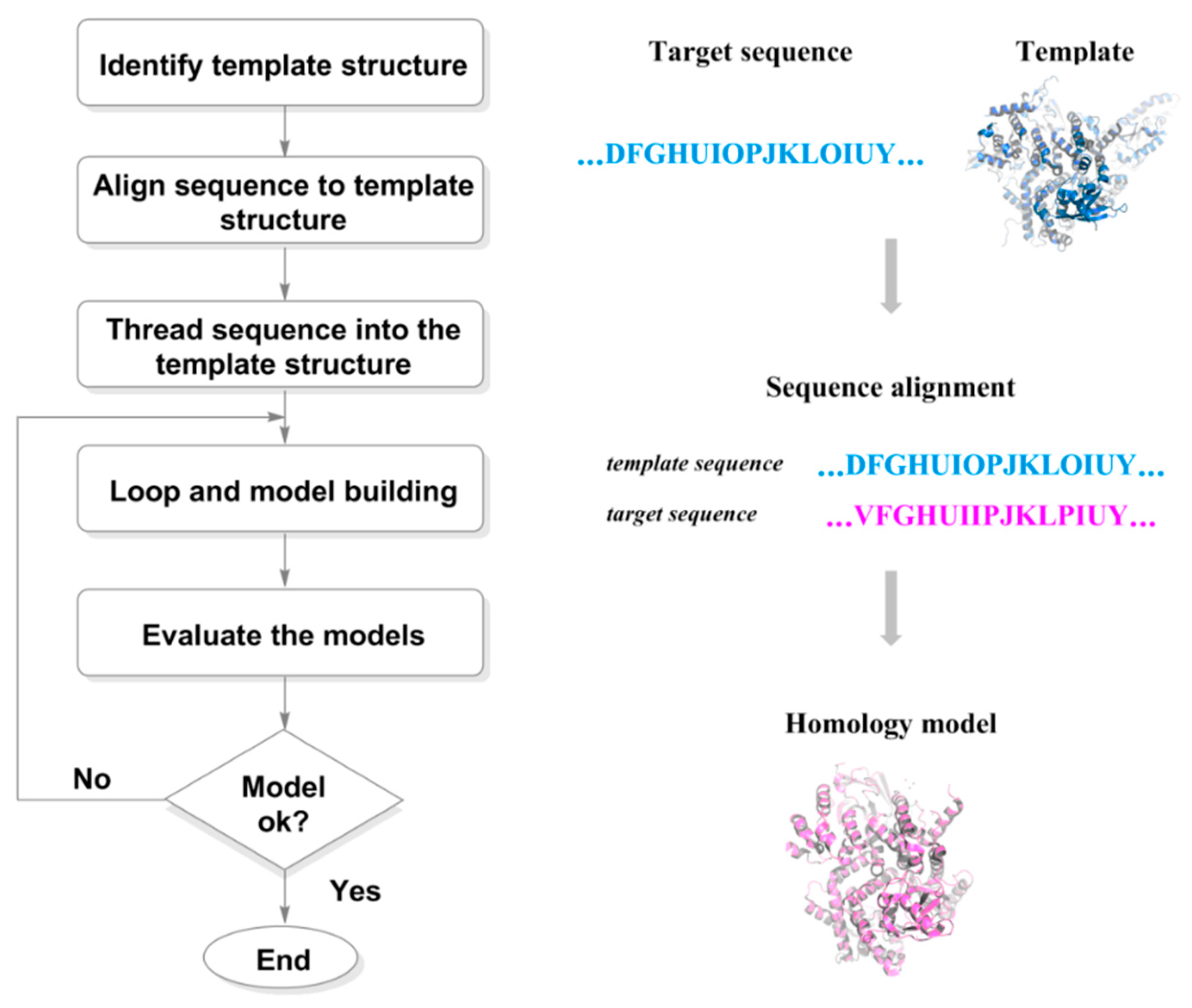
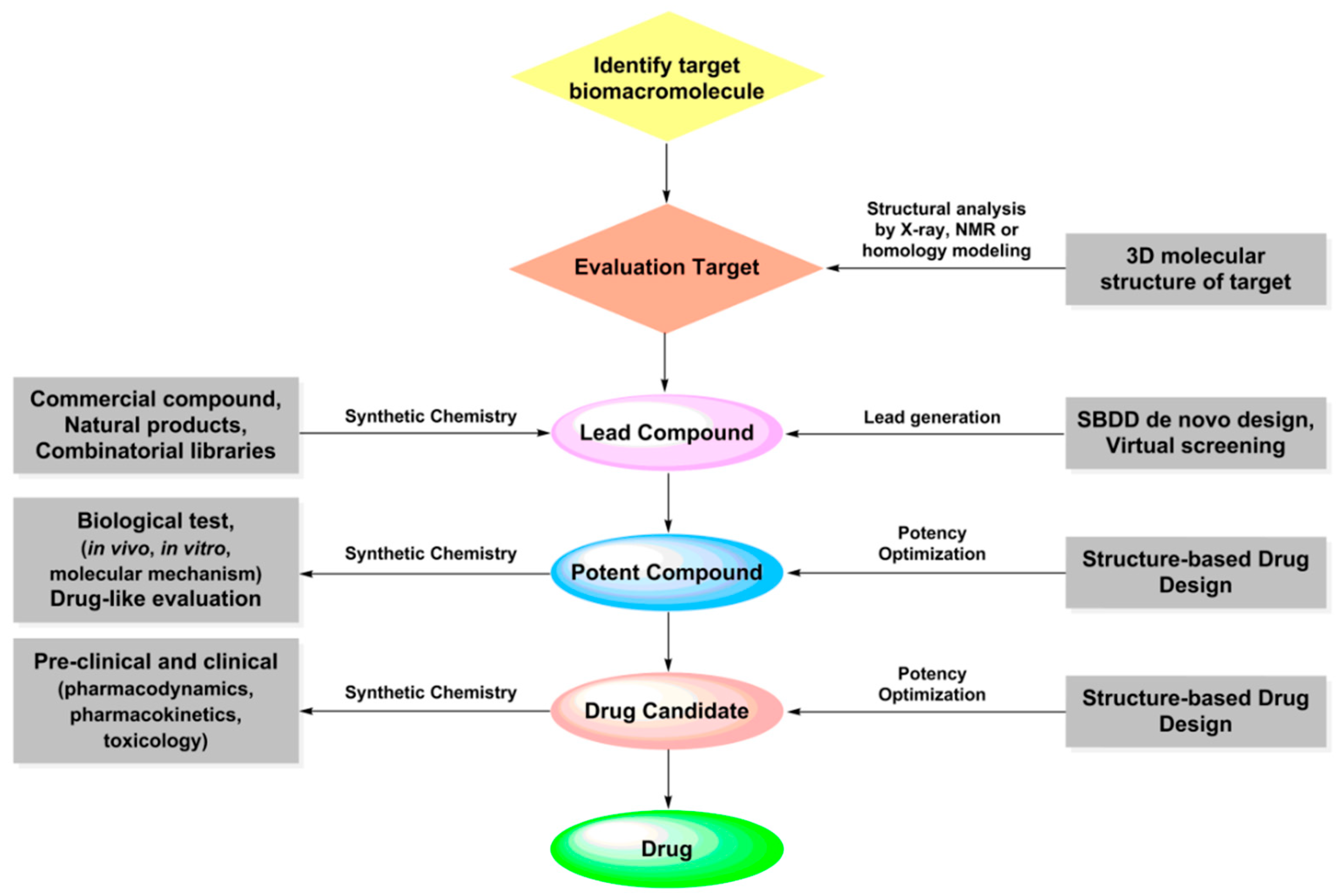
2.1.2. Molecular Docking
| Software | Algorithm | Scoring function | Website |
|---|---|---|---|
| Dock | Fragment growth | Force field, Surface matching score, Environment matching score | http://dock.compbio.ucsf.edu/DOCK_6/ |
| AutoDock | Genetic algorithm | Environment matching score | http://autodock.scripps.edu/ |
| GOLD | Genetic algorithm | Empirical | http://www.biosolveit.de/FlexX/ |
| FlexX | Fragment growth | Empirical | https://github.com/flexxui/flexx |
| Z-Dock | Geometric matching/Molecular dynamics | CAPRI+ | http://zdock.umassmed.edu/ |
| Hex | Geometric matching | CAPRI+ | http://www.csd.abdn.ac.uk/hex/ |
| SLIDE | Systematic | Force field, Empirical | http://www.bmb.msu.edu/~kuhn/software/slide/ |
| Fred | Systematic | Empirical | http://www.eyesopen.com/oedocking |
| LeDock | Annealing-Genetic algorithm | Physics/knowledge hybrid | http://www.lephar.com/software.htm |
| Glide | Systematic | XP/SP/HTVS | https://www.schrodinger.com |
| Surflex-Dock | Hammerhead | Empirical | http://www.tripos.com |
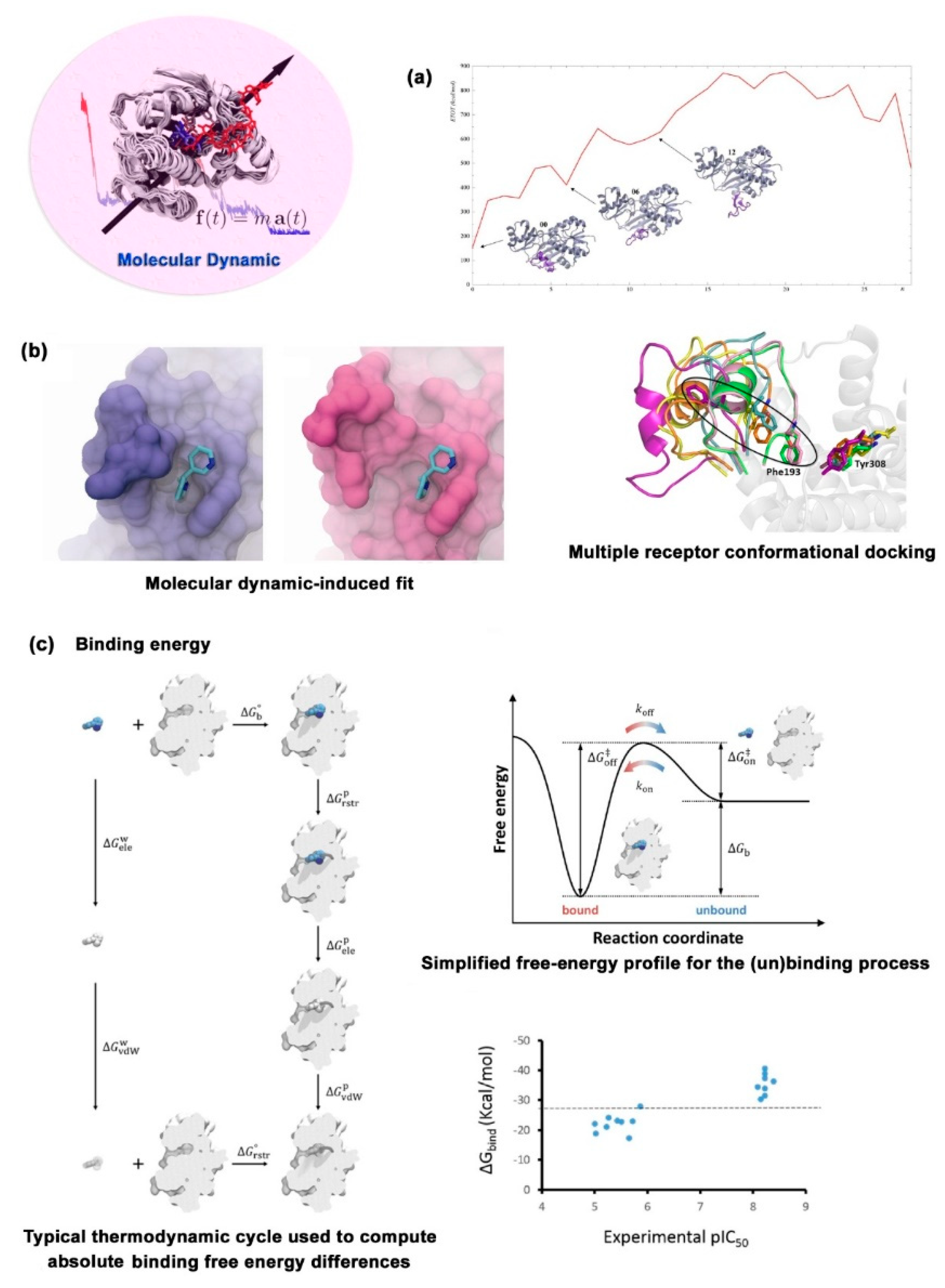
2.1.3. Molecular Dynamic
| Software | Scoring function | Charge | Website |
|---|---|---|---|
| Amber | Mainly for biological system | AmberTools Free | http://ambermd.org/ |
| CPMD | Biological and chemical system | Free | http://www.cpmd.org/ |
| NAMD | Biological and chemical system | Free | http://www.ks.uiuc.edu/Research/namd |
| Lammps | Materials and solid state physical systems | Free | https://www.lammps.org/ |
| Gromacs | Mainly for biological system | Free | https://www.gromacs.org/ |
| Charmm | Mainly for biological system | Free | https://www.charmm.org/ |
| Tinker | Mainly for biological system | Free | http://dasher.wustl.edu/tinker/ |
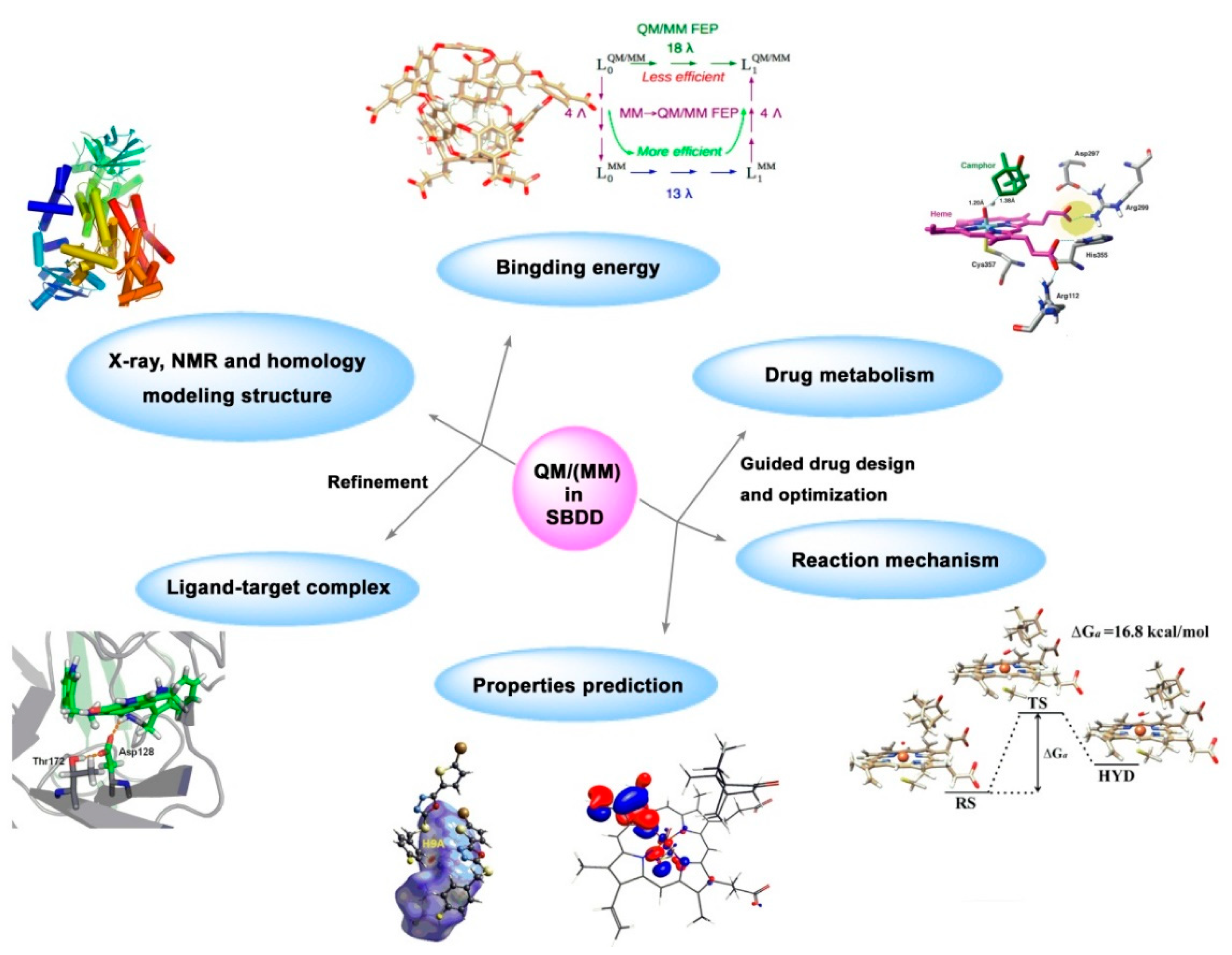
2.1.4. Quantum Chemistry
2.1.5. Molecular Docking-Molecular Dynamic-Quantum Chemistry
2.1.6. Virtual Screening
2.2. Ligand-Based Drug Design
2.2.1. Quantitative Structure-Activity Relationship
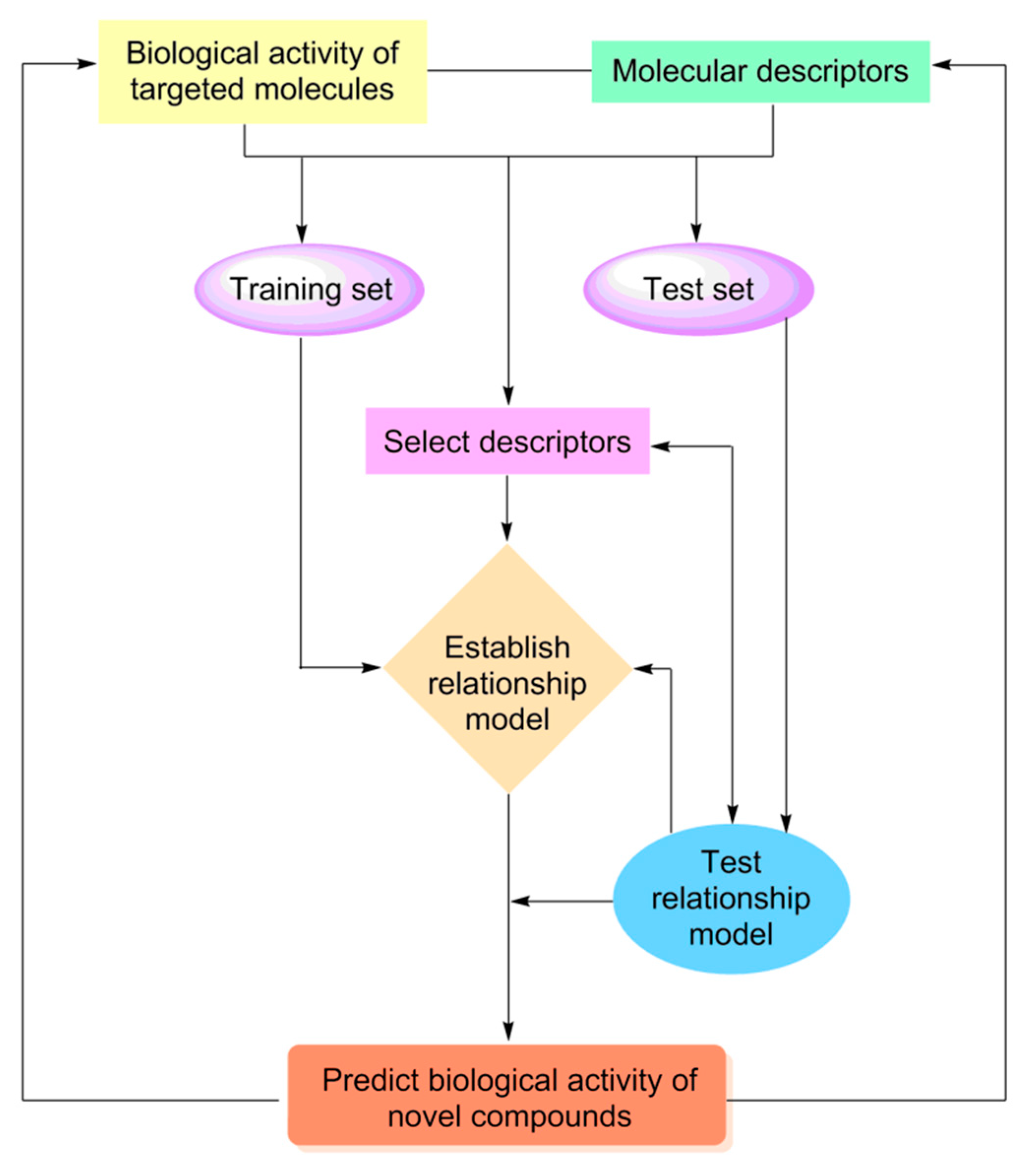
2.2.2. DFT-Based Quantitative Structure-Activity Relationship
| Definition | Name |
|---|---|
| Charges | |
| QA | net atomic charge on atom A |
| Qmin, Qmax | net charges of the most negative and most positive atoms |
| QAB | net group charge on atoms A, B |
| QT, QA | sum of absolute values of the charges of all the atoms in a given molecule |
| QT2, QA2 | sum of squares of the charges of all the atoms in a given molecule or functional group |
| Qm | mean absolute atomic charge (i.e. the average of the absolute values of the charges on all atoms) |
| HOMO and LUMO Energies | |
| EHOMO, ELUMO | energies of the highest occupied (HOMO) and (LUMO) molecular orbitals lowest unoccupied |
| ∆ELUMO-HOMO | HOMO and LUMO orbital energy difference |
| η = (ELUMO - EHOMO)/2 | hardness |
| S = 1/(ELUMO - EHOMO). | softness |
| ∆η = ηR - ηT | activation hardness, R and T stand for reactant and transition state |
| Molecular Polarizabilities | |
| α | molecular polarizability |
| α = (αxx + αyy + αzz)/3 | mean polarizability of the molecule |
| β2 = [(αxx - αyy)2 + (αyy - αzz)2 + (αzz - αxx)2] | anisotropy of the polarizability |
| Dipole Moments and Polarity Indices | |
| µ | molecular dipole moment |
| µchar, µ | charge and hybridization components of the dipole moment |
| µ2 | square of the molecular dipole moment |
| DX, DY, DZ | components of dipole moment along inertia axes |
| ∆ | submolecular polarity parameter (largest difference in electron charges between two atoms) |
| τ | quadrupole moment tensor |
| Energies | |
| E | total energy |
| H | Enthalpy |
| G | Gibbs free energy |
| S | entropy |
| IP | ionization potential |
| EA | electron affinity, difference in total energy between the neutral and anion radical species |
| Orbital Electron Densities | |
| qA, σ, qA, π | σ- and π-electron densities of the atom A |
| QA,H, QA,L | HOMO/LUMO electron densities on the atom A |
| FrE = frE/EHOMO | electrophilic atomic frontier electron densities |
| FrN = frN/ELUMO | |
| Atom-Atom Polarizabilities | |
| πAA, πAB | self-atom polarizabilities and atom-atom polarizabilities |
| Superdelocalizabilities | |
| SE, A, SN, A | electrophilic and nucleophilic superdel ocalizabilities |
2.2.3. Pharmacophore Modeling
2.2.4. Molecular Similarity
3. Kinase
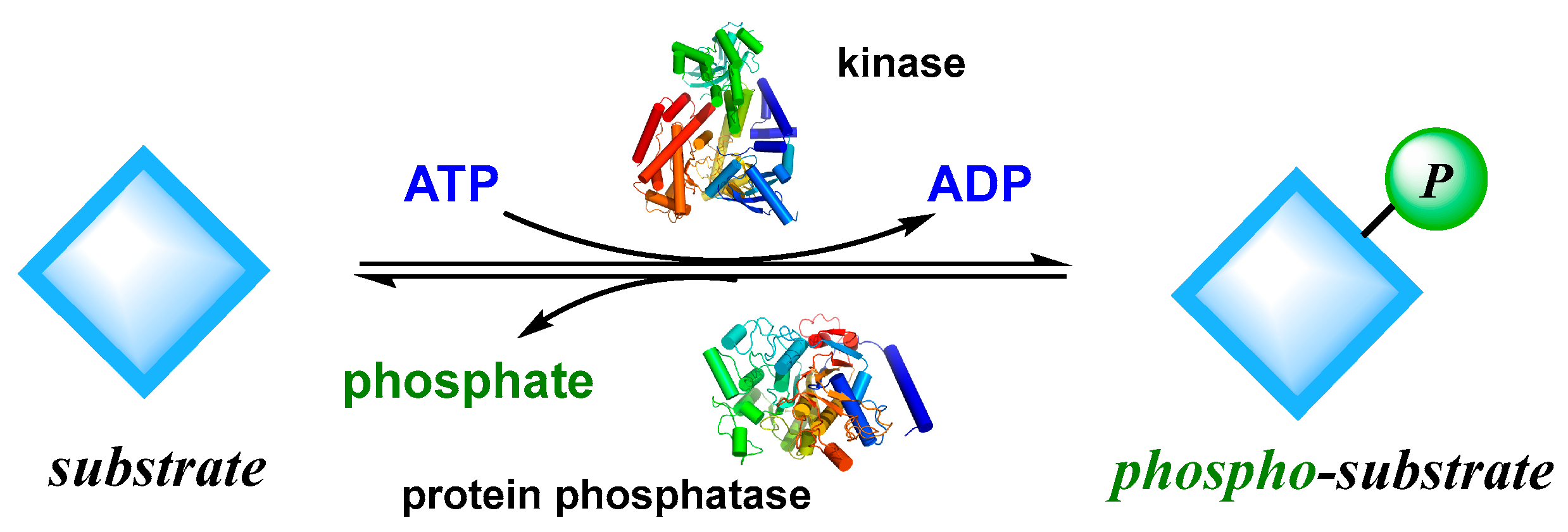
3.1. Structure and Function of Kinase
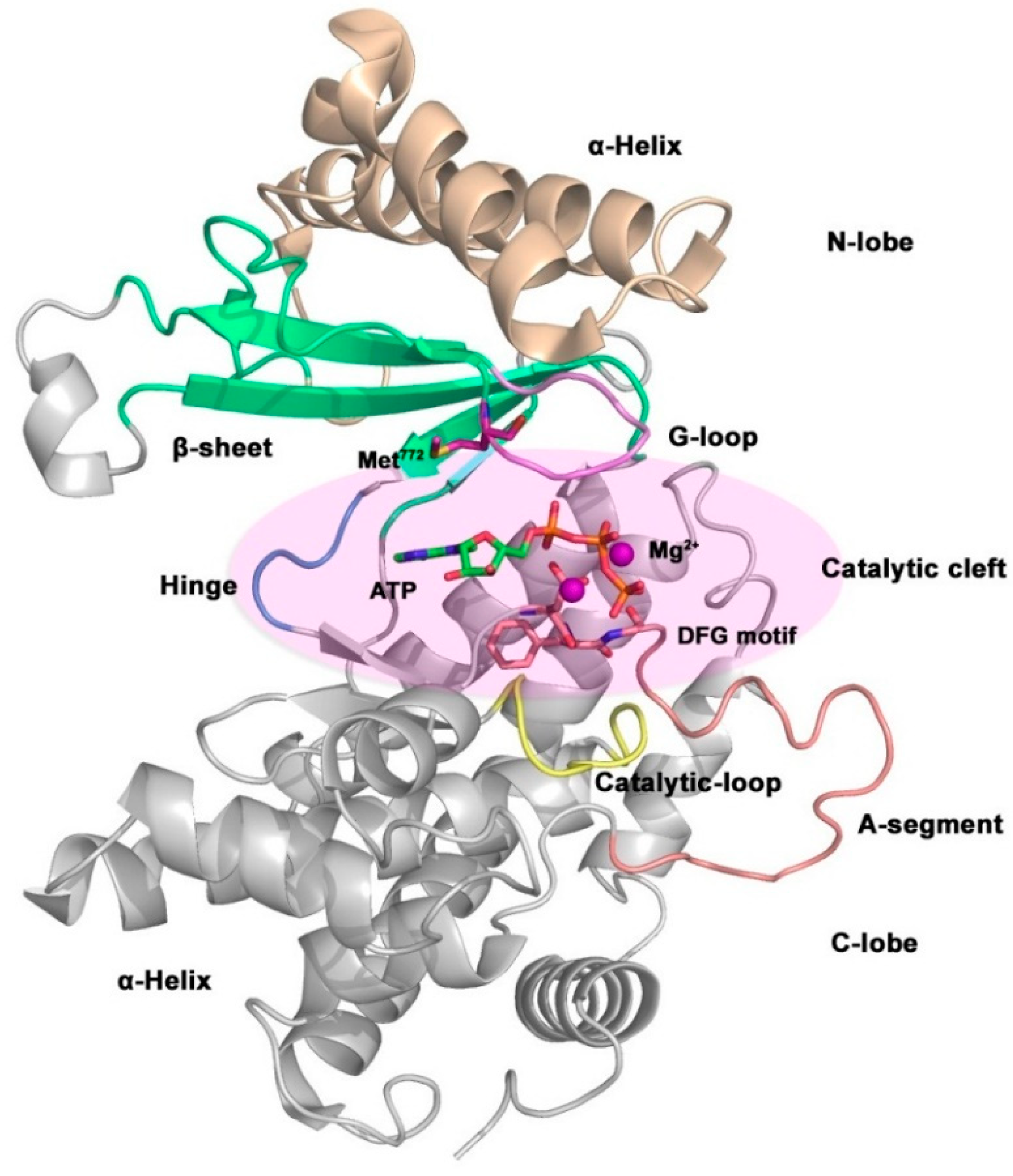
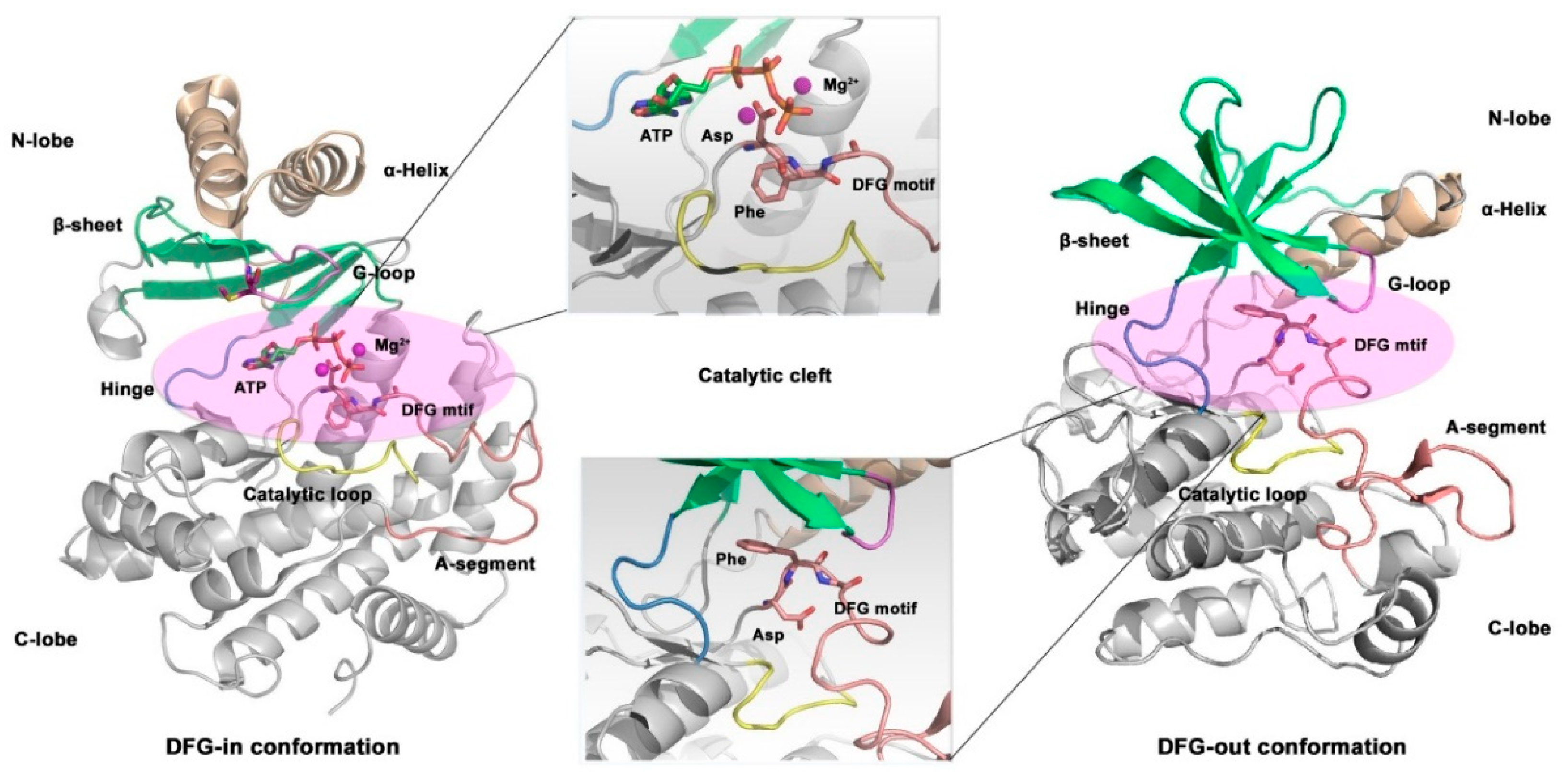
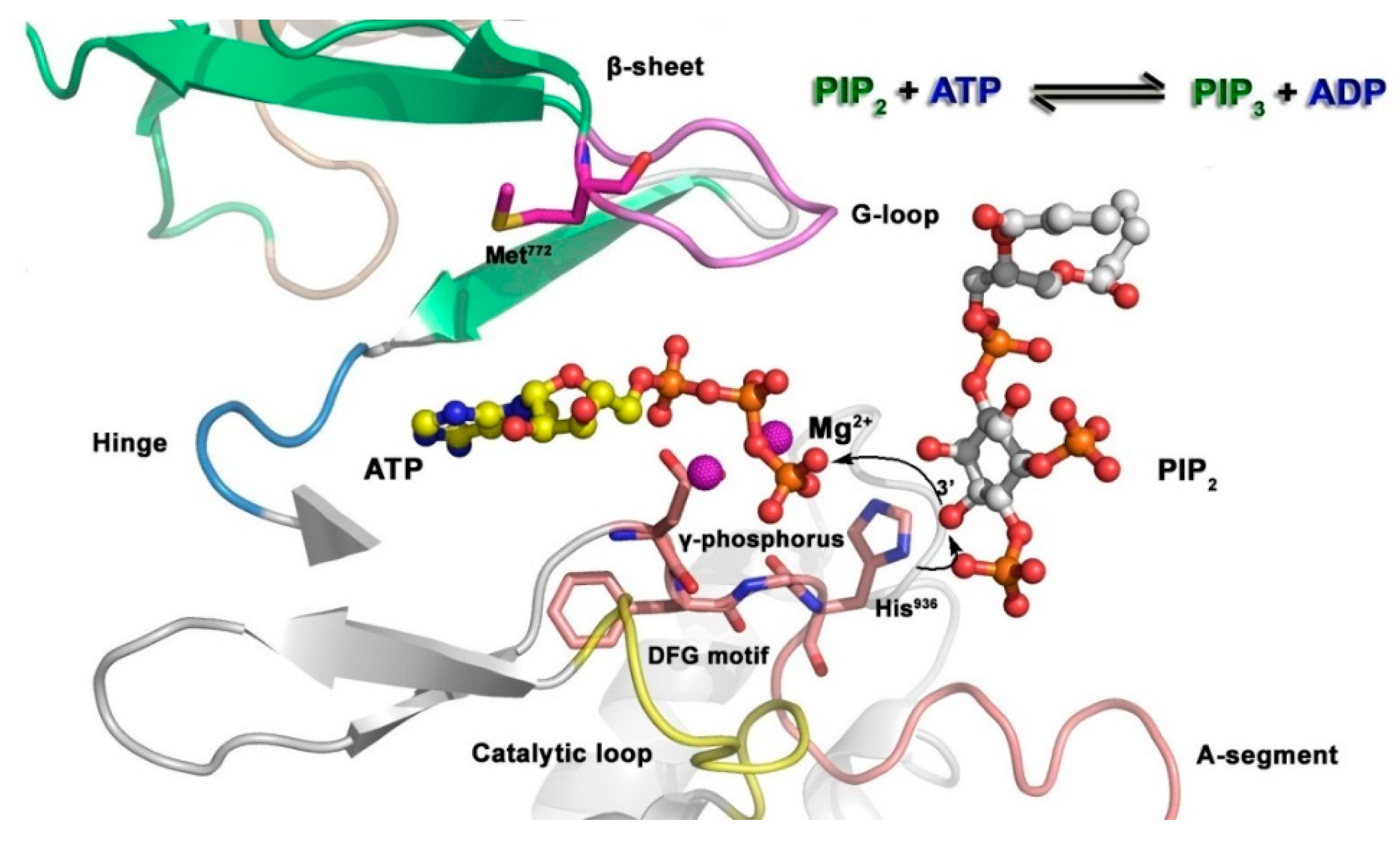
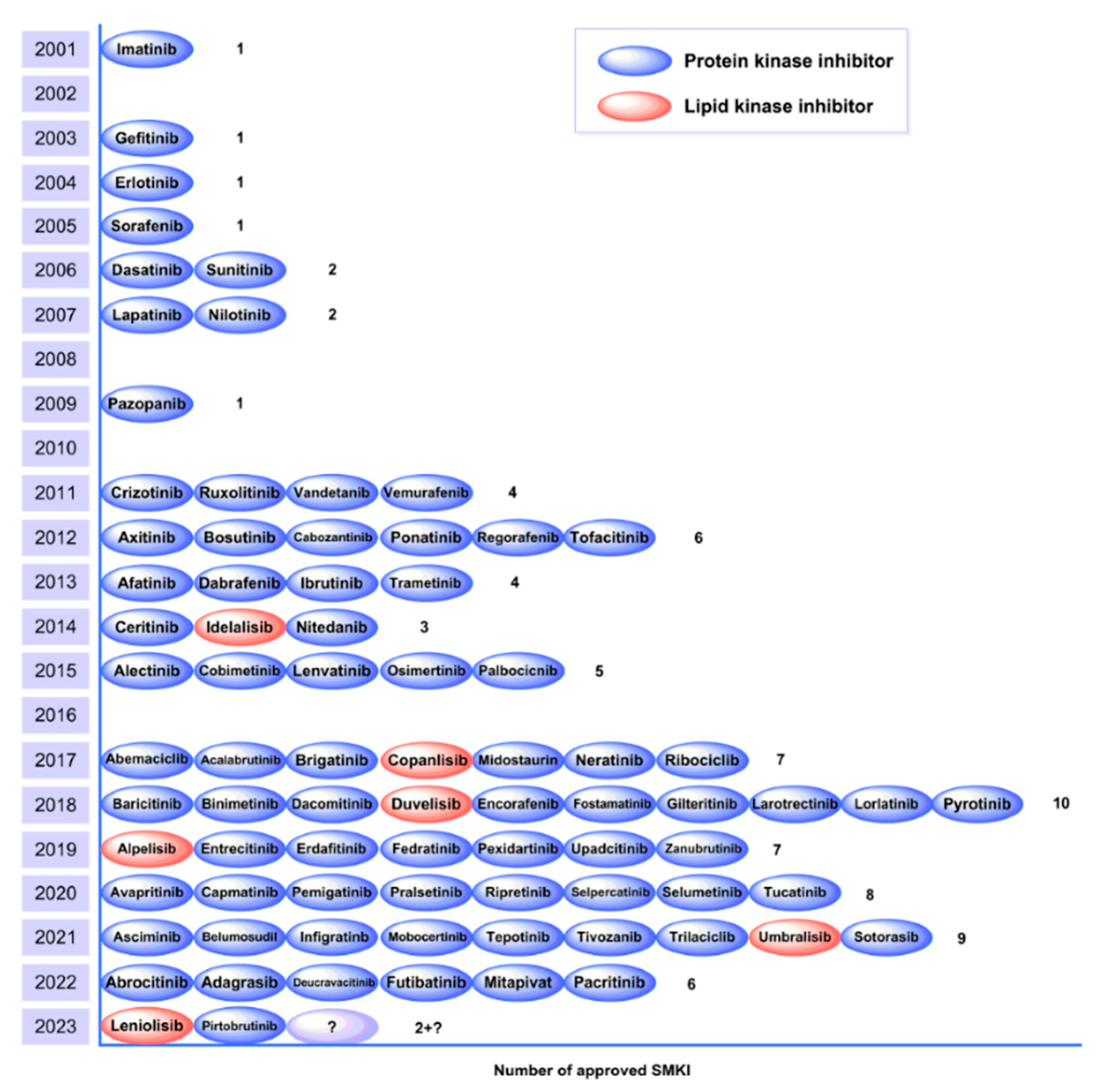
3.2. Small Molecule Kinase Inhibitors
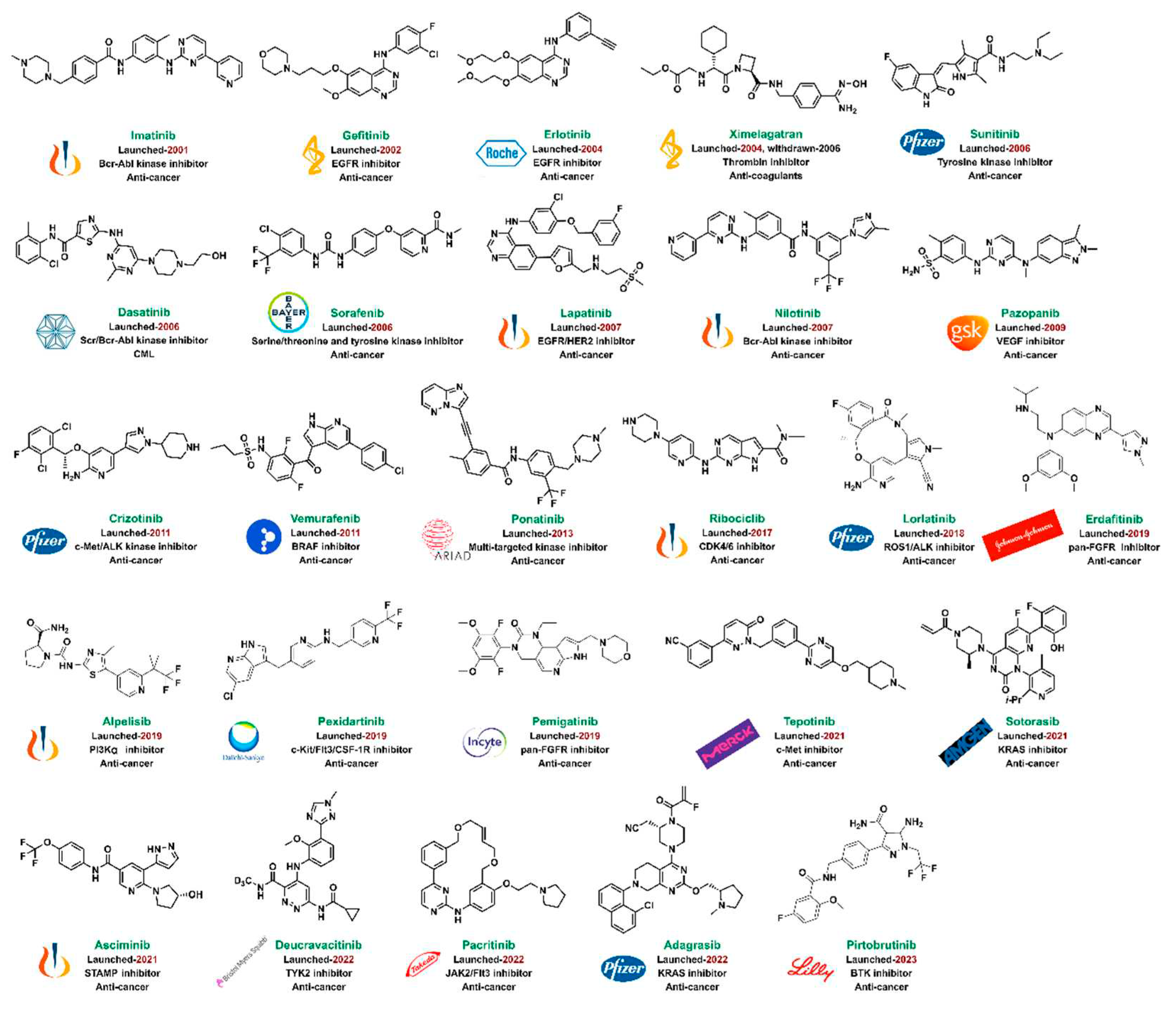
4. Small Molecule Kinase Inhibitors Discovered by CADD
5. Future Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Zhao, L.; Ciallella, H.L.; Aleksunes, L.M.; et al. Advancing computer-aided drug discovery (CADD) by big data and data-driven machine learning modeling. Drug discovery today 2020, 25, 1624–1638. [Google Scholar] [CrossRef]
- Gomeni, R.; Bani, M.; D'angeli, C.; et al. Computer-assisted drug development (CADD): an emerging technology for designing first-time-in-man and proof-of-concept studies from preclinical experiments. European Journal of Pharmaceutical Sciences 2001, 13, 261–270. [Google Scholar] [CrossRef] [PubMed]
- Finn. Application of SBDD to the discovery of new antibacterial drugs. In <i>Structure-Based Drug, Discovery</i>; Tari, L. (Eds.) Finn. Application of SBDD to the discovery of new antibacterial drugs. In Structure-Based Drug Discovery; Tari, L., Eds.; Humana Press: 2012.
- Acharya, C.; Coop, A.; Polli, J.E.; et al. Recent Advances in Ligand-Based Drug Design: Relevance and Utility of the Conformationally Sampled Pharmacophore Approach. Curr Comput Aided Drug Des 2011, 7, 10–22. [Google Scholar] [CrossRef] [PubMed]
- Sabe, V.T.; Ntombela, T.; Jhamba, L.A.; et al. Current trends in computer aided drug design and a highlight of drugs discovered via computational techniques: A review. European Journal of Medicinal Chemistry 2021, 224, 113705. [Google Scholar] [CrossRef] [PubMed]
- Foster, J.S.; Wimalasena, J. Estrogen regulates activity of cyclin-dependent kinases and retinoblastoma protein phosphorylation in breast cancer cells. Molecular Endocrinology 1996, 10, 488–498. [Google Scholar]
- Haldar, S.; Chintapalli, J.; Croce, C.M. Taxol Induces bcl-2 Phosphorylation and Death of Prostate Cancer Cells. Cancer Research 1996, 56, 1253–1255. [Google Scholar]
- Itoh, N.; Semba, S.; Ito, M.; et al. Phosphorylation of Akt/PKB is required for suppression of cancer cell apoptosis and tumor progression in human colorectal carcinoma. Cancer 2010, 94, 3127–3134. [Google Scholar] [CrossRef]
- Mcdonald, P.C.; Oloumi, A.; Mills, J.; et al. Rictor and integrin-linked kinase interact and regulate Akt phosphorylation and cancer cell survival. Cancer Research 2008, 68, 1618–1624. [Google Scholar] [CrossRef]
- Cordwell, S.J.; White, M.Y. Targeted proteomics for determining phosphorylation site-specific associations in cardiovascular disease. Circulation 2012, 126, 1803–1807. [Google Scholar] [CrossRef]
- Nishida, M.; Saiki, S.; Kitajima, N.; et al. ChemInform Abstract: Regulation of Cardiovascular Functions by the Phosphorylation of TRPC Channels. Cheminform 2011, 42, no. [Google Scholar] [CrossRef]
- Streeter, J.; Schickling, B.; Thiel, W.; et al. Nox1 Phosphorylation in Cardiovascular Disease. Free Radical Biology & Medicine 2012, 53, S175–S175. [Google Scholar]
- Wieland, T.; Attwood, P.V. Alterations in reversible protein histidine phosphorylation as intracellular signals in cardiovascular disease. Frontiers in Pharmacology 2015, 6, 173–173. [Google Scholar] [CrossRef] [PubMed]
- Ja, P.; et al. Targeted Deletion of AIF Decreases Mitochondrial Oxidative Phosphorylation and Protects from Obesity and Diabetes. Cell 2007, 131, 476–491. [Google Scholar]
- Liu, Y.; Liu, F.; Grundkeiqbal, I.; et al. Brain glucose transporters, O-GlcNAcylation and phosphorylation of tau in diabetes and Alzheimer's disease. Journal of Neurochemistry 2010, 111, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Szendroedi, J.; Schmid, A.I.; Chmelik, M.; et al. Muscle mitochondrial ATP synthesis and glucose transport/phosphorylation in type 2 diabetes. Plos Medicine 2007, 4, 154–154. [Google Scholar] [CrossRef]
- Liu, B.; Yang, Y.; Chernishof, V.; et al. Proinflammatory stimuli induce IKKalpha-mediated phosphorylation of PIAS1 to restrict inflammation and immunity. Cell 2007, 129, 903–914. [Google Scholar] [CrossRef]
- Caudle, R.M.; Perez, F.M.; Vallepinero AY, D.; et al. Spinal cord NR1 serine phosphorylation and NR2B subunit suppression following peripheral inflammation. Molecular Pain 2005, 1, 25–25. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, M.; Mimuro, H.; Kiga, K.; et al. Helicobacter pylori CagA Phosphorylation-Independent Function in Epithelial Proliferation and Inflammation. Cell Host & Microbe 2009, 5, 23–34. [Google Scholar]
- Wertz, I.E.; Newton, K.; Seshasayee, D.; et al. Phosphorylation and linear ubiquitin direct A20 inhibition of inflammation. Nature 2015, 528, 370–375. [Google Scholar] [CrossRef]
- Zhang, Y.; Qiu, J.; Wang, X.; et al. AMP-activated protein kinase suppresses endothelial cell inflammation through phosphorylation of transcriptional coactivator p300. Arterioscler Thromb Vasc Biol 2011, 31, 2897–2908. [Google Scholar] [CrossRef]
- BuãE, L.; BussiãRe, T.; BuãE-Scherrer V, et al. Tau protein isoforms, phosphorylation and role in neurodegenerative disorders. Brain Res Brain Res Rev 2000, 33, 95–130. [Google Scholar] [CrossRef] [PubMed]
- Hanger, D.P.; Anderton, B.H.; Noble, W. Tau phosphorylation: the therapeutic challenge for neurodegenerative disease. Trends in Molecular Medicine 2009, 15, 112–119. [Google Scholar] [CrossRef] [PubMed]
- Mp, M.; Pm, F. Untangling tau hyperphosphorylation in drug design for neurodegenerative diseases. Nature Reviews Drug Discovery 2007, 6, 464–479. [Google Scholar]
- Davis, M.I.; Hunt, J.P.; Herrgard, S.; et al. Comprehensive analysis of kinase inhibitor selectivity. Nature Biotechnology 2011, 29, 1046–1051. [Google Scholar] [CrossRef] [PubMed]
- Attwood, M.M.; Fabbro, D.; Sokolov, A.V.; et al. Trends in kinase drug discovery: targets, indications and inhibitor design. Nature Reviews Drug Discovery 2021, 20, 839–861. [Google Scholar] [CrossRef]
- Macalino, S.Y.; Vijayakumar, G.; Sunhye, H.; et al. Role of computer-aided drug design in modern drug discovery. Archives of Pharmacal Research 2015, 38, 1686–1701. [Google Scholar] [CrossRef]
- Dimasi, J.A.; Grabowski, H.G.; Hansen, R.W. Innovation in the pharmaceutical industry: New estimates of R&D costs. Journal of Health Economics 2016, 47, 20–33. [Google Scholar]
- Patani, G.A.; Lavoie, E. Bioisosterism: A Rational Approach in Drug Design. Chemical Reviews 1996, 96, 3147–3176. [Google Scholar] [CrossRef]
- Brown, F.K.; Sherer, E.C.; Johnson, S.A.; et al. The evolution of drug design at Merck Research Laboratories. Journal of Computer-Aided Molecular Design 2017, 31, 1–12. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef]
- Cavasotto, C.N.; Phatak, S.S. Homology modeling in drug discovery: current trends and applications. Drug Discovery Today 2009, 14, 676–683. [Google Scholar] [CrossRef] [PubMed]
- Morris, G.M.; Lim-Wilby, M. Molecular docking. Methods in Molecular Biology 2008, 443, 365–382. [Google Scholar] [PubMed]
- Jones, G.; Willett, P.; Glen, R.C. A genetic algorithm for flexible molecular overlay and pharmacophore elucidation. Journal of Computer-Aided Molecular Design 1995, 9, 532–532. [Google Scholar] [CrossRef] [PubMed]
- Hopfinger, A.J.; Wang, S.; Tokarski, J.S.; Jin, B.; Albuquerque, M.; Madhav, P.J.; Duraiswami, C. Construction of 3D-QSAR Models Using the 4D-QSAR Analysis Formalism. Journal of the American Chemical Society 1997, 119, 10509–10524. [Google Scholar] [CrossRef]
- Durrant, J.D.; Andrew, M.C. Molecular dynamics simulations and drug discovery. BMC Biology 2011, 9, 71–71. [Google Scholar] [CrossRef] [PubMed]
- Keseru, G.M.; Kolossváry, I. Molecular mechanics and conformational analysis in drug design[M]. 1999.
- Gravenstein, S.; Johnston, S.L.; Loeschel, E.; et al. Zanamivir. Drug Safety 2001, 24, 1113–1125. [Google Scholar] [CrossRef]
- Noble, S.; Faulds, D. Saquinavir. Drugs 1996, 52, 93–112. [Google Scholar] [CrossRef]
- Silver, R.T. Imatinib mesylate (Gleevec (TM)) reduces phlebotomy requirements in polycythemia vera. Leukemia 2003, 17, 1186–1187. [Google Scholar] [CrossRef]
- Chirikjian, G.S. Conformational Modeling of Continuum Structures in Robotics and Structural Biology: A Review. Advanced Robotics the International Journal of the Robotics Society of Japan 2015, 29, 817–829. [Google Scholar] [CrossRef]
- Congreve, M.; Murray, C.W.; Blundell, T.L. Keynote review: Structural biology and drug discovery. Drug Discovery Today 2005, 10, 895–907. [Google Scholar] [CrossRef]
- Garratt, R. Structural biology and cancer. Bmc Proceedings 2013, 7, 1–1. [Google Scholar] [CrossRef]
- Holler, T.P.; Evdokimov, A.G.; Narasimhan, L. Structural biology approaches to antibacterial drug discovery. Expert Opinion on Drug Discovery 2007, 2, 1085–1101. [Google Scholar] [CrossRef] [PubMed]
- Penin, F.; Dubuisson, J.; Rey, F.A.; et al. Structural biology of hepatitis C virus. Clinics in Liver Disease 2003, 7, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Tomita, T.; Iwatsubo, T. Structural Biology of Presenilins and Signal Peptide Peptidases. Journal of Biological Chemistry 2013, 288, 14673–14680. [Google Scholar] [CrossRef]
- Fenalti, G.; Buckle, A.M. Structural biology of the GAD autoantigen. Autoimmunity Reviews 2010, 9, 148–152. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, L.G.; Dos Santos, R.N.; Oliva, G.; et al. Molecular docking and structure-based drug design strategies. Molecules 2015, 20, 13384–13421. [Google Scholar] [CrossRef]
- Meng, X.Y.; Zhang, H.X.; Mezei, M.; et al. Molecular docking: a powerful approach for structure-based drug discovery. Current Computer-Aided Drug Design 2016, 7, 146–157. [Google Scholar] [CrossRef]
- Yuriev, E.; Holien, J.; Ramsland, P.A. Improvements, trends, and new ideas in molecular docking: 2012-2013 in review. Journal of Molecular Recognition 2015, 28, 581–604. [Google Scholar] [CrossRef] [PubMed]
- Yuriev, E.; Ramsland, P.A. Latest developments in molecular docking: 2010-2011 in review. Journal of Molecular Recognition Jmr 2013, 26, 215–239. [Google Scholar] [CrossRef]
- Chen, H.; Lyne, P.D.; Giordanetto, F.; et al. On evaluating molecular-docking methods for pose prediction and enrichment factors. Journal of Chemical Information & Modeling 2006, 46, 401–415. [Google Scholar]
- Dias, R.; Timmers, L.F.; Caceres, R.A.; et al. Evaluation of molecular docking using polynomial empirical scoring functions. Current Drug Targets 2008, 9, 1062–1070. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.D.; Jewsbury, P.J.; Essex, J.W. A review of protein-small molecule docking methods. Journal of Computer-Aided Molecular Design 2002, 16, 151–166. [Google Scholar] [CrossRef] [PubMed]
- Yadav, D.K.; Kumar, S.; et al. Molecular docking, QSAR and ADMET studies of withanolide analogs against breast cancer. Drug Design Development & Therapy 2017, 11, 1859–1870. [Google Scholar]
- Nair, V.; Ma, X.; Shu, Q.; Zhang, F.; Uchil, V.; Cherukupalli, G.R. Impdh As A Biological Probe For Rna Antiviral Drug Discovery: Synthesis, Enzymology, Molecular Docking, And Antiviral Activity Of New Ribonucleosides With Surrogate Bases. Cheminform 2007, 26, 651–654. [Google Scholar]
- Ding, W.; Gu, J.; Cao, L.; et al. Traditional Chinese herbs as chemical resource library for drug discovery of anti-infective and anti-inflammatory. Journal of Ethnopharmacology 2014, 155, 589–598. [Google Scholar] [CrossRef]
- Balamurugan, R.; Stalin, A.; Ignacimuthu, S. Molecular docking of γ-sitosterol with some targets related to diabetes. European Journal of Medicinal Chemistry 2012, 47, 38–43. [Google Scholar] [CrossRef]
- Rayalu, D.J.; Selvaraj, C.; Singh, S.K.; et al. Homology modeling, active site prediction, and targeting the anti hypertension activity through molecular docking on endothelin-B receptor domain. Bioinformation 2012, 8, 81–86. [Google Scholar] [CrossRef]
- Mathew, B. Molecular Docking Studies of Some Novel Antidepressant 5-Substituted Phenyl-3-(Thiophen-2-yl)-4, 5-Dihydro-1h-Pyrazole-1-Carboxamides against Monoamine Oxidase Isoforms. Central Nervous System Agents in Medicinal Chemistry(Formerly Current Medicinal 2015, 16, 75–80. [Google Scholar] [CrossRef]
- Abdel-Aziz, A.A.; Eltahir, K.E.; Asiri, Y.A. Synthesis, anti-inflammatory activity and COX-1/COX-2 inhibition of novel substituted cyclic imides. Part 1: Molecular docking study. European Journal of Medicinal Chemistry 2011, 46, 1648–1655. [Google Scholar] [CrossRef]
- Cheng, K.; Zheng, Q.Z.; Qian, Y.; et al. Synthesis, antibacterial activities and molecular docking studies of peptide and Schiff bases as targeted antibiotics. Bioorg Med Chem 2009, 17, 7861–7871. [Google Scholar] [CrossRef]
- Ma, D.L.; Chan, S.H.; Leung, C.H. Molecular docking for virtual screening of natural product databases. Chemical Science 2011, 2, 1656–1665. [Google Scholar] [CrossRef]
- Ruyck, J.D.; Brysbaert, G.; Blossey, R.; et al. Molecular docking as a popular tool in drug design, an in silico travel. Advances & Applications in Bioinformatics & Chemistry Aabc 2016, 9, 1–11. [Google Scholar]
- Stark, J.L.; Powers, R. Application of NMR and molecular docking in structure-based drug discovery. Topics in Current Chemistry 2012, 326, 1–34. [Google Scholar] [PubMed]
- Stark, J.L.; Powers, R. Application of NMR and molecular docking in structure-based drug discovery. Topics in Current Chemistry 2012, 326, 1–34. [Google Scholar]
- Foroutan, M.; Fatemi, S.M.; Esmaeilian, F. A review of the structure and dynamics of nanoconfined water and ionic liquids via molecular dynamics simulation. European Physical Journal E 2017, 40, 19. [Google Scholar] [CrossRef]
- Komanduri; Raff, M. L. A review on the molecular dynamics simulation of machining at the atomic scale. Proceedings of the Institution of Mechanical Engineers Part B Journal of Engineering Manufacture 2001, 215, 1639–1672. [Google Scholar] [CrossRef]
- Car, R.; Parrinello, M. Unified approach for molecular dynamics and density-functional theory. Physical Review Letters 1985, 55, 2471. [Google Scholar] [CrossRef]
- Vivo, M.D.; Masetti, M.; Bottegoni, G.; et al. Role of Molecular Dynamics and Related Methods in Drug Discovery. Journal of Medicinal Chemistry 2016, 59, 4035–4061. [Google Scholar] [CrossRef]
- Galeazzi, R. Molecular Dynamics as a Tool in Rational Drug Design: Current Status and Some Major Applications. Current Computer-Aided Drug Design 2009, 5, 225–240. [Google Scholar] [CrossRef]
- Mortier, J.; Rakers, C.; Bermudez, M.; et al. The impact of molecular dynamics on drug design: applications for the characterization of ligand–macromolecule complexes. Drug Discovery Today 2015, 20, 686–702. [Google Scholar] [CrossRef]
- Perryman, A.L.; Lin, J.H.; Mccammon, J.A. Restrained molecular dynamics simulations of HIV-1 protease: the first step in validating a new target for drug design. Biopolymers 2010, 82, 272–284. [Google Scholar] [CrossRef]
- Durrant, J.D.; MxCammon, J.A. Molecular dynamic simulations and drug discovery. BMC Biology.
- Wereszczynski, J.; Mccammon, J.A. Accelerated molecular dynamics in computational drug design. Methods in Molecular Biology 2012, 819, 515. [Google Scholar]
- Zhao, H.; Caflisch, A. Molecular dynamics in drug design. European Journal of Medicinal Chemistry 2015, 91, 4–14. [Google Scholar] [CrossRef]
- Lin, J.H.; Perryman, A.L.; Schames, J.R.; et al. Computational drug design accommodating receptor flexibility: the relaxed complex scheme. Journal of the American Chemical Society 2002, 124, 5632. [Google Scholar] [CrossRef] [PubMed]
- Peters, J.H.; De Groot, B.L. Ubiquitin dynamics in complexes reveal molecular recognition mechanisms beyond induced fit and conformational selection. Plos Computational Biology 2012, 8, e1002704. [Google Scholar] [CrossRef]
- Sotriffer, C.A.; Krämer, O.; Klebe, G. Probing flexibility and "induced-fit" phenomena in aldose reductase by comparative crystal structure analysis and molecular dynamics simulations. Proteins Structure Function & Bioinformatics 2010, 56, 52–66. [Google Scholar]
- Biswa Ranjan, M.; Yixuan, W. Interaction of I50V mutant and I50L/A71V double mutant HIV-protease with inhibitor TMC114 (darunavir): molecular dynamics simulation and binding free energy studies. Journal of Physical Chemistry B 2012, 116, 1884–1900. [Google Scholar]
- Zhou, T.; Georgeon, S.; Moser, R.; et al. Specificity and mechanism-of-action of the JAK2 tyrosine kinase inhibitors ruxolitinib and SAR302503 (TG101348). Leukemia 2014, 28, 404–407. [Google Scholar] [CrossRef]
- Spiliotopoulos, D.; Caflisch, A. Molecular Dynamics Simulations of Bromodomains Reveal Binding-Site Flexibility and Multiple Binding Modes of the Natural Ligand Acetyl-Lysine. Israel Journal of Chemistry 2015, 54, 1084–1092. [Google Scholar] [CrossRef]
- De, V.M.; Masetti, M.; Bottegoni, G.; et al. The Role of Molecular Dynamics and Related Methods in Drug Discovery. Journal of Medicinal Chemistry 2016, 59, 4035–406. [Google Scholar]
- Johnson, K.H. Quantum Chemistry. Annual Review of Physical Chemistry 1975, 26, 39–57. [Google Scholar] [CrossRef]
- Arnold, A.; Weigend, F.; Evers, F. Quantum chemistry calculations for molecules coupled to reservoirs: formalism, implementation, and application to benzenedithiol. Journal of Chemical Physics 2007, 126, 174101–174101. [Google Scholar] [CrossRef] [PubMed]
- Chernev, P.; Zaharieva, I.; Rossini, E.; et al. Merging Structural Information from X-ray Crystallography, Quantum Chemistry and EXAFS Spectra: The Oxygen Evolving Complex in PSII. Journal of Physical Chemistry B 2016, 120, 10899–10922. [Google Scholar] [CrossRef]
- Hernándezvaldés, D.; Alberto, R.; Jáureguihaza, U. Quantum chemistry calculations of technetium and rhenium compounds with application in radiopharmacy: review. Rsc Advances 2016, 6, 107127–107140. [Google Scholar] [CrossRef]
- Kim, W.Y.; Choi, Y.C.; Min, S.K.; et al. Application of quantum chemistry to nanotechnology: electron and spin transport in molecular devices. Cheminform 2009, 38, 2319–2333. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Min, G.; Zheng, X.; et al. Application of Quantum Chemistry Method in the Performance Evaluation and Mechanism Study of Corrosion Inhibitors. Corrosion & Protection 2017, 38, 829–833. [Google Scholar]
- Mannhold, R.; Kubinyi, H.; Folkers, G. Quantum Medicinal Chemistry[M]. 2005.
- Lyne, P.D.; Hodoscek, M.; Karplus, M. A Hybrid QM-MM Potential Employing Hartree-Fock or Density Functional Methods in the Quantum Region. Journal of Physical Chemistry A 1999, 103, 3462–3471. [Google Scholar] [CrossRef]
- Dinner, A.R.; Lopez, X.; Karplus, M. A charge-scaling method to treat solvent in QM/MM simulations. Theoretical Chemistry Accounts 2003, 109, 118–124. [Google Scholar] [CrossRef]
- Reuter, N.; Dejaegere, A.; Maigret, B.; Karplus, M. Frontier Bonds in QM/MM Methods: A Comparison of Different Approaches. Journal of Physical Chemistry A 2011, 104, 1720–1735. [Google Scholar] [CrossRef]
- Pezeshki, S.; Lin, H. Adaptive-Partitioning QM/MM for Molecular Dynamics Simulations: 4. Proton Hopping in Bulk Water. Journal of Chemical Theory & Computation 2015, 11, 2398–2411. [Google Scholar]
- Zhou, Y.Q. Martin Karplus Feeling of winning. Journal of Seeking Knowledge Guide 2013, 150–151. [Google Scholar]
- Cui, Q.; Karplus, M. Quantum Mechanical/Molecular Mechanical Studies of the Triosephosphate Isomerase-Catalyzed Reaction: Verification of Methodology and Analysis of Reaction Mechanisms. Journal of Physical Chemistry B 2002, 106, 1768–1798. [Google Scholar] [CrossRef]
- Chung, L.W.; Sameera, W.M.; Ramozzi, R.; et al. The ONIOM Method and Its Applications. Chemical Reviews 2015, 115, 5678–5796. [Google Scholar] [CrossRef]
- Svensson, M.; Humbel, S.; Froese RD, J.; et al. ONIOM: A Multilayered Integrated MO+MM Method for Geometry Optimizations and Single Point Energy Predictions. A Test for Diels-Alder Reactions and Pt(P(t-Bu)3)2+H2 Oxidative Addition. Journal of Physical Chemistry 1996, 100, 174–186. [Google Scholar] [CrossRef]
- Vreven, T.; Byun, K.S.; Komáromi, I.; et al. Combining Quantum Mechanics Methods with Molecular Mechanics Methods in, O.N.I.O.M. Journal of Chemical Theory & Computation 2006, 2, 815–826. [Google Scholar]
- Vreven, T.; Morokuma, K.; Farkas, O.; et al Geometry optimization with, Q.M.; et al. /.M.M.; ONIOM; other combined methods, I. Microiterations and constraints. Journal of Computational Chemistry 2003, 24, 760–769. [Google Scholar] [CrossRef] [PubMed]
- Adeniyi, A.A.; Soliman, M.E.S. Implementing QM in docking calculations: is it a waste of computational time? Drug Discovery Today 2017, 22, 1216–1223. [Google Scholar] [CrossRef] [PubMed]
- Chaskar, P.; Zoete, V.; Röhrig, U.F. On-the-fly QM/MM Docking with Attracting Cavities. Journal of Chemical Information & Modeling 2016, 57, 73–84. [Google Scholar]
- Lu, J.; Zhang, Z.; Zhong, N.; et al. QM/MM–PB/SA scoring of the interaction strength between Akt kinase and apigenin analogues. Computational Biology & Chemistry 2014, 52, 25–33. [Google Scholar]
- Raha K, Peters M B, Wang B, et al. The role of quantum mechanics in structure-based drug design. Drug Discovery Today 2007, 12, 725–731. [Google Scholar] [CrossRef]
- Kelly, E.B.; Tuszynski, J.A.; Klobukowski, M. QM and QM/MD simulations of the Vinca alkaloids docked to tubulin. Journal of Molecular Graphics & Modelling 2011, 30, 54–66. [Google Scholar]
- Adam, P.; Martin, L.; Jan, E.; et al. QM/MM calculations reveal the different nature of the interaction of two carborane-based sulfamide inhibitors of human carbonic anhydrase, I.I. Journal of Physical Chemistry B 2013, 117, 16096–16104. [Google Scholar]
- Alzate-Morales, J. Caballero Computational study of the interactions between guanine derivatives cyclin-dependent kinase 2 (CDK2) by, C.o.M.F.A.; QM/MM. Journal of Chemical Information & Modeling 2010, 50, 110–122. [Google Scholar]
- Cebriã, N.-P.A.; Rovira, T.; Saura, P.; et al. On the Inhibition of Mammalian 15-Lipoxygenase by Three Ebselen-like Drugs. A QM/MM and MM/PBSA Comparative Study. Journal of Physical Chemistry A 2017, 121, 9752–9763. [Google Scholar]
- Jung-Hsin, L.; Perryman, A.L.; Schames, J.R.; et al. Computational drug design accommodating receptor flexibility: the relaxed complex scheme. Journal of the American Chemical Society 2002, 124, 5632–5633. [Google Scholar]
- Nie, F.; Kunciw, D.L.; Wilcke, D.; et al. A Multidimensional Diversity-Oriented Synthesis Strategy for Structurally Diverse and Complex Macrocycles. Angew Chem Int Ed Engl 2016, 55, 11139–11143. [Google Scholar] [CrossRef]
- Lehn, J.M. Dynamic Combinatorial Chemistry and Virtual Combinatorial Libraries. Chemistry – A European Journal 2015, 5, 2455–2463. [Google Scholar] [CrossRef]
- Szymkuć, S.; Gajewska, E.P.; Klucznik, T.; et al. Computer-Assisted Synthetic Planning: The End of the Beginning. Angewandte Chemie 2016, 55, 5904–5937. [Google Scholar] [CrossRef]
- Shah, S.; Reddy, S.; Sardeshmukh, A.; et al. Application of Machine Learning Techniques for Inverse Prediction in Manufacturing Process Chains[M]. Springer International Publishing, 2015.
- Butler, K.T.; Davies, D.W.; Cartwright, H.; et al. Machine learning for molecular and materials science. Nature 2018, 559, 547–555. [Google Scholar] [CrossRef]
- Granda, J.M.; Donina, L.; Dragone, V.; et al. Controlling an organic synthesis robot with machine learning to search for new reactivity. Nature 2018, 559, 377–381. [Google Scholar] [CrossRef]
- Segler, M.H.S.; Preuss, M.; Waller, M.P. Planning chemical syntheses with deep neural networks and symbolic AI. Nature 2018, 555, 604–610. [Google Scholar] [CrossRef] [PubMed]
- Schneider, G. Virtual screening: an endless staircase? Nature Reviews Drug Discovery 2010, 9, 273–276. [Google Scholar] [CrossRef] [PubMed]
- Miles, J.A.; Ross, B.P. Recent Advances in Virtual Screening for Cholinesterase Inhibitors. Acs Chemical Neuroscience 2021, 12, 30–41. [Google Scholar] [CrossRef] [PubMed]
- Pal, S.; Kuma, V.; Kundu, B.; et al. Ligand-based Pharmacophore Modeling, Virtual Screening and Molecular Docking Studies for Discovery of Potential Topoisomerase I Inhibitors. Computational and Structural Biotechnology Journal 2019, 17, 291–310. [Google Scholar] [CrossRef]
- Gahlawat, A.; Kumar, N.; Kumar, R.; et al. Structure-Based Virtual Screening to Discover Potential Lead Molecules for the SARS-CoV-2 Main Protease. Journal of Chemical Information and Modeling 2020, 60, 5781–5793. [Google Scholar] [CrossRef]
- Dudek, A.Z.; Tomasz, A.; Jorge, G. Computational methods in developing quantitative structure-activity relationships (QSAR): a review. Combinatorial Chemistry & High Throughput Screening 2006, 9, 213–228. [Google Scholar]
- Chen, B. Development of quantitative structure activity relationship (QSAR) model for disinfection byproduct (DBP) research: A review of methods and resources. Journal of Hazardous Materials.
- Lill, M.A. Multi-dimensional QSAR in drug discovery. Drug Discovery Today 2007, 12, 1013–1017. [Google Scholar] [CrossRef]
- Ghafourian, T.; Zandasrar, P.; Hamishekar, H.; et al. The effect of penetration enhancers on drug delivery through skin: a QSAR study. Journal of Controlled Release 2004, 99, 113–125. [Google Scholar] [CrossRef]
- Vilar, S.; Cozza, G. ; S Medicinal chemistry and the molecular operating environment (MOE): application of QSAR and molecular docking to drug discovery. Current Topics in Medicinal Chemistry 2008, 8, 1555–1572. [Google Scholar] [CrossRef]
- Devillers. Neural Networks in QSAR and Drug Design[M]. City: Harcourt Brace, 1996.
- Low, Y.; Uehara, T.; Minowa, Y.; et al. Predicting Drug-induced Hepatotoxicity Using QSAR and Toxicogenomics Approaches. Chemical Research in Toxicology 2011, 24, 1251–1262. [Google Scholar] [CrossRef]
- Yoshida, F.; Topliss, J.G. QSAR model for drug human oral bioavailability. Journal of Medicinal Chemistry 2000, 43, 2575–2585. [Google Scholar] [CrossRef]
- Viskupicova, J.; Danihelova, M.; Majekova, M.; et al. Polyphenol fatty acid esters as serine protease inhibitors: a quantum-chemical QSAR analysis. J Enzyme Inhib Med Chem 2012, 27, 800–809. [Google Scholar] [CrossRef] [PubMed]
- Nongonierma, A.; Fitzgerald, D. Learnings from quantitative structure activity relationship (QSAR) studies with respect to food protein-derived bioactive peptides: A review. Rsc Advances 2016, 6, 75400–75413. [Google Scholar] [CrossRef]
- Mccoy, E.F.; Sykes, M. Quantum-mechanical QSAR/QSPR descriptors from momentum-space wave functions. Journal of Chemical Information & Computer Sciences 2003, 43, 545–553. [Google Scholar]
- Gozalbes, R.; Doucet, J.P.; Derouin, F. Application of topological descriptors in QSAR and drug design: history and new trends. Current Drug Targets - Infectious Disorders 2002, 2, 93–102. [Google Scholar] [CrossRef]
- Karelson, M.; Lobanov, V.S.; Katritzky, A.R. Quantum-Chemical Descriptors in QSAR/QSPR Studies. Chemical Reviews 1996, 96, 1027–1044. [Google Scholar] [CrossRef] [PubMed]
- Borota A, Mracec M, Gruia A, et al. A QSAR study using MTD method and Dragon descriptors for a series of selective ligands of αC adrenoceptor. European Journal of Medicinal Chemistry 2011, 46, 877–884. [Google Scholar] [CrossRef]
- Davood, A.; Nematollahi, A.; Iman, M.; et al. Computational studies of new 1,4-dihydropyridines containing 4-(5)-chloro-2-ethyl-5-(4)-imidazolyl substituent: QSAR and docking. Medicinal Chemistry Research 2010, 19, 58–70. [Google Scholar] [CrossRef]
- Milan, M.; Mirjana, M.; Desanka, B.; et al. In VitroAntioxidant Activity of Selected 4-Hydroxy-chromene-2-one Derivatives-SAR, QSAR and DFT Studies. International Journal of Molecular Sciences 2011, 12, 2822–2841. [Google Scholar]
- Zhang, L.; Wan, J.; Yang, G. A DFT-based QSARs study of protoporphyrinogen oxidase inhibitors: phenyl triazolinones. Bioorganic & Medicinal Chemistry 2004, 12, 6183–6191. [Google Scholar]
- Morales-Bayuelo, A.; Matute, R.A.; Caballero, J. Understanding the comparative molecular field analysis (CoMFA) in terms of molecular quantum similarity and DFT-based reactivity descriptors. Journal of Molecular Modeling 2015, 21, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Ran, T.; Lu, T.; Yuan, H.; et al. A selectivity study on mTOR/PI3Kα inhibitors by homology modeling, 3.D.-Q.S.A.R. Journal of Molecular Modeling 2012, 18, 171–186. [Google Scholar] [CrossRef] [PubMed]
- Bharate, S.B.; Singh, B.; Bharate, J.B.; et al. QSAR and Pharmacophore Modeling of N-Acetyl-2-aminobenzothiazole Class of Phosphoinositide-3-kinase-α Inhibitors. Medicinal Chemistry Research 2012, 22, 890–899. [Google Scholar] [CrossRef]
- Wrã³bel, T.M.; Kieå‚Bus, M.; Kaczor, A.A.; et al. Discovery of nitroaryl urea derivatives with antiproliferative properties. J Enzyme Inhib Med Chem 2016, 31, 608–618. [Google Scholar] [CrossRef]
- Yang, S.Y. Pharmacophore modeling and applications in drug discovery: challenges and recent advances. Drug Discovery Today 2010, 15, 444–450. [Google Scholar] [CrossRef]
- Schaller, D.; Šribar, D.; Noonan, T.; et al. Next generation 3D pharmacophore modeling. Wiley Interdisciplinary Reviews 2020, 10, e1468. [Google Scholar] [CrossRef]
- Giordano, D.; Biancaniello, C.; Argenio, M.A.; et al. Drug Design by Pharmacophore and Virtual Screening Approach. Pharmaceuticals 2022, 15, 646. [Google Scholar] [CrossRef]
- Seidel, T.; Wieder, O.; Garon, A.; et al. Applications of the Pharmacophore Concept in Natural Product inspired Drug Design. Molecular Informatics 2020, 39, 2000059. [Google Scholar] [CrossRef]
- Muhammed, M.T.; Akl-Yalcin, E. Pharmacophore Modeling in Drug Discovery: Methodology and Current Status. Journal of the Turkish Chemical Society Section A; Chemistry 2021, 8, 748–762. [Google Scholar] [CrossRef]
- Seidel, T.; Schuetz, D.A.; Garon, A.; et al. The Pharmacophore Concept and Its Applications in Computer-Aided Drug Design. Progress in the Chemistry of Organic Natural Products 2019, 110, 99–141. [Google Scholar]
- Levinson, A.D.; Oppermann, H.; Levintow, L.; et al. Evidence that the transforming gene of avian sarcoma virus encodes a protein kinase associated with a phosphoprotein. Cell 1978, 15, 561–572. [Google Scholar] [CrossRef] [PubMed]
- Lerea, K.M.; Tonks, N.K.; Krebs, E.G.; et al. Vanadate and molybdate increase tyrosine phosphorylation in a 50-kilodalton protein and stimulate secretion in electropermeabilized platelets. Biochemistry 1989, 28, 9286–9292. [Google Scholar] [CrossRef] [PubMed]
- Erickson, A.K.; Payne, D.M.; Martino, P.A.; et al. Identification by mass spectrometry of threonine 97 in bovine myelin basic protein as a specific phosphorylation site for mitogen-activated protein kinase. Journal of Biological Chemistry 1990, 265, 19728–19735. [Google Scholar] [CrossRef] [PubMed]
- Endicott, J.A.; Johnson, L.N. Protein Kinase Inhibitors: Insights into Drug Design from Structure. Science 2004, 303, 1800–1805. [Google Scholar]
- Hiles, I.D.; Otsu, M.; Volinia, S.; et al. Phosphatidylinositol 3-kinase: structure and expression of the 110 kd catalytic subunit. Cell 1992, 70, 419–429. [Google Scholar] [CrossRef]
- Wymann, M.P.; Pirola, L. Structure and function of phosphoinositide 3-kinases. Biochim Biophys Acta 1998, 1436, 127–150. [Google Scholar] [CrossRef]
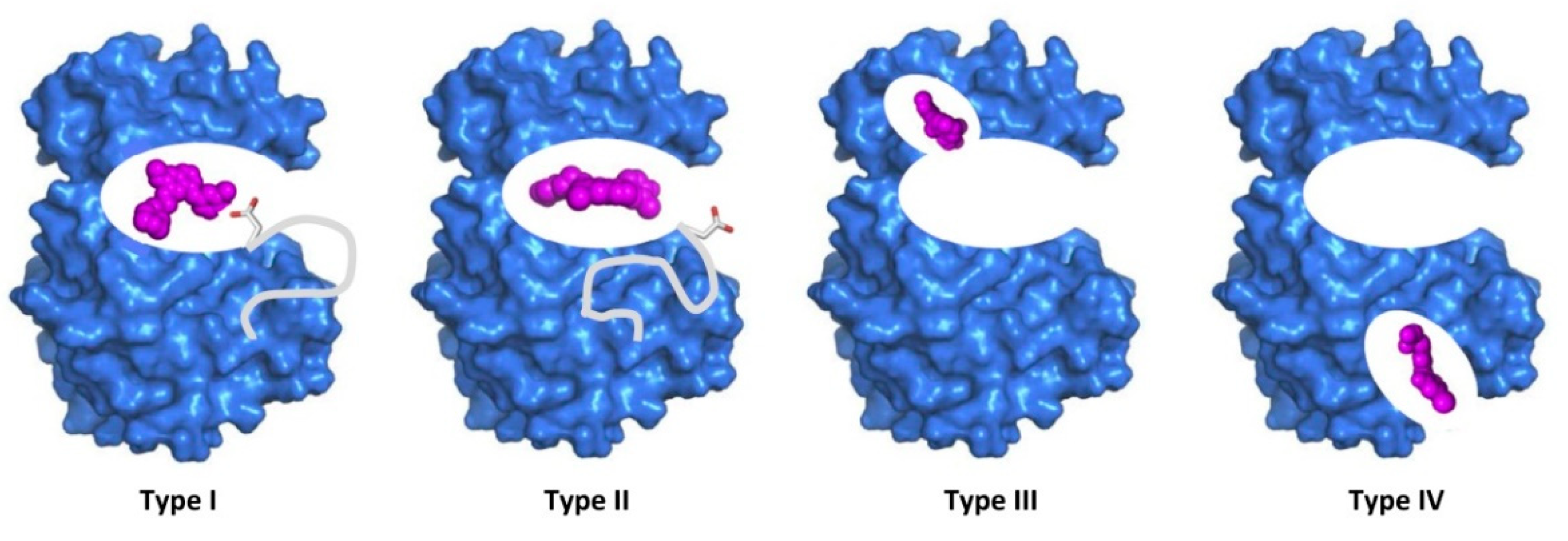
- Liu, Y.; Gray, N.S. Liu, Y. & Gray, N.S. Rational design of inhibitors that bind to inactive kinase conformations. Nat. Chem. Biol. 2, 358-364. Nature Chemical Biology 2006, 2, 358–364. [Google Scholar]
- Tengholm, A.; Meyer, T. A PI3-Kinase Signaling Code for Insulin-Triggered Insertion of Glucose Transporters into the Plasma Membrane. Current Biology 2002, 12, 1871–1876. [Google Scholar] [CrossRef]
- Ung, P.M.; Schlessinger, A. DFGmodel: predicting protein kinase structures in inactive states for structure-based discovery of type-II inhibitors. Acs Chemical Biology 2015, 10, 269–278. [Google Scholar] [CrossRef]
- Mk, A.; Jd, V. Small molecule inhibitors of phosphoinositide 3-kinase (PI3K) delta and gamma. Current Topics in Medicinal Chemistry 2009, 9, 738–753. [Google Scholar]
- Johnson, L.N.; Lewis, R. Structural basis for control by phosphorylation. Chemical Reviews 2001, 101, 2209–2209. [Google Scholar] [CrossRef] [PubMed]
- Roskoski, R. Properties of FDA-approved small molecule protein kinase inhibitors. Pharmacological Research 2023, 187, 106552. [Google Scholar] [CrossRef] [PubMed]
- Peng, W.; Nielsen, T.E.; Clausen, M.H. FDA-approved small-molecule kinase inhibitors. Trends in Pharmacological Sciences 2015, 36, 422–439. [Google Scholar]
- Noble ME, M.; Endicott, J.A.; Johnson, L.N. Protein kinase inhibitors: insights into drug design from structure. Science 2004, 303, 1800–1805. [Google Scholar] [CrossRef]
- Norman, R.A.; Toader, D.; Ferguson, A.D. Structural approaches to obtain kinase selectivity. Trends in Pharmacological Sciences 2012, 33, 273–278. [Google Scholar] [CrossRef]
- Cox, K.J.; Shomin, C.D.; Ghosh, I. Tinkering outside the kinase ATP box: allosteric (type IV) and bivalent (type V) inhibitors of protein kinases. Future Medicinal Chemistry 2011, 3, 29–43. [Google Scholar] [CrossRef]
- Lamba, V.; Ghosh, I. New directions in targeting protein kinases: focusing upon true allosteric and bivalent inhibitors. Current Pharmaceutical Design 2012, 18, 2936–2945. [Google Scholar] [CrossRef]
- Gazit, A.; Yaish, P.; Gilon, C.; et al. Tyrphostins I: synthesis and biological activity of protein tyrosine kinase inhibitors. Journal of Medicinal Chemistry 1989, 32, 2344–2352. [Google Scholar] [CrossRef]
- Yaish, P.; Gazit, A.; Gilon, C.; et al. Blocking of EGF-dependent cell proliferation by EGF receptor kinase inhibitors. Science 1988, 242, 933–935. [Google Scholar] [CrossRef]
- Capdeville, R.; Buchdunger, E.; Zimmermann, J.; et al. Glivec (STI571, imatinib), a rationally developed, targeted anticancer drug. Nature Reviews Drug Discovery 2002, 1, 493–502. [Google Scholar] [CrossRef]
- Wu, P.; Nielsen, T.E.; Clausen, M.H. Small-molecule kinase inhibitors: an analysis of FDA-approved drugs. Drug Discovery Today 2016, 21, 5–10. [Google Scholar] [CrossRef]
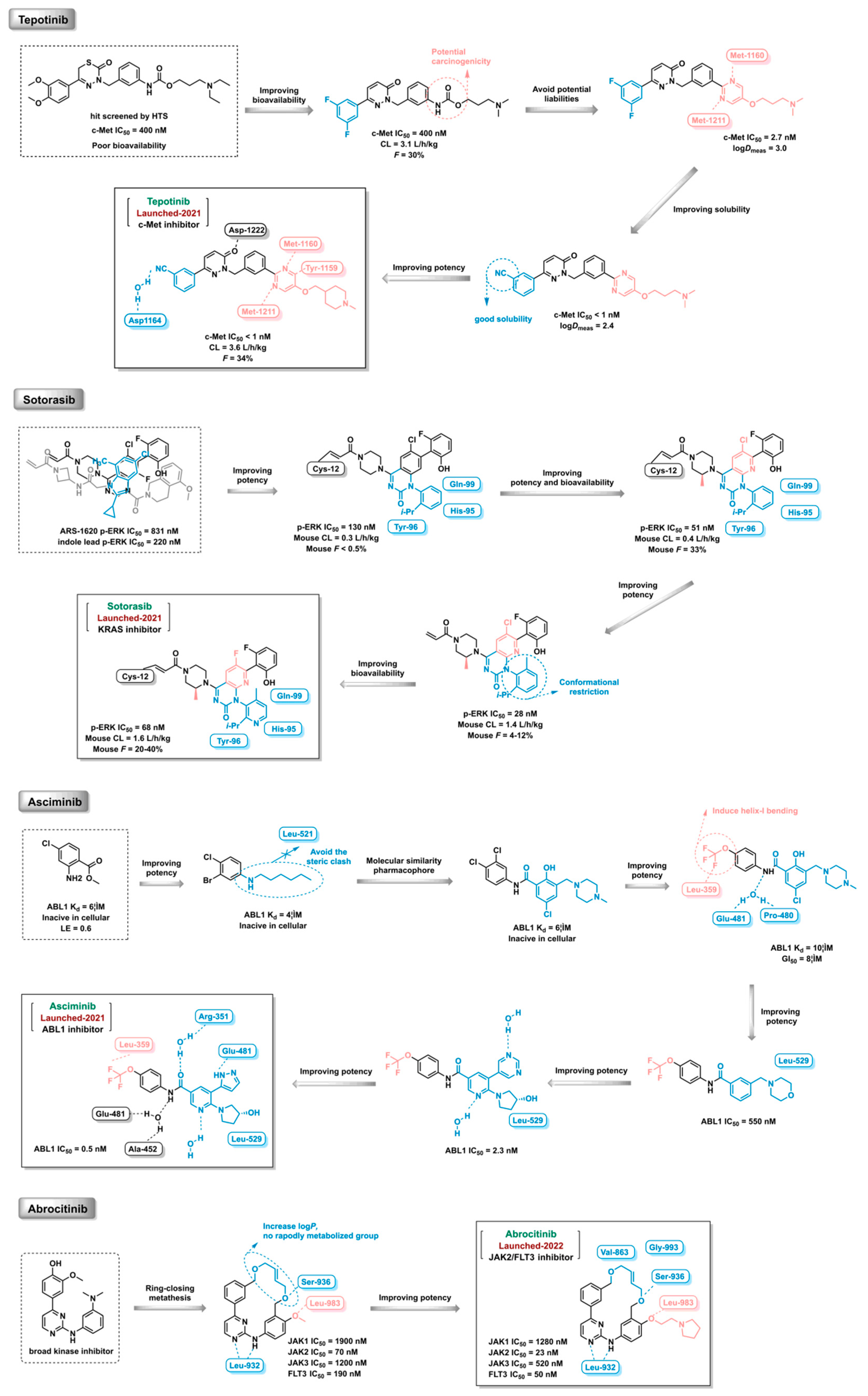
- Dorsch, D.; Schadt, O.; Stieber, F.; et al. Identification and optimization of pyridazinones as potent and selective c-Met kinase inhibitors. Bioorganic & Medicinal Chemistry Letters 2015, 25, 1597–1602. [Google Scholar]
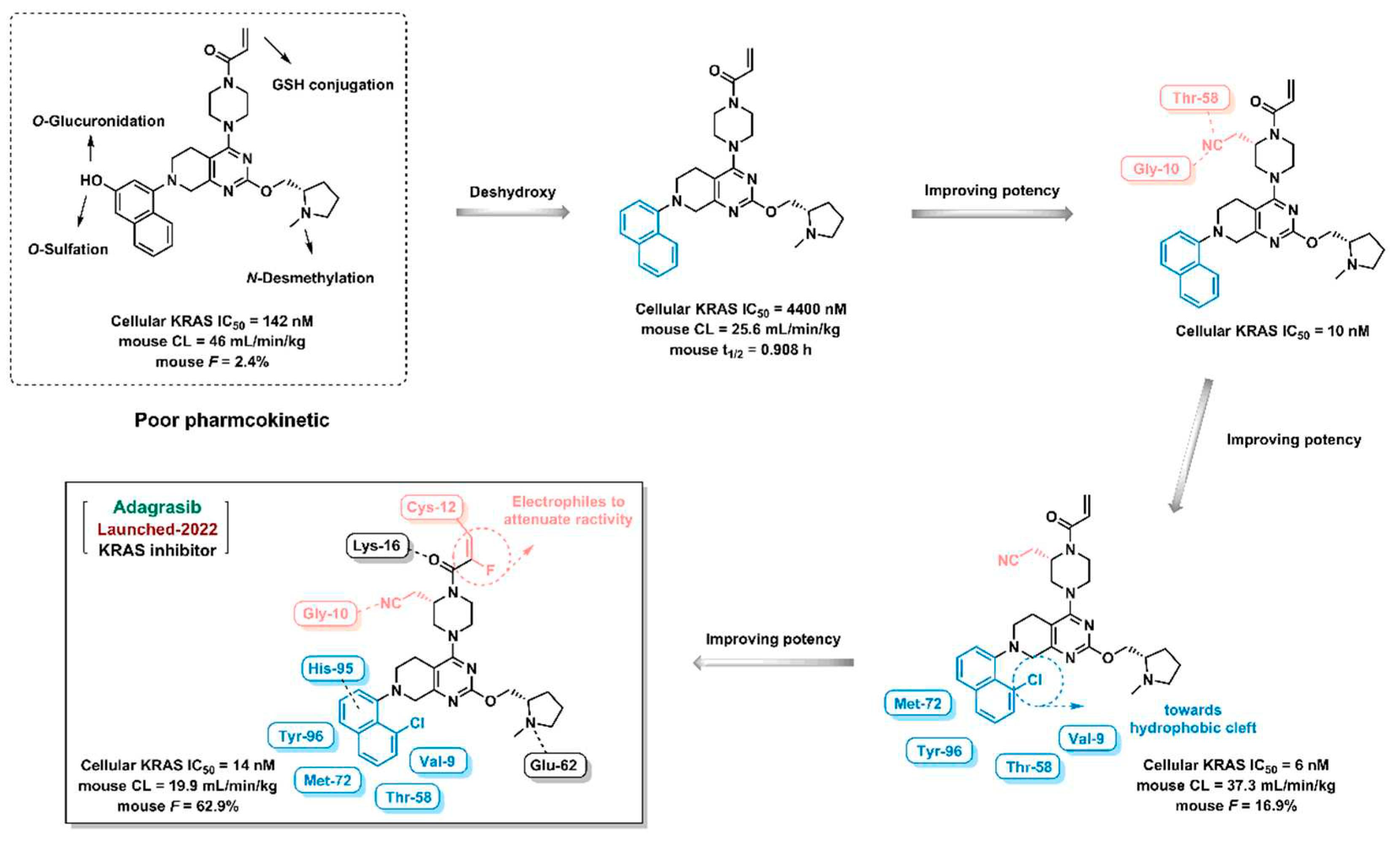
- Lanman, B.A.; Allen, J.R.; Allen, J.G.; et al. Discovery of a covalent inhibitor of KRASG12C (AMG 510) for the treatment of solid tumors. Journal of Medicinal Chemistry 2020, 63, 52–65. [Google Scholar] [CrossRef] [PubMed]
- Schoepfer, J.; Jahnke, W.; Berellini, G.; et al. Discovery of asciminib (ABL001), an allosteric inhibitor of the tyrosine kinase activity of BCR-ABL1. Journal of Medicinal Chemistry 2018, 61, 8120–8135. [Google Scholar] [CrossRef] [PubMed]
- William, A.D.; Lee AC, H.; Blanchard, S.; et al. Discovery of the Macrocycle 11-(2-Pyrrolidin-1-yl-ethoxy)-14,19-dioxa-5,7,26-triaza-tetracyclo[19.3.1.1(2,6).1(8,12)]heptacosa-1(25),2(26),3,5,8,10,12(27),16,21,23-decaene (SB1518), a Potent Janus Kinase 2/Fms-Like Tyrosine Kinase-3 (JAK2/FLT3) Inhibitor for the Treatment of Myelofibrosis and Lymphoma. Journal of Medicinal Chemistry 2011, 54, 4638–4658. [Google Scholar]
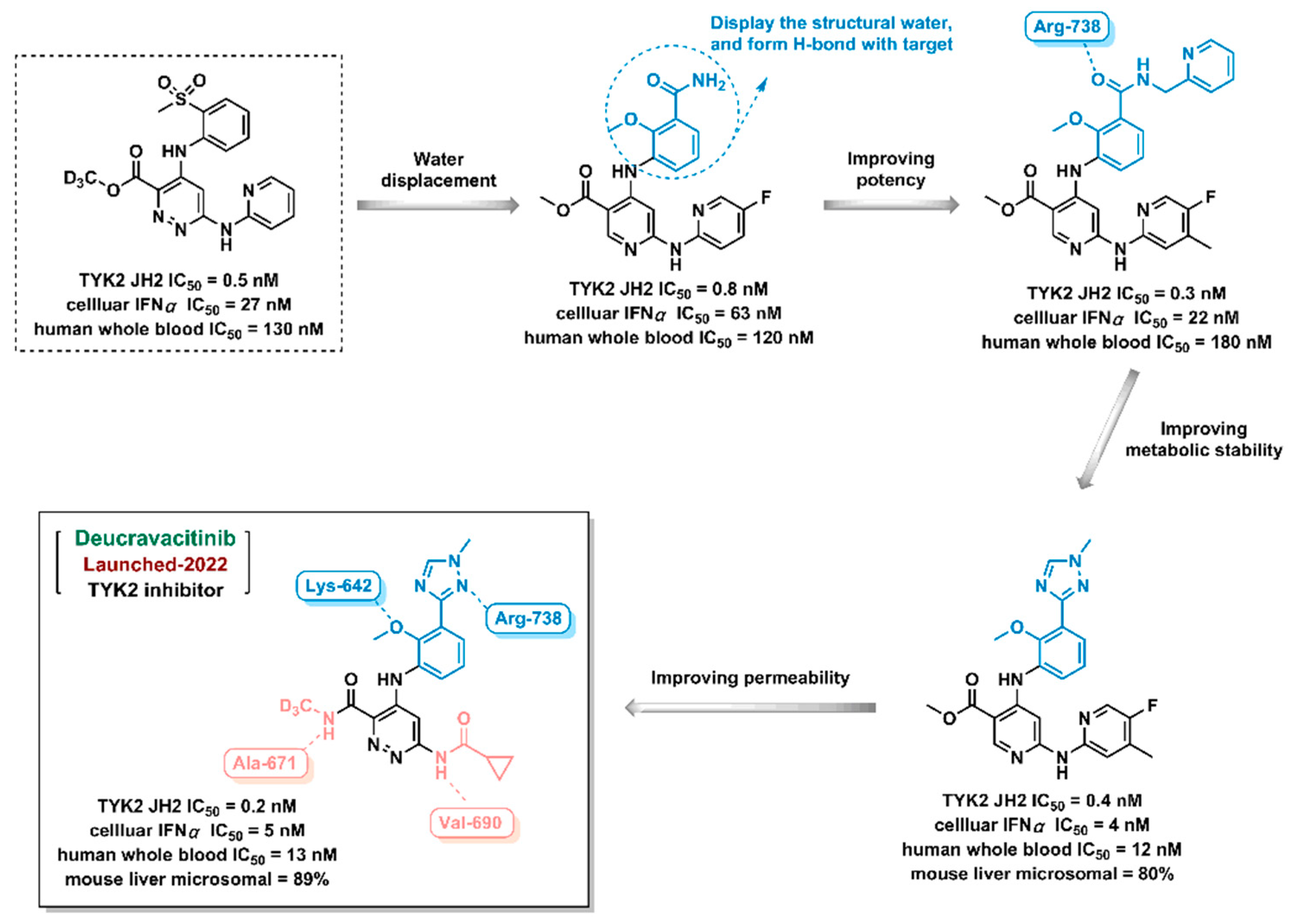
- Wrobleski, S.T.; Moslin, R.; Lin, S.; et al. Highly Selective Inhibition of Tyrosine Kinase 2 (TYK2) for the Treatment of Autoimmune Diseases: Discovery of the Allosteric Inhibitor BMS-986165. Journal of Medicinal Chemistry 2019, 62, 8973–8995. [Google Scholar] [CrossRef]
- Fell, J.B.; Fischer, J.P.; Baer, B.R.; et al. Identification of the Clinical Development Candidate MRTX849, a Covalent KRASG12C Inhibitor for the Treatment of Cancer. Journal of Medicinal Chemistry 2020, 63, 6679–6693. [Google Scholar] [CrossRef]
- Nivolas, G. Compounds useful as kinase inhibitors. WO201710 3611.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




