Submitted:
11 July 2023
Posted:
12 July 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
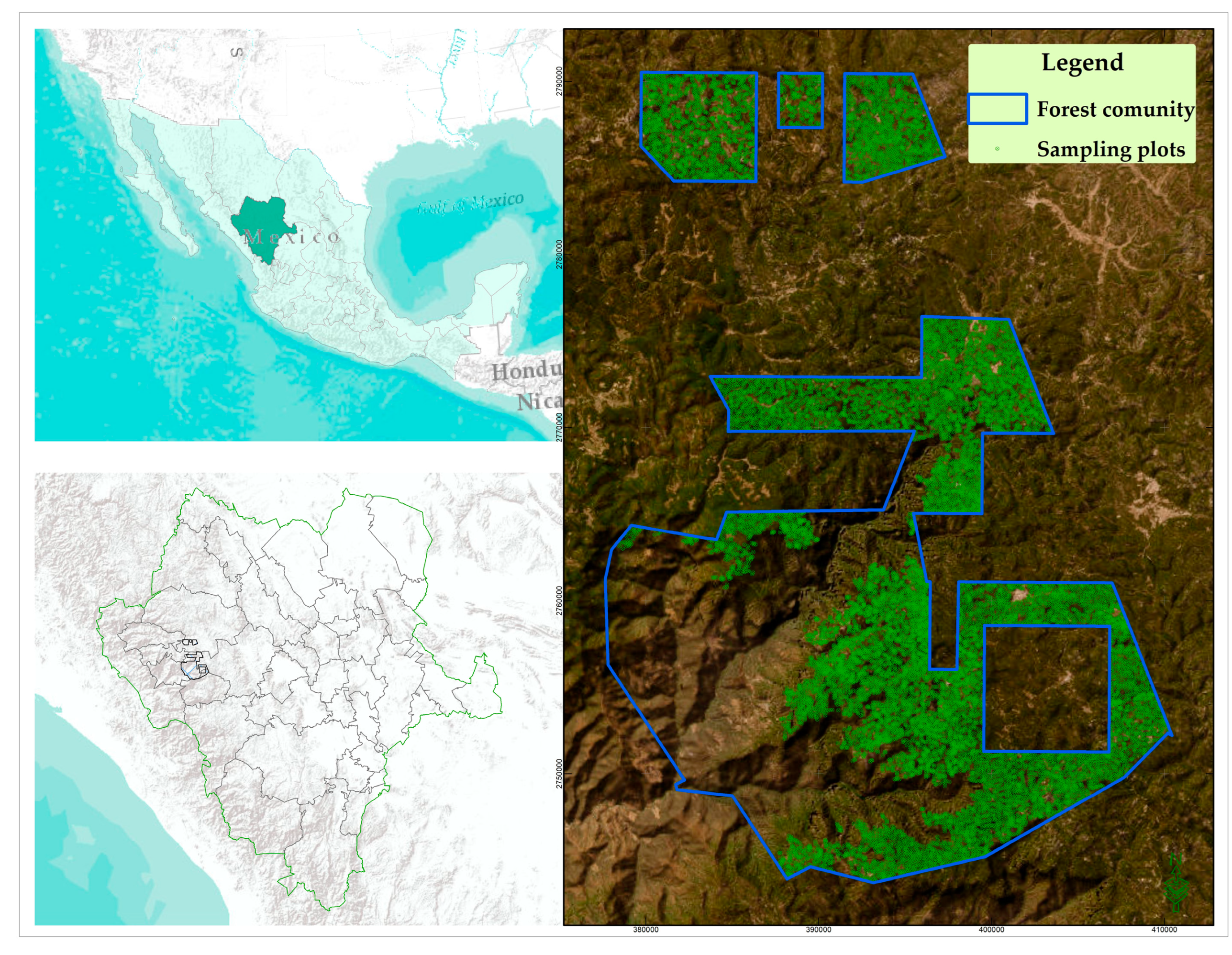
2.1. Study area
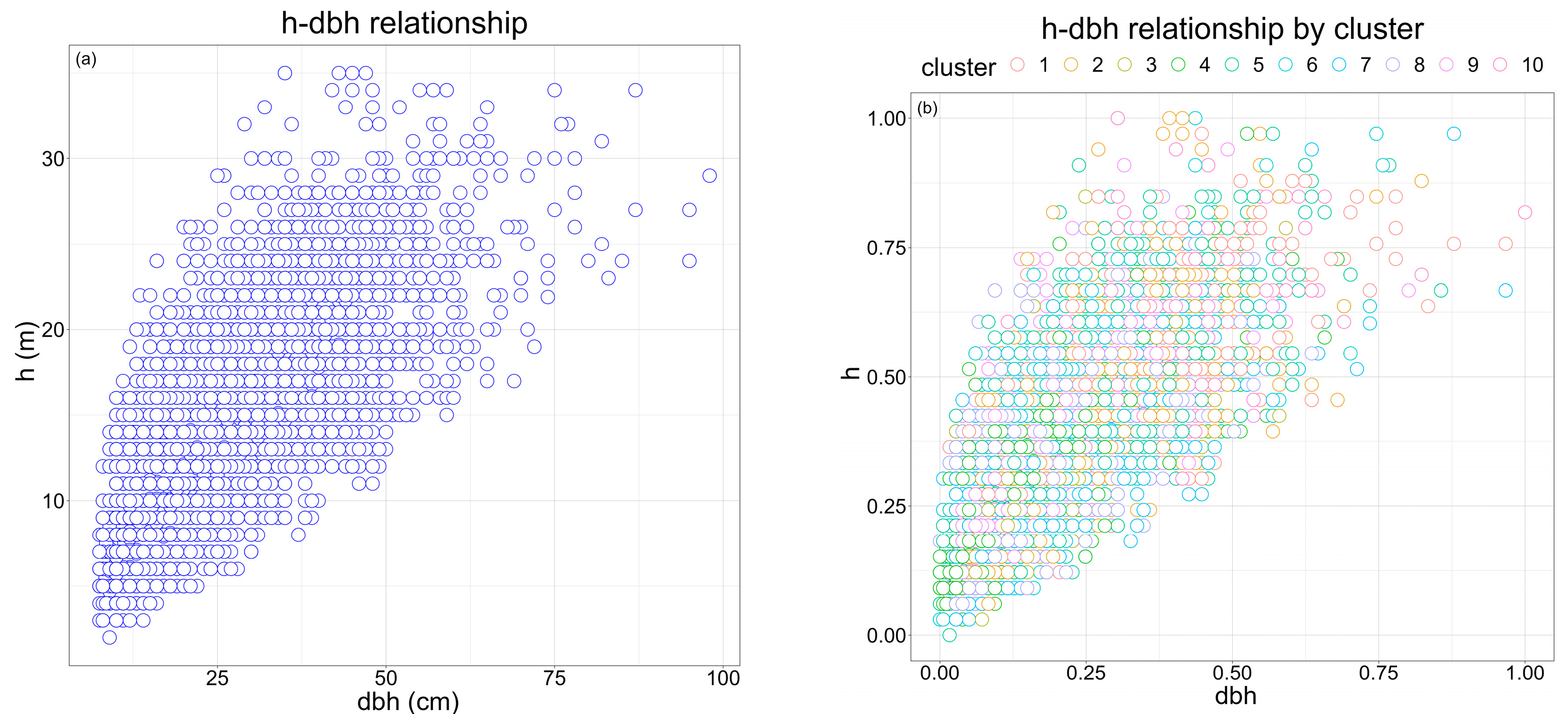
2.2. Dataset description
2.3. Nonlinear mixed effect modeling (NLMEM)
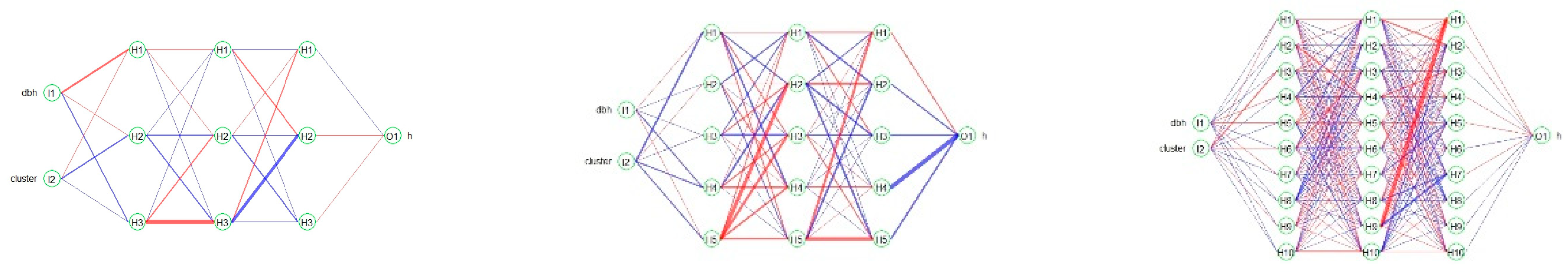
2.4. Artificial neural network (ANN)
2.5. Fitting modeling
2.6. Models performance criteria
3. Results
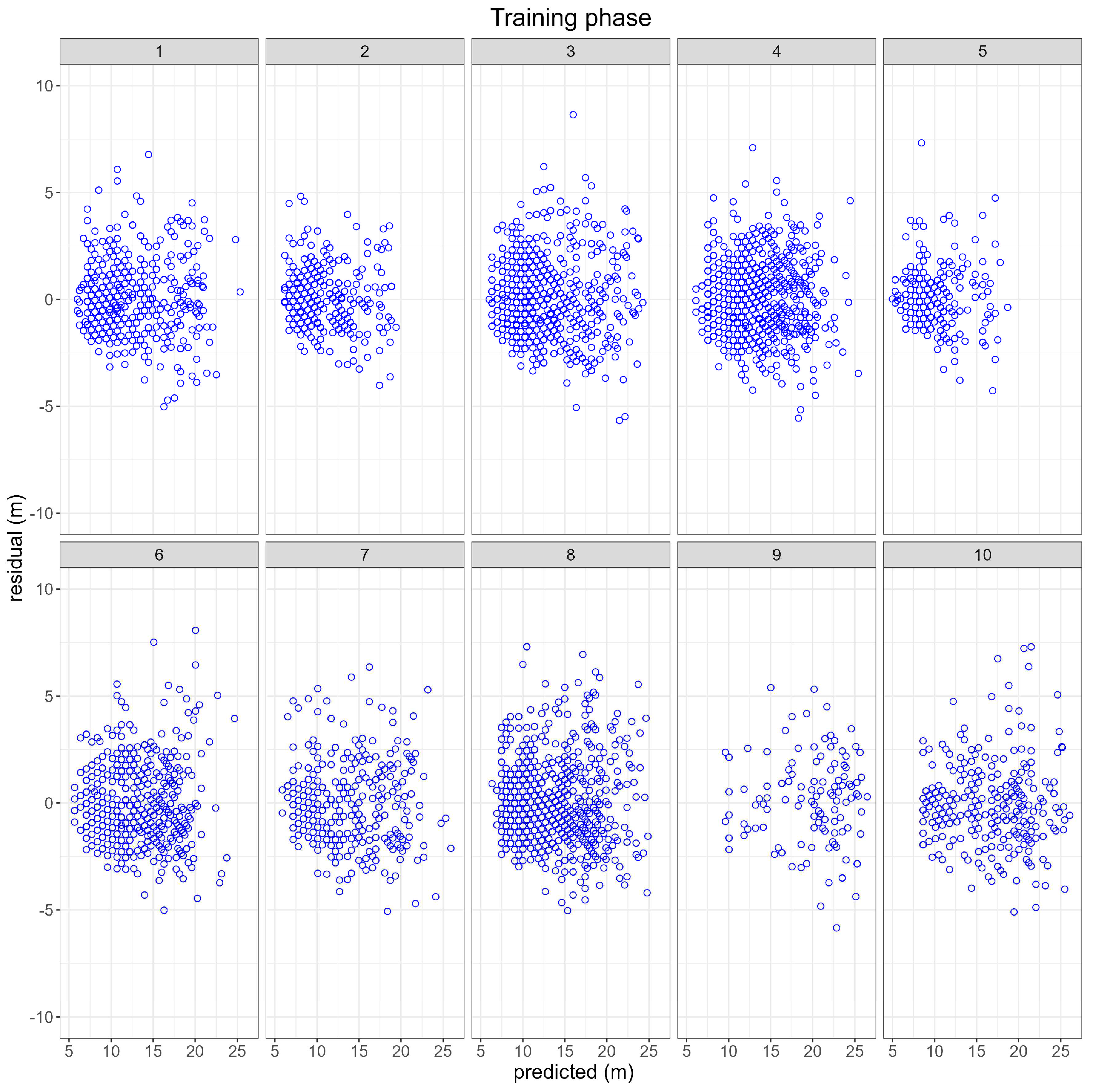
3.1. Training phase
3.1.1. NLMEM
3.1.1. RBPANN
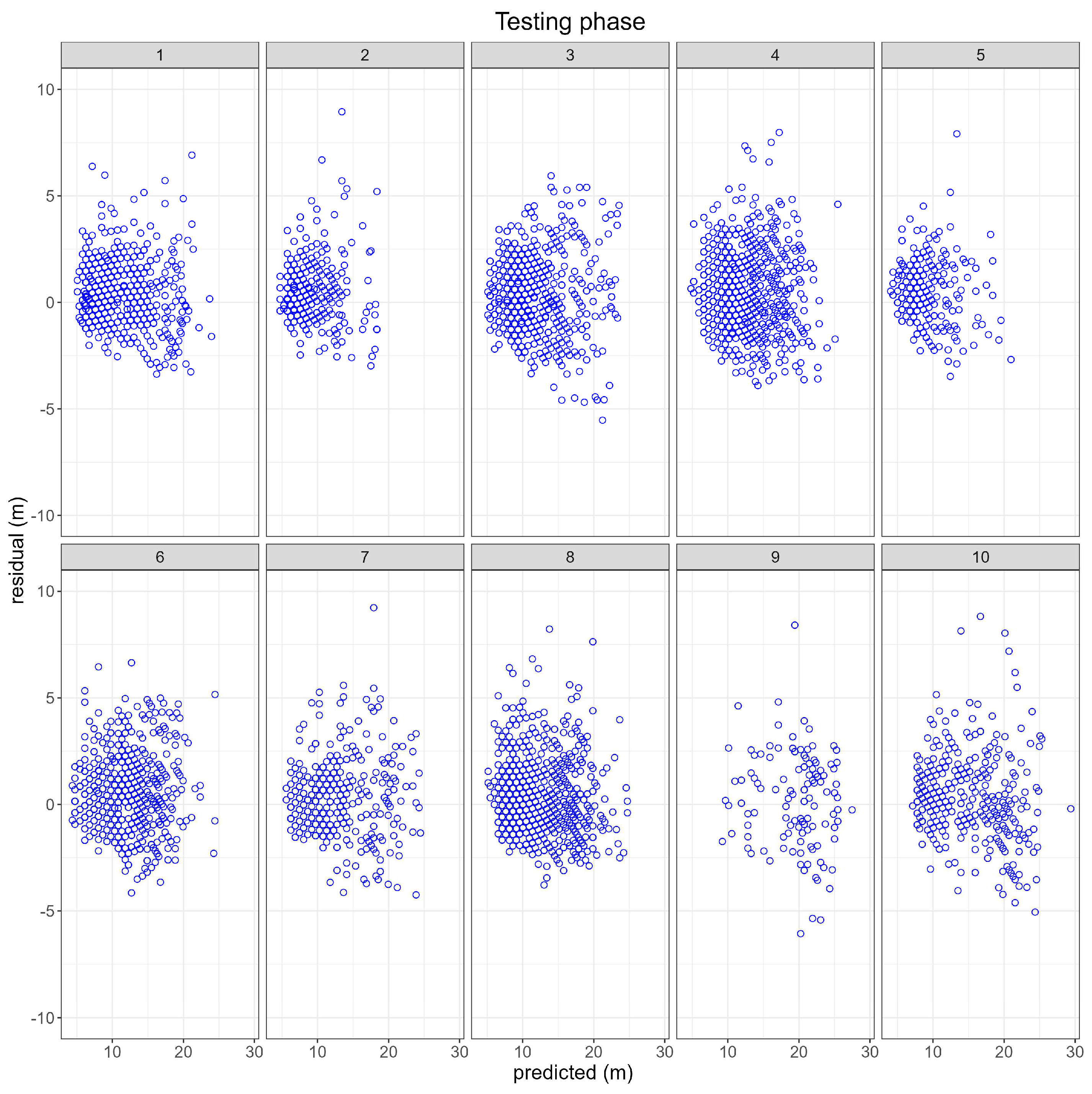
3.1. Testing or Validation phase
3.1.1. NLMEM
3.1.1. RBPANN
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wodecki, A. Artificial Intelligence Methods and Techniques. In Artificial Intelligence in Value Creation; Wodecki, A., Ed. Springer International Publishing: Cham, 2019. [CrossRef]
- Holzinger, A.; Keiblinger, K.; Holub, P.; Zatloukal, K.; Müller, H. AI for life: Trends in artificial intelligence for biotechnology. New Biotechnology 2023, 74, 16–24. [Google Scholar] [CrossRef]
- Riedmiller, M.; Braun, H. A direct adaptive method for faster backpropagation learning: The RPROP algorithm; 0780309995; IEEE, 1993; pp. 586–591. [Google Scholar]
- Riedmiller, M.; Braun, H. Rprop-description and implementation details; Technical Report. Univer, 1994. [Google Scholar]
- Hecht-Nielsen, R. Hecht-Nielsen, R. III.3 - Theory of the Backpropagation Neural Network**Based on “nonindent” by Robert Hecht-Nielsen, which appeared in Proceedings of the International Joint Conference on Neural Networks 1, 593–611, June 1989. © 1989 IEEE. In Neural Networks for Perception; Wechsler, H., Ed. Academic Press, 1992. [Google Scholar] [CrossRef]
- Chen, W.; Shirzadi, A.; Shahabi, H.; Ahmad, B.B.; Zhang, S.; Hong, H.; Zhang, N. A novel hybrid artificial intelligence approach based on the rotation forest ensemble and naïve Bayes tree classifiers for a landslide susceptibility assessment in Langao County, China. Geomatics, Natural Hazards and Risk 2017, 8, 1955–1977. [Google Scholar] [CrossRef]
- Özçelik, R.; Diamantopoulou, M.J.; Crecente-Campo, F.; Eler, U. Estimating Crimean juniper tree height using nonlinear regression and artificial neural network models. Forest Ecology and Management 2013, 306, 52–60. [Google Scholar] [CrossRef]
- Russell, S.J. Artificial intelligence a modern approach; Pearson Education, Inc., 2010. [Google Scholar]
- Gui, Y.; Li, D.; Fang, R. A fast adaptive algorithm for training deep neural networks. Applied Intelligence 2023, 53, 4099–4108. [Google Scholar] [CrossRef]
- Mehtätalo, L.; de-Miguel, S.; Gregoire, T.G. Modeling height-diameter curves for prediction. Canadian Journal of Forest Research 2015, 45, 826–837. [Google Scholar] [CrossRef]
- Sharma, M.; Parton, J. Height–diameter equations for boreal tree species in Ontario using a mixed-effects modeling approach. Forest Ecology and Management 2007, 249, 187–198. [Google Scholar] [CrossRef]
- Santiago-García, W.; Jacinto-Salinas, A.H.; Rodríguez-Ortiz, G.; Nava-Nava, A.; Santiago-García, E.; Ángeles-Pérez, G.; Enríquez-del Valle, J.R. Generalized height-diameter models for five pine species at Southern Mexico. Forest Science and Technology 2020, 16, 49–55. [Google Scholar] [CrossRef]
- Szurszewski, J.H. Mechanism of action of pentagastrin and acetylcholine on the longitudinal muscle of the canine antrum. J Physiol 1975, 252, 335–361. [Google Scholar] [CrossRef] [PubMed]
- Elliott, R.J.; Gardner, D.L. Proline determination with isatin, in the presence of amino acids. Anal Biochem 1976, 70, 268–273. [Google Scholar] [CrossRef] [PubMed]
- Ogana, F.N.; Ercanli, I. Modelling height-diameter relationships in complex tropical rain forest ecosystems using deep learning algorithm. Journal of Forestry Research 2021, 33, 883–898. [Google Scholar] [CrossRef]
- Raptis, D.I.; Kazana, V.; Kazaklis, A.; Stamatiou, C. Mixed-effects height–diameter models for black pine (Pinus nigra Arn.) forest management. Trees 2021, 35, 1167–1183. [Google Scholar] [CrossRef]
- Shen, J.; Hu, Z.; Sharma, R.P.; Wang, G.; Meng, X.; Wang, M.; Wang, Q.; Fu, L. Modeling height–diameter relationship for poplar plantations using combined-optimization multiple hidden layer back propagation neural network. Forests 2020, 11, 442. [Google Scholar] [CrossRef]
- Diamantopoulou, M.J. Predicting fir trees stem diameters using Artificial Neural Network models. The Southern African Forestry Journal 2005, 205, 39–44. [Google Scholar] [CrossRef]
- Barbosa, L.O.; Costa, E.A.; Schons, C.T.; Finger, C.A.G.; Liesenberg, V.; Bispo, P.d.C. Individual Tree Basal Area Increment Models for Brazilian Pine (Araucaria angustifolia) Using Artificial Neural Networks. Forests 2022, 13, 1108. [Google Scholar] [CrossRef]
- Qin, Y.; Wu, B.; Lei, X.; Feng, L. Prediction of tree crown width in natural mixed forests using deep learning algorithm. Forest Ecosystems 2023, 10, 100109. [Google Scholar] [CrossRef]
- Vahedi, A.A. Artificial neural network application in comparison with modeling allometric equations for predicting above-ground biomass in the Hyrcanian mixed-beech forests of Iran. Biomass and Bioenergy 2016, 88, 66–76. [Google Scholar] [CrossRef]
- Diamantopoulou, M.J.; Milios, E. Modelling total volume of dominant pine trees in reforestations via multivariate analysis and artificial neural network models. Biosystems Engineering 2010, 105, 306–315. [Google Scholar] [CrossRef]
- Wu, Z.; Wang, B.; Li, M.; Tian, Y.; Quan, Y.; Liu, J. Simulation of forest fire spread based on artificial intelligence. Ecological Indicators 2022, 136, 108653. [Google Scholar] [CrossRef]
- Richards, M.; McDonald, A.J.S.; Aitkenhead, M.J. Optimisation of competition indices using simulated annealing and artificial neural networks. Ecological Modelling 2008, 214, 375–384. [Google Scholar] [CrossRef]
- Öhman, K.; Lämås, T. Clustering of harvest activities in multi-objective long-term forest planning. Forest Ecology and Management 2003, 176, 161–171. [Google Scholar] [CrossRef]
- Akay, A.O.; AKGÜL, M.; Esin, A.I.; Demir, M.; ŞENTÜRK, N.; ÖZTÜRK, T. Evaluation of occupational accidents in forestry in Europe and Turkey by k-means clustering analysis. Turkish Journal of Agriculture and Forestry 2021, 45, 495–509. [Google Scholar] [CrossRef]
- Phillips, P.D.; Yasman, I.; Brash, T.E.; van Gardingen, P.R. Grouping tree species for analysis of forest data in Kalimantan (Indonesian Borneo). Forest Ecology and Management 2002, 157, 205–216. [Google Scholar] [CrossRef]
- Corral-Rivas, S.; Álvarez-González, J.; Crecente-Campo, F.; Corral-Rivas, J. Local and generalized height-diameter models with random parameters for mixed, uneven-aged forests in Northwestern Durango, Mexico. Forest Ecosystems 2014, 1, 6. [Google Scholar] [CrossRef]
- Castillo-Gallegos, E.; Jarillo-Rodríguez, J.; Escobar-Hernández, R. Diameter-height relationships in three species grown together in a commercial forest plantation in eastern tropical Mexico. Revista Chapingo serie ciencias forestales y del ambiente 2018, 24, 33–48. [Google Scholar] [CrossRef]
- Richards, F.J. A Flexible Growth Function for Empirical Use. Journal of Experimental Botany 1959, 10, 290–300. [Google Scholar] [CrossRef]
- Burkhart, H.E.; Tomé, M. Modeling forest trees and stands; Springer Science & Business Media, 2012. [Google Scholar]
- Von Gadow, K.; Nagel, J.; Saborowski, J. Continuous cover forestry: assessment, analysis, scenarios; Springer Science & Business Media, 2002; Volume 4. [Google Scholar]
- Quiñonez-Barraza, G.; Zhao, D.; De Los Santos Posadas, H.M.; Corral-Rivas, J.J. Considering neighborhood effects improves individual dbh growth models for natural mixed-species forests in Mexico. Annals of forest science 2018, 75, 1–11. [Google Scholar] [CrossRef]
- Climate Action Reserve [CAR]. Protocolo Forestal para México. Versión 3.0. CA, USA. Climate Action Reserve: 2022; Vol. Versión 3.0, p 2080 p.
- R Core Team. A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2012. https://www.r-project.org/ 2023.
- Ghoshal, S.; Nohria, N.; Wong, M. A K-means clustering algorithm. Algorithm AS 136. Applied Statistics 1979, 28, 100–108. [Google Scholar]
- Forgy, E.W. Cluster analysis of multivariate data: Efficiency vs. interpretability of classifications. biometrics 1965, 21, 768–769. [Google Scholar]
- Milligan, G.W.; Cooper, M.C. A study of standardization of variables in cluster analysis. Journal of Classification 1988, 5, 181–204. [Google Scholar] [CrossRef]
- Fang, Z.; Bailey, R.L. Height–diameter models for tropical forests on Hainan Island in southern China. Forest Ecology and Management 1998, 110, 315–327. [Google Scholar] [CrossRef]
- Pinheiro, J.C.; Bates, D.M.; Lindstrom, M.J. Model building for nonlinear mixed effects models; University of Wisconsin, Department of Biostatistics Madison, WI, 1995. [Google Scholar]
- Krogh, A. What are artificial neural networks? Nat Biotechnol 2008, 26, 195–197. [Google Scholar] [CrossRef] [PubMed]
- Anastasiadis, A.D.; Magoulas, G.D.; Vrahatis, M.N. New globally convergent training scheme based on the resilient propagation algorithm. Neurocomputing 2005, 64, 253–270. [Google Scholar] [CrossRef]
- Cartwright, H. Artificial neural networks: Methods in Molecular Biology, Third Edition. New Yor, NY USA ed.; Springer Human Press: New Yor, NY USA, 2021. [Google Scholar]
- Han, W.; Nan, L.; Su, M.; Chen, Y.; Li, R.; Zhang, X. Research on the prediction method of centrifugal pump performance based on a double hidden layer BP neural network. Energies 2019, 12, 2709. [Google Scholar] [CrossRef]
- Fausett, L.V. Fundamentals of neural networks: architectures, algorithms and applications; Pearson Education India, 2006. [Google Scholar]
- Kozak, A.; Smith, J.H.G. Standards for evaluating taper estimating systems. The Forestry Chronicle 1993, 69, 438–444. [Google Scholar] [CrossRef]
- Bicego, M. K-Random Forests: a K-means style algorithm for Random Forest clustering. Proceedings of 2019 International Joint Conference on Neural Networks (IJCNN), 14-19 July 2019; pp. 1–8. [Google Scholar]
- Ercanlı, İ. Innovative deep learning artificial intelligence applications for predicting relationships between individual tree height and diameter at breast height. Forest Ecosystems 2020, 7, 12. [Google Scholar] [CrossRef]
- Skudnik, M.; Jevšenak, J. Artificial neural networks as an alternative method to nonlinear mixed-effects models for tree height predictions. Forest Ecology and Management 2022, 507, 120017. [Google Scholar] [CrossRef]
- Thanh, T.N.; Tien, T.D.; Shen, H.L. Height-diameter relationship for Pinus koraiensis in Mengjiagang Forest Farm of Northeast China using nonlinear regressions and artificial neural network models. Journal of Forest Science 2019, 65, 134–143. [Google Scholar] [CrossRef]

| Statistic | |||||
|---|---|---|---|---|---|
| Variable | n | Minimum | Mean | Maximum | SD |
| N | 1000 | 1.0000 | 11.4720 | 57.0000 | 8.7717 |
| BA | 1000 | 0.0007 | 0.0193 | 0.0924 | 0.0137 |
| Dm | 1000 | 8.5000 | 22.9636 | 75.0000 | 7.6788 |
| Hm | 1000 | 4.0000 | 12.8062 | 35.0000 | 3.9813 |
| QMD | 1000 | 8.5147 | 24.7478 | 75.0000 | 8.0708 |
| A | 1000 | 2032.0000 | 2588.2170 | 2978.0000 | 137.3215 |
| S | 1000 | 0.0000 | 43.0499 | 96.0000 | 20.0551 |
| As | 1 | 5 | 9 | 2 | |
| Dataset | Statistic | |||||
|---|---|---|---|---|---|---|
| Variable | n | Minimum | Mean | Maximum | SD | |
| Training | h | 5736 | 7.5000 | 21.5362 | 95.0000 | 11.4394 |
| dbh | 5736 | 3.0000 | 12.3900 | 35.0000 | 5.3217 | |
| Testing | h | 5736 | 7.5000 | 21.3846 | 98.0000 | 11.5267 |
| dbh | 5736 | 2.0000 | 12.1742 | 35.0000 | 5.2871 | |
| Parameter | Estimate | SE | DF | t-value | p-value | lower | upper |
|---|---|---|---|---|---|---|---|
| 26.409060 | 1.100113 | 5724 | 24.005770 | <0.00001 | 24.252985 | 28.565134 | |
| 0.029786 | 0.002534 | 5724 | 11.754320 | <0.00001 | 0.024820 | 0.034752 | |
| 1.083133 | 0.040518 | 5724 | 26.732200 | <0.00001 | 1.003723 | 1.162543 | |
| 1.928939 | 0.583997 | 5724 | 3.302992 | 0.000962 | 1.210579 | 3.073574 | |
| 3.110839 | 0.029338 | 5724 | 106.033502 | <0.00001 | 3.054379 | 3.168342 | |
| -3.371745 | 0.106547 | 407 | -31.645570 | <0.00001 | -3.580578 | -3.162913 | |
| -2.840826 | 0.089770 | 320 | -31.645570 | <0.00001 | -3.016775 | -2.664877 | |
| -0.601879 | 0.019019 | 631 | -31.645570 | <0.00001 | -0.639157 | -0.564601 | |
| 3.572580 | 0.112894 | 133 | 31.645570 | <0.00001 | 3.351309 | 3.793851 | |
| 0.773802 | 0.024452 | 925 | 31.645570 | <0.00001 | 0.725876 | 0.821729 | |
| 0.478565 | 0.015123 | 1109 | 31.645570 | <0.00001 | 0.448925 | 0.508206 | |
| 0.945549 | 0.029879 | 364 | 31.645570 | <0.00001 | 0.886985 | 1.004113 | |
| -0.308794 | 0.009758 | 876 | -31.645570 | <0.00001 | -0.327919 | -0.289668 | |
| 0.012327 | 0.000390 | 654 | 31.645570 | <0.00001 | 0.011564 | 0.013091 | |
| 1.340420 | 0.042357 | 317 | 31.645570 | <0.00001 | 1.257400 | 1.423440 |
| Dataset | n | RMSE | SEE | RSEE | FI | E | RE | AIC | BIC | logLik |
| All-dataset | 5736 | 3.1085 | 3.1123 | 25.1193 | 0.6588 | -0.0005 | -0.0042 | 13039.75 | 13139.57 | -13009.75 |
| C1 | 631 | 2.4735 | 2.4889 | 6.4322 | 0.6182 | -0.0324 | -0.3161 | 8834.14 | 772.23 | -736.18 |
| C2 | 407 | 2.5289 | 2.5489 | 6.1485 | 0.5378 | 0.0050 | 0.0515 | 7113.33 | 627.39 | -592.78 |
| C3 | 925 | 2.9631 | 2.9749 | 8.4267 | 0.6525 | -0.1111 | -0.9526 | 16437.91 | 1408.51 | -1369.83 |
| C4 | 1109 | 3.8688 | 3.9442 | 2.9876 | 0.5610 | 0.3416 | 1.7378 | 4306.53 | 388.22 | -358.88 |
| C5 | 320 | 3.1341 | 3.1426 | 10.6971 | 0.6312 | -0.0072 | -0.0610 | 25348.12 | 2153.32 | -2112.34 |
| C6 | 654 | 2.8727 | 2.8792 | 10.3014 | 0.6134 | 0.0555 | 0.4528 | 28074.42 | 2381.60 | -2339.53 |
| C7 | 364 | 3.4345 | 3.4584 | 6.5911 | 0.6329 | -0.0640 | -0.4879 | 10767.05 | 932.64 | -897.25 |
| C8 | 876 | 3.3842 | 3.3939 | 10.2592 | 0.6015 | 0.0210 | 0.1632 | 25618.61 | 2175.54 | -2134.88 |
| C9 | 133 | 3.2098 | 3.2221 | 8.8088 | 0.6138 | -0.0229 | -0.1862 | 18292.63 | 1563.28 | -1524.39 |
| C10 | 317 | 3.6168 | 3.6458 | 5.1452 | 0.6196 | -0.0058 | -0.0353 | 9768.73 | 848.61 | -814.06 |
| ANN | Error | Reached Threshold | Steps | AIC | BIC |
| RBPANN-tanh | 27.8455 | 0.0775 | 301 | 577.69 | 2314.52 |
| RBPANN-softplus | 27.3939 | 0.0838 | 1885 | 576.79 | 2313.62 |
| RBPANN-logistic | 28.4113 | 0.0994 | 88 | 578.82 | 2315.65 |
| Dataset | RMSE | SEE | RSEE | FI | E | RE | AIC | BIC | logLik |
|---|---|---|---|---|---|---|---|---|---|
| RBPANN-tanh | |||||||||
| All-dataset | 2.8122 | 2.8134 | 22.7071 | 0.7208 | 0.0001 | -0.0003 | 11872.59 | 11912.52 | -11860.59 |
| C1 | 2.6322 | 2.6427 | 8.0184 | 0.7258 | 0.0002 | 0.0018 | 14644.89 | 1259.09 | -1220.41 |
| C2 | 2.2127 | 2.2264 | 6.1634 | 0.6945 | 0.0073 | 0.0717 | 7745.69 | 681.53 | -645.47 |
| C3 | 2.8170 | 2.8246 | 10.2988 | 0.7020 | -0.0087 | -0.0741 | 22979.70 | 1955.95 | -1914.98 |
| C4 | 2.5795 | 2.5853 | 9.9082 | 0.6883 | 0.0636 | 0.5191 | 25209.16 | 2142.83 | -2100.76 |
| C5 | 2.4178 | 2.4370 | 6.2966 | 0.5776 | -0.6268 | -6.4583 | 6768.31 | 598.64 | -564.03 |
| C6 | 2.8907 | 2.9019 | 8.4977 | 0.6867 | 0.2559 | 2.0805 | 16649.44 | 1426.35 | -1387.45 |
| C7 | 3.1340 | 3.1558 | 6.4423 | 0.6943 | 0.3677 | 2.8037 | 9967.09 | 865.97 | -830.59 |
| C8 | 3.0652 | 3.0740 | 9.9535 | 0.6731 | -0.3691 | -2.8636 | 23537.45 | 2002.11 | -1961.45 |
| C9 | 3.7470 | 3.8200 | 3.0995 | 0.5882 | 1.5107 | 7.6864 | 4204.43 | 379.71 | -350.37 |
| C10 | 3.2490 | 3.2751 | 4.9509 | 0.6930 | -0.1391 | -0.8426 | 8952.96 | 780.63 | -746.08 |
| RBPANN-softplus | |||||||||
| All-dataset | 2.8431 | 2.8443 | 22.9565 | 0.7143 | 0.7146 | -0.0013 | -0.0107 | 11997.88 | 12037.81 |
| C1 | 2.6516 | 2.6621 | 8.0772 | 0.7218 | 0.0489 | 0.4190 | 14755.60 | 1268.32 | -1229.63 |
| C2 | 2.4591 | 2.4744 | 6.8498 | 0.6226 | -0.9459 | -9.2408 | 8777.19 | 767.49 | -731.43 |
| C3 | 2.8529 | 2.8606 | 10.4301 | 0.6944 | 0.4200 | 3.5697 | 23260.85 | 1979.38 | -1938.40 |
| C4 | 2.6003 | 2.6062 | 9.9882 | 0.6833 | 0.3109 | 2.5351 | 25423.15 | 2160.66 | -2118.60 |
| C5 | 2.4956 | 2.5154 | 6.4993 | 0.5499 | -0.9140 | -9.4178 | 7011.62 | 618.91 | -584.30 |
| C6 | 2.8841 | 2.8952 | 8.4781 | 0.6882 | -0.1042 | -0.8472 | 16613.23 | 1423.33 | -1384.44 |
| C7 | 3.1030 | 3.1246 | 6.3787 | 0.7003 | 0.1339 | 1.0213 | 9880.37 | 858.75 | -823.36 |
| C8 | 3.0748 | 3.0837 | 9.9847 | 0.6711 | -0.3712 | -2.8799 | 23603.31 | 2007.59 | -1966.94 |
| C9 | 3.8187 | 3.8931 | 3.1588 | 0.5723 | 1.6461 | 8.3754 | 4264.92 | 384.75 | -355.41 |
| C10 | 3.2550 | 3.2811 | 4.9600 | 0.6919 | 0.0992 | 0.6009 | 8966.90 | 781.80 | -747.24 |
| RBPANN-logistic | |||||||||
| All-dataset | 2.8486 | 2.8498 | 23.0012 | 0.7135 | -0.0052 | -0.0420 | 12020.19 | 12060.12 | -12008.19 |
| C1 | 2.6570 | 2.6676 | 8.0938 | 0.7206 | 0.0785 | 0.6729 | 14786.65 | 1270.90 | -1232.22 |
| C2 | 2.4675 | 2.4828 | 6.8732 | 0.6201 | -0.9254 | -9.0403 | 8810.45 | 770.26 | -734.20 |
| C3 | 2.8586 | 2.8664 | 10.4510 | 0.6932 | 0.4146 | 3.5237 | 23305.34 | 1983.09 | -1942.11 |
| C4 | 2.6024 | 2.6083 | 9.9962 | 0.6827 | 0.2868 | 2.3386 | 25444.48 | 2162.44 | -2120.37 |
| C5 | 2.5183 | 2.5382 | 6.5583 | 0.5417 | -0.9468 | -9.7561 | 7081.01 | 624.69 | -590.08 |
| C6 | 2.9055 | 2.9167 | 8.5411 | 0.6835 | -0.1415 | -1.1503 | 16729.32 | 1433.01 | -1394.11 |
| C7 | 3.0957 | 3.1172 | 6.3636 | 0.7017 | 0.1066 | 0.8129 | 9859.77 | 857.03 | -821.65 |
| C8 | 3.0740 | 3.0828 | 9.9818 | 0.6712 | -0.3751 | -2.9095 | 23597.18 | 2007.08 | -1966.43 |
| C9 | 3.8192 | 3.8936 | 3.1592 | 0.5721 | 1.6756 | 8.5253 | 4265.34 | 384.79 | -355.45 |
| C10 | 3.2584 | 3.2845 | 4.9652 | 0.6912 | 0.1835 | 1.1115 | 8974.85 | 782.46 | -747.90 |
| Model | Dataset | RMSE | SEE | RSEE | FI | E | RE | AIC | BIC | logLik | Rank |
|---|---|---|---|---|---|---|---|---|---|---|---|
| NLMEM | Overall | 4 | 4 | 4 | 4 | 3 | 2 | 4 | 4 | 4 | 33 (4) |
| RBPANN-tanh | Overall | 1 | 1 | 1 | 1 | 4 | 4 | 2 | 1 | 2 | 17 (2) |
| RBPANN-softplus | Overall | 2 | 2 | 2 | 2 | 1 | 3 | 1 | 2 | 1 | 16 (1) |
| RBPANN-logistic | Overall | 3 | 3 | 3 | 3 | 2 | 1 | 3 | 3 | 3 | 24 (3) |
| NLMEM | C1 | 1 | 1 | 1 | 4 | 2 | 2 | 1 | 1 | 1 | 14 |
| RBPANN-tanh | C1 | 2 | 2 | 2 | 1 | 1 | 1 | 2 | 2 | 2 | 15 |
| RBPANN-softplus | C1 | 3 | 3 | 3 | 2 | 3 | 3 | 3 | 3 | 3 | 26 |
| RBPANN-logistic | C1 | 4 | 4 | 4 | 3 | 4 | 4 | 4 | 4 | 4 | 35 |
| NLMEM | C2 | 4 | 4 | 1 | 4 | 1 | 1 | 1 | 1 | 1 | 18 |
| RBPANN-tanh | C2 | 1 | 1 | 2 | 1 | 2 | 2 | 2 | 2 | 2 | 15 |
| RBPANN-softplus | C2 | 2 | 2 | 3 | 2 | 4 | 4 | 3 | 3 | 3 | 26 |
| RBPANN-logistic | C2 | 3 | 3 | 4 | 3 | 3 | 3 | 4 | 4 | 4 | 31 |
| NLMEM | C3 | 4 | 4 | 1 | 4 | 2 | 2 | 1 | 1 | 1 | 20 |
| RBPANN-tanh | C3 | 1 | 1 | 2 | 1 | 1 | 1 | 2 | 2 | 2 | 13 |
| RBPANN-softplus | C3 | 2 | 2 | 3 | 2 | 4 | 4 | 3 | 3 | 3 | 26 |
| RBPANN-logistic | C3 | 3 | 3 | 4 | 3 | 3 | 3 | 4 | 4 | 4 | 31 |
| NLMEM | C4 | 4 | 4 | 1 | 4 | 4 | 2 | 1 | 1 | 1 | 22 |
| RBPANN-tanh | C4 | 1 | 1 | 2 | 1 | 1 | 1 | 2 | 2 | 2 | 13 |
| RBPANN-softplus | C4 | 2 | 2 | 3 | 2 | 3 | 4 | 3 | 3 | 3 | 25 |
| RBPANN-logistic | C4 | 3 | 3 | 4 | 3 | 2 | 3 | 4 | 4 | 4 | 30 |
| NLMEM | C5 | 4 | 4 | 4 | 1 | 1 | 1 | 4 | 4 | 4 | 27 |
| RBPANN-tanh | C5 | 1 | 1 | 1 | 2 | 2 | 2 | 1 | 1 | 1 | 12 |
| RBPANN-softplus | C5 | 2 | 2 | 2 | 3 | 3 | 3 | 2 | 2 | 2 | 21 |
| RBPANN-logistic | C5 | 3 | 3 | 3 | 4 | 4 | 4 | 3 | 3 | 3 | 30 |
| NLMEM | C6 | 1 | 1 | 4 | 4 | 1 | 1 | 4 | 4 | 4 | 24 |
| RBPANN-tanh | C6 | 3 | 3 | 2 | 2 | 4 | 4 | 2 | 2 | 2 | 24 |
| RBPANN-softplus | C6 | 2 | 2 | 1 | 1 | 2 | 2 | 1 | 1 | 1 | 13 |
| RBPANN-logistic | C6 | 4 | 4 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 29 |
| NLMEM | C7 | 4 | 4 | 4 | 4 | 1 | 1 | 4 | 4 | 4 | 30 |
| RBPANN-tanh | C7 | 3 | 3 | 3 | 3 | 4 | 4 | 3 | 3 | 3 | 29 |
| RBPANN-softplus | C7 | 2 | 2 | 2 | 2 | 3 | 3 | 2 | 2 | 2 | 20 |
| RBPANN-logistic | C7 | 1 | 1 | 1 | 1 | 2 | 2 | 1 | 1 | 1 | 11 |
| NLMEM | C8 | 4 | 4 | 4 | 4 | 1 | 1 | 4 | 4 | 4 | 30 |
| RBPANN-tanh | C8 | 1 | 1 | 1 | 1 | 2 | 2 | 1 | 1 | 1 | 11 |
| RBPANN-softplus | C8 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 27 |
| RBPANN-logistic | C8 | 2 | 2 | 2 | 2 | 4 | 4 | 2 | 2 | 2 | 22 |
| NLMEM | C9 | 1 | 1 | 4 | 1 | 1 | 1 | 4 | 4 | 4 | 21 |
| RBPANN-tanh | C9 | 2 | 2 | 1 | 2 | 2 | 2 | 1 | 1 | 1 | 14 |
| RBPANN-softplus | C9 | 3 | 3 | 2 | 3 | 3 | 3 | 2 | 2 | 2 | 23 |
| RBPANN-logistic | C9 | 4 | 4 | 3 | 4 | 4 | 4 | 3 | 3 | 3 | 32 |
| NLMEM | C10 | 4 | 4 | 4 | 4 | 1 | 1 | 4 | 4 | 4 | 30 |
| RBPANN-tanh | C10 | 1 | 1 | 1 | 1 | 3 | 3 | 1 | 1 | 1 | 13 |
| RBPANN-softplus | C10 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 18 |
| RBPANN-logistic | C10 | 3 | 3 | 3 | 3 | 4 | 4 | 3 | 3 | 3 | 29 |
| Dataset | n | RMSE | SEE | RSEE | FI | E | RE | AIC | BIC | logLik |
|---|---|---|---|---|---|---|---|---|---|---|
| All-dataset | 5736 | 3.1438 | 3.1476 | 25.8549 | 0.6464 | -0.1611 | -1.3229 | 13169.29 | 13269.10 | -13139.29 |
| C1 | 631 | 2.7037 | 2.7141 | 24.1931 | 0.6893 | -0.0987 | -0.8795 | 15671.32 | 1344.87 | -1305.94 |
| C2 | 407 | 2.9045 | 2.9228 | 29.6973 | 0.4680 | 1.2223 | 12.4194 | 10249.76 | 890.11 | -854.15 |
| C3 | 925 | 3.0941 | 3.1029 | 27.5443 | 0.6136 | -0.7206 | -6.3964 | 23870.14 | 2029.86 | -1989.18 |
| C4 | 1109 | 3.0509 | 3.0578 | 25.0690 | 0.6102 | -0.0942 | -0.7725 | 29971.18 | 2539.72 | -2497.60 |
| C5 | 320 | 2.7563 | 2.7797 | 28.6992 | 0.5175 | 0.9194 | 9.4926 | 7263.59 | 639.50 | -605.30 |
| C6 | 654 | 3.1718 | 3.1834 | 25.4080 | 0.6085 | 0.1428 | 1.1398 | 19047.71 | 1626.51 | -1587.31 |
| C7 | 364 | 3.4512 | 3.4752 | 26.2781 | 0.6545 | -0.7996 | -6.0462 | 10839.28 | 938.67 | -903.27 |
| C8 | 876 | 3.0615 | 3.0704 | 24.6303 | 0.6290 | -0.1456 | -1.1680 | 23216.40 | 1975.28 | -1934.70 |
| C9 | 133 | 5.0061 | 5.1138 | 25.4219 | 0.2128 | -2.4332 | -12.0959 | 4665.31 | 417.55 | -388.78 |
| C10 | 317 | 3.8668 | 3.8957 | 24.6024 | 0.5555 | -0.7956 | -5.0245 | 10991.37 | 950.90 | -915.95 |
| Dataset | RMSE | SEE | RSEE | FI | E | RE | AIC | BIC | logLik |
|---|---|---|---|---|---|---|---|---|---|
| RBPANN-tanh | |||||||||
| All-dataset | 2.8693 | 2.8706 | 23.5793 | 0.7055 | 0.6603 | 5.4241 | 12103.39 | 12143.32 | -12091.39 |
| C1 | 2.5090 | 2.5186 | 8.1057 | 0.7324 | 0.6015 | 5.3617 | 14492.43 | 1246.63 | -1207.70 |
| C2 | 2.4793 | 2.4949 | 7.1292 | 0.6124 | 0.8309 | 8.4421 | 8726.27 | 763.15 | -727.19 |
| C3 | 2.7294 | 2.7372 | 10.1704 | 0.6994 | 0.4046 | 3.5917 | 21218.12 | 1808.86 | -1768.18 |
| C4 | 2.8842 | 2.8907 | 11.1931 | 0.6516 | 0.8969 | 7.3531 | 28460.68 | 2413.85 | -2371.72 |
| C5 | 2.3880 | 2.4083 | 6.0227 | 0.6378 | 0.0825 | 0.8517 | 6234.38 | 553.73 | -519.53 |
| C6 | 3.0888 | 3.1001 | 9.1436 | 0.6287 | 1.2080 | 9.6415 | 18609.70 | 1590.01 | -1550.81 |
| C7 | 3.1665 | 3.1885 | 6.4643 | 0.7091 | 0.8613 | 6.5128 | 10085.03 | 875.82 | -840.42 |
| C8 | 2.7710 | 2.7790 | 9.2458 | 0.6961 | 0.1519 | 1.2183 | 21146.63 | 1802.80 | -1762.22 |
| C9 | 4.2322 | 4.3232 | 3.2613 | 0.4374 | 1.8156 | 9.0259 | 4177.60 | 376.91 | -348.13 |
| C10 | 3.4732 | 3.4992 | 5.7063 | 0.6414 | 0.5225 | 3.2997 | 10118.01 | 878.12 | -843.17 |
| RBPANN-softplus | |||||||||
| All-dataset | 2.8764 | 2.8776 | 23.6371 | 0.7040 | 0.6578 | 5.4029 | 12131.48 | 12171.41 | -12119.48 |
| C1 | 2.5181 | 2.5278 | 8.1353 | 0.7305 | 0.6756 | 6.0219 | 14550.00 | 1251.43 | -1212.50 |
| C2 | 2.3388 | 2.3536 | 6.7253 | 0.6551 | -0.0814 | -0.8266 | 8164.99 | 716.38 | -680.42 |
| C3 | 2.8312 | 2.8393 | 10.5500 | 0.6765 | 0.8131 | 7.2176 | 21992.85 | 1873.42 | -1832.74 |
| C4 | 2.9657 | 2.9724 | 11.5095 | 0.6317 | 1.1148 | 9.1400 | 29209.97 | 2476.29 | -2434.16 |
| C5 | 2.4000 | 2.4204 | 6.0529 | 0.6342 | -0.2679 | -2.7660 | 6270.23 | 556.72 | -522.52 |
| C6 | 2.9772 | 2.9881 | 8.8135 | 0.6550 | 0.7866 | 6.2785 | 18002.41 | 1539.40 | -1500.20 |
| C7 | 3.1029 | 3.1244 | 6.3344 | 0.7207 | 0.6147 | 4.6485 | 9907.23 | 861.00 | -825.60 |
| C8 | 2.7748 | 2.7828 | 9.2584 | 0.6952 | 0.1450 | 1.1632 | 21174.89 | 1805.15 | -1764.57 |
| C9 | 4.3045 | 4.3971 | 3.3171 | 0.4180 | 2.0025 | 9.9547 | 4226.81 | 381.01 | -352.23 |
| C10 | 3.5266 | 3.5530 | 5.7940 | 0.6303 | 0.8555 | 5.4026 | 10242.05 | 888.46 | -853.50 |
| RBPANN-logistic | |||||||||
| All-dataset | 2.8820 | 2.8832 | 23.6832 | 0.7029 | 0.6484 | 5.3263 | 12153.85 | 12193.77 | -12141.85 |
| C1 | 2.5071 | 2.5168 | 8.0998 | 0.7328 | 0.6480 | 5.7760 | 14481.07 | 1245.68 | -1206.76 |
| C2 | 2.3476 | 2.3624 | 6.7505 | 0.6525 | -0.0800 | -0.8123 | 8200.94 | 719.38 | -683.41 |
| C3 | 2.8287 | 2.8368 | 10.5405 | 0.6771 | 0.8418 | 7.4726 | 21973.90 | 1871.84 | -1831.16 |
| C4 | 2.9718 | 2.9785 | 11.5331 | 0.6302 | 1.1385 | 9.3342 | 29264.97 | 2480.87 | -2438.75 |
| C5 | 2.3833 | 2.4035 | 6.0108 | 0.6392 | -0.2199 | -2.2702 | 6220.20 | 552.55 | -518.35 |
| C6 | 2.9674 | 2.9783 | 8.7844 | 0.6573 | 0.8301 | 6.6255 | 17947.97 | 1534.87 | -1495.66 |
| C7 | 3.1024 | 3.1240 | 6.3335 | 0.7208 | 0.6342 | 4.7956 | 9905.96 | 860.90 | -825.50 |
| C8 | 2.7763 | 2.7843 | 9.2634 | 0.6949 | 0.1438 | 1.1539 | 21186.18 | 1806.09 | -1765.51 |
| C9 | 4.2538 | 4.3453 | 3.2780 | 0.4316 | 1.9452 | 9.6698 | 4192.42 | 378.14 | -349.37 |
| C10 | 3.4913 | 3.5174 | 5.7360 | 0.6377 | 0.7841 | 4.9519 | 10160.28 | 881.65 | -846.69 |
| Model | Dataset | RMSE | SEE | RSEE | FI | E | RE | AIC | BIC | logLik | Rank |
|---|---|---|---|---|---|---|---|---|---|---|---|
| NLMEM | Overall | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 36 (4) |
| RBPANN-tanh | Overall | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 9 (1) |
| RBPANN-softplus | Overall | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 18 (2) |
| RBPANN-logistic | Overall | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 27 (3) |
| NLMEM | C1 | 4 | 4 | 4 | 4 | 1 | 1 | 4 | 4 | 4 | 30 |
| RBPANN-tanh | C1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 18 |
| RBPANN-softplus | C1 | 3 | 3 | 3 | 3 | 4 | 4 | 3 | 3 | 3 | 29 |
| RBPANN-logistic | C1 | 1 | 1 | 1 | 1 | 3 | 3 | 1 | 1 | 1 | 13 |
| NLMEM | C2 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 36 |
| RBPANN-tanh | C2 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 27 |
| RBPANN-softplus | C2 | 1 | 1 | 1 | 1 | 2 | 2 | 1 | 1 | 1 | 11 |
| RBPANN-logistic | C2 | 2 | 2 | 2 | 2 | 1 | 1 | 2 | 2 | 2 | 16 |
| NLMEM | C3 | 4 | 4 | 4 | 4 | 2 | 2 | 4 | 4 | 4 | 32 |
| RBPANN-tanh | C3 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 9 |
| RBPANN-softplus | C3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 27 |
| RBPANN-logistic | C3 | 2 | 2 | 2 | 2 | 4 | 4 | 2 | 2 | 2 | 22 |
| NLMEM | C4 | 4 | 4 | 4 | 4 | 1 | 1 | 4 | 4 | 4 | 30 |
| RBPANN-tanh | C4 | 1 | 1 | 1 | 1 | 2 | 2 | 1 | 1 | 1 | 11 |
| RBPANN-softplus | C4 | 2 | 2 | 2 | 2 | 3 | 3 | 2 | 2 | 2 | 20 |
| RBPANN-logistic | C4 | 3 | 3 | 3 | 3 | 4 | 4 | 3 | 3 | 3 | 29 |
| NLMEM | C5 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 36 |
| RBPANN-tanh | C5 | 2 | 2 | 2 | 2 | 1 | 1 | 2 | 2 | 2 | 16 |
| RBPANN-softplus | C5 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 27 |
| RBPANN-logistic | C5 | 1 | 1 | 1 | 1 | 2 | 2 | 1 | 1 | 1 | 11 |
| NLMEM | C6 | 4 | 4 | 4 | 4 | 1 | 1 | 4 | 4 | 4 | 30 |
| RBPANN-tanh | C6 | 3 | 3 | 3 | 3 | 4 | 4 | 3 | 3 | 3 | 29 |
| RBPANN-softplus | C6 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 18 |
| RBPANN-logistic | C6 | 1 | 1 | 1 | 1 | 3 | 3 | 1 | 1 | 1 | 13 |
| NLMEM | C7 | 4 | 4 | 4 | 4 | 3 | 3 | 4 | 4 | 4 | 34 |
| RBPANN-tanh | C7 | 3 | 3 | 3 | 3 | 4 | 4 | 3 | 3 | 3 | 29 |
| RBPANN-softplus | C7 | 2 | 2 | 2 | 2 | 1 | 1 | 2 | 2 | 2 | 16 |
| RBPANN-logistic | C7 | 1 | 1 | 1 | 1 | 2 | 2 | 1 | 1 | 1 | 11 |
| NLMEM | C8 | 4 | 4 | 4 | 4 | 3 | 3 | 4 | 4 | 4 | 34 |
| RBPANN-tanh | C8 | 1 | 1 | 1 | 1 | 4 | 4 | 1 | 1 | 1 | 15 |
| RBPANN-softplus | C8 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 18 |
| RBPANN-logistic | C8 | 3 | 3 | 3 | 3 | 1 | 1 | 3 | 3 | 3 | 23 |
| NLMEM | C9 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 36 |
| RBPANN-tanh | C9 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 9 |
| RBPANN-softplus | C9 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 27 |
| RBPANN-logistic | C9 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 18 |
| NLMEM | C10 | 4 | 4 | 4 | 4 | 3 | 3 | 4 | 4 | 4 | 34 |
| RBPANN-tanh | C10 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 9 |
| RBPANN-softplus | C10 | 3 | 3 | 3 | 3 | 4 | 4 | 3 | 3 | 3 | 29 |
| RBPANN-logistic | C10 | 3 | 3 | 3 | 3 | 4 | 4 | 3 | 3 | 3 | 29 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




