Submitted:
15 June 2023
Posted:
19 June 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
- We propose a new concept of alternating different deep learning blocks to construct a unique architecture that demonstrates how the aggregation of different block types could help learn better representations.
- We propose a joint self-supervised method of inpainting and shuffling as a pretraining task. That enforces the models to learn a challenging proxy task during pretraining, which helps improve the finetuning performance.
- We introduce a new clinical problem to the medical image analysis community; i.e., the segmentation of the RIMA and perivascular space, which can be helpful in a range of clinical studies for cardiovascular disease prognosis.
- Finally, we provide a thorough clinical evaluation on external datasets through intra-observer variability, inter-observer variability, model-versus-clinician analysis, and post-model segmentation refinement analysis with expert clinicians.
2. Materials and Methods
2.1. Building Blocks
2.1.1. SE block
2.1.2. Swin block
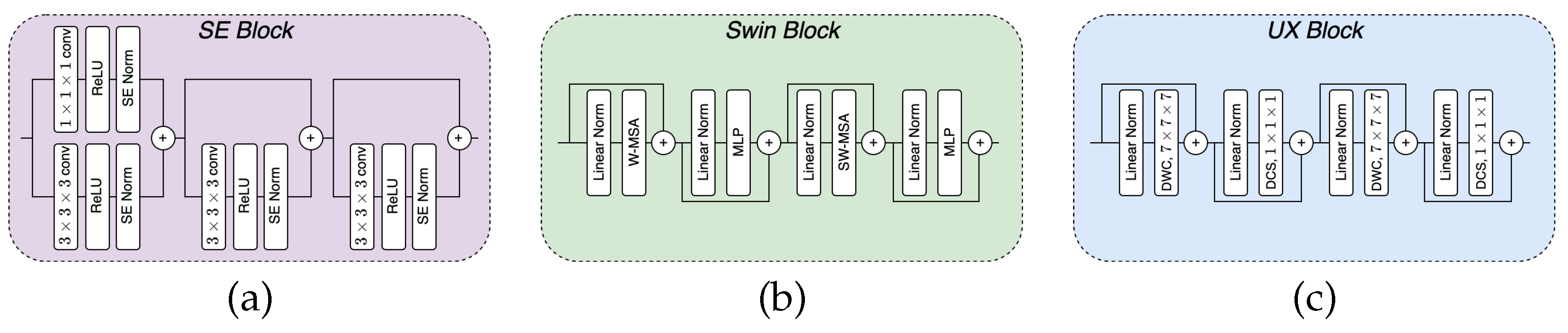
2.1.3. UX block
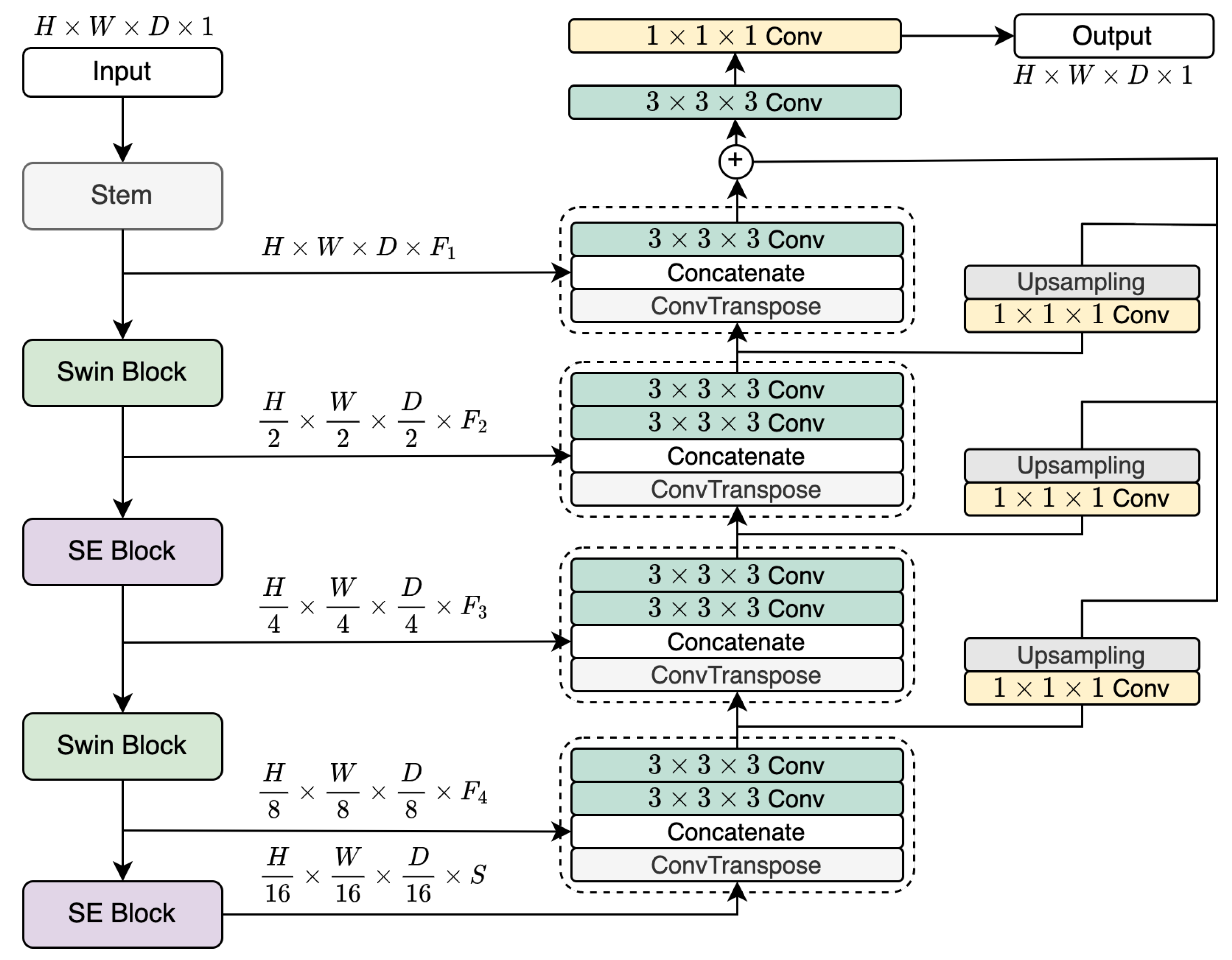
2.2. LegoNet Architecture
2.3. Alternating Composition of LegoNet
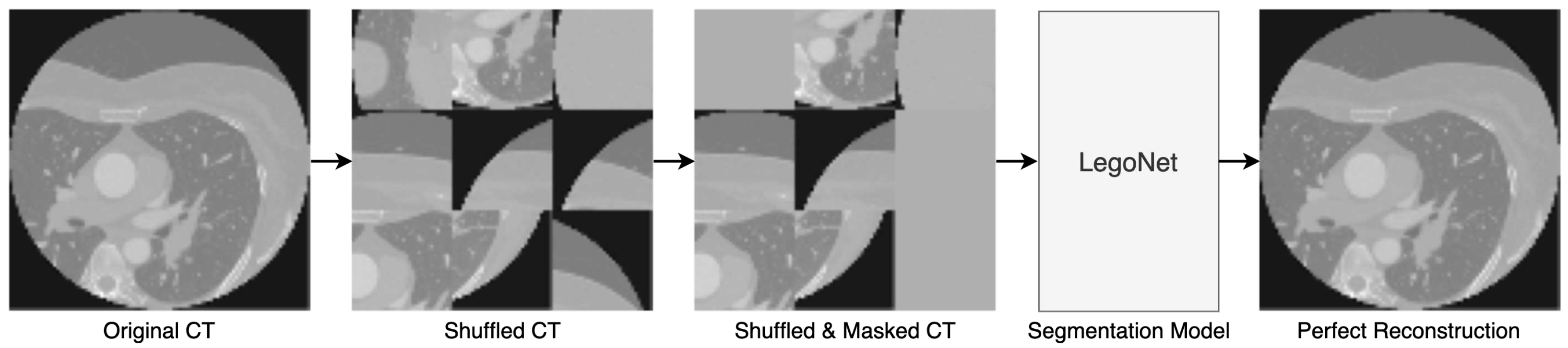
2.4. Joint SSL Pretraining
3. Dataset and Preprocessing
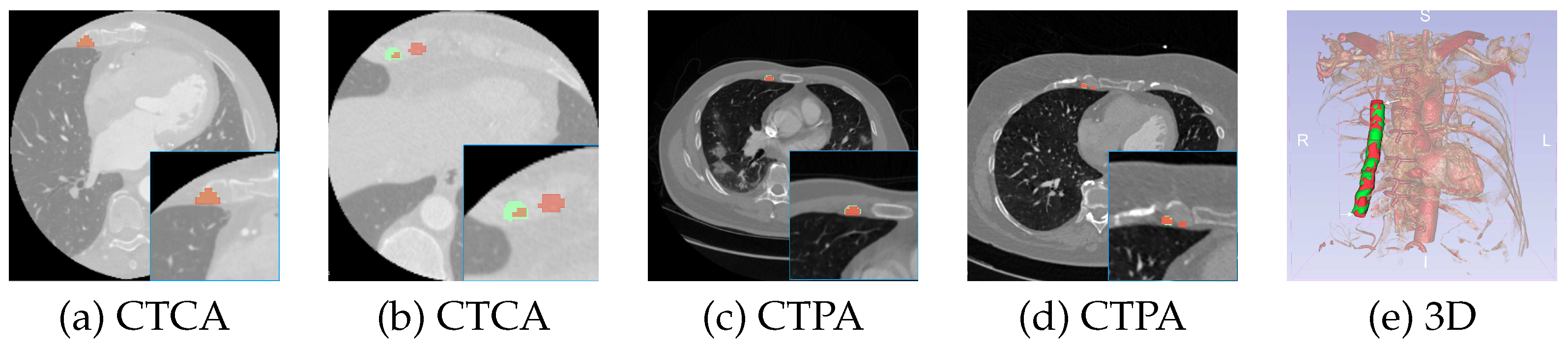
3.1. CTCA
3.2. CTPA
3.3. External Datasets
4. Experimental Setup
5. Results
5.1. CTCA
5.2. CTPA
5.3. Inter- and Intra-observer Variability
5.4. External Cohorts
| Variability | Years of XP | DSC | External | Number of cases | DSC |
| Intra | 3 | 0.804 | Cohort 1 | 40 | 0.935 |
| Inter | 1 | 0.761 | Cohort 2 | 40 | 0.942 |
| Model vs Human | NA | 0.733 | Cohort 3 | 60 | 0.947 |
5.5. SSL
6. Discussion
7. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CNN | Convolutional neural networks |
| CTA | Computed tomography angiography |
| CTCA | Computed tomography coronary angiography |
| CTPA | Computed tomography pulmonary angiography |
| DCS | Depth-wise convolutional scaling |
| DL | Deep learning |
| DSC | Dice similarity coefficient |
| DWC | Depth-wise convolution |
| FLOPs | Floating point operations |
| IN | Instance normalization |
| MSA | Multi-head self-attention |
| ORFAN | The Oxford risk factors and non-invasive imaging study |
| ReLU | Rectified linear activation unit |
| RIMA | Right internal mammary artery |
| SE | Squeeze-and-excitation |
| ViT | Vision transformer |
References
- Çiçek, Ö.; Abdulkadir, A.; Lienkamp, S.S.; Brox, T.; Ronneberger, O. 3D U-Net: learning dense volumetric segmentation from sparse annotation. Medical Image Computing and Computer-Assisted Intervention–MICCAI 2016: 19th International Conference, Athens, Greece, October 17-21, 2016, Proceedings, Part II 19. Springer, 2016, pp. 424–432.
- Hatamizadeh, A.; Tang, Y.; Nath, V.; Yang, D.; Myronenko, A.; Landman, B.; Roth, H.R.; Xu, D. Unetr: Transformers for 3d medical image segmentation. Proceedings of the IEEE/CVF winter conference on applications of computer vision, 2022, pp. 574–584.
- Hatamizadeh, A.; Nath, V.; Tang, Y.; Yang, D.; Roth, H.R.; Xu, D. Swin unetr: Swin transformers for semantic segmentation of brain tumors in mri images. Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries: 7th International Workshop, BrainLes 2021, Held in Conjunction with MICCAI 2021, Virtual Event, September 27, 2021, Revised Selected Papers, Part I. Springer, 2022, pp. 272–284.
- Liu, H.; Simonyan, K.; Yang, Y. Darts: Differentiable architecture search. arXiv preprint arXiv:1806.09055 2018.
- Tan, M.; Chen, B.; Pang, R.; Vasudevan, V.; Sandler, M.; Howard, A.; Le, Q.V. Mnasnet: Platform-aware neural architecture search for mobile. Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 2019, pp. 2820–2828.
- Zela, A.; Elsken, T.; Saikia, T.; Marrakchi, Y.; Brox, T.; Hutter, F. Understanding and robustifying differentiable architecture search. arXiv preprint arXiv:1909.09656 2019.
- Yang, X.; Zhou, D.; Liu, S.; Ye, J.; Wang, X. Deep model reassembly. Advances in neural information processing systems 2022, 35, 25739–25753. [Google Scholar]
- Chen, J.; Lu, Y.; Yu, Q.; Luo, X.; Adeli, E.; Wang, Y.; Lu, L.; Yuille, A.L.; Zhou, Y. Transunet: Transformers make strong encoders for medical image segmentation. arXiv preprint arXiv:2102.04306 2021.
- Zhang, Y.; Liu, H.; Hu, Q. Transfuse: Fusing transformers and cnns for medical image segmentation. Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, September 27–October 1, 2021, Proceedings, Part I 24. Springer, 2021, pp. 14–24.
- Kotanidis, C.P.; Xie, C.; Alexander, D.; Rodrigues, J.C.; Burnham, K.; Mentzer, A.; O’Connor, D.; Knight, J.; Siddique, M.; Lockstone, H.; others. Constructing custom-made radiotranscriptomic signatures of vascular inflammation from routine CT angiograms: a prospective outcomes validation study in COVID-19. The Lancet Digital Health 2022, 4, e705–e716. [Google Scholar] [CrossRef] [PubMed]
- Otsuka, F.; Yahagi, K.; Sakakura, K.; Virmani, R. Why is the mammary artery so special and what protects it from atherosclerosis? Annals of cardiothoracic surgery 2013, 2, 519. [Google Scholar] [PubMed]
- Akoumianakis, I.; Sanna, F.; Margaritis, M.; Badi, I.; Akawi, N.; Herdman, L.; Coutinho, P.; Fagan, H.; Antonopoulos, A.S.; Oikonomou, E.K.; others. Adipose tissue–derived WNT5A regulates vascular redox signaling in obesity via USP17/RAC1-mediated activation of NADPH oxidases. Science translational medicine 2019, 11, eaav5055. [Google Scholar] [CrossRef] [PubMed]
- Iantsen, A.; Jaouen, V.; Visvikis, D.; Hatt, M. Squeeze-and-excitation normalization for brain tumor segmentation. Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries: 6th International Workshop, BrainLes 2020, Held in Conjunction with MICCAI 2020, Lima, Peru, October 4, 2020, Revised Selected Papers, Part II 6. Springer, 2021, pp. 366–373.
- Ulyanov, D.; Vedaldi, A.; Lempitsky, V. Instance normalization: The missing ingredient for fast stylization. arXiv preprint arXiv:1607.08022 2016.
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Lin, S.; Guo, B. Swin transformer: Hierarchical vision transformer using shifted windows. Proceedings of the IEEE/CVF international conference on computer vision, 2021, pp. 10012–10022.
- Lee, H.H.; Bao, S.; Huo, Y.; Landman, B.A. 3D UX-Net: A Large Kernel Volumetric ConvNet Modernizing Hierarchical Transformer for Medical Image Segmentation. arXiv preprint arXiv:2209.15076 2022.
- Pathak, D.; Krahenbuhl, P.; Donahue, J.; Darrell, T.; Efros, A.A. Context encoders: Feature learning by inpainting. Proceedings of the IEEE conference on computer vision and pattern recognition, 2016, pp. 2536–2544.
- Noroozi, M.; Favaro, P. Unsupervised learning of visual representations by solving jigsaw puzzles. Computer Vision–ECCV 2016: 14th European Conference, Amsterdam, The Netherlands, October 11-14, 2016, Proceedings, Part VI. Springer, 2016, pp. 69–84.
- Kerfoot, E.; Clough, J.; Oksuz, I.; Lee, J.; King, A.P.; Schnabel, J.A. Left-ventricle quantification using residual U-Net. Statistical Atlases and Computational Models of the Heart. Atrial Segmentation and LV Quantification Challenges: 9th International Workshop, STACOM 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, September 16, 2018, Revised Selected Papers 9. Springer, 2019, pp. 371–380.
- Myronenko, A. 3D MRI brain tumor segmentation using autoencoder regularization. Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries: 4th International Workshop, BrainLes 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, September 16, 2018, Revised Selected Papers, Part II 4. Springer, 2019, pp. 311–320.
- Yu, X.; Yang, Q.; Zhou, Y.; Cai, L.Y.; Gao, R.; Lee, H.H.; Li, T.; Bao, S.; Xu, Z.; Lasko, T.A.; others. UNesT: Local Spatial Representation Learning with Hierarchical Transformer for Efficient Medical Segmentation. arXiv preprint arXiv:2209.14378 2022.

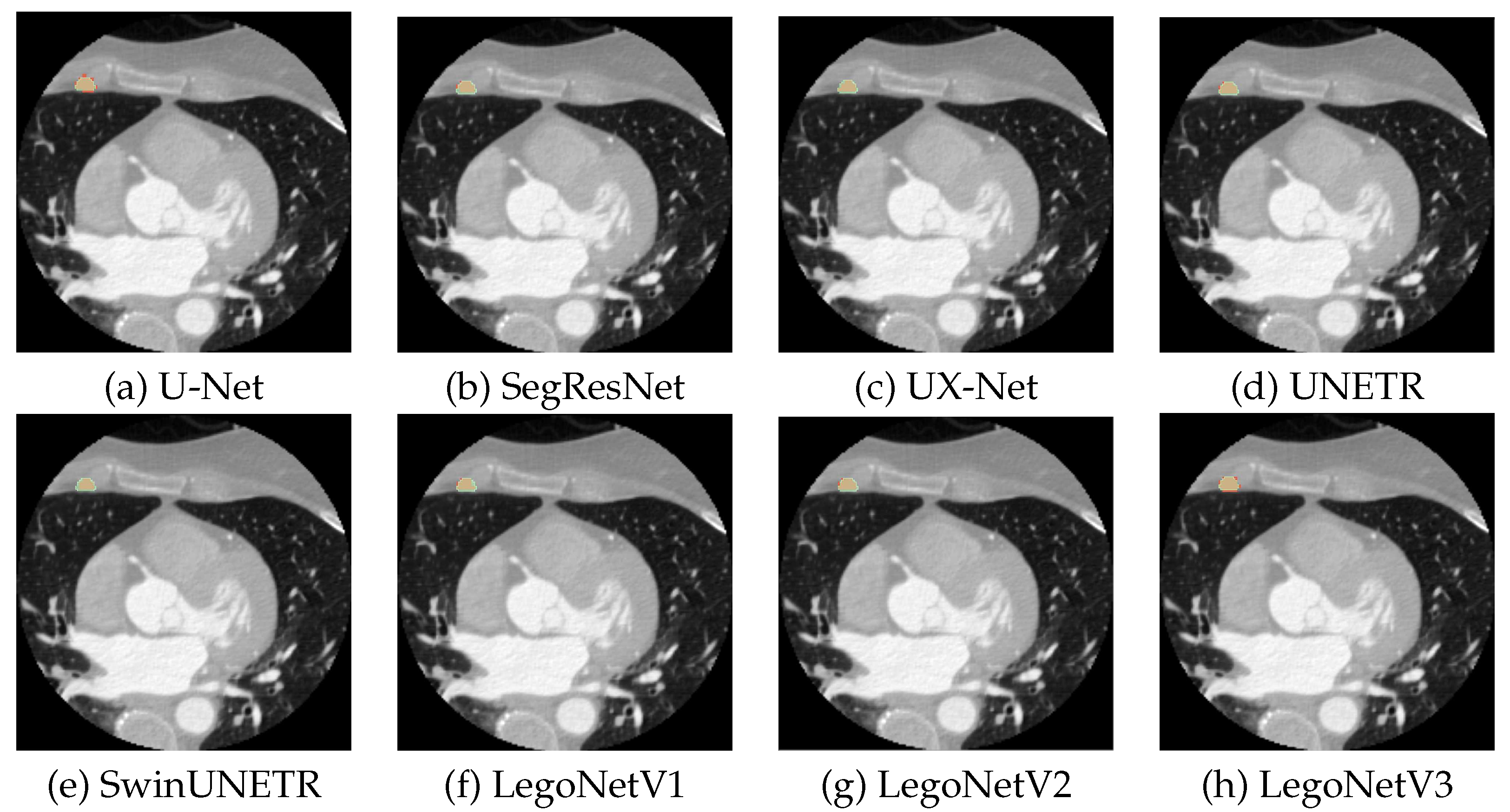
| Network | Used blocks | Hidden size | Feature size |
|---|---|---|---|
| LegoNetV1 | SE→UX→SE→UX | 768 | (24, 48, 96, 192) |
| LegoNetV2 | SE→Swin→SE→Swin | 768 | (24, 48, 96, 192) |
| LegoNetV3 | Swin→UX→Swin→UX | 768 | (24, 48, 96, 192) |
| Models | DSC↑ | Precision↑ | Recall↑ | HD95↓ | Params (M)↓ | FLOPs (G)↓ |
|---|---|---|---|---|---|---|
| UNet [1,19] | 0.686±0.03 | 0.72±0.04 | 0.69±0.03 | 2.70 | 3.99 | 27.64 |
| SegResNet [20] | 0.732±0.01 | 0.75±0.02 | 0.74±0.03 | 2.50 | 1.18 | 15.58 |
| UX-Net [16] | 0.695±0.03 | 0.73±0.06 | 0.70±0.01 | 3.17 | 27.98 | 164.17 |
| UNETR [2] | 0.690±0.02 | 0.72±0.03 | 0.69±0.03 | 3.00 | 92.78 | 82.48 |
| SwinUNETR [3] | 0.713±0.02 | 0.74±0.02 | 0.71±0.04 | 2.46 | 62.83 | 384.20 |
| UNesT [21] | 0.555±0.04 | 0.59±0.06 | 0.55±0.05 | 4.35 | 87.20 | 257.91 |
| LegoNetV1 | 0.747±0.02 | 0.75±0.02 | 0.77±0.03 | 2.34 | 50.58 | 175.77 |
| LegoNetV2 | 0.749±0.02 | 0.77±0.01 | 0.76±0.04 | 2.11 | 50.71 | 188.02 |
| LegoNetV3 | 0.741±0.02 | 0.76±0.02 | 0.75±0.03 | 2.34 | 11.14 | 173.41 |
| Models | DSC↑ | Precision↑ | Recall↑ |
|---|---|---|---|
| UNet [1,19] | 0.611±0.10 | 0.66±0.10 | 0.61±0.10 |
| SegResNet [20] | 0.638±0.09 | 0.68±0.10 | 0.65±0.08 |
| UX-Net [16] | 0.608±0.08 | 0.65±0.10 | 0.62±0.06 |
| UNETR [2] | 0.581±0.10 | 0.62±0.12 | 0.61±0.07 |
| LegoNetV1 | 0.642±0.08 | 0.66±0.11 | 0.67±0.07 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




