1. Introduction
Molecular clockworks consisting of feedback loops of core clock genes drive cell-autonomous circadian oscillation in various species (1). In mammals, the transcription factors (TFs) CLOCK and BMAL1 dimerize to activate the transcription of Per1/2 and Cry1/2, whose protein products are repressors that inhibit CLOCK/BMAL1 action through negative feedback (2). While post-translational regulation of clock proteins play critical roles in setting clock pace (3, 4), the prime mover of circadian oscillation is thought to be transcription (5). High throughput technologies such as microarray (6), RNA-seq (7, 8) and ChIP-seq (9) enable detailed characterization of gene rhythms and genomic binding of clock proteins, allowing in depth analyses of circadian rhythm generation at the level of transcription.
Binding sites of clock proteins are located within open chromatin regions established by tissue-specific pioneer TFs (tsTFs), thus typically tissue-specific (10). Chromatin is known to be a barrier to transcription, and DNA sequences are often not accessible to many TFs, with the exception of tsTFs that are sufficient to trigger enhancer competency within chromatin. Furthermore, tsTFs allow subsequent binding by other TFs, including clock proteins. Some tsTFs (e.g., HNF4a (11, 12)) and ubiquitous TFs (u-TFs. E.g., RELA/p65. (13, 14)) interact with and recruit clock proteins to their cis-elements. CLOCK/BMAL1 can also facilitate the binding of some tsTFs, leading to the suggestion that CLOCK/BMAL1 acts like a pioneer-like TF (10, 15). Like many TFs (16), clock proteins recruit cofactors to modify histone and remodel nucleosome to regulate transcription. Clock proteins and their cofactors form a complex with a M.W. over 1 MDa, and deficiencies in some cofactors alter clock dynamics (17). For example, clock proteins in both Drosophila and mammals recruit the TIP60 complex to regulate clock oscillation (18-20). To control Pol II transcription at the transcription start site (TSS), TFs require the mediator complex (21) to interact with general transcription factors (GTFs) present at gene promoter (22). The mediator subunits interacting with the clock protein complex remain to be determined.
Traditional studies addressed how TFs and cofactors direct the mediator complex to assemble the pre-initiation complex (PIC) for transcription initiation and reinitiation at the TSS (23). Distal enhancers bound by TFs and promoters are thought to be brought to proximity via chromatin looping, a process assisted by proteins such as cohesin and CTCF (24). Traditional view of transcription, however, has difficulties in explaining new findings such as transcription bursting, which represents Pol II initiation and multiple rounds of reinitiation (25). Imaging studies in single cells revealed that the transcription of many genes, including clock genes (26), is stochastic and of low frequency. Transcription often toggles between active and inactive states within a cell, and the active state is characterized by transcription bursting followed by prolonged dormancy of the inactive state (27). Bursting could be readily explained by the formation of transcription hub and molecular condensate of TFs, cofactors, mediators and Pol IIs that permit multiple rounds of Pol II initiation (28, 29). Recent studies revealed that TFs often contain intrinsically disordered low complexity domains, whose interactions induce the formation of transcription hubs and even molecular condensates, the latter via“lipid-lipid phase separation” (28, 29). Transcription bursting also requires pause release (30), which refers to P-TEFb-licensed Pol II elongation to overcome the +1 nucleosome barrier and transcribe into the gene body (31). Initiated Pol II travels only a short distance; it then enters the state of pausing, wherein Pol II stays paused downstream of the TSS via actions of pausing factors (DSIF and NELF) and the +1 nucleosome (31, 32). P-TEFb, a component of the super elongation complex (SEC) (33, 34), releases paused Pol II for elongation, permitting Pol II reinitiation to achieve transcription bursting.
Compared to other aspects of transcription regulation of circadian rhythms (35), Pol II recruitment, initiation, and pausing just begin to attract attention from the circadian rhythm field (2, 19). Recently, we performed analyses of Pol II recruitment and pausing during daily transcription in mouse liver, and revealed unique characteristics of those regulatory steps (36). In this article, we first reviewed the roles of rhythmic transcription in clock oscillation. We also provided a brief account of recent progress on transcription regulation by clock proteins and other TFs. We then discussed our findings on Pol II recruitment and pausing during daily transcription and the implications of their regulation to rhythmic gene transcription.
2. Transcription regulation is the main driving force for gene expression rhythms
First demonstrated for Per in flies (37), core clock genes exhibit robust daily changes in their mRNA expression. Owing to rapid co-transcriptional splicing, pre-mRNA level can be used as the surrogate for transcription activity. RNAse-protection assays against Per pre-mRNA and mRNA showed that the Per mRNA rhythm in Drosophila is mainly driven at the level of transcription (38). Post-transcriptional regulation of mRNA stability also contributes to Per mRNA rhythm, and is sufficient to confer rhythmic mRNA expression to other genes (39, 40). In mammals, core clock genes such as Per1&2 also exhibit robust daily changes in mRNA levels (41, 42). pre-mRNA measurement implicated rhythmic transcription as the driving force for mRNA rhythms of core clock genes and many other genes (43, 44). Deep sequencing studies evaluated the contribution of transcription regulation to mRNA rhythm generation in a genomewide manner. One study estimated that 22% of mRNA rhythms are driven by rhythmic transcription (9). Later studies with high sequencing depth and kinetic modeling increased the estimate to about 70-80%, whereas rhythmic degradation contributes to the mRNA rhythms of 30%-35% genes (45, 46). Nuclear export, another post-transcriptional regulation step, contributes to rhythm generation for 10% of rhythmic transcriptome (47). Overall, rhythmic transcription is deemed as the main driving force for gene rhythms (5).
3. Both the intrinsic tissue clock and extrinsic cues can regulate gene expression rhythms
Clock genes typically harbor multiple cis-elements for clock proteins, which also have numerous other binding sites across the genome. Clock proteins thus also regulate many other genes. Clock genes and other genes are also influenced by extrinsic cues, which are often rhythmic in wildtype animals. Such cues include body temperature (Tb) (48), feeding (49), and communicating signals from other tissues (including the autonomous nervous system) (50). The extrinsic cues can engage TFs as well as post-transcriptional mechanisms to regulate gene rhythms, including those of clock genes. For example, daily changes in Tb drive rhythmic HSF1 expression to regulate gene transcription (51). The Tb rhythm also drives Cirbp expression to post-transcriptionally regulate clock dynamics (52). Besides the Tb rhythm, blood-borne cues also regulate clock dynamics: serum and plasma can activate multiple signaling pathways to impact clock genes (53-55). For example, rhythmic cues in plasma activate SRF, which regulates transcription of the clock gene mPer2 (55). Certain blood-borne cues impacting clock dynamics are heat-labile, implying that they are proteins (56). Lipids can also serve as inter-tissue communicating cues. For example, phosphatidylcholine is synthesized by liver and released into plasma to activate PPARα in muscles (57).
Overall, clock proteins and many other TFs exhibit daily changes in their actions. Like clock proteins, other TFs also have thousands of genomic binding sites in various tissues. Therefore, they potentially can regulate numerous genes besides clock genes. Gene expression rhythms are thus driven by both clock proteins and other TFs. How clock proteins and other TFs work together to control gene rhythms was the focus of recent studies in various peripheral mouse tissues.
4. Clock proteins typically collaborate with other TFs to regulate transcription rhythms
Clock proteins and other TFs often collaborate to regulate target genes (10). The independent contribution of the clock to gene rhythms is rather limited (58, 59). In studies that reconstitute clock oscillation (RE) in specific tissues of Bmal1 deficient mice (58, 59), it was shown that only 10% of the rhythmic transcriptome can be restored in livers of the liver-RE mice (58). However, that is not to say that the liver clock regulates only 10% of the rhythmic transcriptome in wildtype mice. In fact, disruption of the liver clock disturbs about 90% of gene rhythms in mouse liver (60, 61). Overall, those results indicated that the majority of gene rhythms are regulated in a combinatorial manner by both the intrinsic clock and TFs engaged by extrinsic cues.
By comparing liver gene rhythms in Bmal1 KO, liver-RE and wildtype mice under ad libitum versus nighttime restricted feeding, it was shown that the mRNA rhythms in livers of wildtype mice can be partitioned into four parts based on their modes of regulation (62). Some rhythms can be driven by the intrinsic liver clock alone (13.7%); some can be driven by rhythmic feeding cues alone (17.5%); some require not only the intrinsic clock but also rhythmic feeding cues (34.5%), while the rest (34.4%) require both the intrinsic clock and rhythmic cues from other tissues (and their clocks). Those results indicate that, for the regulation of a majority of gene rhythms, there is a mandatory requirement for clock proteins to collaborate with other TFs. For example, feeding engages the TF CEBPB to coregulate BMAL1 target genes, and CEBPB deficiency disrupts the rhythms of some BMAL1 target genes that are also regulated by feeding (62).
5. The need to study Pol II pausing regulation near the TSS
Clock proteins and other TFs occupying distinct enhancers of the same gene can collaborate through chromatin looping to regulate transcription. Techniques such as Hi-C and CHIA-PET revealed daily changes in long-range interactions between distinct enhancers bound by clock proteins and other TFs respectively, and between those enhancers and gene promoters (10, 63). The collaboration between clock proteins and other TFs can also occur at the same enhancers. Indeed, TFs often exhibit cooperative binding at the same enhancers to increase the affinities of the two factors to their respective motifs. However, cooperative binding does not necessarily leads to coactivation. For example, HNF4a and RELA/p65 can recruit CLOCK/BMAL1 for genomic binding (11, 13), but transrepress its transcription activation (12, 14). Such interactions between TFs at same and/or distinct enhancers pose serious challenges in elucidating how clock proteins contribute to the final transcription output at the TSS. Indeed, genomic binding of CLOCK/BMAL1 at enhancers is often not sufficient to specify rhythm phase and amplitude, and even could not confer rhythmicity to some target genes (35).
Another aspect of complexity of transcription regulation is the lack of consensus on how TFs and their cofactors at distal enhancers regulate transcription near the TSS (64). The textbook model of chromatin looping posits that stable contact is formed between distal enhancers and promoters. A variation of this classical model is the “kiss-and-run” model of transient contact between distal enhancers and promoters. However, the nature of such long-range genomic interactions and its relevance to transcription are recently questioned (64). The alternative TAG (TF activity gradient) model (64) emphasizes on contact-independent “communication by diffusion” of TFs and its cofactors between enhancers and promoters. However, given the diversity of interacting TFs and their multitudes of cofactors, it would be difficult, if not impossible, to dissect out the specific contribution of an individual TF and/or cofactor to transcription output. On the other hand, Pol II recruitment, transcription initiation and Pol II pausing represent the final regulatory outcome by a plethora of TFs and cofactors. Those regulatory steps are directly related to final transcription output. Information about them is thus critical for understanding the logic of transcription regulation. Surprisingly, such information is lacking in circadian rhythm research.
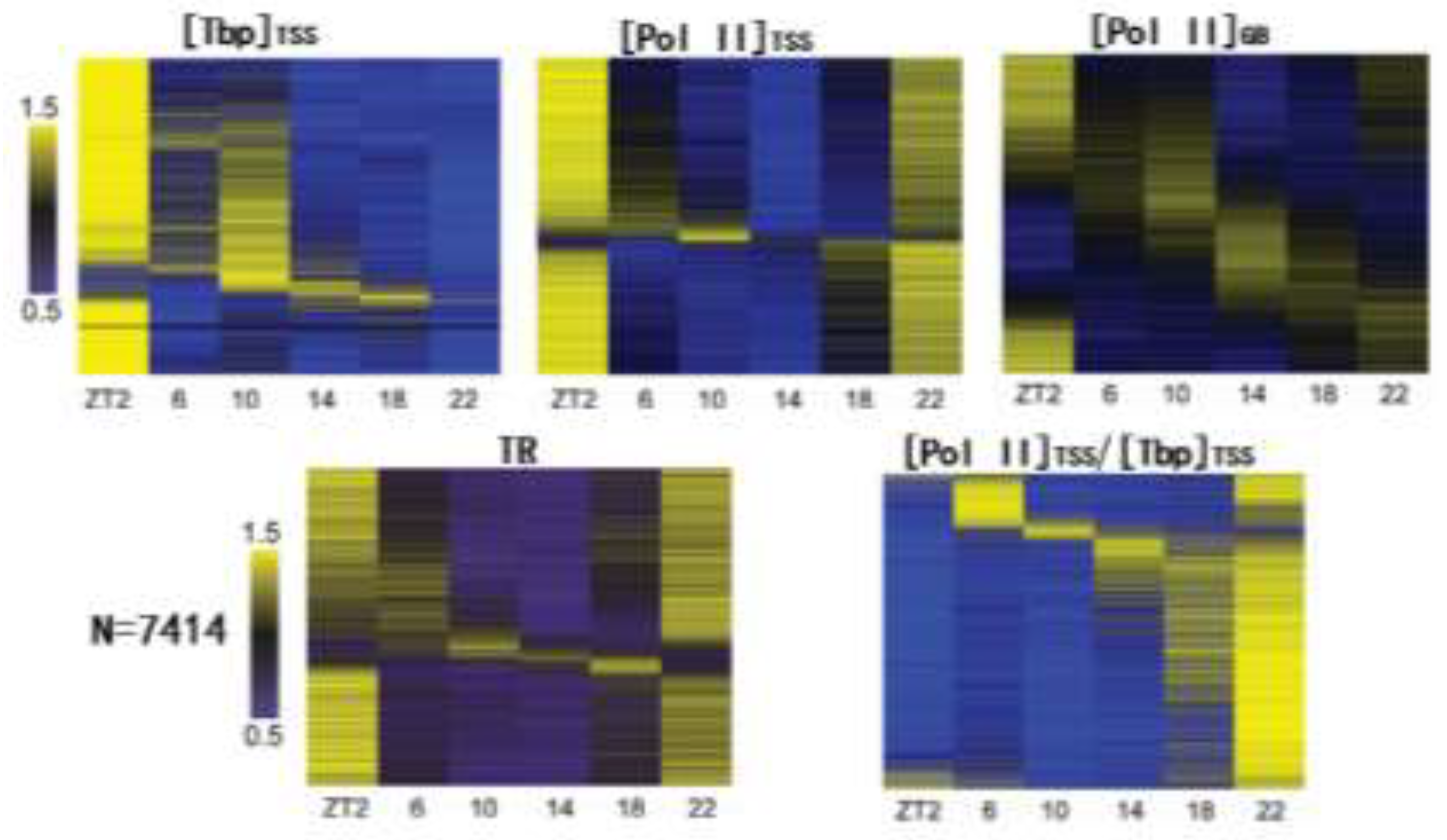
Against this backdrop, we performed a ChIP-seq study of Tbp (TATA-binding protein. A TFII D subunit) and Pol II during daily transcription in mouse liver (36). Tbp ([Tbp]TSS) and Pol II ([Pol II]TSS) signals in the TSS region (defined as -50 to +300 bp to TSS (36, 65)) and gene body ([Pol II]GB) of 7414 genes were quantitatively analyzed. Pol II traveling ratios (TR: [Pol II]TSS:[Pol II]GB), a quantitative measure of Pol II pausing (65, 66), were also calculated. By systematic characterization of Pol II recruitment and pausing during daily transcription, our study provided the first glimpse of their genomewide characteristics.
6. Global characteristics of Pol II recruitment and pausing during daily transcription in the mouse liver
The results of our study are summarized in
Figure 1. As can be seen, Pol II recruitment measured by [Tbp]
TSS is typically low during nighttime, especially at ZT22 (zeitgeber time 22. ZT0 corresponds to lights on during the 12:12 light/dark cycle). [Tbp]
TSS exhibits great rebound at ZT2. Most genes also had higher [Pol II]
TSS near ZT2. Moreover, numerous genes’ transcription rates, as measured by [Pol II]
GB, are higher at ZT2, concurrent with the increase in [Pol II]
TSS and [Tbp]
TSS (
Figure 1). Together, those results indicate global upregulation of gene transcription at ZT2. Nonetheless, [Pol II]
GB exhibits more gene-specific changes across the day than [Pol II]
TSS and [Tbp]
TSS, with many genes’ transcription peaking at other time points than ZT2, especially near ZT14. The overall bimodal distribution of peak gene transcription is consistent with pre-mRNA analysis results by others (46).
The characteristics of daily changes in [Tbp]
TSS, [Pol II]
TSS and [Pol II]
GB (
Figure 1) could be related to the cell cycle, whose progression is known to impact transcription (67). In particular, transcription is generally inhibited during mitosis and reactivated upon mitotic exit (68, 69). While the liver is typically considered a non-dividing organ, it still exhibits daily changes in cell-cycle related activities that interplay with the clock (70, 71). For example, the activity of CDK1, which is critical for mitosis entry, exhibits daily changes controlled by the clock (72) and also regulates clock oscillation in return (73). CDK1 activity peaks before ZT0 in mouse liver (73, 74). CDK1 is known to phosphorylate TFII D to inhibit PIC formation (75). The late night rise of CDK1 activity in mouse liver could induce global diminishment of [Tbp]
TSS , as indeed observed at ZT22. On the other hand, the rebound of [Tbp]
TSS and [Pol II]
TSS at ZT2 could be analogous to gene reactivation upon mitotic exit (68, 69).
Our results showed that Pol II TRs of all genes exhibit daily changes (
Figure 1). TRs of most genes are high near ZT0, and their nadirs are near ZT12, especially at ZT14 (
Figure 1). [Pol II]
TSS also reaches its genomewide nadir at ZT14 (
Figure 1). Pause release lowers Pol II TR by decreasing [Pol II]
TSS and increasing [Pol II]
GB. The patterns of daily changes in [Pol II]
TSS and TR in our results suggest a global rhythm of pause release affecting most liver genes. Against this global trend, however, some genes’ TRs peak near ZT14. The possible causes for such exceptions are discussed in section 8.
The results of our ChIP-seq study are population averages lacking single-cell resolution. In single cells, the inherent noises of stochastic gene expression lead to phenotypic variations, such as period heterogeneity among clonal cell populations (76-78). More specifically to our study, Pol II recruitment (and subsequent pausing) and pause release could be distinct events occurring randomly among cells. However, single cell imaging studies showed that Pol II recruitment and bursting are not mutually independent, but are sequential events occurring in close succession (79). This permits cross analyses of Tbp and Pol II signals to infer the daily rules of transcription regulation. Below we discuss our analysis results and their implications.
7. Pol II recruitment is not a direct determinant of gene transcription rate
Traditionally, Pol II recruitment is thought to be the determinant of transcription output. However, our results showed that [Tbp]
TSS does not correlate well with [Pol II]
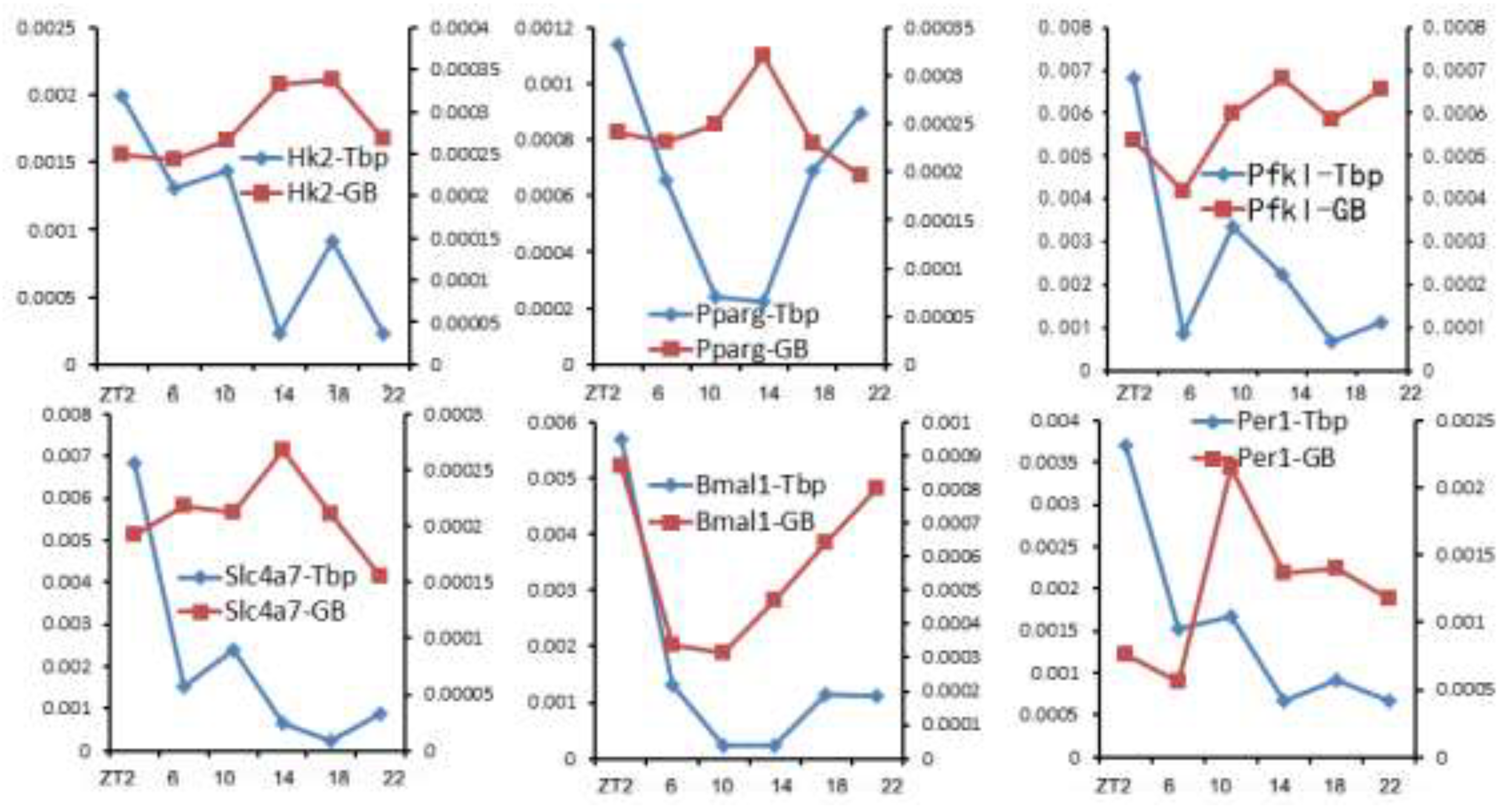
GB for numerous genes (36). Six example genes are shown in
Figure 2. While [Tbp]
TSS values of all 6 genes are highest at ZT2 and lowest at ZT22, transcription rates ([Pol II]
GB) of those genes peak at different phases. For example, [Tbp]
TSS and [Pol II]
GB values of
Bmal1 are highest at ZT2, but several genes’ transcription rates peak near ZT14, when their [Tbp]
TSS values are low compared to other time points. Such results clearly indicate that Pol II recruitment does not directly determine transcription rate. This appears at odds with the current view of coordinated regulation of Pol II recruitment and transcription bursting (79). Nonetheless, the paradox could be reconciled if pause release, which is required for transcription bursting, is regulated independently from PIC formation. Such a scenario has been reported (80). Upon acute depletion of the mediator complex to limit Pol II recruitment and initiation, cell-type specific genes’ transcription is lowered (80). However, the transcription of many other genes is maintained due to a compensatory rise of pause release (80). Such results indicate that pause release could affect transcription in a manner independent of Pol II recruitment.
Transcription bursting consists two parameters, burst frequency and burst size, which often respond differentially to biological stimuli and experimental manipulations (81). Burst frequency is primarily determined by Pol II recruitment, which leads to PIC formation and transcription initiation (82). On the other hand, burst size is mainly affected by pause release, which permits rounds of reinitiation. If pause release is low during Pol II recruitment, then Pol II from the first initiation round would stay paused downstream of the TSS, creating a barrier for trailing Pol II. Eventually it could hinder downstream movement of Pol II in the PIC. Indeed, Pol II pausing inhibits initiation (83, 84). By contrast, P-TEFb-mediated release of paused Poll promotes Pol II reinitiation to increase burst size. It might also reduce the dwelling time of the Pol II “hub” and/or condensate near the TSS to facilitate new PIC formation and initiation. Thus, pause release might increase burst frequency.
With regard to daily transcription in mouse liver, we suspect that genes with highest [Tbp]TSS, [Pol II]TSS and [Pol II]GB at ZT2 have high burst frequency but low burst size at this time point. By contrast, the upregulation of some genes’ transcription near ZT14 could be due to increased burst sizes associated with high pause release activity. Such a scenario is supported by other evidence. For example, a genomewide increase of H3K4me3 at early night is evident in mouse liver (9, 85, 86). During pause release, P-TEFb recruits the PAF1 complex (PAF1c) (87) to stimulate the activity of SET1 (88, 89), which deposits H3K4me3 downstream of the TSSs of genes. The increase of H3K4me3 at early night is consistent with a global rise in pause release activity at that time.
TFs, their cofactors, and the mediator complex are known to regulate pause release through interactions with P-TEFb. TFs such as c-Myc (66) can directly recruit P-TEFb. The cofactor BRD4 also binds P-TEFb (90, 91), and the TIP60 complex is suggested to acetylate BMAL1 to recruit BRD4-P-TEFb (19). The mediator complex not only regulates PIC formation but also downstream transcription events, including Pol II initiation and pause release (25). The MED23 and MED26 subunits and the CDK8 mediator kinase module (MKM) of the mediator complex have been shown to interact with P-TEFb/SEC (92, 93). However, we want to emphasize that, in the molecular interactions controlling pause release, P-TEFb is probably a rate-limiting factor. P-TEFb can be sequestered into the 7SK snRNP, where it remains inactive and unable to elicit pause release. Signaling pathways can activate pause release via inducing 7SK snRNP disassembly to release P-TEFb (80, 94, 95). As shown in (80), the disassembly of 7SK snRNP boosts P-TEFb availability to increase pause release. This could be a potential mechanism accounting for daily changes in pause release activity in mouse liver.
8. Premature transcription termination at the 5’end of genes contribute to the regulation of Pol II pausing
Pause release decreases Pol II stability in the TSS region by enabling Pol II outflux for elongation. Footprinting and imaging studies revealed rapid turnover of Pol II near the TSS (96, 97). However, blocking pause release via P-TEFb inhibition only partially increases Pol II stability (96-98). Those results implicate premature termination as the major determinant of Pol II stability near the TSS (99, 100). We used the ratio of [Pol II]
TSS to [Tbp]
TSS as the index of Pol II stability within the TSS region. As evident in
Figure 1, Pol II stability is lowest at ZT2, when pause release activity appears low. On the other hand, Pol II stability at ZT14, a time point presumably of high pause release activity, is intermediate among the 6 daily time points (
Figure 1). Those results indicate a critical role of premature termination in lowering Pol II stability (99, 100). Because premature termination decreases [Pol II]
TSS (thus Pol II TR) in a manner independent of P-TEFb, it leads to inaccurate estimate of pause release. For example, Pol II TRs of most genes are low at ZT14, most probably due to a global increase in pause release. However, some genes’ TRs peak at ZT14 (
Figure 1). We suspect that, for those outlier genes, premature termination significantly lower their TR values at other time points to confound the estimate of pause release activity at ZT14.
The TSS region can only harbor a limited number of Pol IIs, and such space limitation is suggested to subject Pol II to collisions that promote premature termination (101). However, the target of premature termination needs clarification. In ChIP-seq studies with high sequencing depth, two Pol II peaks can be observed in the TSS region (102). The 1st peak centers on the TSS and represents initiated Pol II before pausing. The 2nd peak is 110 bp downstream of the TSS and represents canonical pausing. Existing evidence suggests that paused Pol II (the 2nd Pol II peak) is stable. For example, following triptolide treatment to inhibit Pol II initiation, the half-lives of Pol IIs near the TSS are typically minutes to even above an hour (103, 104), indicating that paused Pol IIs are not prone to rapid turnover via premature termination. While such long half-lives were questioned based on efficacy of triptolide treatment (96, 97), we suspect that premature termination mainly targets initiated Pol II before pausing. The capping of nascent pre-mRNAs starts upon transcription initiation and is completed when Pol II enters the pausing state (105). 5’-capped nascent RNAs are very stable, with only about 1% subject to premature termination, indicating that Pol II pausing is stable (106).
The pausing factor NELF recruits the cap-binding complex (CBC) to bind the m7G cap (107). Importantly, the m7G cap and its binding by CBC are checkpoints for pre-mRNA splicing and Pol II elongation (108-110). This suggests a quality control role of premature termination to ensure productive Pol II elongation. Indeed, premature termination involves pre-mRNA quality control mechanisms. For example, XRN2 plays a role in premature termination (111). XRN2 acts on uncapped RNAs, and its action is assisted by decapping enzymes such as DXO and DCP2, whose likely targets are inappropriately capped pre-mRNAs (112). The integrator complex (INTS), which cleaves nascent RNAs and recruits PP2A to dephosphorylate Pol II CTD, also functions in premature termination (113). INTS depletion leads to the production of unspliced transcripts by Pol IIs incompetent for productive elongation (114, 115), indicating that INTS functions in pre-mRNA quality control. By contrast, CBC functions in P-TEFb recruitment and INTS exclusion to activate pause release and productive elongation (116, 117).
9. Future perspectives
To regulate gene transcription rhythms, clock proteins and their collaborating TFs at distal enhancers need to gain access to the mediator complex and GTFs to control Pol II recruitment, initiation and pause release near the TSS. Our study revealed that Pol II recruitment and pause release exhibit genomewide changes that peak at distinct clock phases, thus providing new perspectives on the logic of transcription regulation of circadian rhythms. Our results support the notion that, besides Pol II recruitment, Pol II pausing and pause release are also critical regulatory steps to influence final transcription out.
A limitation of our study is the use of [Tbp]TSS to measure Pol II initiation. Tbp and mediator remain promoter-bound during PIC formation, initiation and reinitiation (118, 119). By contrast, GTFs such as TFII B dissociate after Pol II initiation and recycle for Pol II binding during reinitiation (120, 121). Thus, [Tbp]TSS overestimates Pol II initiation and reinitiation rate. GTFs such as TFII B could provide more accurate rate measures. Nonetheless, Tbp and other GTFs could exhibit concordant changes (83, 84), permitting the use of [Tbp]TSS to measure Pol II initiation and reinitiation (122).
Our study and those by others routinely use [Pol II]TSS as the measure of paused Pol II. However, [Pol II]TSS actually contains signals from multiple Pol II forms (102), including PIC, initiated Pol II before pausing, and paused Pol II. [Pol II]TSS thus overestimates paused Pol II. Moreover, both pause release and premature termination can lowers [Pol II]TSS. Those confounding factors can lead to inaccuracies in TR results. Ideally, only signals of paused Pol II should be used for TR calculation to measure Pol II pausing and pause release. However, it is not feasible in practice to find an antibody that exclusively recognizes paused Pol II. Even monoclonal antibodies against a specific Pol II CTD form (e.g., pSer5) show various cross-reactivities to other forms, and different antibodies against the same epitope could yield dramatically different results (123). We suggest alternative targets such as CBC components (107) as surrogates of paused Pol II. This would improve the accuracies in quantitative analyses of Pol II pausing and pause release. Such improvement would help to elucidate the mechanisms for daily changes in pause release and premature termination, and to better pinpoint their roles in shaping transcription rhythms.
Acknowledgments
This work is supported by the National Natural Science Foundation of China (NSFC31971091).
References
- Rosbash M (2009) The implications of multiple circadian clock origins. PLoS Biol 7(3):e62. [CrossRef]
- Takahashi JS (2017) Transcriptional architecture of the mammalian circadian clock. Nat Rev Genet 18(3):164-179. [CrossRef]
- Zhou M, Kim JK, Eng GW, Forger DB, & Virshup DM (2015) A Period2 Phosphoswitch Regulates and Temperature Compensates Circadian Period. Mol Cell 60(1):77-88. [CrossRef]
- Toh KL, et al. (2001) An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291(5506):1040-1043. [CrossRef]
- Montenegro-Montero A & Larrondo LF (2016) In the Driver's Seat: The Case for Transcriptional Regulation and Coupling as Relevant Determinants of the Circadian Transcriptome and Proteome in Eukaryotes. J Biol Rhythms 31(1):37-47. [CrossRef]
- Panda S, et al. (2002) Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109(3):307-320. [CrossRef]
- Zhang R, Lahens NF, Ballance HI, Hughes ME, & Hogenesch JB (2014) A circadian gene expression atlas in mammals: implications for biology and medicine. Proc Natl Acad Sci U S A 111(45):16219-16224. [CrossRef]
- Mure LS, et al. (2018) Diurnal transcriptome atlas of a primate across major neural and peripheral tissues. Science 359(6381):10.1126/science.aao0318. [CrossRef]
- Koike N, et al. (2012) Transcriptional architecture and chromatin landscape of the core circadian clock in mammals. Science 338(6105):349-354. [CrossRef]
- Beytebiere JR, et al. (2019) Tissue-specific BMAL1 cistromes reveal that rhythmic transcription is associated with rhythmic enhancer-enhancer interactions. Genes Dev 33(5-6):294-309. [CrossRef]
- Qu M, Qu H, Jia Z, & Kay SA (2021) HNF4A defines tissue-specific circadian rhythms by beaconing BMAL1::CLOCK chromatin binding and shaping the rhythmic chromatin landscape. Nat Commun 12(1):6350. [CrossRef]
- Qu M, Duffy T, Hirota T, & Kay SA (2018) Nuclear receptor HNF4A transrepresses CLOCK:BMAL1 and modulates tissue-specific circadian networks. Proc Natl Acad Sci U S A 115(52):E12305-E12312. [CrossRef]
- Hong HK, et al. (2018) Requirement for NF-kappaB in maintenance of molecular and behavioral circadian rhythms in mice. Genes Dev 32(21-22):1367-1379. [CrossRef]
- Shen Y, et al. (2021) NF-kappaB modifies the mammalian circadian clock through interaction with the core clock protein BMAL1. PLoS Genet 17(11):e1009933. [CrossRef]
- Menet JS, Pescatore S, & Rosbash M (2014) CLOCK:BMAL1 is a pioneer-like transcription factor. Genes Dev 28(1):8-13. [CrossRef]
- Lemon B & Tjian R (2000) Orchestrated response: a symphony of transcription factors for gene control. Genes Dev 14(20):2551-2569. [CrossRef]
- Aryal RP, et al. (2017) Macromolecular Assemblies of the Mammalian Circadian Clock. Mol Cell 67(5):770-782 e776. [CrossRef]
- Mahesh G, et al. (2020) Proteomic analysis of Drosophila CLOCK complexes identifies rhythmic interactions with SAGA and Tip60 complex component NIPPED-A. Sci Rep 10(1):17951. [CrossRef]
- Petkau N, Budak H, Zhou X, Oster H, & Eichele G (2019) Acetylation of BMAL1 by TIP60 controls BRD4-P-TEFb recruitment to circadian promoters. Elife 8. [CrossRef]
- Ju D, et al. (2020) Chemical perturbations reveal that RUVBL2 regulates the circadian phase in mammals. Sci Transl Med 12(542). [CrossRef]
- Malik S & Roeder RG (2010) The metazoan Mediator co-activator complex as an integrative hub for transcriptional regulation. Nat Rev Genet 11(11):761-772. [CrossRef]
- Thomas MC & Chiang CM (2006) The general transcription machinery and general cofactors. Crit Rev Biochem Mol Biol 41(3):105-178. [CrossRef]
- Roeder RG (2019) 50+ years of eukaryotic transcription: an expanding universe of factors and mechanisms. Nat Struct Mol Biol 26(9):783-791. [CrossRef]
- Hsieh TS, et al. (2022) Enhancer-promoter interactions and transcription are largely maintained upon acute loss of CTCF, cohesin, WAPL or YY1. Nat Genet 54(12):1919-1932. [CrossRef]
- Richter WF, Nayak S, Iwasa J, & Taatjes DJ (2022) The Mediator complex as a master regulator of transcription by RNA polymerase II. Nat Rev Mol Cell Biol 23(11):732-749. [CrossRef]
- Suter DM, et al. (2011) Mammalian genes are transcribed with widely different bursting kinetics. Science 332(6028):472-474. [CrossRef]
- Rodriguez J & Larson DR (2020) Transcription in Living Cells: Molecular Mechanisms of Bursting. Annu Rev Biochem 89:189-212. [CrossRef]
- Lim B & Levine MS (2021) Enhancer-promoter communication: hubs or loops? Curr Opin Genet Dev 67:5-9. [CrossRef]
- McSwiggen DT, Mir M, Darzacq X, & Tjian R (2019) Evaluating phase separation in live cells: diagnosis, caveats, and functional consequences. Genes Dev 33(23-24):1619-1634. [CrossRef]
- Jonkers I & Lis JT (2015) Getting up to speed with transcription elongation by RNA polymerase II. Nat Rev Mol Cell Biol 16(3):167-177. [CrossRef]
- Chiu AC, et al. (2018) Transcriptional Pause Sites Delineate Stable Nucleosome-Associated Premature Polyadenylation Suppressed by U1 snRNP. Mol Cell 69(4):648-663 e647. [CrossRef]
- Yamaguchi Y, Shibata H, & Handa H (2013) Transcription elongation factors DSIF and NELF: promoter-proximal pausing and beyond. Biochim Biophys Acta 1829(1):98-104. [CrossRef]
- Luo Z, Lin C, & Shilatifard A (2012) The super elongation complex (SEC) family in transcriptional control. Nat Rev Mol Cell Biol 13(9):543-547. [CrossRef]
- Chen Y, et al. (2021) Allosteric transcription stimulation by RNA polymerase II super elongation complex. Mol Cell 81(16):3386-3399 e3310. [CrossRef]
- Beytebiere JR, Greenwell BJ, Sahasrabudhe A, & Menet JS (2019) Clock-controlled rhythmic transcription: is the clock enough and how does it work? Transcription 10(4-5):212-221. [CrossRef]
- Zhu J, Li C, Gong C, & Li X (2018) Regulation of Pol II Pausing Is Involved in Daily Gene Transcription in the Mouse Liver. J Biol Rhythms 33(4):350-362. [CrossRef]
- Hardin PE, Hall JC, & Rosbash M (1990) Feedback of the Drosophila period gene product on circadian cycling of its messenger RNA levels. Nature 343(6258):536-540. [CrossRef]
- Hardin PE, Hall JC, & Rosbash M (1992) Circadian oscillations in period gene mRNA levels are transcriptionally regulated. Proc Natl Acad Sci U S A 89(24):11711-11715. [CrossRef]
- Frisch B, Hardin PE, Hamblen-Coyle MJ, Rosbash M, & Hall JC (1994) A promoterless period gene mediates behavioral rhythmicity and cyclical per expression in a restricted subset of the Drosophila nervous system. Neuron 12(3):555-570. [CrossRef]
- So WV & Rosbash M (1997) Post-transcriptional regulation contributes to Drosophila clock gene mRNA cycling. EMBO J 16(23):7146-7155. [CrossRef]
- Sun ZS, et al. (1997) RIGUI, a putative mammalian ortholog of the Drosophila period gene. Cell 90(6):1003-1011. [CrossRef]
- Shearman LP, Zylka MJ, Weaver DR, Kolakowski LF, Jr., & Reppert SM (1997) Two period homologs: circadian expression and photic regulation in the suprachiasmatic nuclei. Neuron 19(6):1261-1269. [CrossRef]
- Preitner N, et al. (2002) The orphan nuclear receptor REV-ERBalpha controls circadian transcription within the positive limb of the mammalian circadian oscillator. Cell 110(2):251-260. [CrossRef]
- Atger F, et al. (2015) Circadian and feeding rhythms differentially affect rhythmic mRNA transcription and translation in mouse liver. Proc Natl Acad Sci U S A 112(47):E6579-6588. [CrossRef]
- Luck S, Thurley K, Thaben PF, & Westermark PO (2014) Rhythmic degradation explains and unifies circadian transcriptome and proteome data. Cell Rep 9(2):741-751. [CrossRef]
- Wang J, et al. (2018) Circadian clock-dependent and -independent posttranscriptional regulation underlies temporal mRNA accumulation in mouse liver. Proc Natl Acad Sci U S A 115(8):E1916-E1925. [CrossRef]
- Hurni C, Weger BD, Gobet C, & Naef F (2022) Comprehensive analysis of the circadian nuclear and cytoplasmic transcriptome in mouse liver. PLoS Genet 18(8):e1009903. [CrossRef]
- Buhr ED, Yoo SH, & Takahashi JS (2010) Temperature as a universal resetting cue for mammalian circadian oscillators. Science 330(6002):379-385. [CrossRef]
- Damiola F, et al. (2000) Restricted feeding uncouples circadian oscillators in peripheral tissues from the central pacemaker in the suprachiasmatic nucleus. Genes Dev 14(23):2950-2961. [CrossRef]
- Koronowski KB & Sassone-Corsi P (2021) Communicating clocks shape circadian homeostasis. Science 371(6530):10.1126/science.abd0951. [CrossRef]
- Reinke H, et al. (2008) Differential display of DNA-binding proteins reveals heat-shock factor 1 as a circadian transcription factor. Genes Dev 22(3):331-345. [CrossRef]
- Gotic I & Schibler U (2017) Posttranscriptional mechanisms controlling diurnal gene expression cycles by body temperature rhythms. RNA Biol 14(10):1294-1298. [CrossRef]
- Balsalobre A, Marcacci L, & Schibler U (2000) Multiple signaling pathways elicit circadian gene expression in cultured Rat-1 fibroblasts. Curr Biol 10(20):1291-1294. [CrossRef]
- Balsalobre A, Damiola F, & Schibler U (1998) A serum shock induces circadian gene expression in mammalian tissue culture cells. Cell 93(6):929-937. [CrossRef]
- Gerber A, et al. (2013) Blood-borne circadian signal stimulates daily oscillations in actin dynamics and SRF activity. Cell 152(3):492-503. [CrossRef]
- Pagani L, et al. (2011) Serum factors in older individuals change cellular clock properties. Proc Natl Acad Sci U S A 108(17):7218-7223. [CrossRef]
- Liu S, et al. (2013) A diurnal serum lipid integrates hepatic lipogenesis and peripheral fatty acid use. Nature 502(7472):550-554. [CrossRef]
- Koronowski KB, et al. (2019) Defining the Independence of the Liver Circadian Clock. Cell 177(6):1448-1462 e1414. [CrossRef]
- Welz PS, et al. (2019) BMAL1-Driven Tissue Clocks Respond Independently to Light to Maintain Homeostasis. Cell 177(6):1436-1447 e1412. [CrossRef]
- Cho H, et al. (2012) Regulation of circadian behaviour and metabolism by REV-ERB-alpha and REV-ERB-beta. Nature 485(7396):123-127. [CrossRef]
- Kornmann B, Schaad O, Bujard H, Takahashi JS, & Schibler U (2007) System-driven and oscillator-dependent circadian transcription in mice with a conditionally active liver clock. PLoS Biol 5(2):e34. [CrossRef]
- Greco CM, et al. (2021) Integration of feeding behavior by the liver circadian clock reveals network dependency of metabolic rhythms. Sci Adv 7(39):eabi7828. [CrossRef]
- Kim YH, et al. (2018) Rev-erbalpha dynamically modulates chromatin looping to control circadian gene transcription. Science 359(6381):1274-1277. [CrossRef]
- Karr JP, Ferrie JJ, Tjian R, & Darzacq X (2022) The transcription factor activity gradient (TAG) model: contemplating a contact-independent mechanism for enhancer-promoter communication. Genes Dev 36(1-2):7-16. [CrossRef]
- Day DS, et al. (2016) Comprehensive analysis of promoter-proximal RNA polymerase II pausing across mammalian cell types. Genome Biol 17(1):120. [CrossRef]
- Rahl PB, et al. (2010) c-Myc regulates transcriptional pause release. Cell 141(3):432-445. [CrossRef]
- Fischer M, Schade AE, Branigan TB, Muller GA, & DeCaprio JA (2022) Coordinating gene expression during the cell cycle. Trends Biochem Sci 47(12):1009-1022. [CrossRef]
- Hsiung CC, et al. (2016) A hyperactive transcriptional state marks genome reactivation at the mitosis-G1 transition. Genes Dev 30(12):1423-1439. [CrossRef]
- Palozola KC, et al. (2017) Mitotic transcription and waves of gene reactivation during mitotic exit. Science 358(6359):119-122. [CrossRef]
- Gaucher J, Montellier E, & Sassone-Corsi P (2018) Molecular Cogs: Interplay between Circadian Clock and Cell Cycle. Trends Cell Biol 28(5):368-379. [CrossRef]
- Bieler J, et al. (2014) Robust synchronization of coupled circadian and cell cycle oscillators in single mammalian cells. Mol Syst Biol 10(7):739. [CrossRef]
- Matsuo T, et al. (2003) Control mechanism of the circadian clock for timing of cell division in vivo. Science 302(5643):255-259. [CrossRef]
- Zhao X, et al. (2016) Circadian Amplitude Regulation via FBXW7-Targeted REV-ERBalpha Degradation. Cell 165(7):1644-1657. [CrossRef]
- Wang J, et al. (2017) Nuclear Proteomics Uncovers Diurnal Regulatory Landscapes in Mouse Liver. Cell Metab 25(1):102-117. [CrossRef]
- Segil N, Guermah M, Hoffmann A, Roeder RG, & Heintz N (1996) Mitotic regulation of TFIID: inhibition of activator-dependent transcription and changes in subcellular localization. Genes Dev 10(19):2389-2400. [CrossRef]
- Li Y, et al. (2020) Noise-driven cellular heterogeneity in circadian periodicity. Proc Natl Acad Sci U S A 117(19):10350-10356. [CrossRef]
- Li Y, et al. (2020) Epigenetic inheritance of circadian period in clonal cells. Elife 9. [CrossRef]
- Nikhil KL, Korge S, & Kramer A (2020) Heritable gene expression variability and stochasticity govern clonal heterogeneity in circadian period. PLoS Biol 18(8):e3000792. [CrossRef]
- Bartman CR, et al. (2019) Transcriptional Burst Initiation and Polymerase Pause Release Are Key Control Points of Transcriptional Regulation. Mol Cell 73(3):519-532 e514. [CrossRef]
- Jaeger MG, et al. (2020) Selective Mediator dependence of cell-type-specifying transcription. Nat Genet 52(7):719-727. [CrossRef]
- Nicolas D, Phillips NE, & Naef F (2017) What shapes eukaryotic transcriptional bursting? Mol Biosyst 13(7):1280-1290. [CrossRef]
- Fukaya T, Lim B, & Levine M (2016) Enhancer Control of Transcriptional Bursting. Cell 166(2):358-368. [CrossRef]
- Shao W & Zeitlinger J (2017) Paused RNA polymerase II inhibits new transcriptional initiation. Nat Genet 49(7):1045-1051. [CrossRef]
- Gressel S, Schwalb B, & Cramer P (2019) The pause-initiation limit restricts transcription activation in human cells. Nat Commun 10(1):3603. [CrossRef]
- Valekunja UK, et al. (2013) Histone methyltransferase MLL3 contributes to genome-scale circadian transcription. Proc Natl Acad Sci U S A 110(4):1554-1559. [CrossRef]
- Baxter M, et al. (2022) Circadian clock function does not require the histone methyltransferase MLL3. FASEB J 36(7):e22356. [CrossRef]
- Yu M, et al. (2015) RNA polymerase II-associated factor 1 regulates the release and phosphorylation of paused RNA polymerase II. Science 350(6266):1383-1386. [CrossRef]
- Kim J, et al. (2009) RAD6-Mediated transcription-coupled H2B ubiquitylation directly stimulates H3K4 methylation in human cells. Cell 137(3):459-471. [CrossRef]
- Kim J, et al. (2013) The n-SET domain of Set1 regulates H2B ubiquitylation-dependent H3K4 methylation. Mol Cell 49(6):1121-1133. [CrossRef]
- Yang Z, et al. (2005) Recruitment of P-TEFb for stimulation of transcriptional elongation by the bromodomain protein Brd4. Mol Cell 19(4):535-545. [CrossRef]
- Jang MK, et al. (2005) The bromodomain protein Brd4 is a positive regulatory component of P-TEFb and stimulates RNA polymerase II-dependent transcription. Mol Cell 19(4):523-534. [CrossRef]
- Conaway RC & Conaway JW (2013) The Mediator complex and transcription elongation. Biochim Biophys Acta 1829(1):69-75. [CrossRef]
- Luyties O & Taatjes DJ (2022) The Mediator kinase module: an interface between cell signaling and transcription. Trends Biochem Sci 47(4):314-327. [CrossRef]
- Li Y, Liu M, Chen LF, & Chen R (2018) P-TEFb: Finding its ways to release promoter-proximally paused RNA polymerase II. Transcription 9(2):88-94. [CrossRef]
- Contreras X, Barboric M, Lenasi T, & Peterlin BM (2007) HMBA releases P-TEFb from HEXIM1 and 7SK snRNA via PI3K/Akt and activates HIV transcription. PLoS Pathog 3(10):1459-1469. [CrossRef]
- Krebs AR, et al. (2017) Genome-wide Single-Molecule Footprinting Reveals High RNA Polymerase II Turnover at Paused Promoters. Mol Cell 67(3):411-422 e414. [CrossRef]
- Steurer B, et al. (2018) Live-cell analysis of endogenous GFP-RPB1 uncovers rapid turnover of initiating and promoter-paused RNA Polymerase II. Proc Natl Acad Sci U S A 115(19):E4368-E4376. [CrossRef]
- Buckley MS, Kwak H, Zipfel WR, & Lis JT (2014) Kinetics of promoter Pol II on Hsp70 reveal stable pausing and key insights into its regulation. Genes Dev 28(1):14-19. [CrossRef]
- Price DH (2018) Transient pausing by RNA polymerase II. Proc Natl Acad Sci U S A 115(19):4810-4812. [CrossRef]
- Kamieniarz-Gdula K & Proudfoot NJ (2019) Transcriptional Control by Premature Termination: A Forgotten Mechanism. Trends Genet 35(8):553-564. [CrossRef]
- Ehrensberger AH, Kelly GP, & Svejstrup JQ (2013) Mechanistic interpretation of promoter-proximal peaks and RNAPII density maps. Cell 154(4):713-715. [CrossRef]
- Quinodoz M, Gobet C, Naef F, & Gustafson KB (2014) Characteristic bimodal profiles of RNA polymerase II at thousands of active mammalian promoters. Genome Biol 15(6):R85. [CrossRef]
- Jonkers I, Kwak H, & Lis JT (2014) Genome-wide dynamics of Pol II elongation and its interplay with promoter proximal pausing, chromatin, and exons. Elife 3:e02407. [CrossRef]
- Chen F, Gao X, & Shilatifard A (2015) Stably paused genes revealed through inhibition of transcription initiation by the TFIIH inhibitor triptolide. Genes Dev 29(1):39-47. [CrossRef]
- Noe Gonzalez M, Sato S, Tomomori-Sato C, Conaway JW, & Conaway RC (2018) CTD-dependent and -independent mechanisms govern co-transcriptional capping of Pol II transcripts. Nat Commun 9(1):3392.
- Henriques T, et al. (2013) Stable pausing by RNA polymerase II provides an opportunity to target and integrate regulatory signals. Mol Cell 52(4):517-528. [CrossRef]
- Aoi Y, et al. (2020) NELF Regulates a Promoter-Proximal Step Distinct from RNA Pol II Pause-Release. Mol Cell 78(2):261-274 e265. [CrossRef]
- Lidschreiber M, Leike K, & Cramer P (2013) Cap completion and C-terminal repeat domain kinase recruitment underlie the initiation-elongation transition of RNA polymerase II. Mol Cell Biol 33(19):3805-3816. [CrossRef]
- Ramanathan A, Robb GB, & Chan SH (2016) mRNA capping: biological functions and applications. Nucleic Acids Res 44(16):7511-7526. [CrossRef]
- Topisirovic I, Svitkin YV, Sonenberg N, & Shatkin AJ (2011) Cap and cap-binding proteins in the control of gene expression. Wiley Interdiscip Rev RNA 2(2):277-298. [CrossRef]
- Cortazar MA, et al. (2022) Xrn2 substrate mapping identifies torpedo loading sites and extensive premature termination of RNA pol II transcription. Genes Dev 36(19-20):1062-1078. [CrossRef]
- Jiao X, Chang JH, Kilic T, Tong L, & Kiledjian M (2013) A mammalian pre-mRNA 5' end capping quality control mechanism and an unexpected link of capping to pre-mRNA processing. Mol Cell 50(1):104-115. [CrossRef]
- Wagner EJ, Tong L, & Adelman K (2023) Integrator is a global promoter-proximal termination complex. Mol Cell. [CrossRef]
- Lykke-Andersen S, et al. (2021) Integrator is a genome-wide attenuator of non-productive transcription. Mol Cell 81(3):514-529 e516. [CrossRef]
- Stein CB, et al. (2022) Integrator endonuclease drives promoter-proximal termination at all RNA polymerase II-transcribed loci. Mol Cell 82(22):4232-4245 e4211. [CrossRef]
- Rambout X, et al. (2023) PGC-1alpha senses the CBC of pre-mRNA to dictate the fate of promoter-proximally paused RNAPII. Mol Cell 83(2):186-202 e111. [CrossRef]
- Lenasi T, Peterlin BM, & Barboric M (2011) Cap-binding protein complex links pre-mRNA capping to transcription elongation and alternative splicing through positive transcription elongation factor b (P-TEFb). J Biol Chem 286(26):22758-22768. [CrossRef]
- Yudkovsky N, Ranish JA, & Hahn S (2000) A transcription reinitiation intermediate that is stabilized by activator. Nature 408(6809):225-229. [CrossRef]
- Hawley DK & Roeder RG (1987) Functional steps in transcription initiation and reinitiation from the major late promoter in a HeLa nuclear extract. J Biol Chem 262(8):3452-3461. [CrossRef]
- Zawel L, Kumar KP, & Reinberg D (1995) Recycling of the general transcription factors during RNA polymerase II transcription. Genes Dev 9(12):1479-1490. [CrossRef]
- Zhang Z, et al. (2015) Chemical perturbation of an intrinsically disordered region of TFIID distinguishes two modes of transcription initiation. Elife 4. [CrossRef]
- Tantale K, et al. (2016) A single-molecule view of transcription reveals convoys of RNA polymerases and multi-scale bursting. Nat Commun 7:12248. [CrossRef]
- Nojima T, et al. (2015) Mammalian NET-Seq Reveals Genome-wide Nascent Transcription Coupled to RNA Processing. Cell 161(3):526-540. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).