Submitted:
23 June 2023
Posted:
26 June 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Data Acquisitions and Preprocessing
2.2. Feature Engineering and Data Sampling
2.3. Machine Learning Modelling and Evaluation
2.4. Interaction Predictions
2.5. Molecular Docking
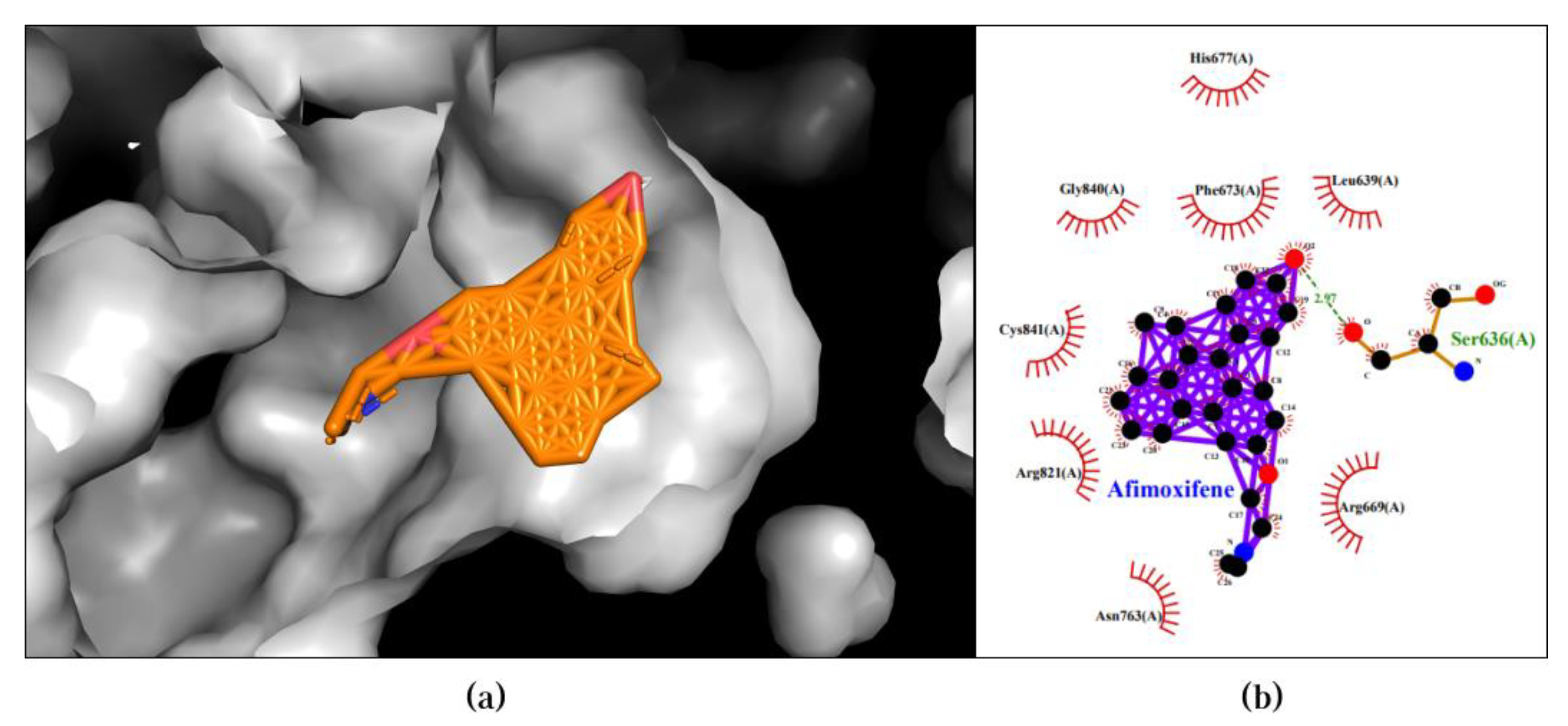
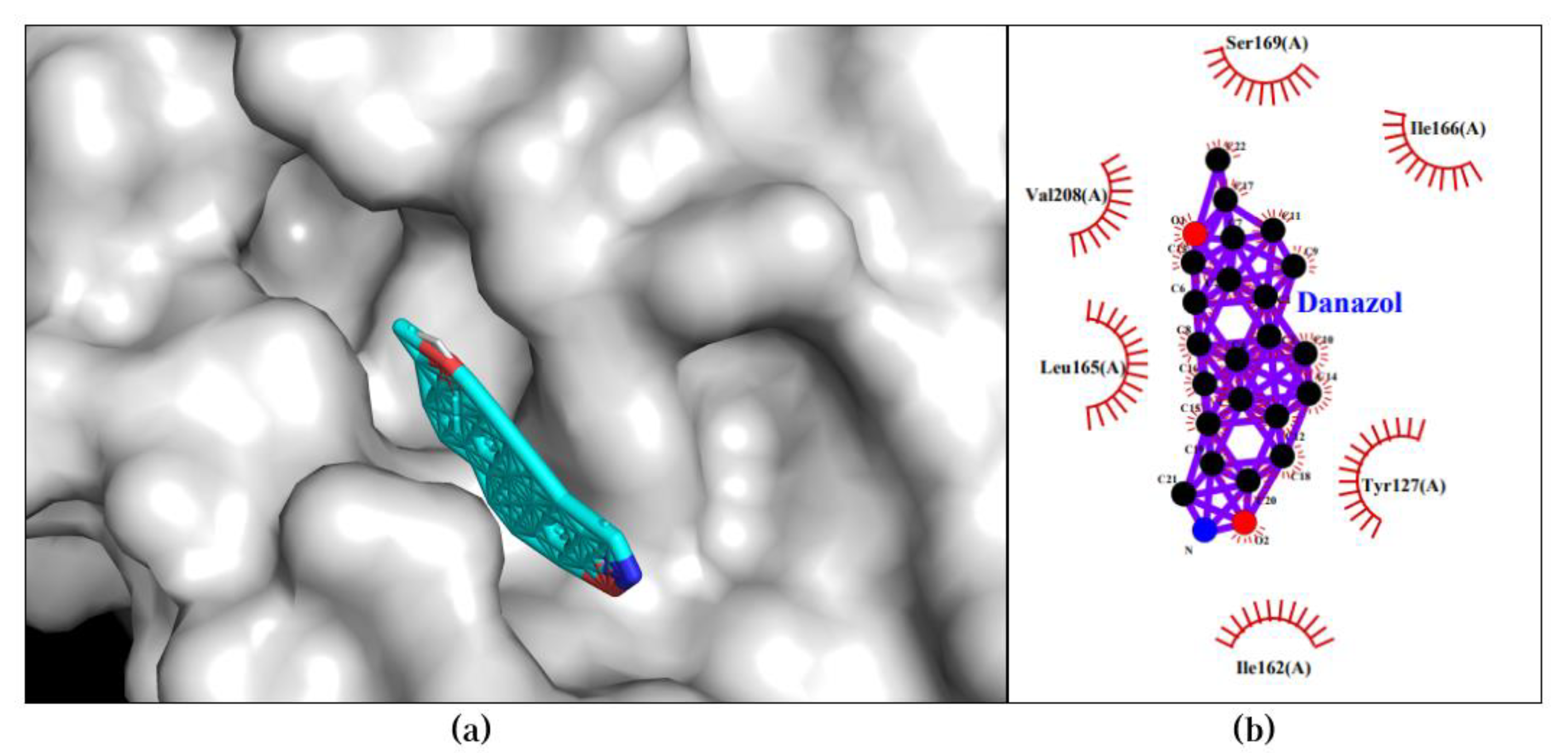
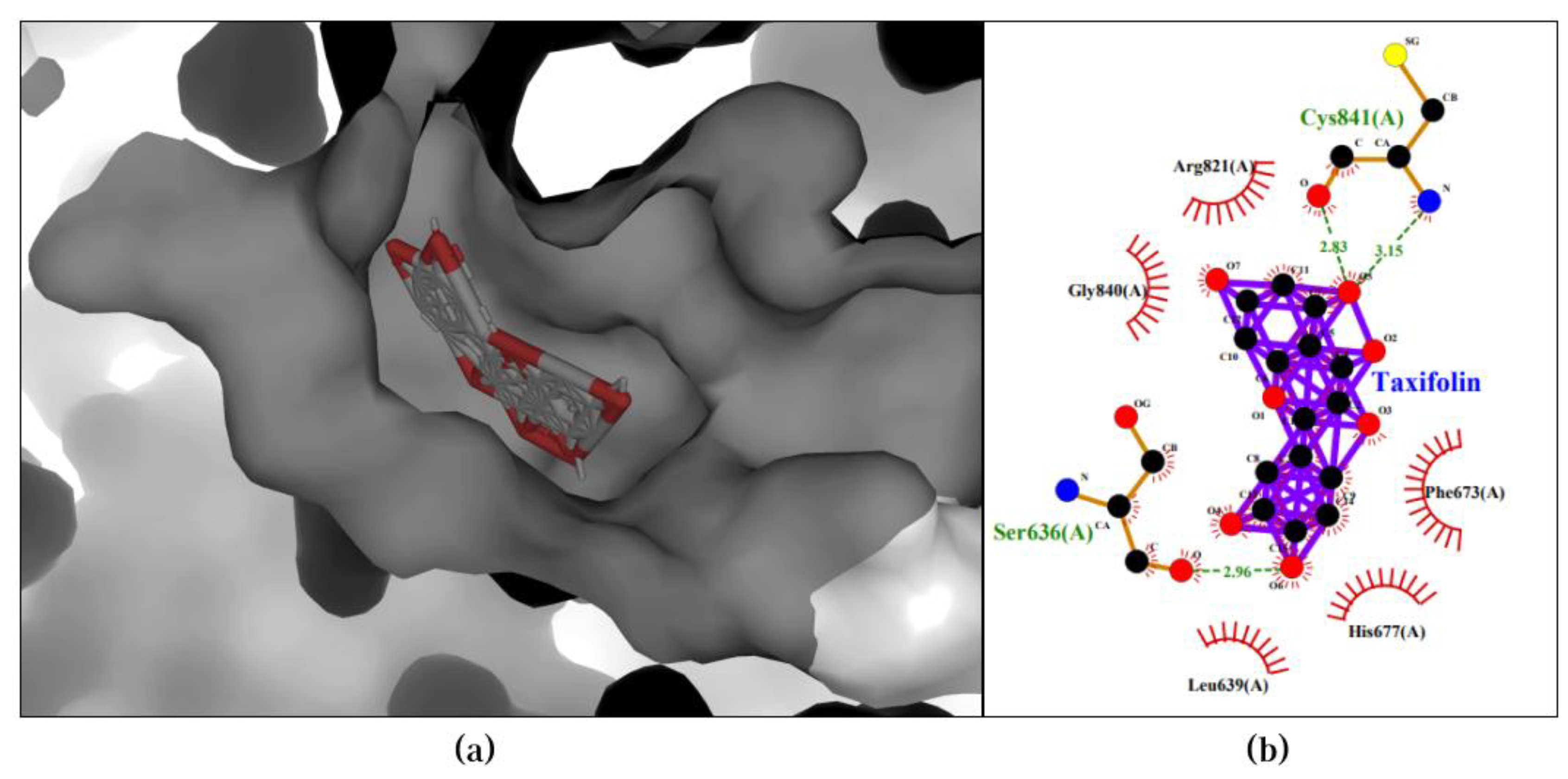
2.6. ADMET Analysis
| Compound (PubChem ID) | Afimoxifene (449459) | Danazol (28417) | Taxifolin (439533) | Terfenadine (5405) |
| Molecular Formula | C26H29NO2 | C22H27NO2 | C15H12O7 | C32H41NO2 |
| Lipinski rule-of-five | Passed | Passed | Passed | Passed |
| hERG Blockers | ++ | +++ | --- | +++ |
| H-HT | - | +++ | --- | -- |
| AMES Toxicity | --- | --- | + | --- |
| Rat Oral Acute Toxicity | + | + | -- | --- |
| Carcinogencity | -- | +++ | --- | --- |
| Eye Corrossion | --- | --- | --- | --- |
| Respiratory Toxicity | ++ | +++ | --- | +++ |
| CaCO2 permeability | -4,46 | -4,88 | -6,06 | -5,274 |
| MDCK permeability | ||||
| Intestinal Absorption | --- | --- | --- | --- |
| PPB | 95,63% | 98,46% | 93,23% | 74,04% |
| VD | 1,745 | 3,01 | 0,56 | 2,28 |
| CYP1A2 inhibitor | ++ | +++ | - | --- |
| CL | 10,08 | 5,265 | 12,29 | 5,11 |
| T1/2 | 0,108 | 0,12 | 0,76 | 0,005 |
3. Discussion
3.1. Machine Learning-Based DTI Analysis Reveals the Superiority of CDF Using ECFP-Aaindex1 as a Feature Combination
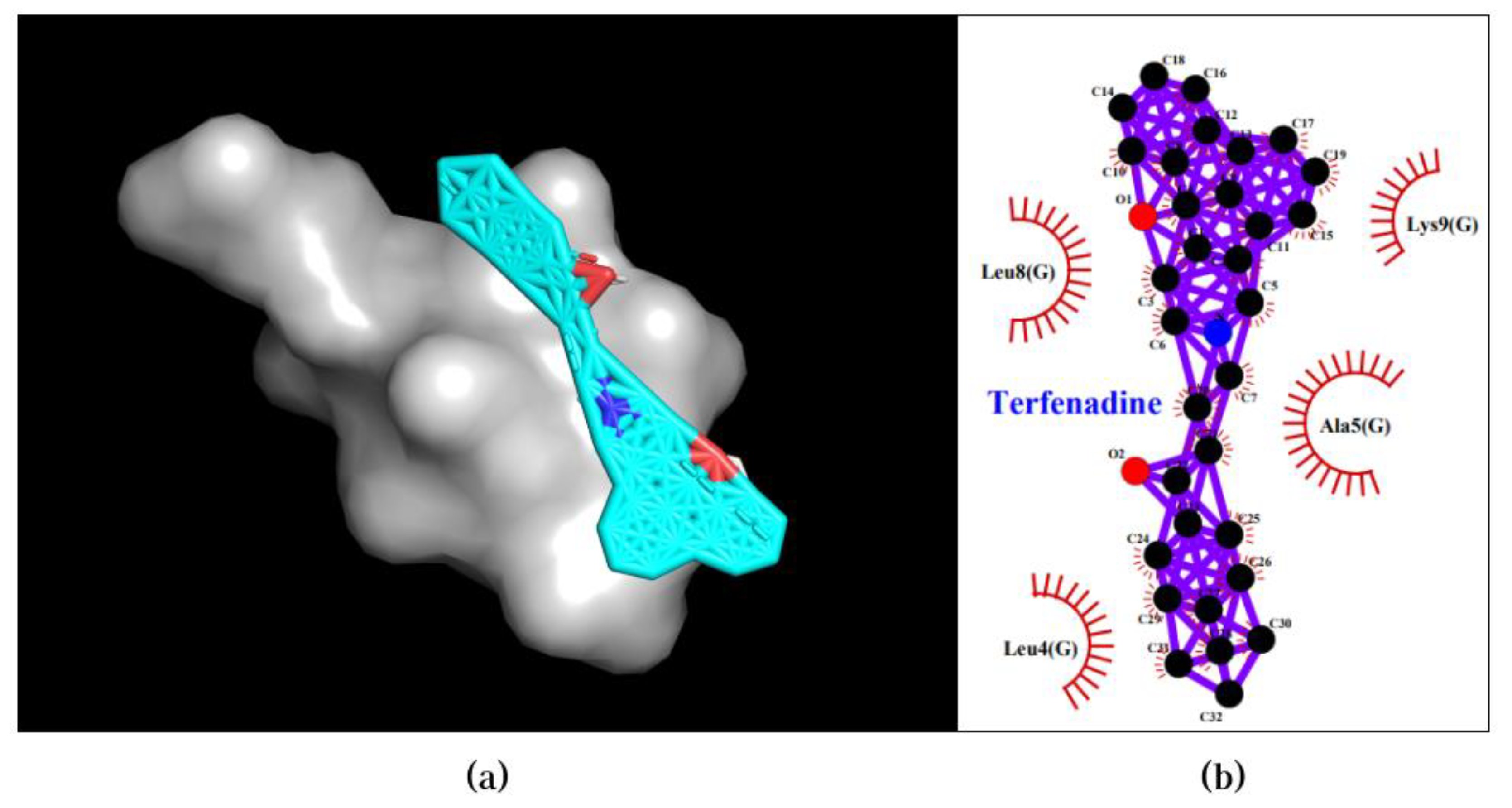
3.2. Molecular Docking Validates Predictions by CDF and Enhances Understanding of Compound-Protein Interaction Mechanisms
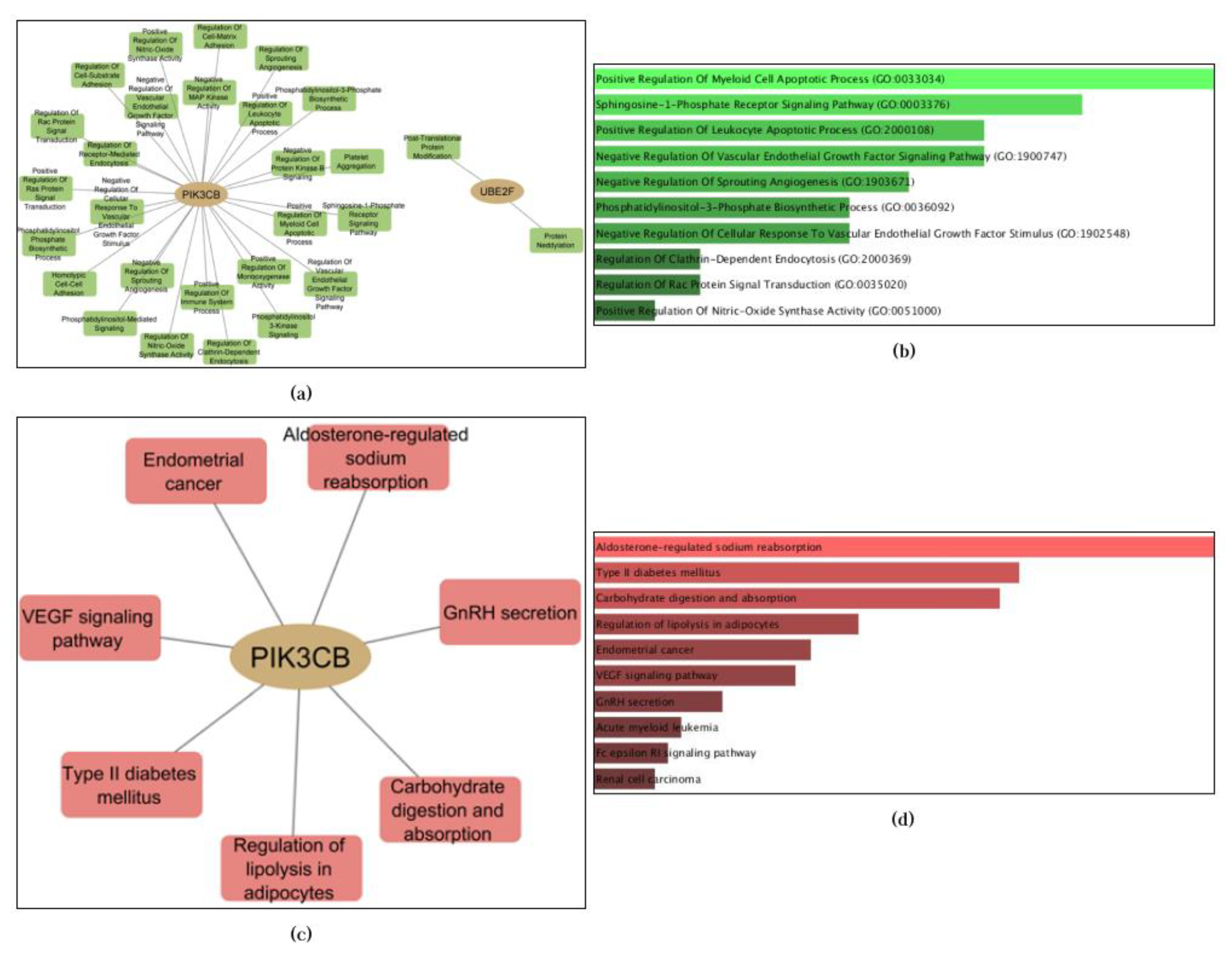
3.3. Enrichment Analysis of Validated Genes/Proteins Reveals Potential Biological Processes and Pathways in Cancer
3.4. ADMET Analysis Reveals Insights into Druglikeliness and Bioavailability of Selected Bioactive Compounds
4. Materials and Methods
4.1. Data Acquisition
4.2. Data Preprocessing
4.3. Feature Engineering
4.4. Data Sampling
4.5. Machine Learning Modelling
4.6. Model Evaluations
4.7. Interaction Predictions
4.8. Molecular Docking
4.9. Absorption, Distribution, Metabolism, dan Toxicity (ADMET) Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- W.-T. Lee et al., “Black sea cucumber (Holothuria atra Jaeger, 1833) rescues Pseudomonas aeruginosa -infected Caenorhabditis elegans via reduction of pathogen virulence factors and enhancement of host immunity,” Food Funct., vol. 10, no. 9, pp. 5759–5767, 2019. [CrossRef]
- R. Pangestuti and Z. Arifin, “Medicinal and health benefit effects of functional sea cucumbers,” J. Tradit. Complement. Med., vol. 8, no. 3, pp. 341–351, Jul. 2018. [CrossRef]
- S. Bordbar, F. Anwar, and N. Saari, “High-Value Components and Bioactives from Sea Cucumbers for Functional Foods—A Review,” Mar. Drugs, vol. 9, no. 10, pp. 1761–1805, Oct. 2011. [CrossRef]
- S. A. Kristina, D. Endarti, and H. Aditama, “Prediction of Productivity Costs Related to Cervical Cancer Mortality in Indonesia 2018,” Malaysian J. Med. Sci., vol. 29, no. 1, pp. 138–144, Feb. 2022. Pusdatin Kemenkes RI, “Beban Kanker di Indonesia,” 2019. [CrossRef]
- A. S. Rifaioglu, H. Atas, M. J. Martin, et al., “Recent applications of deep learning and machine intelligence on in silico drug discovery: methods, tools and databases,” Briefings …, 2019, Available online: https://academic.oup.com/bib/article-abstract/20/5/1878/5062947.
- G. Bocci, E. Carosati, P. Vayer, A. Arrault, S. Lozano, and G. Cruciani, “ADME-Space: a new tool for medicinal chemists to explore ADME properties,” Sci. Rep., vol. 7, no. 1, p. 6359, Jul. 2017. [CrossRef]
- C. Cava and I. Castiglioni, “Integration of Molecular Docking and In Vitro Studies: A Powerful Approach for Drug Discovery in Breast Cancer,” Appl. Sci., vol. 10, no. 19, p. 6981, Oct. 2020. [CrossRef]
- M. T. F. Calangian and V. P. C. Magboo, “Predicting Drug-Target Interaction (DTI) based on Machine Learning with Lasso Dimensionality Reduction and SMOTE from Protein Sequence and Drug Fingerprint,” in 2022 International Conference on Electrical, Computer and Energy Technologies (ICECET), Jul. 2022, pp. 1–6. [CrossRef]
- R. Chen, X. Liu, S. Jin, J. Lin, and J. Liu, “Machine Learning for Drug-Target Interaction Prediction,” Molecules, vol. 23, no. 9, p. 2208, Aug. 2018. [CrossRef]
- A. Ezzat, M. Wu, X. Li, and C.-K. Kwoh, “Computational Prediction of Drug-Target Interactions via Ensemble Learning,” in Computational Methods for Drug Repurposing, 2019, pp. 239–254.
- B. Li, et al. “Machine Learning Models Combined with Virtual Screening and Molecular Docking to Predict Human Topoisomerase I Inhibitors,” Molecules, vol. 24, no. 11, p. 2107, Jun. 2019. [CrossRef]
- N. Srivastava, P. Garg, P. Srivastava, and P. K. Seth, “A molecular dynamics simulation study of the ACE2 receptor with screened natural inhibitors to identify novel drug candidate against COVID-19.,” PeerJ, vol. 9, p. e11171, Apr. 2021. [CrossRef]
- L. L. G. Ferreira and A. D. Andricopulo, “ADMET modeling approaches in drug discovery,” Drug Discov. Today, vol. 24, no. 5, pp. 1157–1165, May 2019. [CrossRef]
- K. E. Knoll, M. M. van der Walt, and D. T. Loots, “In Silico Drug Discovery Strategies Identified ADMET Properties of Decoquinate RMB041 and Its Potential Drug Targets against Mycobacterium tuberculosis,” Microbiol. Spectr., vol. 10, no. 2, Apr. 2022. [CrossRef]
- K. Rajagopal et al., “In Silico Drug Design of Anti-Breast Cancer Agents,” Molecules, vol. 28, no. 10, p. 4175, May 2023. [CrossRef]
- H. R. M. Rashdan, M. El-Naggar, and A. H. Abdelmonsef, “Synthesis, Molecular Docking Studies and In Silico ADMET Screening of New Heterocycles Linked Thiazole Conjugates as Potent Anti-Hepatic Cancer Agents,” Molecules, vol. 26, no. 6, p. 1705, Mar. 2021. [CrossRef]
- Y. Wang et al., “RoFDT: Identification of Drug–Target Interactions from Protein Sequence and Drug Molecular Structure Using Rotation Forest,” Biology (Basel)., vol. 11, no. 5, p. 741, May 2022. [CrossRef]
- Y. Chu et al., “DTI-CDF: a cascade deep forest model towards the prediction of drug-target interactions based on hybrid features,” Brief. Bioinform., vol. 22, no. 1, pp. 451–462, Jan. 2021. [CrossRef]
- K. Nasution, S. H. Wijaya, and W. A. Kusuma, “Prediction of Drug-Target Interaction on Jamu Formulas using Machine Learning Approaches,” in 2019 International Conference on Advanced Computer Science and information Systems (ICACSIS), Oct. 2019, pp. 169–174. [CrossRef]
- A. Fadli, A. Annisa, and W. A. Kusuma, “Prediction of Drug-Target Interaction Using Random Forest in Coronavirus Disease 2019 Case,” Bioinforma. Biomed. Res. J., vol. 4, no. 1, pp. 1–7, Nov. 2021. [CrossRef]
- Y. Ding, J. Tang, F. Guo, and Q. Zou, “Identification of drug–target interactions via multiple kernel-based triple collaborative matrix factorization,” Brief. Bioinform., vol. 23, no. 2, Mar. 2022. [CrossRef]
- Z. Cheng, C. Yan, F. X. Wu, and J. Wang, “Drug-target interaction prediction using multi-head self-attention and graph attention network,” IEEE/ACM Trans. …, 2021, Available online: https://ieeexplore.ieee.org/abstract/document/9425008/.
- H. Lee, J. Palm, S. M. Grimes, and H. P. Ji, “The Cancer Genome Atlas Clinical Explorer: a web and mobile interface for identifying clinical–genomic driver associations,” Genome Med., vol. 7, no. 1, p. 112, Dec. 2015. [CrossRef]
- M. Uhlén et al., “A Human Protein Atlas for Normal and Cancer Tissues Based on Antibody Proteomics,” Mol. Cell. Proteomics, vol. 4, no. 12, pp. 1920–1932, Dec. 2005. [CrossRef]
- V. Premachandran and R. Kakarala, “Measuring the effectiveness of bad pixel detection algorithms using the ROC curve,” IEEE Trans. Consum. Electron., vol. 56, no. 4, pp. 2511–2519, Nov. 2010. [CrossRef]
- J. Jumper et al., “Highly accurate protein structure prediction with AlphaFold,” Nature, vol. 596, no. 7873, pp. 583–589, Aug. 2021. [CrossRef]
- N. M. O’Boyle, M. Banck, C. A. James, C. Morley, T. Vandermeersch, and G. R. Hutchison, “Open Babel: An open chemical toolbox,” J. Cheminform., vol. 3, no. 1, p. 33, Dec. 2011. [CrossRef]
- O. Trott and A. J. Olson, “AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading,” J. Comput. Chem., p. NA-NA, 2009. [CrossRef]
- S. S. Islam, M. S. Haque, M. S. U. Miah, T. Bin Sarwar, and R. Nugraha, “Application of machine learning algorithms to predict the thyroid disease risk: an experimental comparative study,” PeerJ Comput. Sci., vol. 8, p. e898, Mar. 2022. [CrossRef]
- N. Nigar, M. Umar, M. K. Shahzad, S. Islam, and D. Abalo, “A Deep Learning Approach Based on Explainable Artificial Intelligence for Skin Lesion Classification,” IEEE Access, vol. 10, pp. 113715–113725, 2022. [CrossRef]
- V. Malathi, M. P. Gopinath, M. Kumar, S. Bhushan, and S. Jayaprakash, “Enhancing the Paddy Disease Classification by Using Cross-Validation Strategy for Artificial Neural Network over Baseline Classifiers,” J. Sensors, vol. 2023, pp. 1–13, Apr. 2023. [CrossRef]
- L. Nie, C. Li, F. Marzani, H. Wang, F. Thibouw, and A. B. Grayeli, “Classification of Wideband Tympanometry by Deep Transfer Learning With Data Augmentation for Automatic Diagnosis of Otosclerosis,” IEEE J. Biomed. Heal. Informatics, vol. 26, no. 2, pp. 888–897, Feb. 2022. [CrossRef]
- Z.-H. Zhou and J. Feng, “Deep forest,” Natl. Sci. Rev., vol. 6, no. 1, pp. 74–86, Jan. 2019. [CrossRef]
- Y. Liu, X. Yang, J. Gan, S. Chen, Z.-X. Xiao, and Y. Cao, “CB-Dock2: improved protein–ligand blind docking by integrating cavity detection, docking and homologous template fitting,” Nucleic Acids Res., vol. 50, no. W1, pp. W159–W164, Jul. 2022. [CrossRef]
- X.-Y. Meng, H.-X. Zhang, M. Mezei, and M. Cui, “Molecular Docking: A Powerful Approach for Structure-Based Drug Discovery,” Curr. Comput. Aided-Drug Des., vol. 7, no. 2, pp. 146–157, Jun. 2011. [CrossRef]
- N. M. Hassan, A. A. Alhossary, Y. Mu, and C.-K. Kwoh, “Protein-Ligand Blind Docking Using QuickVina-W With Inter-Process Spatio-Temporal Integration,” Sci. Rep., vol. 7, no. 1, p. 15451, Nov. 2017. [CrossRef]
- M.D. Anderson Cancer Center, “Afimoxifene in Reducing the Risk of Breast Cancer in Women With Mammographically Dense Breast,” 2023. https://beta.clinicaltrials.gov/study/NCT03063619 (accessed May 29, 2023).
- Y.-T. Chang et al., “Danazol mediates collateral sensitivity via STAT3/Myc related pathway in multidrug-resistant cancer cells,” Sci. Rep., vol. 9, no. 1, p. 11628, Aug. 2019. [CrossRef]
- D. Turck et al., “Scientific Opinion on taxifolin-rich extract from Dahurian Larch (Larix gmelinii),” EFSA J., vol. 15, no. 2, Feb. 2017. [CrossRef]
- S. Begum, A. Banerjee, and B. De, “Antioxidant and Enzyme Inhibitory Properties of Mangifera indica leaf Extract,” Nat. Prod. J., vol. 10, no. 4, pp. 384–394, Aug. 2020. [CrossRef]
- V. S. Rogovskiĭ et al., “Antiproliferative and antioxidant activity of new dihydroquercetin derivatives,” Eksp. Klin. Farmakol., vol. 73, no. 9, pp. 39–42, Sep. 2010, Available online: http://www.ncbi.nlm.nih.gov/pubmed/21086652.
- J. K. Racha et al., “Substrate Dependent Inhibition Profiles of Fourteen Drugs on CYP3A4 Activity Measured by A High Throughput LCMS/MS Method with Four Probe Drugs, Midazolam, Testosterone, Nifedipine and Terfenadine,” Drug Metab. Pharmacokinet., vol. 18, no. 2, pp. 128–138, 2003. [CrossRef]
- Z. Wu, S.-N. Jang, S.-Y. Park, N. M. Phuc, and K.-H. Liu, “Inhibitory Potential of Bilobetin Against CYP2J2 Activities in Human Liver Microsomes,” Mass Spectrom. Lett., vol. 11, no. 4, pp. 113–117, 2020. UL - https://msletters/v.11/4/113/7217. [CrossRef]
- W. Huang et al., “Terfenadine resensitizes doxorubicin activity in drug-resistant ovarian cancer cells via an inhibition of CaMKII/CREB1 mediated ABCB1 expression,” Front. Oncol., vol. 12, Nov. 2022. [CrossRef]
- P. Shannon et al., “Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks,” Genome Res., vol. 13, no. 11, pp. 2498–2504, Nov. 2003. [CrossRef]
- Williams and, S. Halappanavar, “Application of biclustering of gene expression data and gene set enrichment analysis methods to identify potentially disease causing nanomaterials,” Beilstein J. Nanotechnol., vol. 6, pp. 2438–2448, Dec. 2015. [CrossRef]
- S. G. C. Mestrum et al., “Proliferative and anti-apoptotic fractions in maturing hematopoietic cell lineages and their role in homeostasis of normal bone marrow,” Cytom. Part A, vol. 101, no. 7, pp. 552–563, Jul. 2022. [CrossRef]
- X.-Q. Xu, C.-M. Huang, Y.-F. Zhang, L. Chen, H. Cheng, and J.-M. Wang, “S1PR1 mediates anti-apoptotic/pro-proliferative processes in human acute myeloid leukemia cells,” Mol. Med. Rep., vol. 14, no. 4, pp. 3369–3375, Oct. 2016. [CrossRef]
- A. Lother, F. Jaisser, and U. O. Wenzel, “Emerging fields for therapeutic targeting of the aldosterone–mineralocorticoid receptor signaling pathway,” Br. J. Pharmacol., vol. 179, no. 13, pp. 3099–3102, Jul. 2022. [CrossRef]
- W. Ware, J. J. Harris, T. L. Slatter, H. E. Cunliffe, and F. J. McDonald, “The epithelial sodium channel has a role in breast cancer cell proliferation,” Breast Cancer Res. Treat., vol. 187, no. 1, pp. 31–43, May 2021. [CrossRef]
- E. Ceraudo et al., “Direct evidence that the GPCR CysLTR2 mutant causative of uveal melanoma is constitutively active with highly biased signaling,” J. Biol. Chem., vol. 296, p. 100163, Jan. 2021. [CrossRef]
- D. F. Akin-Bali, “Bioinformatics analysis of GNAQ, GNA11, BAP1, SF3B1,SRSF2, EIF1AX, PLCB4, and CYSLTR2 genes and their role in the pathogenesis of Uveal Melanoma,” Ophthalmic Genet., vol. 42, no. 6, pp. 732–743, Nov. 2021. [CrossRef]
- M. Venerito et al., “Leukotriene receptor expression in esophageal squamous cell cancer and non-transformed esophageal epithelium: a matched case control study,” BMC Gastroenterol., vol. 16, no. 1, p. 85, Dec. 2016. [CrossRef]
- W. Liu, C. Zhang, X. Gong, W. Liao, J. Xu, and X. Zhang, “Prognostic value of immune-related genes in laryngeal squamous cell carcinoma,” Transl. Cancer Res., vol. 9, no. 10, pp. 6287–6302, Oct. 2020. [CrossRef]
- P. M. Natarajan, V. R. Umapathy, A. Murali, and B. Swamikannu, “Computational simulations of identified marine-derived natural bioactive compounds as potential inhibitors of oral cancer,” Futur. Sci. OA, vol. 8, no. 3, Mar. 2022. [CrossRef]
- M. R. R. Rahardhian, Y. Susilawati, I. Musfiroh, R. M. Febriyanti, Muchtaridi, and S. A. Sumiwi, “In Silico Study of Bioactive Compounds from Sungkai (Peronema Canescens) As Immunomodulator,” Int. J. Appl. Pharm., vol. 14, no. 4, pp. 135–141, Nov. 2022. [CrossRef]
- N. S. Aini et al., “In Silico Screening of Bioactive Compounds from Garcinia mangostana L. Against SARS-CoV-2 via Tetra Inhibitors,” Pharmacogn. J., vol. 14, no. 5, pp. 575–579, Nov. 2022. [CrossRef]
- M. J. Windley, W. Lee, J. I. Vandenberg, and A. P. Hill, “The Temperature Dependence of Kinetics Associated with Drug Block of hERG Channels Is Compound-Specific and an Important Factor for Proarrhythmic Risk Prediction,” Mol. Pharmacol., vol. 94, no. 1, pp. 760–769, Jul. 2018. [CrossRef]
- J. M. Kratz, U. Grienke, O. Scheel, S. A. Mann, and J. M. Rollinger, “Natural products modulating the hERG channel: heartaches and hope,” Nat. Prod. Rep., vol. 34, no. 8, pp. 957–980, 2017. [CrossRef]
- H. Park, B. Kim, J.-H. Oh, Y. Kim, and Y.-J. Lee, “First-pass Metabolism of Decursin, a Bioactive Compound of Angelica gigas, in Rats,” Planta Med., vol. 78, no. 09, pp. 909–913, Jun. 2012. [CrossRef]
- J. Kim, M. Han, and W. K. Jeon, “Acute and Subacute Oral Toxicity of Mumefural, Bioactive Compound Derived from Processed Fruit of Prunus mume Sieb. et Zucc., in ICR Mice,” Nutrients, vol. 12, no. 5, p. 1328, May 2020. [CrossRef]
- J. Yu and S.-J. Choi, “Particle Size and Biological Fate of ZnO Do Not Cause Acute Toxicity, but Affect Toxicokinetics and Gene Expression Profiles in the Rat Livers after Oral Administration,” Int. J. Mol. Sci., vol. 22, no. 4, p. 1698, Feb. 2021. [CrossRef]
- Y. H. Bai et al., “Assessment of a Bioactive Compound for Its Potential Antiinflammatory Property by Tight Junction Permeability,” Phyther. Res., vol. 19, no. 12, pp. 1009–1012, Dec. 2005. [CrossRef]
- K. S. Roy et al., “Optimizing a High-Throughput Solid-Phase Microextraction System to Determine the Plasma Protein Binding of Drugs in Human Plasma,” Anal. Chem., vol. 93, no. 32, pp. 11061–11065, Aug. 2021. [CrossRef]
- C. Fărcaș et al., “An Update Regarding the Bioactive Compound of Cereal By-Products: Health Benefits and Potential Applications,” Nutrients, vol. 14, no. 17, p. 3470, Aug. 2022. [CrossRef]
- F. Bachmann et al., “Metamizole is a Moderate Cytochrome P450 Inducer Via the Constitutive Androstane Receptor and a Weak Inhibitor of CYP1A2,” Clin. Pharmacol. Ther., vol. 109, no. 6, pp. 1505–1516, Jun. 2021. [CrossRef]
- T. J. K. Kaartinen et al., “Effect of High-Dose Esomeprazole on CYP1A2, CYP2C19, and CYP3A4 Activities in Humans: Evidence for Substantial and Long-lasting Inhibition of CYP2C19,” Clin. Pharmacol. Ther., vol. 108, no. 6, pp. 1254–1264, Dec. 2020. [CrossRef]
- I. R. Fadhlia, M. Nurilmala, and T. Nurhayati, “Karakterisasi Konsentrat dan Hidrolisat Protein Teripang Keling (Holothuria atra) dan Potensinya sebagai Imunomodulator,” Institut Pertanian Bogor, 2017.
- T. Liu, Y. Lin, X. Wen, R. N. Jorissen, and M. K. Gilson, “BindingDB: a web-accessible database of experimentally determined protein-ligand binding affinities.,” Nucleic Acids Res., vol. 35, no. Database issue, pp. D198-201, Jan. 2007. [CrossRef]
- B. Zhao et al., “DescribePROT: database of amino acid-level protein structure and function predictions,” Nucleic Acids Res., vol. 49, no. D1, pp. D298–D308, Jan. 2021. [CrossRef]
- S. Kawashima, P. Pokarowski, M. Pokarowska, A. Kolinski, T. Katayama, and M. Kanehisa, “AAindex: amino acid index database, progress report 2008.,” Nucleic Acids Res., vol. 36, no. Database issue, pp. D202-5, Jan. 2008. [CrossRef]
- K.-C. Chou, “Prediction of protein cellular attributes using pseudo-amino acid composition,” Proteins Struct. Funct. Genet., vol. 43, no. 3, pp. 246–255, May 2001. [CrossRef]
- R. Kumar et al., “An in silico platform for predicting, screening and designing of antihypertensive peptides,” Sci. Rep., vol. 5, no. 1, p. 12512, Jul. 2015. [CrossRef]
- D. Rogers and M. Hahn, “Extended-Connectivity Fingerprints,” J. Chem. Inf. Model., vol. 50, no. 5, pp. 742–754, May 2010. [CrossRef]
- J. Klekota and F. P. Roth, “Chemical substructures that enrich for biological activity,” Bioinformatics, vol. 24, no. 21, pp. 2518–2525, Nov. 2008. [CrossRef]
- H. L. Morgan, “The Generation of a Unique Machine Description for Chemical Structures-A Technique Developed at Chemical Abstracts Service.,” J. Chem. Doc., vol. 5, no. 2, pp. 107–113, May 1965. [CrossRef]
- Y. Wang, J. Xiao, T. O. Suzek, J. Zhang, J. Wang, and S. H. Bryant, “PubChem: a public information system for analyzing bioactivities of small molecules,” Nucleic Acids Res., vol. 37, no. Web Server, pp. W623–W633, Jul. 2009. [CrossRef]
- F. Pedregosa et al., “Scikit-learn: Machine Learning in Python,” J. Mach. Learn. Res., vol. 12, no. 85, pp. 2825–2830, 2011, Available online: http://jmlr.org/papers/v12/pedregosa11a.html.
- T. Chen and C. Guestrin, “XGBoost,” in Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Aug. 2016, pp. 785–794. [CrossRef]
- G. Ke et al., “LightGBM: A Highly Efficient Gradient Boosting Decision Tree,” in Advances in Neural Information Processing Systems, 2017, vol. 30, Available online: https://proceedings.neurips.cc/paper_files/paper/2017/file/6449f44a102fde848669bdd9eb6b76fa-Paper.pdf.
- G. M. Morris et al., “AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility,” J. Comput. Chem., vol. 30, no. 16, pp. 2785–2791, Dec. 2009. [CrossRef]
- Schrödinger, LLC, “The {PyMOL} Molecular Graphics System, Version~1.8,” Nov. 2015.
- R. A. Laskowski and M. B. Swindells, “LigPlot+: Multiple Ligand–Protein Interaction Diagrams for Drug Discovery,” J. Chem. Inf. Model., vol. 51, no. 10, pp. 2778–2786, Oct. 2011. [CrossRef]
- S. Kumari and P. Kumar, “Design and Computational Analysis of an MMP9 Inhibitor in Hypoxia-Induced Glioblastoma Multiforme.” ACS omega, vol. 8, no. 11, pp. 10565–10590, Mar. 2023. [CrossRef]
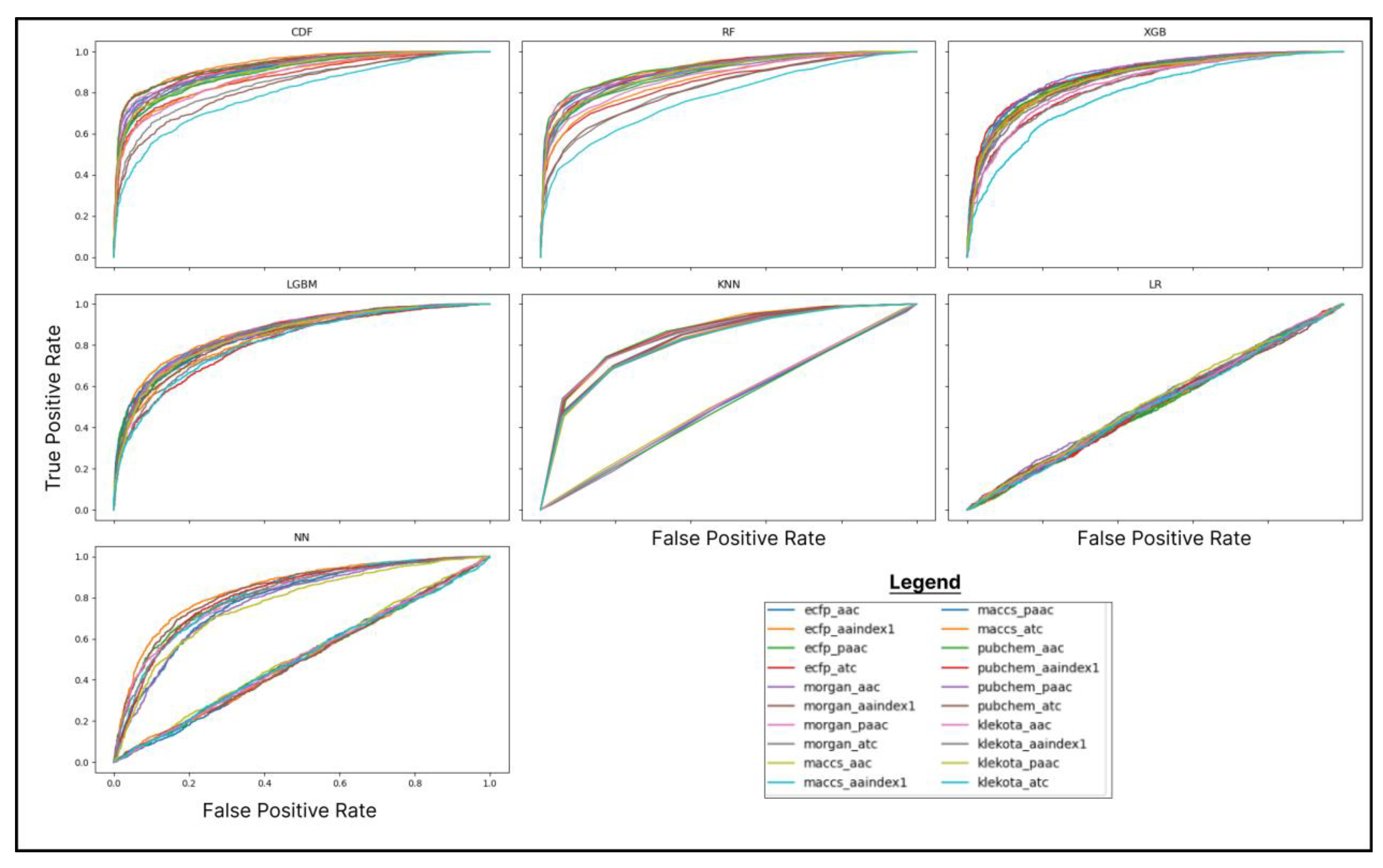
| Compound Name (PubChem ID) |
Protein/Gene Name (Uniprot ID) |
Average Confidence Score |
|---|---|---|
| Meclizine (4034) | UBE2F (Q969M7) | 0.8445 |
| Taxifolin (439533) | PIK3CB (P42338) | 0.790625 |
| Terfenadine (5405) | UBE2F (Q969M7) | 0.856 |
| Afimoxifene (449459) | PIK3CB (P42338) | 0.892125 |
| Selegiline (26757) | UBE2F (Q969M7) | 0.87525 |
| Phencyclidine (6468) | UBE2F (Q969M7) | 0.89625 |
| Danazol (28417) | CYSLTR2 (Q9NS75) | 0.80275 |
| Compound Name | Protein/Gene Name | Best Binding Affinity |
|---|---|---|
| Afimoxifene | PIK3CB | -12.7 |
| Danazol | CYSLTR2 | -12.3 |
| Taxifolin | PIK3CB | -10.0 |
| Terfenadine | UBE2F | -6.6 |
| Phencyclidine | UBE2F | -4.6 |
| Meclizine | UBE2F | -4.3 |
| Selegiline | UBE2F | -3.4 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




