Submitted:
26 May 2023
Posted:
30 May 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sampling
2.2. Microbiological analyses
2.2.1. Identification of bacterial strains
2.2.2. Biofilm formation
2.2.3. Antibiotic susceptibility of isolates
2.3. Molecular characterization of isolates
2.3.1. DNA extraction
2.3.2. Search for alleles of genes encoding virulence
2.4. Data analysis and processing
3. Results
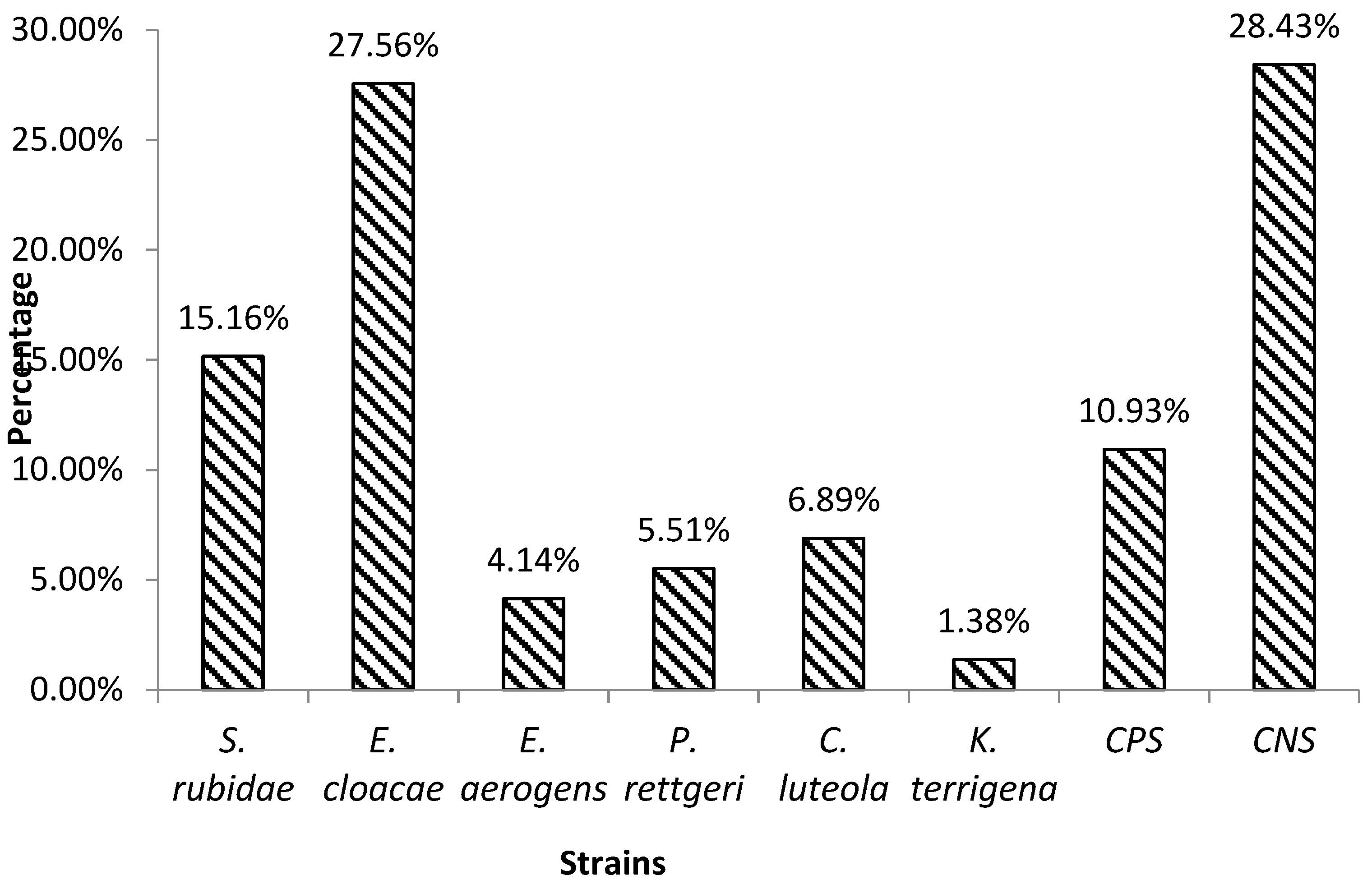
3.1. Identification of pathogenic bacteria isolated
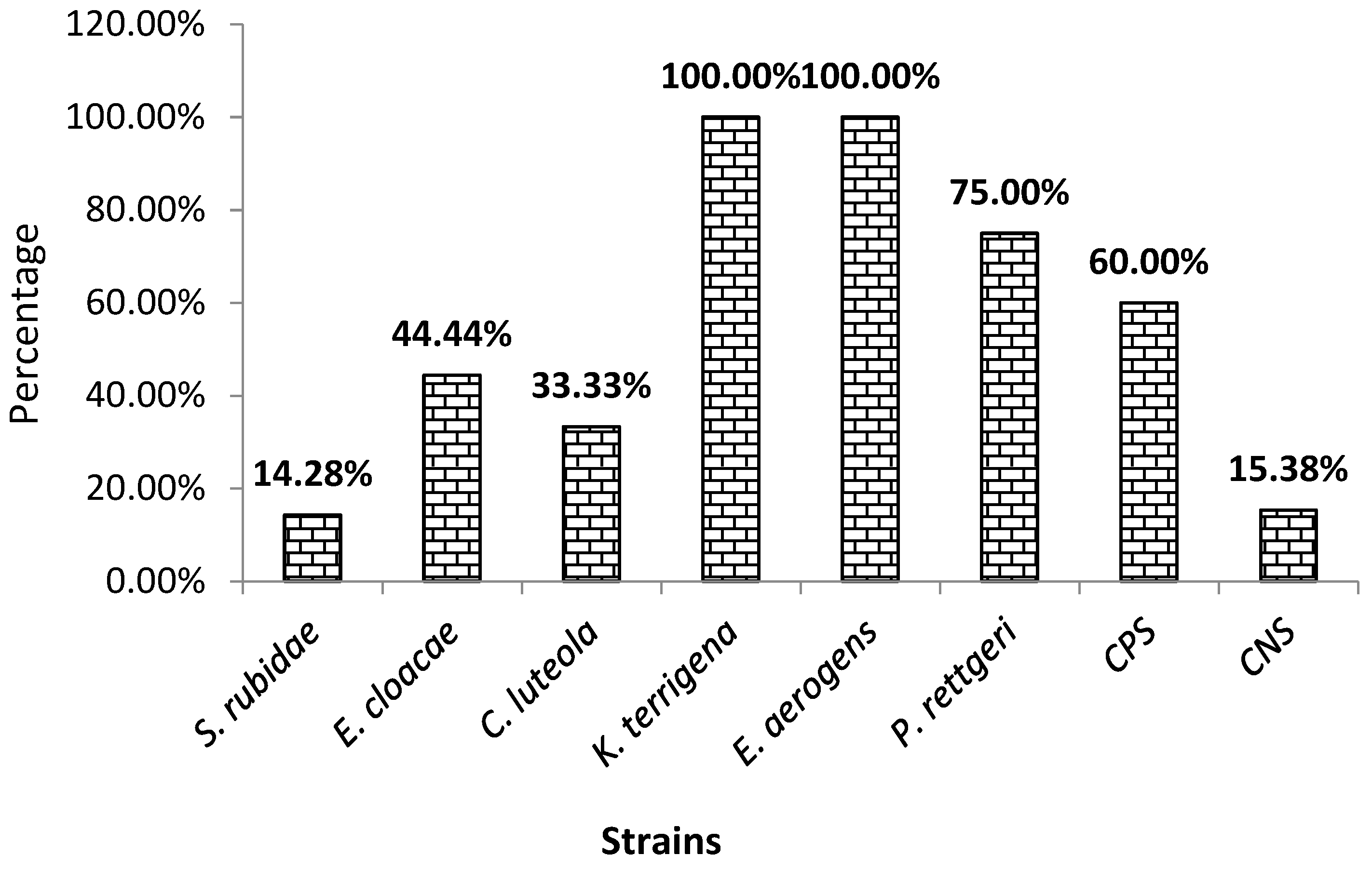
3.2. Biofilm production
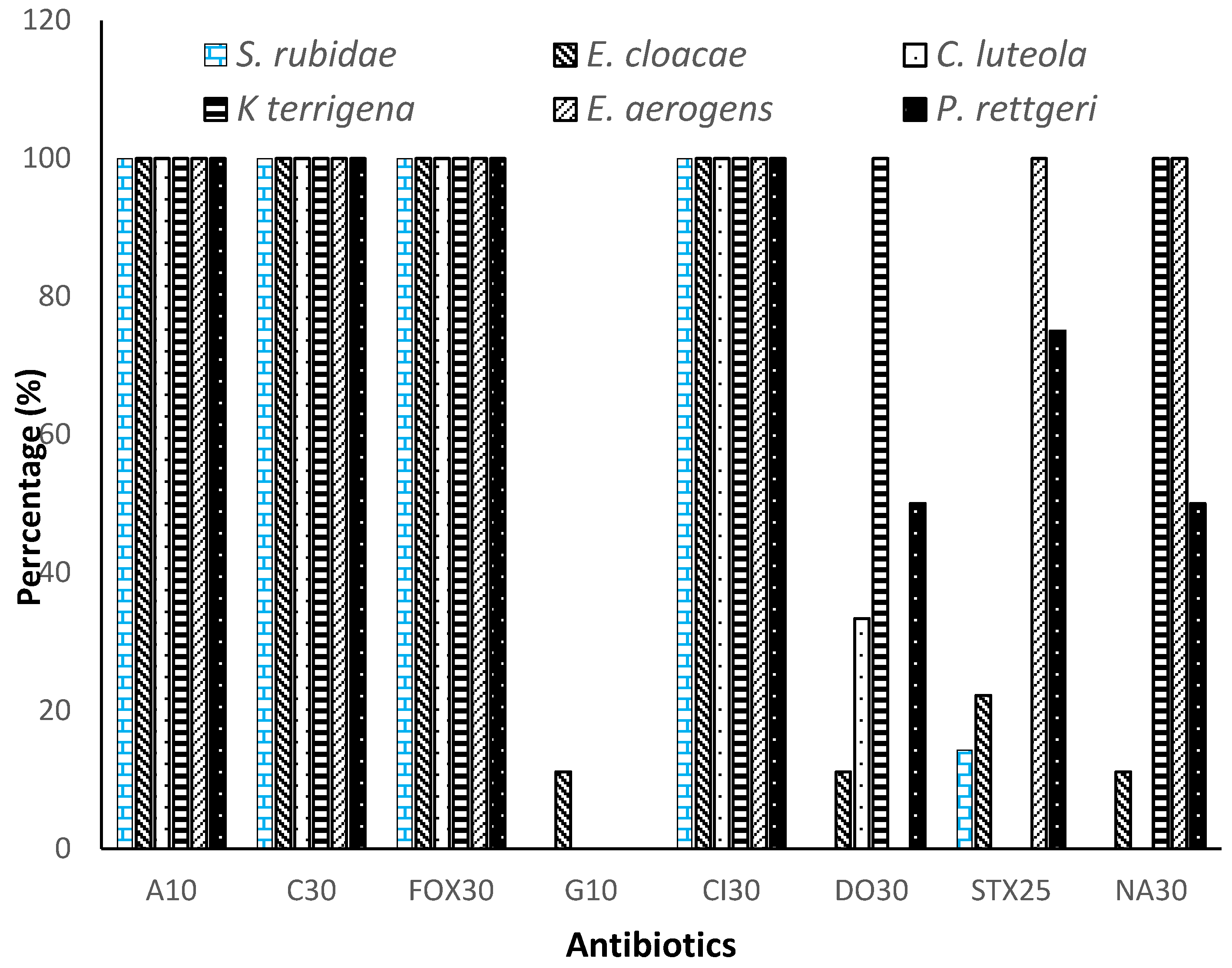
3.3. Resistance profile of Enterobacteriaceae to antibiotics by species
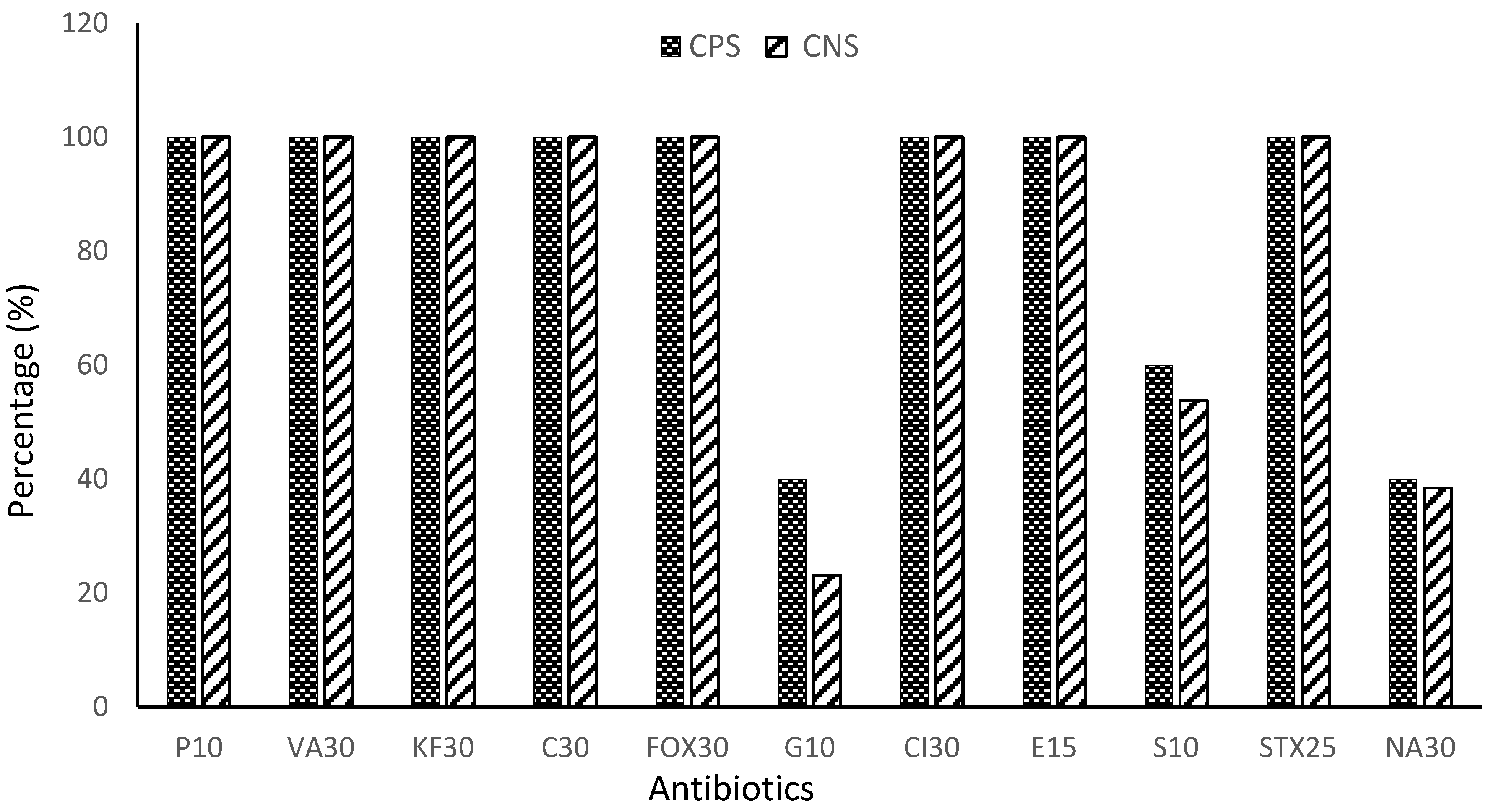
3.4. Resistance profile of Enterobacteriaceae to antibiotics by species
3.5. Virulence genes and Resistance gene macrolides
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- ayodé, APP; Linmemann, A.R.; Nout, M.J.R.; Hounhouigan, D.J.; Stomph, T.J.; Smulders, M.J.M.; “Diversity and food quality properties of farmers’ varieties of sorghum from Benin,” Journal of Science of Food and Agriculture, 2006, vol. 86, pp. 1032–1039.
- Missihoun, A.A.; Agbangla, C.; Sagbadja, H. A.; Ahanhanzo, C.; Vodouhe, R. ; “Gestion traditionnelle et statut des ressources génétiques du sorgho (Sorghum bicolor L. Moench) au Nord-Ouest du Bénin,” International Journal of Biological and Chemical Sciences, 2012, vol. 6, no. 3, pp. 1003-1018.
- Aholou, C. C. “Construction identitaire et urbaine autour du cabaret à Lomé (Togo),” Hommes et migrations, 2010, vol. 1283, pp. 32-41. [CrossRef]
- Osseyi, E. G. , Tagba, P.; Karou, S. D. “Stabilization of the Traditional Sorghum Beer “Tchoukoutou” using Rustic Wine- Making Method,” Advance Journal of Food Science and Technology, 2011, vol. 3, no. 4, pp. 254-258.
- Baba-Moussa, L. ; Bokossa, Y. I. ; Baba-Moussa, F. ; Ahisou, H. ; Adeoti, Z. ; Yehouenou, B. ; Mamadou, A. Toukourou, F. ; Sanni, A. “Etude des possibilités de contamination des aliments de rues au Bénin : Cas de la ville de Cotonou,” Journal de Recherche Scientifique de l’Université de Lomé (Togo) série A, 2006, vol. 8, no. 2, pp. 149-156.
- WHO. Rapport OMS : 420 000 personnes tuées par l’intoxication alimentaire. 2018.
- Adesetan, T. O., Efuntoye, M. O., Babalola, O. O. “Profiling of Bacillus Cereus Enterotoxigenic Genes from Retailed Foods and Detection of the Nhe and Hbl Toxins with Immunological Assay”. Journal of Applied and Natural Science, 2022, 14, 254-267. [CrossRef]
- Imade, E.E. , Omonigho, S.E., Babalola, O.O., Enagbonma, B.J. “Lactic acid bacterial bacteriocins and their bioactive properties against food-associated antibiotic-resistant bacteria.” Annals of Microbiology, 2021, vol. 71, n°44. [CrossRef]
- Chuah, L.O.; Shamila-Syuhada, A.K. ; Liong, MT; Rosma, A.; Thong, K. L.; Rusul, G. “Physio-chemical, microbiological properties of tempoyak and molecular characterisation of lactic acid bacteria isolated from tempoyak,” Food Microbiology,2016, vol. 58, pp. 95–104. [CrossRef]
- Egbuta, M. A. , Mwanza, M., Babalola, O. O. “Health Risks Associated with Exposure to Filamentous Fungi.” International Journal of environmental research and public health, 2017, vol. 14, n°7, 719. [CrossRef]
- Achi, O. K.; Ukwuru, M. “Cereal-based fermented foods of Africa as functional foods,” International Journal of Microbiology Applied, 2015, vol. 2, pp. 71–83.
- Adesetan, O., T. , Efuntoye, M. O., Babalola, O. O. “Biochemical Characterization and Antimicrobial Susceptibility of Bacillus Cereus Isolates from Some Retailed Foods in Ogun State, Nigeria. Journal of Microbiology, Biotechnology and Food Sciences, 2019, 9(3), 616–621. [CrossRef]
- Kimmons, J. E.; Brown, K. H.; Lartey, A.; Collisone, M. P. P. ; Dewey. K. G. “The effects of fermentation and/or vacuum flask storage on the presence of coliforms in complementary foods prepared in Ghana,” International Journal of food Sciences and Nutrition, 1999, vol. 50, pp.195-201.
- Akaki, K. D.; Aw, S.; Loiseau, G.; Guyot, J. P. “Etude du comportement des souches de Bacillus cereus ATCC 9139 et d’Escherichia coli ATCC 25922 par la méthode des challenge-tests lors de la confection de bouillies à base de pâte de mil fermentée en provenance de Ouagadougou (Burkina-Faso),” Revue Ivoirienne des Sciences et Technologie, 2008, vol. 11, pp. 103-117.
- N’tcha, C.; Adéyèmi, A. D.; Kayodé, A. P. P.; Vieira-Dalodé, G.; Agbobatinkpo, B. P.; Codjia, J. C.; Baba-Moussa, L. “Indigenous knowledge associated with the production of starters culture used to produce Beninese opaque sorghum beers,” Journal of Applied Biosciences,2015, vol. 88, pp. 8223-8234. [CrossRef]
- Lyhs, U.; Björkroth, J.; Hyytiä, E.; Korkeala, H. “The spoilage flora of vacuumpackaged, sodium nitrite or potassium nitrate treated, cold-smoked rainbow trout stored at 4ºC or 8ºC,” International Journal of food microbiology,1998, vol. 45, pp. 135-142. [CrossRef]
- Sundaramoorthy, N.; Yogesh, P.; Dhandapani, R. ; “Production of prodigiosin from Serratia marcescens isolated from soil,” Indian Journal of Sciences and Technology, 2009, vol. 2, no. 10, pp. 32-34.
- Buchanan R., E.; Gibbons, N. E. “Bergey's Manual of Determinative Bacteriology” 8th Edition, Baltimore, USA, Williams and Wilkcns Company, 1974.
- Hounsa, A.; Kouadio, L.; De Mol, P. “Self-medication with antibiotics obtained from private pharmacies in Abidjan, Ivory Coast,” Médecine et maladies infectieuses 2009, vol. 40, no. 6, pp. 333-340. [CrossRef]
- Alsan, M.; Morden, N.; Gottlieb, J. D.; Weiping, Z.; Skinner, J. “Antibiotic use in cold and flu season and prescribing quality: a retrospective cohort study,” Medical care,2015, vol. 53, no. 12, pp. 1066. [CrossRef]
- Singh, A.; Gautam, P. K.; Verma, A.; Singh, V.; Shivapriya, P. M.; Shivalkar, S.; Sahoo, A K.; Samanta, S. K. “Green synthesis of metallic nanoparticles as effective alternatives to treat antibiotics resistant bacterial infections: A review,” Biotechnology Reports, 2020, vol. 25, e00427. [CrossRef]
- Akindolire, M. A. , Babalola, O. O., Ateba, C. N. “Detection of Antibiotic Resistant Staphylococcus aureus from Milk: A Public Health Implication.” International Journal of Environmental Research and Public Health, 2015, 12(9), 10254–10275. [CrossRef]
- Muylaert, A.; Mainil, J. G. “Résistances aux fluoroquinolones : la situation actuelle,” Annales de Medecine Veterinaire, 2013, vol. 157, pp. 15-26.
- Christensen, G.D.; Simpson, W.A.; Bisno, A.L. ; Beachy, EH "Adherence of biofilm producing strains of Staphylococci epidermidis to smooth surfaces,” Infections and Immunity, 1985, vol. 37, pp. 318-326.
- Stepanović, S.; Vuković, D.; Dakić, I.; Savić, B.; Švabić-Vlahović, M. “A modified microtiter-plate test for quantification of staphylococcal biofilm formation” Journal of Microbiology Methods,2000, vol. 40, no. 2, pp. 175-179. [CrossRef]
- CASFM “Comité de l’antibiogramme de la Société Française de Microbiologie : Recommandations 2020 V.1.1,” 181 p, 2020. Avaible on https://www.sfm-microbiologie.org/wp-content/uploads/2020/04/CASFM2020_Avril2020_V1.1.pdf.
- Rasmussen R., S.; Morrissey, M. T. “DNA-based methods for the identification of commercial fish and seafood species.” Comprehensive Reviws in Food Science and Food Safetay, 2008, vol. 7, no. 3, pp. 280-295. [CrossRef]
- Lamy, M. C.; Dramsi, S.; Billoët, A.; Réglier-Poupet, H.; Tazi, A.; Raymond, J.; Guérin, F.; Couvé, E.; Kunst, F.; Glaser, P. “Rapid detection of the highly virulent group B Straptococcus ST-17 clone,” Microbial and Infection, 2006, vol. 8, no. 7, pp. 1714-1722. [CrossRef]
- Kaczmarek, A.; Budzyńska, A.; Gospodarek, E. “Prevalence of genes encoding virulence factors among Escherichia coli wiyh K1 antigen and-K1 E. coli strains,” Journal of Medical Microbiology, 2012, vol. 61, no. 10, pp. 1360-1365. [CrossRef]
- Moulin-Schouleur, M.; Schouleur, C.; Tailliez, P.; Kao, M. R.; Brée, A.; Germon, P.; Oswald, E.; Mainil, J.; Blanco, M.; Blanco, J. “Common Virulence factors and genetic relatonships between O18 : K1 : H7 Escherichia coli isolates of human and avian origin,” Journal of Clinical Microbiology, 2006, vol. 44, no. 10, pp. 3484-3492. [CrossRef]
- Germon, P.; Chen, Y. H.; He, L.; Blanco, J.E.; Bree, A.; Schouler, C.; Huang, S. H.; Moulin-Schouleur, M. “IbeA, a virulence factors of avian pathogenenic Escherichia coli, Microbiology, 2005, vol. 151, no. 4, pp. 1179-1186. [CrossRef]
- Desjardins, M.; Delgaty, K. L.; Ramotar, K.; Toye, B. “Prevalence and mechanisms of erythromycin resistance in group A and group B Streptococcus: Implications for reporting susceptibility results,” Journal of Clinical Microbiology, 2004, vol. 42, no. 12, pp. 5620-5623. [CrossRef]
- Griffin, P. ; Findlay,G. E. “Process and engineering improvements to rotating biological contactor design,” Water Sciences Technology,2000, vol. 41, no. 1, pp. 137–144. [CrossRef]
- Tankoano, A.; Diop, M. B.; Sawadogo-Lingani, H.; Kaoré, D.; Savadogo, A. ; “Les aspects technologiques, microbiogiques et nutritionnels de saliments fermentés à base de lait de mil en Afrique de l’ouest,” International Journal of Advanced Research, 2017, vol. 5, no. 8, pp. 1509-1526.
- Cason, D. E.; Mahlomaholo, J. B.; Taole, M. M.; Abong, O. G. ; Vermeulen, J, G.; de Smidt, O.; Vermeulen, M.; Steyn, L.; Valverde A.; Vijoen, B. “Bacterial and Fungal Dynamics during the Fermentation Process of Sesotho, a Traditional Beer of Southern Africa,” Frontiers in microbiology,2020, vol. 11, pp. 1451.
- Escalante, A.; Maria, E. R.; Martinez, A.; Munguia, A. L. “Characterization of bacterial diversity in Pulque, a traditional Mexican alcoholic fermented beverage, as determined by 16S rDNA analysis,” FEMS Microbiology Letters,2004, vol. 235, no. 2, pp. 273. [CrossRef]
- Bokulich, N. A.; Bamforth, C. W.; Mills, D. A. “Brewhouse-resident microbiota is responsible for multistage fermentation of American coolshipale,” Journal Plos One,2012, 7: e35507. [CrossRef]
- Taylor, G.; Keane, P. M. “Cross-infection with Serratia marcescens,” Journal of Clinical Pathology,1962, vol. 15, pp. 145–147. [CrossRef]
- Kuzina, L.; Peloquin, J.; Vacek, D. “Isolation and identification of bacteria associated with adult laboratory Mexican fruit flies, Anastrepha ludens (Diptera: Tephritidae),” Current Microbiology; 2001, vol. 42, no. 4, pp. 290–294. [CrossRef]
- Bensakhria. A. Biologie: Staphylococcus aureus (staphylocoques), 2017 www.magazinescience.com. Accessed on 05 may 2019.
- Kostaki, M.; Chorianopoulos, N.; Braxou, E.; Nychas, G.J.; Efstathios, G. “Differential biofilm formation and chemical disinfection resistance of sessile cells of Listeria monocytogenes strains under monospecies and dual-species (with Salmonella enterica) conditions,” Applied and Environmental Microbiology,2012, vol. 8, no. 8, pp. 2586–2595. [CrossRef]
- Jefferson, K. “What drives bacteria to produce a biofilm?” FEMS microbiology letters, 2004, vol. 236, no. 2, pp. 163-173. [CrossRef]
- Balestrino, D.; Ghigo, J.M.; Charbonnel, N.; Haagensen, J. A.; Forestier, C. “The characterization of functions involved in the establishment and maturation of Klebsiella pneumoniae in vitro biofilm reveals dual roles for surface exopolysaccharides,” Environmental Microbiology,2008, vol. 10, pp. 685–701. [CrossRef]
- Ahouandjinou, H. Qualité microbiologique, antibiorésistance des souches de Staphylococcus spp et d’Escherichia coli isolées des carcasses bovines au Bénin, PhD dissertation University of Abomey- Calavi (Benin). 2016, 175p.
- Lerebour, G.; Cupferman, S.; Bellon-Fontaine, M.N. “Adhesion of Staphylococcus aureus and Staphylococcus epidermidis to the Episkin reconstructed epidermis model and to an inert 304 stainless steel substrate” Journal of Applied Microbiology, 2004, vol. 97, no. 1, pp. 7-16. [CrossRef]
- Marques, S. C.; Silva-Rezende, J. G.; de Freitas-Alves, L. A. “Formation of biofilms by Staphylococcus aureus on stainless steel and glass surfaces and its resistance to some selected chemical sanitiziers” Brazilian Journal of Microbiology,2007, vol. 38, no. 3, pp. 538–543. [CrossRef]
- Mah, T. F. “Biofilm-specific antibiotic resistance” Future Microbiology, 2012,vol. 7, no. 9, pp. 1061–1072. [CrossRef]
- Hama, H. , Recherche de bactéries associées aux mammites subcliniques dans le lait de chèvre en Mauritanie et au Togo et détermination de leur antibiosensibilité. Doctorate dissertation Veterinary Medicine, Dakar (Senegal), 2004,31.
- Anago, E.; Ayi-Fanou, L.; Akpovi, C. D.; Hounkpe, W. B.; Tchibozo, M.; Bankole, H. S. “Antibiotic resistance and genotype of beta-lactamase producing Escherichia coli in nosocomial infections in Cotonou, Benin” Annals of Clinical Microbiology and Antimicrobials, 2015,vol. 14, no. 1, pp. 1-6. [CrossRef]
- Touati, A.; Brasme, L.; Benallaoua, S.; Gharout, A.; Madoux, J.; De Champs, C. “First report of qnrB-producing Enterobacter cloacae and qnrA-producing Acinetobacter baumannii recovered from Algerian hospitals,” Diagnostical Microbiology Infectious Diseases, 2008, vol. 60, pp. 287-290. [CrossRef]
- Soussy, C. J. Quinolones et bactéries à Gram négatif. Dans: Patrice Courvalin, Roland Leclercq, Edouard Bingen eds. Antibiogramme, 2006,2ème édition ESKA, Paris. 277p.
- . Kadja, M. C.; Kane, Y.; Viban, B. V.; Kaboret, Y.; Alambedji, B. Sensibilité aux antibiotiques des Bactéries associées aux mammites Clinique des petits ruminants dans la région de Dakar, Annales des Sciences Agronomique,2013, vol. 17, no. 2, pp. 205-216.
- Davin-Regli, A.; Bolla, J. M.; James, C. E.; Lavigne, J. P. Chevalier J.; Garnotel, E. “Membrane permeability and regulation of drug “influx and efflux” in enterobacterial pathogens,” Current drug Targets,,2008, vol. 9, pp. 750–759.
- Wilson, K. Shafiee, M.; Hasso, M.; Sharif, U. “Providencia rettgeri: an unexpected case of Gram-negative cellulitis”, Case Report, Wounds International,2015, vol 6, no. 4.
- Sina, H.; Attien, P.; Wélé, M.; Socohou, A.; Boukary-Chabi, A.; Dougnon, V. T.; Baba-Moussa, F.; Adjanohoun, A.; and Baba-Moussa, L. “Sanitary Risk Factors and Microbial Contamination of Grilled Meats Sold in Cotonou, Benin”, Journal of Food Security,2019, vol. 7, no. 5, pp. 175-182.
- S. Schwarz, and E. Chaslus-Dancla, “Use of antimicrobials in veterinary medicine and mechanisms of resistance,” Veterinary Research, 2001, vol. 32, pp. 201-225. [CrossRef]
- Chiquet, C.; Max, M.; Altayrac, J.; Aptel, F.; Boisset, S.; Vandenesch, F.; Carricajo, A. “Correlation between clinical data and antibiotic resistance in coagulase negative Staphylococcus species isolated from 68 patients with acute post-cataract endophthalmitis,” Clinical Microbiology and Infection, 2015,vol. 21, no. 6, 592.el-592.e8. [CrossRef]
- Bonnel, A. S.; Quinque, K.; Le Roux, P.; Le Luyer, B. Aspergillose nécrossante semi invasive chez un enfant de 14 ans atteint de mucoviscidose, Revue des maladies Respiratoires,2006 vol. 23, no. 4, pp. 343-347.
- Rodrigues, M. X.; Silva, C. C. N.; Trevillin, H. J.; Cruzado, B. M.; Mui, S. T.; Sanches Duarte, R. F.; Castillo, C. C.; Canniatti-Brazaca G., S.; Porto, E. “Antibiotic resistance and molecular characterization of Staphylococcus species from mastitic milk” African Journal of Microbiology Research,2017, vol. 11, no. 3, pp. 84-91.
- Lesch, C.; Itokazu, G.; Danziger, L.; Weinstein, R. “Multi-hospital analysis of antimicrobial usage and resistance trends,” Diagnostic microbiology and infectious disease, 2001, vol. 41, no. 3, pp. 149-154. [CrossRef]
- Trivedi, K.; Cupakova, S.; Karpiskova, R. “Virulence factors and antibiotic resistance in enterococci isolated food-stuffs,” Veterinarni Medicina, 2011, vol. 56, no. 7, pp. 352-357. [CrossRef]
- Clegg, S. , Hughes, K. T. “FimZ is a molecular link between sticking and swimming in Salmonella enterica serovar Typhimurium” Journal of Bacteriology,”2002, vol. 184, pp. 1209–1213.
- Le Trong, I. “Donor strand exchange and conformational changes during E. coli fimbrial formation,” Journal of Structural Biology, 2008, vol. 172, pp. 380–388.
- Soria-Bustos, J.; Ares, M.A.; Gómez-Aldapa, C.A.; González-y-Merchand, J.A.; Girón, J.A.; De la Cruz, M.A. “Two Type VI Secretion Systems of Enterobacter cloacae Are Required for Bacterial Competition, Cell Adherence, and Intestinal Colonization” Frontier in Microbiology,2020, vol. 11: 560488.
- Lockman, H. A. “Yersinia pseudotuberculosis produces a cytotoxic necrotizing factor” Infection and Immunity, 2002, vol. 70, pp. 2708–2714.
- Jensen, L. B.; Frimodt-Moller, N.; Aarestrup, F. M. Presence of erm gene classes in Gram-positive bacteria of animal and human origin in Denmark. FEMS Microbiology Letters, 1999, vol. 170, pp. 151–158. [CrossRef]
- Luthje, P.; Schwarz, S. “Antimicrobial resistance of coagulase-negative staphylococci from bovine subclinical mastitis with particular reference to macrolide lincosamide resistance phenotypes and genotypes,” Journal of Antimicrobial Chemotherapy, 2006, vol. 57, pp. 966–969. [CrossRef]
- Donlan R., M.; Costerton, J. W. “Biofilms: survival mechanisms of clinically relevant microorganisms,” Clinical Microbiology Reviews, 2002, vol. 15, pp. 167–193. [CrossRef]
- Zhang, L.; Mah, T. F. “Involvement of a novel efflux system in biofilm-specific resistance to antibiotics,” Journal of bacteriology, 2008 vol. 190, pp. 4447–4452. [CrossRef]
| Enterobacteriaceae | P value | ||
|---|---|---|---|
| Antibiotics | Percentage of resistance | ||
| Ampicillin (A10) | 100.00% | >0.9999 | < 0.0001 |
| Ceftriaxone (Cl30) | 100.00% | ||
| Cefoxitin (FOX30) | 100.00% | ||
| Chloramphenicol (C30) | 100.00% | ||
| Gentamicin (G10) | 1.85% | >0.9999 | |
| Doxycycline (DO30) | 32.40% | ||
| Trimethoprim-sulfamethoxazole (STX25) | 32.85% | ||
| Nalidixic acid (NA30) | 43.51% | ||
| Staphylococci | P value | ||
|---|---|---|---|
| Antibiotics | Percentage of resistance | ||
| Penicillin G (P10) | 100.00% | >0.9999 | <0.0001 |
| Ceftriaxone (Cl30) | 100.00% | ||
| Cefoxitin (FOX30) | 100.00% | ||
| Chloramphenicol (C30) | 100.00% | ||
| Cephalothin (KF30) | 100.00% | ||
| Vancomycin (VA30) | 100.00% | ||
| Erythromycin (E15) | 100.00% | ||
| Trimethoprim-sulfamethoxazole (STX25) | 100.00% | ||
| Gentamicin (G10) | 31.54% | >0.9999 | |
| Nalidixic Acid (NA30) | 39.23% | ||
| Streptomycin (S10) | 59.92% | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




