Submitted:
07 July 2023
Posted:
10 July 2023
You are already at the latest version
Abstract
Keywords:
Introduction
Structure of HTT and expression of mutant HTT
HTT interacting proteins
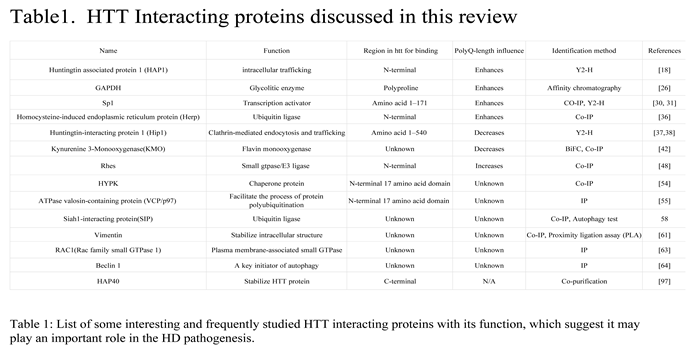
1.1. Proteins binding to HTT enhanced by Poly Q expansion
Huntingtin-associated protein 1 (HAP1)
Glyceraldehyde-3-Phosphate Dehydrogenase (GAPDH)
Transcription factor Sp1 (Sp1)
Homocysteine-induced endoplasmic reticulum protein (Herp)
2.2. Proteins binding to HTT inhibited by PolyQ expansion
Huntingtin-interacting protein 1 (Hip1)
Kynurenine 3-Monooxygenase (KMO)
Interacting protein that can enhance the mHTT toxicity
HTT interacting proteins that can alter mHtt aggregates
Proteins with impaired function when interacting with mHtt
New insights in HAP1
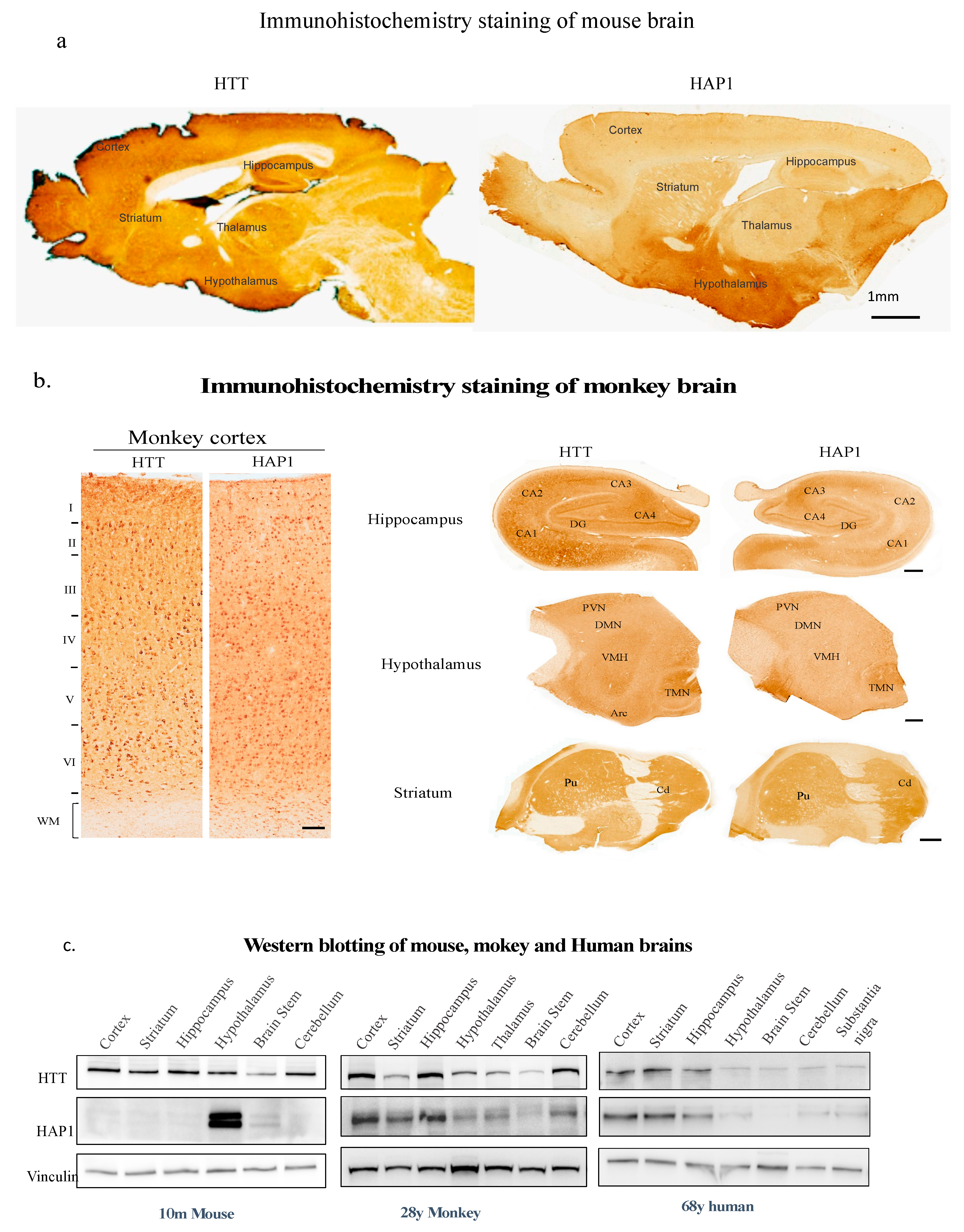
(1) HAP1 in the primate brains
(1) HAP1 is neuroprotective in monkey tissue
(1) Other functions of HAP1 in human
Conclusion and future remarks
References
- Tabrizi, S.J.; Flower, M.D.; Ross, C.A.; Wild, E.J. Huntington disease: new insights into molecular pathogenesis and therapeutic opportunities. Nat. Rev. Neurol. 2020, 16, 529–546. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, R.; Tabrizi, S.J. Clinical Features of Huntington’s Disease. In Polyglutamine Disorders. Advances in Experimental Medicine and Biology; Nóbrega, C., Pereira de Almeida, L., Eds.; Springer: Cham, Switzerland, 2018; Volume 1049, pp. 1–28. [Google Scholar] [CrossRef]
- Martino, E.; Chiarugi, S.; Margheriti, F.; Garau, G. Mapping, Structure and Modulation of PPI. Front. Chem. 2021, 9, 718405. [Google Scholar] [CrossRef] [PubMed]
- Wanker, E.E.; Ast, A.; Schindler, F.; Trepte, P.; Schnoegl, S. The pathobiology of perturbed mutant huntingtin protein–protein interactions in Huntington's disease. J. Neurochem. 2019, 151, 507–519. [Google Scholar] [CrossRef] [PubMed]
- Saudou, F.; Humbert, S. The Biology of Huntingtin. Neuron 2016, 89, 910–926. [Google Scholar] [CrossRef] [PubMed]
- Parsons, M.P.; Raymond, L.A. Huntington Disease. Neurobiol. Brain Disord. 2015, 303–320. [Google Scholar] [CrossRef]
- Ochaba, J.; Lukacsovich, T.; Csikos, G.; Zheng, S.; Margulis, J.; Salazar, L.; Mao, K.; Lau, A.L.; Yeung, S.Y.; Humbert, S.; et al. Potential function for the Huntingtin protein as a scaffold for selective autophagy. Proc. Natl. Acad. Sci. 2014, 111, 16889–16894. [Google Scholar] [CrossRef]
- Pandey, N.K.; Isas, J.M.; Rawat, A.; Lee, R.V.; Langen, J.; Pandey, P.; Langen, R. The 17-residue-long N terminus in huntingtin controls stepwise aggregation in solution and on membranes via different mechanisms. J. Biol. Chem. 2018, 293, 2597–2605. [Google Scholar] [CrossRef]
- Pigazzini, M.L.; Lawrenz, M.; Margineanu, A.; Schierle, G.S.K.; Kirstein, J. An Expanded Polyproline Domain Maintains Mutant Huntingtin Soluble in vivo and During Aging. Front. Mol. Neurosci. 2021, 14, 721749. [Google Scholar] [CrossRef]
- Gao, Y.-G.; Yan, X.-Z.; Song, A.-X.; Chang, Y.-G.; Gao, X.-C.; Jiang, N.; Zhang, Q.; Hu, H.-Y. Structural Insights into the Specific Binding of Huntingtin Proline-Rich Region with the SH3 and WW Domains. Structure 2006, 14, 1755–1765. [Google Scholar] [CrossRef]
- Arbez, N.; Ratovitski, T.; Roby, E.; Chighladze, E.; Stewart, J.C.; Ren, M.; Wang, X.; Lavery, D.J.; Ross, C.A. Post-translational modifications clustering within proteolytic domains decrease mutant huntingtin toxicity. J. Biol. Chem. 2017, 292, 19238–19249. [Google Scholar] [CrossRef]
- Neueder, A.; Landles, C.; Ghosh, R.; Howland, D.; Myers, R.H.; Faull, R.L.M.; Tabrizi, S.J.; Bates, G.P. The pathogenic exon 1 HTT protein is produced by incomplete splicing in Huntington’s disease patients. Sci. Rep. 2017, 7, 1307. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Yang, S.; Jing, L.; Huang, L.; Chen, L.; Zhao, X.; Yang, W.; Pan, Y.; Yin, P.; Qin, Z.S.; et al. Truncation of mutant huntingtin in knock-in mice demonstrates exon1 huntingtin is a key pathogenic form. Nat. Commun. 2020, 11, 2582. [Google Scholar] [CrossRef] [PubMed]
- Sathasivam, K.; Neueder, A.; Gipson, T.A.; Landles, C.; Benjamin, A.C.; Bondulich, M.K.; Smith, D.L.; Faull, R.L.M.; Roos, R.A.C.; Howland, D.; et al. Aberrant splicing of HTT generates the pathogenic exon 1 protein in Huntington disease. Proc. Natl. Acad. Sci. 2013, 110, 2366–2370. [Google Scholar] [CrossRef] [PubMed]
- Schulte, J.; Littleton, J.T. The biological function of the Huntingtin protein and its relevance to Huntington’s Disease pathology. Curr. Trends Neurol. 2011, 5, 65–78. [Google Scholar] [PubMed]
- Li, S.-H.; Li, X.-J. Huntington and its Role in Neuronal Degeneration. Neurosci. 2004, 10, 467–475. [Google Scholar] [CrossRef] [PubMed]
- Caron, N.S.; Banos, R.; Yanick, C.; Aly, A.E.; Byrne, L.M.; Smith, E.D.; Xie, Y.; Smith, S.E.; Potluri, N.; Black, H.F.; et al. Mutant Huntingtin Is Cleared from the Brain via Active Mechanisms in Huntington Disease. J. Neurosci. 2021, 41, 780–796. [Google Scholar] [CrossRef]
- Li, X.-J.; Li, S.-H.; Sharp, A.H.; Nucifora, F.C.; Schilling, G.; Lanahan, A.; Worley, P.; Snyder, S.H.; Ross, C.A. A huntingtin-associated protein enriched in brain with implications for pathology. Nature 1995, 378, 398–402. [Google Scholar] [CrossRef]
- Wu, L.L.-Y.; Zhou, X.-F. Huntingtin associated protein 1 and its functions. Cell Adhes. Migr. 2009, 3, 71–76. [Google Scholar] [CrossRef]
- Li, X.-J.; Li, S.-H. HAP1 and intracellular trafficking. Trends Pharmacol. Sci. 2005, 26, 1–3. [Google Scholar] [CrossRef]
- Liu, Q.; Cheng, S.; Yang, H.; Zhu, L.; Pan, Y.; Jing, L.; Tang, B.; Li, S.; Li, X.-J. Loss of Hap1 selectively promotes striatal degeneration in Huntington disease mice. Proc. Natl. Acad. Sci. 2020, 117, 20265–20273. [Google Scholar] [CrossRef]
- Zhao, X.; Chen, A.; Wang, Z.; Xu, X.-H.; Tao, Y. Biological functions and potential therapeutic applications of huntingtin-associated protein 1: progress and prospects. Clin. Transl. Oncol. 2022, 24, 203–214. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.K.; Kang, S.; Wang, X.; Rosales, J.L.; Gao, X.; Byun, H.-G.; Jin, Y.; Fu, S.; Wang, J.; Lee, K.-Y. HAP1 loss confers l-asparaginase resistance in ALL by downregulating the calpain-1-Bid-caspase-3/12 pathway. Blood 2019, 133, 2222–2232. [Google Scholar] [CrossRef] [PubMed]
- Tisdale, E.J. Glyceraldehyde-3-phosphate Dehydrogenase Is Phosphorylated by Protein Kinase Cι/λ and Plays a Role in Microtubule Dynamics in the Early Secretory Pathway. J. Biol. Chem. 2002, 277, 3334–3341. [Google Scholar] [CrossRef] [PubMed]
- Tristan, C.; Shahani, N.; Sedlak, T.W.; Sawa, A. The diverse functions of GAPDH: Views from different subcellular compartments. Cell. Signal. 2011, 23, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Burke, J.R.; Enghild, J.J.; Martin, M.E.; Jou, Y.-S.; Myers, R.M.; Roses, A.D.; Vance, J.M.; Strittmatter, W.J. Huntingtin and DRPLA proteins selectively interact with the enzyme GAPDH. Nat. Med. 1996, 2, 347–350. [Google Scholar] [CrossRef] [PubMed]
- Bae, B.-I.; Hara, M.R.; Cascio, M.B.; Wellington, C.L.; Hayden, M.R.; Ross, C.A.; Ha, H.C.; Li, X.-J.; Snyder, S.H.; Sawa, A. Mutant Huntingtin: Nuclear translocation and cytotoxicity mediated by GAPDH. Proc. Natl. Acad. Sci. 2006, 103, 3405–3409. [Google Scholar] [CrossRef]
- Hwang, S.; Disatnik, M.-H.; Mochly-Rosen, D. Impaired GAPDH -induced mitophagy contributes to the pathology of Huntington's disease. EMBO Mol. Med. 2015, 7, 1307–1326. [Google Scholar] [CrossRef]
- Vizcaíno, C.; Mansilla, S.; Portugal, J. Sp1 transcription factor: A long-standing target in cancer chemotherapy. Pharmacol. Ther. 2015, 152, 111–124. [Google Scholar] [CrossRef]
- Li, S.-H.; Cheng, A.L.; Zhou, H.; Lam, S.; Rao, M.; Li, H.; Li, X.-J. Interaction of Huntington Disease Protein with Transcriptional Activator Sp1. Mol. Cell. Biol. 2002, 22, 1277–1287. [Google Scholar] [CrossRef]
- Dunah, A.W.; Jeong, H.; Griffin, A.; Kim, Y.-M.; Standaert, D.G.; Hersch, S.M.; Mouradian, M.M.; Young, A.B.; Tanese, N.; Krainc, D. Sp1 and TAFII130 Transcriptional Activity Disrupted in Early Huntington's Disease. Science 2002, 296, 2238–2243. [Google Scholar] [CrossRef]
- Bradford, J.; Shin, J.-Y.; Roberts, M.; Wang, C.-E.; Li, X.-J.; Li, S. Expression of mutant huntingtin in mouse brain astrocytes causes age-dependent neurological symptoms. Proc. Natl. Acad. Sci. 2009, 106, 22480–22485. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Luo, Y.; Ly, P.T.; Cai, F.; Zhou, W.; Zou, H.; Song, W. Sp1 regulates human huntingtin gene expression. J Mol Neurosci. 2012, 47, 311–321. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Z.; Norflus, F.; Singh, B.; Swindell, M.K.; Buzescu, R.; Bejarano, M.; Chopra, R.; Zucker, B.; Benn, C.L.; DiRocco, D.P.; et al. Sp1 Is Up-regulated in Cellular and Transgenic Models of Huntington Disease, and Its Reduction Is Neuroprotective. J. Biol. Chem. 2006, 281, 16672–16680. [Google Scholar] [CrossRef]
- Kokame, K.; Agarwala, K.L.; Kato, H.; Miyata, T. Herp, a New Ubiquitin-like Membrane Protein Induced by Endoplasmic Reticulum Stress. J. Biol. Chem. 2000, 275, 32846–32853. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.; Cao, L.; Liang, X.; Du, A.; Peng, T.; Li, H. Herp Promotes Degradation of Mutant Huntingtin: Involvement of the Proteasome and Molecular Chaperones. Mol. Neurobiol. 2018, 55, 7652–7668. [Google Scholar] [CrossRef]
- Kalchman, M.A.; Koide, H.B.; McCutcheon, K.; Graham, R.K.; Nichol, K.; Nishiyama, K.; Kazemi-Esfarjani, P.; Lynn, F.C.; Wellington, C.; Metzler, M.; et al. HIP1, a human homologue of S. cerevisiae Sla2p, interacts with membrane-associated huntingtin in the brain. Nat. Genet. 1997, 16, 44–53. [Google Scholar] [CrossRef]
- Wanker, E.E.; Rovira, C.; Scherzinger, E.; Hasenbank, R.; Wälter, S.; Tait, D.; Colicelli, J.; Lehrach, H. HIP-I: A huntingtin interacting protein isolated by the yeast two-hybrid system. Hum. Mol. Genet. 1997, 6, 487–495. [Google Scholar] [CrossRef]
- Waelter, S.; Scherzinger, E.; Hasenbank, R.; Nordhoff, E.; Lurz, R.; Goehler, H.; Gauss, C.; Sathasivam, K.; Bates, G.P.; Lehrach, H.; et al. The huntingtin interacting protein HIP1 is a clathrin and alpha-adaptin-binding protein involved in receptor-mediated endocytosis. Hum. Mol. Genet. 2001, 10, 1807–1817. [Google Scholar] [CrossRef]
- Laragione, T.; Brenner, M.; Lahiri, A.; Gao, E.; Harris, C.; Gulko, P.S. Huntingtin-interacting protein 1 (HIP1) regulates arthritis severity and synovial fibroblast invasiveness by altering PDGFR and Rac1 signalling. Ann. Rheum. Dis. 2018, 77, 1627–1635. [Google Scholar] [CrossRef]
- Nicita, F.; Garone, G.; Spalice, A.; Savasta, S.; Striano, P.; Pantaleoni, C.; Spartà, M.V.; Kluger, G.; Capovilla, G.; Pruna, D.; et al. Epilepsy is a possible feature in Williams-Beuren syndrome patients harboring typical deletions of the 7q11.23 critical region. Am. J. Med Genet. Part A 2016, 170, 148–155. [Google Scholar] [CrossRef]
- Swaih, A.M.; Breda, C.; Sathyasaikumar, K.V.; Allcock, N.; Collier, M.E.W.; Mason, R.P.; Feasby, A.; Herrera, F.; Outeiro, T.F.; Schwarcz, R.; et al. Kynurenine 3-Monooxygenase Interacts with Huntingtin at the Outer Mitochondrial Membrane. Biomedicines 2022, 10, 2294. [Google Scholar] [CrossRef] [PubMed]
- Sathyasaikumar, K.V.; de la Cruz, V.P.; Pineda, B.; Cervantes, G.I.V.; Ortega, D.R.; Donley, D.W.; Severson, P.L.; West, B.L.; Giorgini, F.; Fox, J.H.; et al. Cellular Localization of Kynurenine 3-Monooxygenase in the Brain: Challenging the Dogma. Antioxidants 2022, 11, 315. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Shao, M.; Wu, T. Kynurenine-3-monooxygenase: A new direction for the treatment in different diseases. Food Sci. Nutr. 2020, 8, 711–719. [Google Scholar] [CrossRef] [PubMed]
- Zwilling, D.; Huang, S.-Y.; Sathyasaikumar, K.V.; Notarangelo, F.M.; Guidetti, P.; Wu, H.-Q.; Lee, J.; Truong, J.; Andrews-Zwilling, Y.; Hsieh, E.W.; et al. Kynurenine 3-Monooxygenase Inhibition in Blood Ameliorates Neurodegeneration. Cell 2011, 145, 863–874. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Zhang, A.; Binnie, M.; McGuire, K.; Webster, S.P.; Hughes, J.; Howie, S.E.M.; Mole, D.J. Kynurenine 3-monooxygenase is a critical regulator of renal ischemia–reperfusion injury. Exp. Mol. Med. 2019, 51, 1–14. [Google Scholar] [CrossRef]
- Park, S.-Y.; Nam, J.-S. Kynurenine pathway enzyme KMO in cancer progression: A tip of the Iceberg. EBioMedicine 2020, 55, 102762. [Google Scholar] [CrossRef]
- Subramaniam, S.; Sixt, K.M.; Barrow, R.; Snyder, S.H. Rhes, a Striatal Specific Protein, Mediates Mutant-Huntingtin Cytotoxicity. Science 2009, 324, 1327–1330. [Google Scholar] [CrossRef]
- Mealer, R.G.; Subramaniam, S.; Snyder, S.H. Rhes Deletion Is Neuroprotective in the 3-Nitropropionic Acid Model of Huntington's Disease. J. Neurosci. 2013, 33, 4206–4210. [Google Scholar] [CrossRef]
- Ramirrez-Jarquin, U.N.; Manish Sharmaa, M.; Zhou, W.; Shahani, N.; Subramaniam, S. Deletion of SUMO1 attenuates behavioral andanatomical deficits by regulating autophagic activities in Huntington disease. Proc. Natl. Acad. Sci. USA 2022, 119, e2107187119. [Google Scholar] [CrossRef]
- Mealer, R.G.; Murray, A.J.; Shahani, N.; Subramaniam, S.; Snyder, S.H. Rhes, a Striatal-selective Protein Implicated in Huntington Disease, Binds Beclin-1 and Activates Autophagy. J. Biol. Chem. 2014, 289, 3547–3554. [Google Scholar] [CrossRef]
- DiGiovanni, L.F.; Mocle, A.J.; Xia, J.; Truant, R. Huntingtin N17 domain is a reactive oxygen species sensor regulating huntingtin phosphorylation and localization. Hum. Mol. Genet. 2016, 25, 3937–3945. [Google Scholar] [CrossRef] [PubMed]
- Raychaudhuri, S.; Sinha, M.; Mukhopadhyay, D.; Bhattacharyya, N.P. HYPK, a Huntingtin interacting protein, reduces aggregates and apoptosis induced by N-terminal Huntingtin with 40 glutamines in Neuro2a cells and exhibits chaperone-like activity. Hum. Mol. Genet. 2008, 17, 240–255. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, K.R.; Bhattacharyya, N.P. Chaperone protein HYPK interacts with the first 17 amino acid region of Huntingtin and modulates mutant HTT-mediated aggregation and cytotoxicity. Biochem. Biophys. Res. Commun 2015, 456, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, D.K.; Roy, A.; Ranjan, A. The ATP ase VCP /p97 functions as a disaggregase against toxic Huntingtin-exon1 aggregates. FEBS Lett. 2018, 592, 2680–2692. [Google Scholar] [CrossRef] [PubMed]
- Topolska-Woś, A.M.; Chazin, W.J.; Filipek, A. CacyBP/SIP—Structure and variety of functions. Biochimica et Biophysica Acta (BBA)-General Subjects 2016, 1860, 79–85. [Google Scholar] [CrossRef]
- Santelli, E.; Leone, M.; Li, C.; Fukushima, T.; Preece, N.E.; Olson, A.J.; Ely, K.R.; Reed, J.C.; Pellecchia, M.; Liddington, R.C.; et al. Structural Analysis of Siah1-Siah-interacting Protein Interactions and Insights into the Assembly of an E3 Ligase Multiprotein Complex. J. Biol. Chem. 2005, 280, 34278–34287. [Google Scholar] [CrossRef]
- Latoszek, E.; Wiweger, M.; Ludwiczak, J.; Dunin-Horkawicz, S.; Kuznicki, J.; Czeredys, M. Siah-1-interacting protein regulates mutated huntingtin protein aggregation in Huntington’s disease models. Cell Biosci. 2022, 12. [Google Scholar] [CrossRef]
- Xu, H.; Bensalel, J.; Raju, S.; Capobianco, E.; Lu, M.L.; Wei, J. Characterization of huntingtin interactomes and their dynamic responses in living cells by proximity proteomics. J. Neurochem. 2023, 164, 512–528. [Google Scholar] [CrossRef]
- Morrow, C.S.; Porter, T.J.; Xu, N.; Arndt, Z.P.; Ako-Asare, K.; Heo, H.J.; Thompson, E.A.; Moore, D.L. Vimentin Coordinates Protein Turnover at the Aggresome during Neural Stem Cell Quiescence Exit. Cell Stem Cell 2020, 26, 558–568. [Google Scholar] [CrossRef]
- Bauer, P.; Hudec, R.; Goswami, A.; Kurosawa, M.; Matsumoto, G.; Mikoshiba, K.; Nukina, N. ROCK-phosphorylated vimentin modifies mutant huntingtin aggregation via sequestration of IRBIT. Mol. Neurodegener. 2012, 7, 43–43. [Google Scholar] [CrossRef]
- Chi, X.; Wang, S.; Huang, Y.; Stamnes, M.; Chen, J.-L. Roles of Rho GTPases in Intracellular Transport and Cellular Transformation. Int. J. Mol. Sci. 2013, 14, 7089–7108. [Google Scholar] [CrossRef] [PubMed]
- Tousley, A.; Iuliano, M.; Weisman, E.; Sapp, E.; Zhang, N.; Vodicka, P.; Alexander, J.; Aviolat, H.; Gatune, L.; Reeves, P.; et al. Rac1 Activity Is Modulated by Huntingtin and Dysregulated in Models of Huntington’s Disease. J. Huntington's Dis. 2019, 8, 53–69. [Google Scholar] [CrossRef] [PubMed]
- Ashkenazi, A.; Bento, C.F.; Ricketts, T.; Vicinanza, M.; Siddiqi, F.; Pavel, M.; Squitieri, F.; Hardenberg, M.C.; Imarisio, S.; Menzies, F.M.; et al. Polyglutamine tracts regulate beclin 1-dependent autophagy. Nature 2017, 545, 108–111. [Google Scholar] [CrossRef] [PubMed]
- Shibata, M.; Lu, T.; Furuya, T.; Degterev, A.; Mizushima, N.; Yoshimori, T.; MacDonald, M.; Yankner, B.; Yuan, J. Regulation of Intracellular Accumulation of Mutant Huntingtin by Beclin 1. Perspect. Surg. 2006, 281, 14474–14485. [Google Scholar] [CrossRef]
- Pircs, K.; Drouin-Ouellet, J.; Horváth, V.; Gil, J.; Rezeli, M.; Garza, R.; A Grassi, D.; Sharma, Y.; St-Amour, I.; Harris, K.; et al. Distinct subcellular autophagy impairments in induced neurons from patients with Huntington's disease. Brain 2022, 145, 3035–3057. [Google Scholar] [CrossRef]
- Croce, K.R.; Yamamoto, A. A role for autophagy in Huntington's disease. Neurobio. Dis. 2019, 122, 16–22. [Google Scholar] [CrossRef]
- Valionyte, E.; Yang, Y.; Roberts, S.L.; Kelly, J.; Lu, B.; Luo, S. Lowering Mutant Huntingtin Levels and Toxicity: Autophagy-Endolysosome Pathways in Huntington's Disease. J. Mol. Bio. 2020, 432, 2673–2691. [Google Scholar] [CrossRef]
- Yang, W.; Chen, X.; Li, S.; Li, X.-J. Genetically modified large animal models for investigating neurodegenerative diseases. Cell Biosci. 2021, 11. [Google Scholar] [CrossRef]
- Yin, P.; Li, S.; Li, X.-J.; Yang, W. New pathogenic insights from large animal models of neurodegenerative diseases. Protein Cell 2022, 13, 707–720. [Google Scholar] [CrossRef]
- Li, C.; Li, J.; Lai, L.; Li, S.; Yan, S. Genetically engineered pig models of neurological diseases. Ageing Neurodegener. Dis. 2022, 2, 13. [Google Scholar] [CrossRef]
- Chen, X.; Sun, Y.; Chen, L.; Chen, X.-S.; Pan, M.; Zhang, Y.; Wang, Q.; Yang, W.; Yin, P.; He, D.; et al. Differential expression and roles of Huntingtin and Huntingtin-associated protein 1 in the mouse and primate brains. Cell. Mol. Life Sci. 2022, 79, 554. [Google Scholar] [CrossRef] [PubMed]
- Li, S.-H.; Hosseini, S.H.; Gutekunst, C.-A.; Hersch, S.M.; Ferrante, R.J.; Li, X.-J. A Human HAP1 Homologue. J. Biol. Chem. 1998, 273, 19220–19227. [Google Scholar] [CrossRef] [PubMed]
- Bertaux, F.; Sharp, A.H.; A Ross, C.; Lehrach, H.; Bates, G.P.; Wanker, E. HAP1-huntingtin interactions do not contribute to the molecular pathology in Huntington's disease transgenic mice. FEBS Lett. 1998, 426, 229–232. [Google Scholar] [CrossRef] [PubMed]
- A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. The Huntington's Disease Collaborative Research Group. Cell 1993, 72, 971–983. [CrossRef] [PubMed]
- Li, S.H.; Schilling, G.; Young, W.S.; Li, X.; Margolis, R.L.; Stine, O.C.; Wagster, M.V.; Abbott, M.H.; Franz, M.L.; Ranen, N.G.; et al. Huntington's disease gene (IT15) is widely expressed in human and rat tissues. Neuron 1993, 11, 985–993. [Google Scholar] [CrossRef] [PubMed]
- Strong, T.V.; Tagle, D.A.; Valdes, J.M.; Elmer, L.W.; Boehm, K.; Swaroop, M.; Kaatz, K.W.; Collins, F.S.; Albin, R.L. Widespread expression of the human and rat Huntington's disease gene in brain and nonneural tissues. Nat. Genet. 1993, 5, 259–265. [Google Scholar] [CrossRef] [PubMed]
- Sharp, A.H.; Loev, S.J.; Schilling, G.; Li, S.-H.; Li, X.-J.; Bao, J.; Wagster, M.V.; Kotzuk, J.A.; Steiner, J.P.; Lo, A.; et al. Widespread expression of Huntington's disease gene (IT15) protein product. Neuron 1995, 14, 1065–1074. [Google Scholar] [CrossRef]
- Landwehrmeyer, G.B.; McNeil, S.M.; Dure, L.S.; Ge, P.; Aizawa, H.; Huang, Q.; Ambrose, C.M.; Duyao, M.P.; Bird, E.D.; Bonilla, E.; et al. Huntington's disease gene: Regional and cellular expression in brain of normal and affected individuals. Ann. Neurol. 1995, 37, 218–230. [Google Scholar] [CrossRef]
- Harjes, P.; Wanker, E. The hunt for huntingtin function: interaction partners tell many different stories. Trends Biochem. 2003, 28, 425–433. [Google Scholar] [CrossRef]
- Kaltenbach, L.S.; Romero, E.; Becklin, R.R.; Chettier, R.; Bell, R.; Phansalkar, A.; Strand, A.; Torcassi, C.; Savage, J.; Hurlburt, A.; et al. Huntingtin Interacting Proteins Are Genetic Modifiers of Neurodegeneration. PLOS Genet. 2007, 3, e82. [Google Scholar] [CrossRef]
- Miller, J and Hughes, R.E. Protein Interactions and Target Discovery in Huntington’s Disease. Neurobiology of Huntington Disease (Applications to Drug Discovery) Frontiers in Neuroscience 2011, ISBN: 978-0-8493-9000.
- Li, X.-J.; Li, S. Influence of Species Differences on the Neuropathology of Transgenic Huntington's Disease Animal Models. J. Genet. Genom. 2012, 39, 239–245. [Google Scholar] [CrossRef] [PubMed]
- Eaton, S.L.; Wishart, T.M. Bridging the gap: large animal models in neurodegenerative research. Mamm. Genome 2017, 28, 324–337. [Google Scholar] [CrossRef] [PubMed]
- Sheng, G.; Xu, X.; Lin, Y.-F.; Wang, C.-E.; Rong, J.; Cheng, D.; Peng, J.; Jiang, X.; Li, S.-H.; Li, X.-J. Huntingtin-associated protein 1 interacts with Ahi1 to regulate cerebellar and brainstem development in mice. J. Clin. Investig. 2008, 118, 2785–2795. [Google Scholar] [CrossRef] [PubMed]
- Vonsattel, J.-P.; Myers, R.H.; Stevens, T.J.; Ferrante, R.J.; Bird, E.D.; Richardson, E.P., Jr. Neuropathological Classification of Huntington's Disease. J. Neuropathol. Exp. Neurol. 1985, 44, 559–577. [Google Scholar] [CrossRef]
- Vonsattel, J.P.G.; Keller, C.; Pilar Amaya, M.d. Neuropathology of Huntington's Disease. Handbook of clinical neurology; Elsevier: 2008; Volume 89, pp. 599-618.
- Yan, S.; Tu, Z.C.; Liu, Z.M.; Fan, N.N.; Yang, H.M.; Yang, S.; Yang, W.L.; Zhao, Y.; Ouyang, Z.; Lai, C.D.; et al. A Huntingtin Knockin Pig Model Recapitulates Features of Selective Neurodegeneration in Huntington’s Disease. Cell 2018, 173, 989–1002. [Google Scholar] [CrossRef] [PubMed]
- Caviston, J.P.; Holzbaur, E.L. Huntingtin as an essential integrator of intracellular vesicular trafficking. Trends Cell Biol. 2009, 19, 147–155. [Google Scholar] [CrossRef]
- Millecamps, S.; Julien, J.-P. Axonal transport deficits and neurodegenerative diseases. Nat. Rev. Neurosci. 2013, 14, 161–176. [Google Scholar] [CrossRef]
- Page, K.J.; Potter, L.; Aronni, S.; Everitt, B.J.; Dunnett, S.B. The expression of Huntingtin-associated protein (HAP1) mRNA in developing, adult and ageing rat CNS: implications for Huntington's disease neuropathology. Eur. J. Neurosci. 1998, 10, 1835–1845. [Google Scholar] [CrossRef]
- Li, T.; Li, S.; Gao, X.; Cai, Q.; Li, X.-J. Expression and Localization of Huntingtin-Associated Protein 1 (HAP1) in the Human Digestive System. Dig. Dis. Sci. 2019, 64. [Google Scholar] [CrossRef]
- Qu, Y.-M.; Chen, A.; Zhao, X.; Wang, Z.; Guo, D.; Shao, S.-L.; Tao, Y.-Y.; Li, Q.-J.; Wang, M.-Y.; Ma, W.-S. Huntingtin-associated protein 1 is a potential tumor suppressor for gastric cancer. Mol. Biol. Rep. 2023, 50, 1517–1531. [Google Scholar] [CrossRef]
- Zhu, L.; Song, X.; Tang, J.; Wu, J.; Ma, R.; Cao, H.; Ji, M.; Jing, C.; Wang, Z. Huntingtin-associated protein 1: A potential biomarker of breast cancer. Oncol. Rep. 2013, 29, 1881–1887. [Google Scholar] [CrossRef]
- Wu, J.; Zhang, J.-y.; Yin, L.; Wu, J.-z.; Guo, W.-j.; Wu, J.-f.; Chen, M.; Xia, Y.-y.; Tang, J.-h.; Ma, Y.-c.; et al. HAP1 gene expression is associated with radiosensitivity in breast cancer cells. Biochem. Biophys. Res. 2015, 456, 162–166. [Google Scholar] [CrossRef]
- Wang, W.; Yan, H.; Zhang, Q.; Song, W.; Li, H.; Xu, J. Evaluating the association of polymorphisms in the HAP1 gene with lung cancer risk: a meta-analysis. Tumor Biol. 2014, 35, 10825–10831. [Google Scholar] [CrossRef]
- Peters, M.F.; Ross, C.A. Isolation of a 40-kDa Huntingtin-associated Protein. J. Biol. Chem. 2001, 276, 3188–3194. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.; Huang, B.; Cheng, J.; Seefelder, M.; Engler, T.; Pfeifer, G.; Oeckl, P.; Otto, M.; Moser, F.; Maurer, M.; et al. The cryo-electron microscopy structure of huntingtin. Nature 2018, 555, 117–120. [Google Scholar] [CrossRef] [PubMed]
- Huang, B.; Seefelder, M.; Buck, E.; Engler, T.; Lindenberg, K.S.; Klein, F.; Landwehrmeyer, G.B.; Kochanek, S. HAP40 protein levels are huntingtin-dependent and decrease in Huntington disease. Neurobiol. Dis. 2021, 158, 105476. [Google Scholar] [CrossRef] [PubMed]
- Harding, R.J.; Deme, J.C.; Hevler, J.F.; Tamara, S.; Lemak, A.; Cantle, J.P.; Szewczyk, M.M.; Begeja, N.; Goss, S.; Zuo, X.; et al. Huntingtin structure is orchestrated by HAP40 and shows a polyglutamine expansion-specific interaction with exon 1. Commun. Biol. 2021, 4, 1374. [Google Scholar] [CrossRef]
- Metzger, S.; Rong, J.; Nguyen, H.-P.; Cape, A.; Tomiuk, J.; Soehn, A.S.; Propping, P.; Freudenberg-Hua, Y.; Freudenberg, J.; Tong, L.; et al. Huntingtin-associated protein-1 is a modifier of the age-at-onset of Huntington's disease. Hum. Mol. Genet. 2008, 17, 1137–1146. [Google Scholar] [CrossRef]
- Lee, J.-M.; Huang, Y.; Orth, M.; Gillis, T.; Siciliano, J.; Hong, E.; Mysore, J.S.; Lucente, D.; Wheeler, V.C.; Seong, I.S.; et al. Genetic modifiers of Huntington disease differentially influence motor and cognitive domains. Am. J. Hum. Genet. 2022, 109, 885–899. [Google Scholar] [CrossRef]
- Bhide, P.G.; Day, M.; Sapp, E.; Schwarz, C.; Sheth, A.; Kim, J.; Young, A.B.; Penney, J.; Golden, J.; Aronin, N.; et al. Expression of Normal and Mutant Huntingtin in the Developing Brain. J. Neurosci. 1996, 16, 5523–5535. [Google Scholar] [CrossRef]
- Barnat, M.; Capizzi, M.; Aparicio, E.; Boluda, S.; Wennagel, D.; Kacher, R.; Kassem, R.; Lenoir, S.; Agasse, F.; Braz, B.Y.; et al. Huntington’s disease alters human neurodevelopment. Science 2020, 369, 787–793. [Google Scholar] [CrossRef] [PubMed]
- Molero, A.E.; Arteaga-Bracho, E.E.; Chen, C.H.; Gulinello, M.; Winchester, M.L.; Pichamoorthy, N.; Gokhan, S.; Khodakhah, K.; Mehler, M.F. Selective expression of mutant huntingtin during development recapitulates characteristic features of Huntington’s disease. Proc. Natl. Acad. Sci. 2016, 113, 5736–5741. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



