Submitted:
04 May 2023
Posted:
23 May 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Cell culture.
Materials
Immunoblot
Lentivector preparation and transduction
Establishment of tagged-proteasome cell lines
Osmotic stress protocol
Generation of PSMD1-KD cells
Generation of PSMA3 C-terminal truncated cells
In-gel analysis of fluorescently tagged proteins
Live cell imaging
Data Processing and Analysis
Proteasome modeling
3. Results
Intracellular monitoring of the 26S proteasome
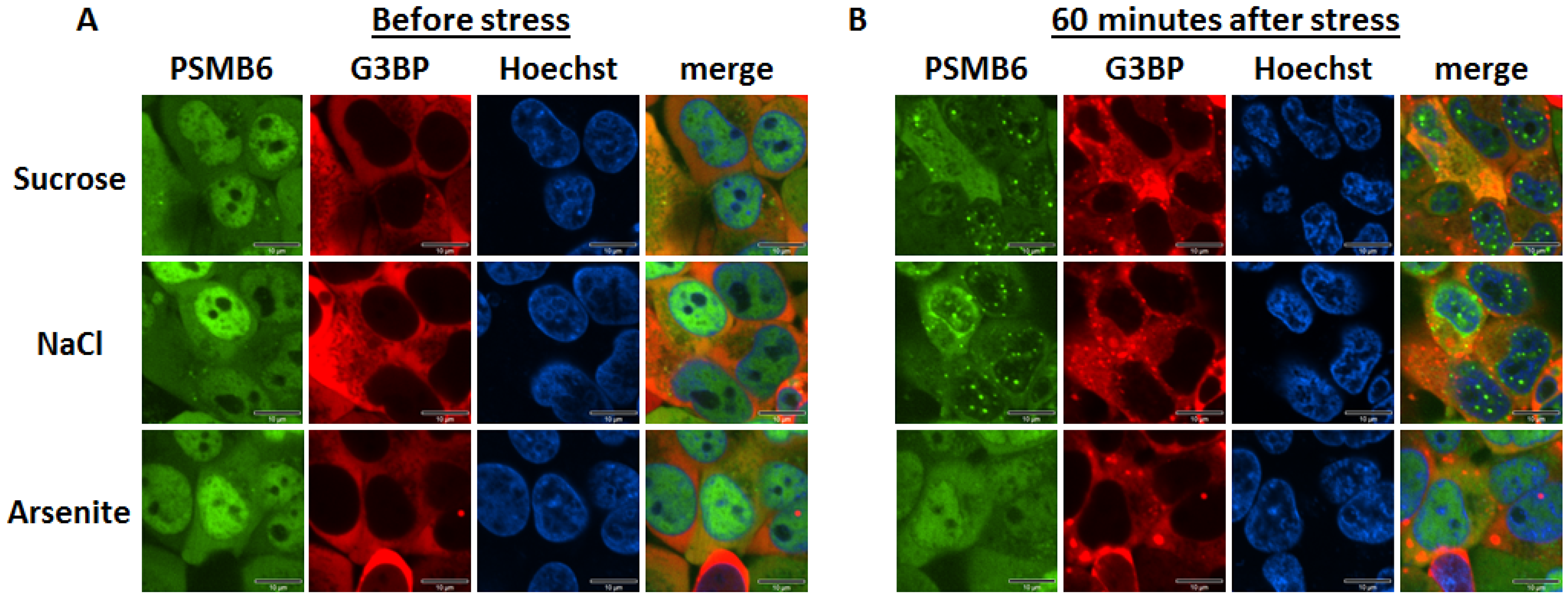
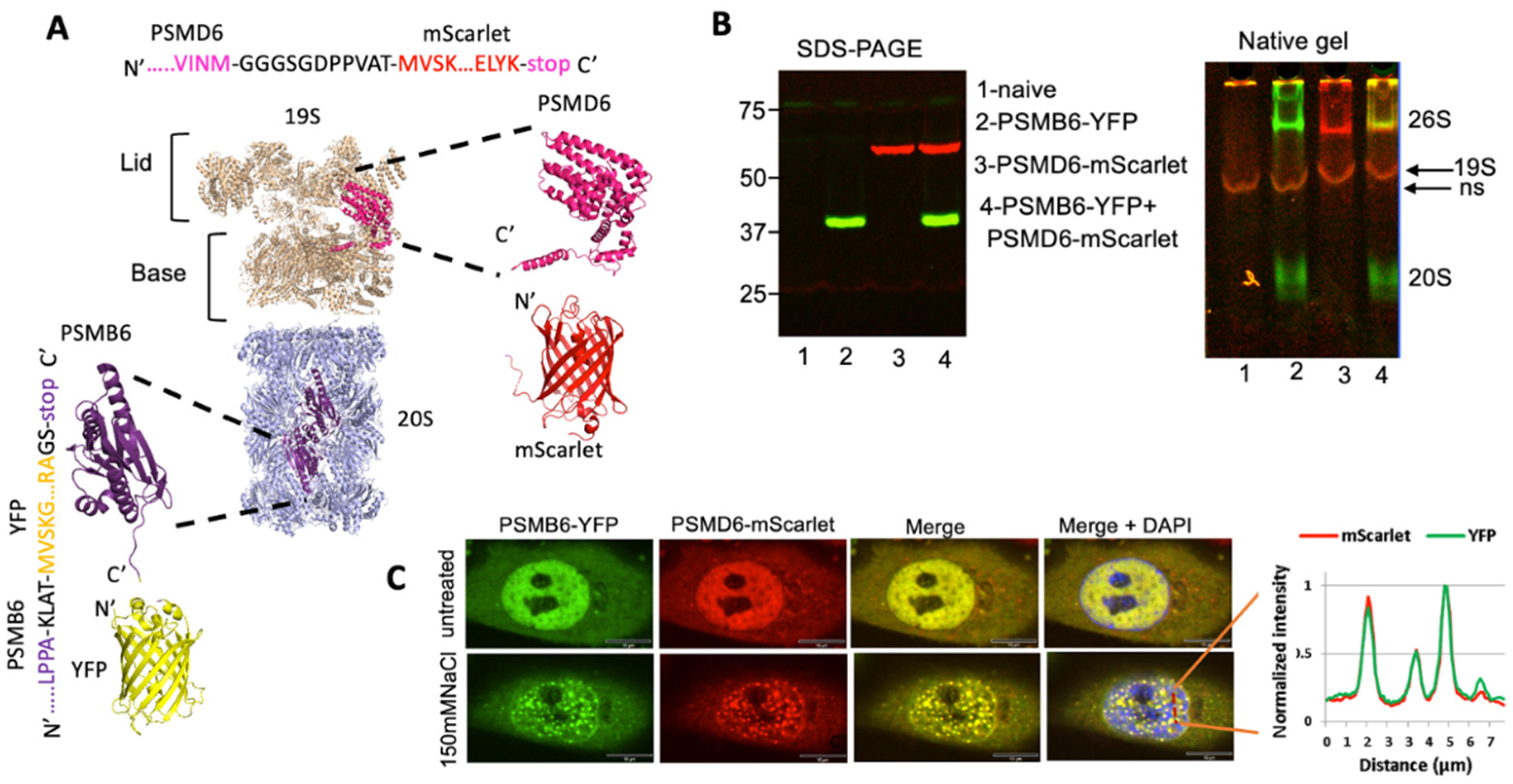
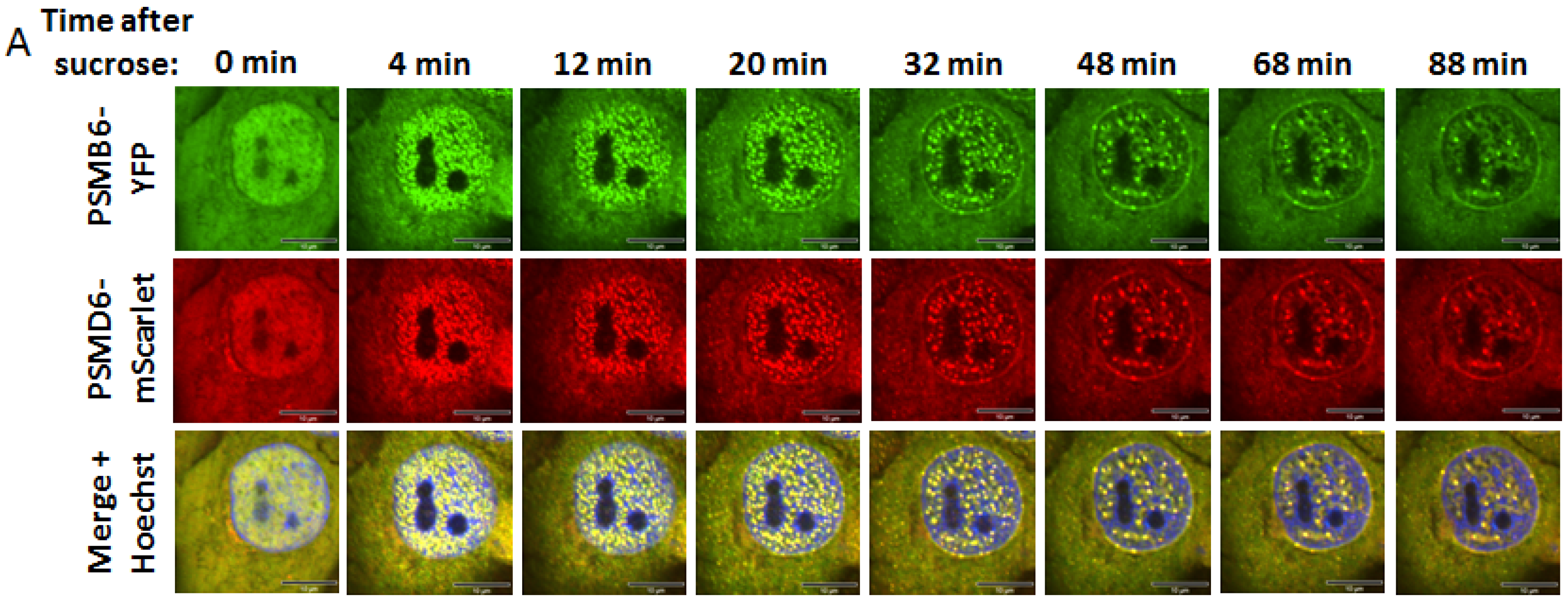
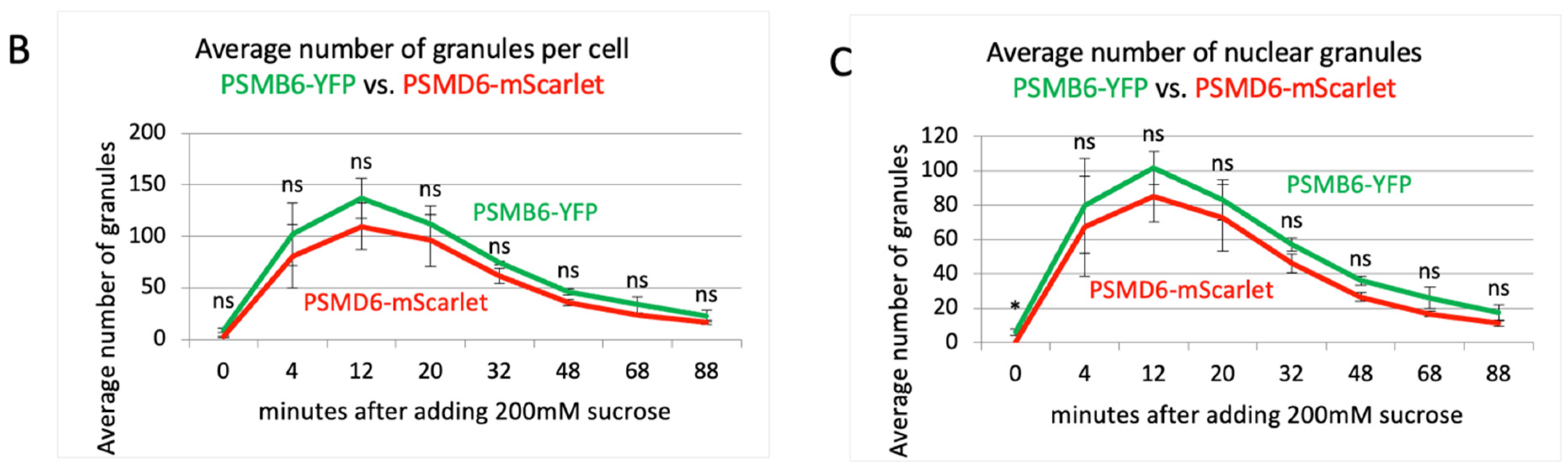
Time kinetics of the 26S proteasome granules in response to osmotic stress
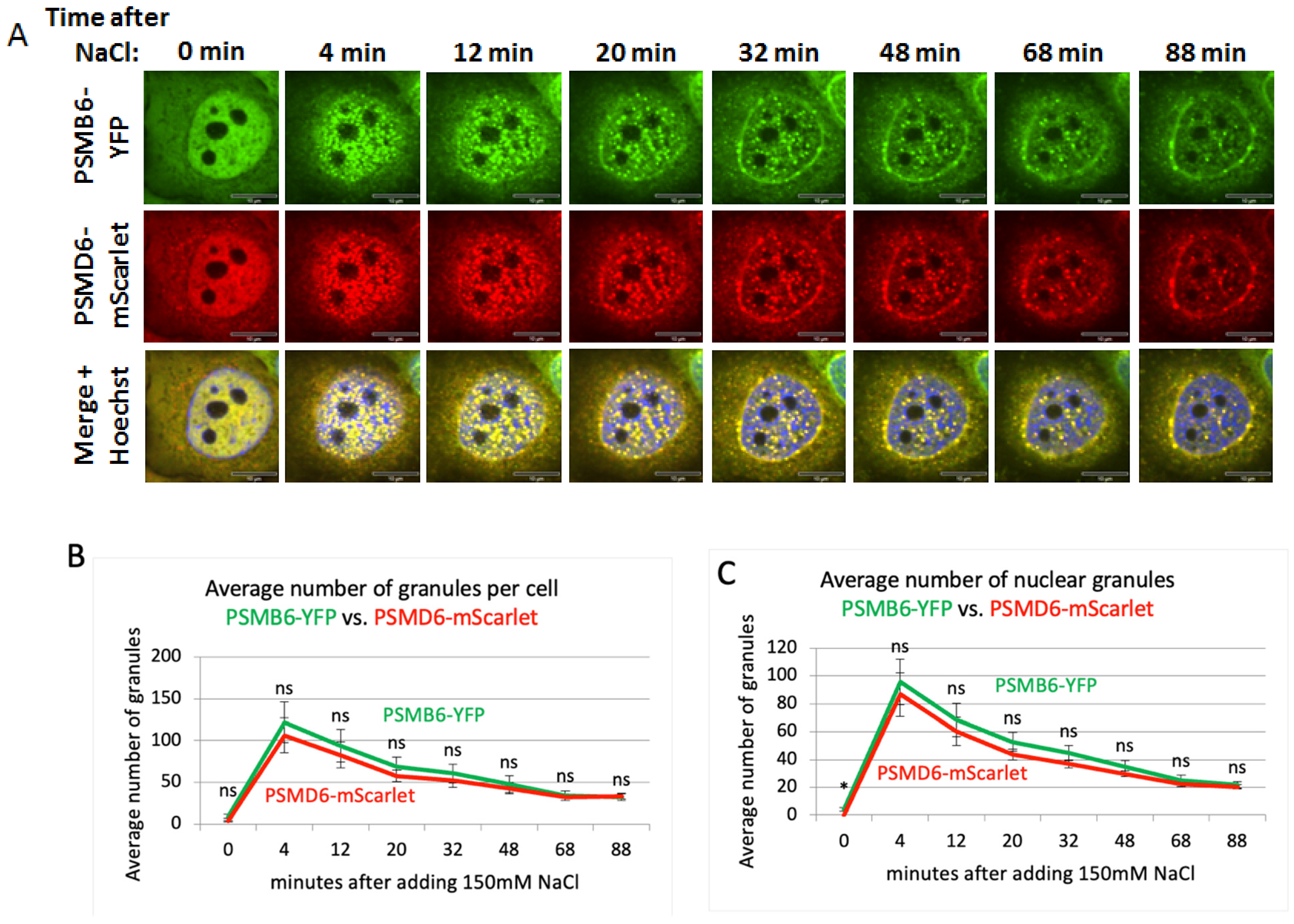
Nuclear localization of the 20S proteasome in 19S knockdown cells
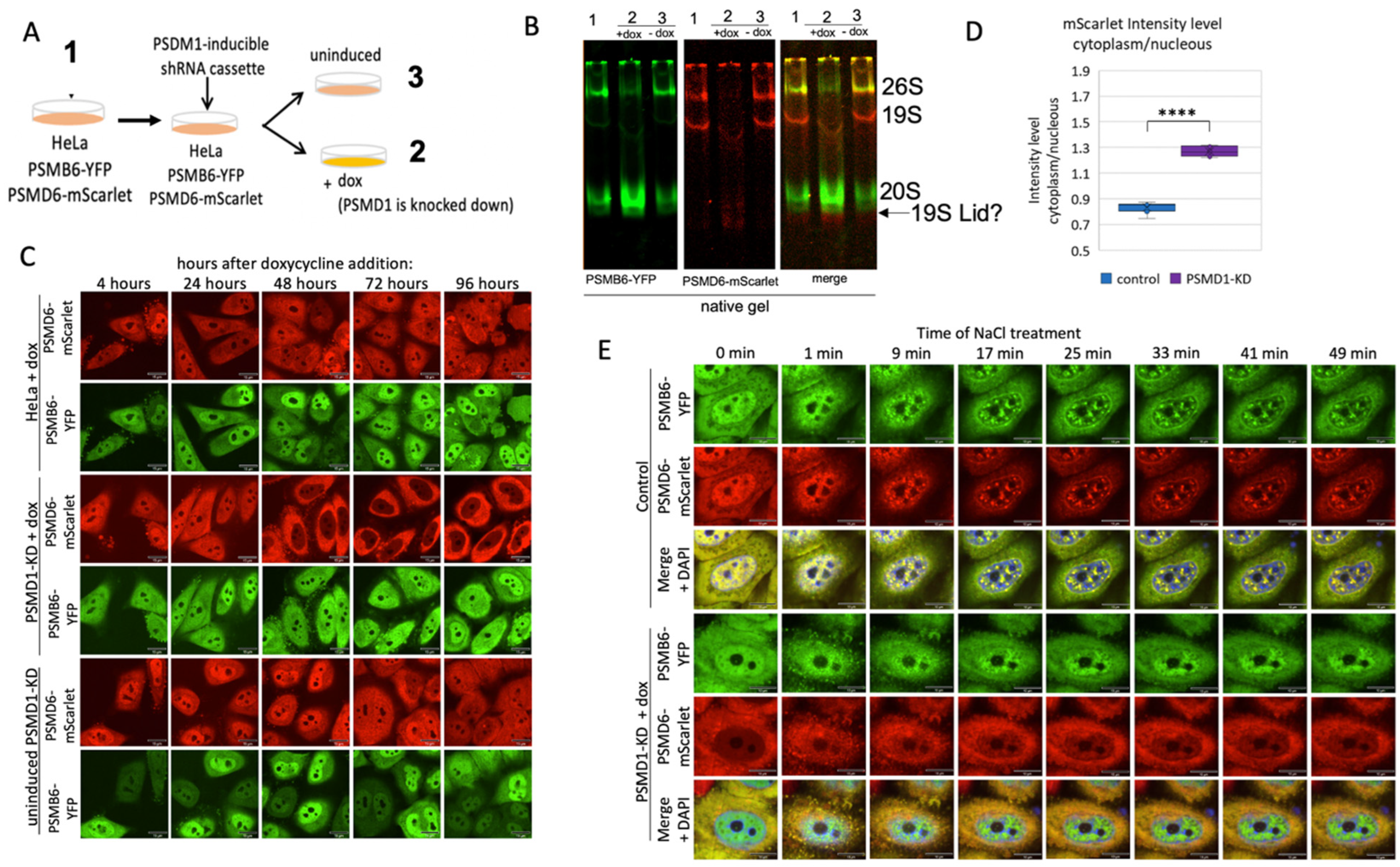
PSMA3 C-terminus regulates 26S proteasome integrity.
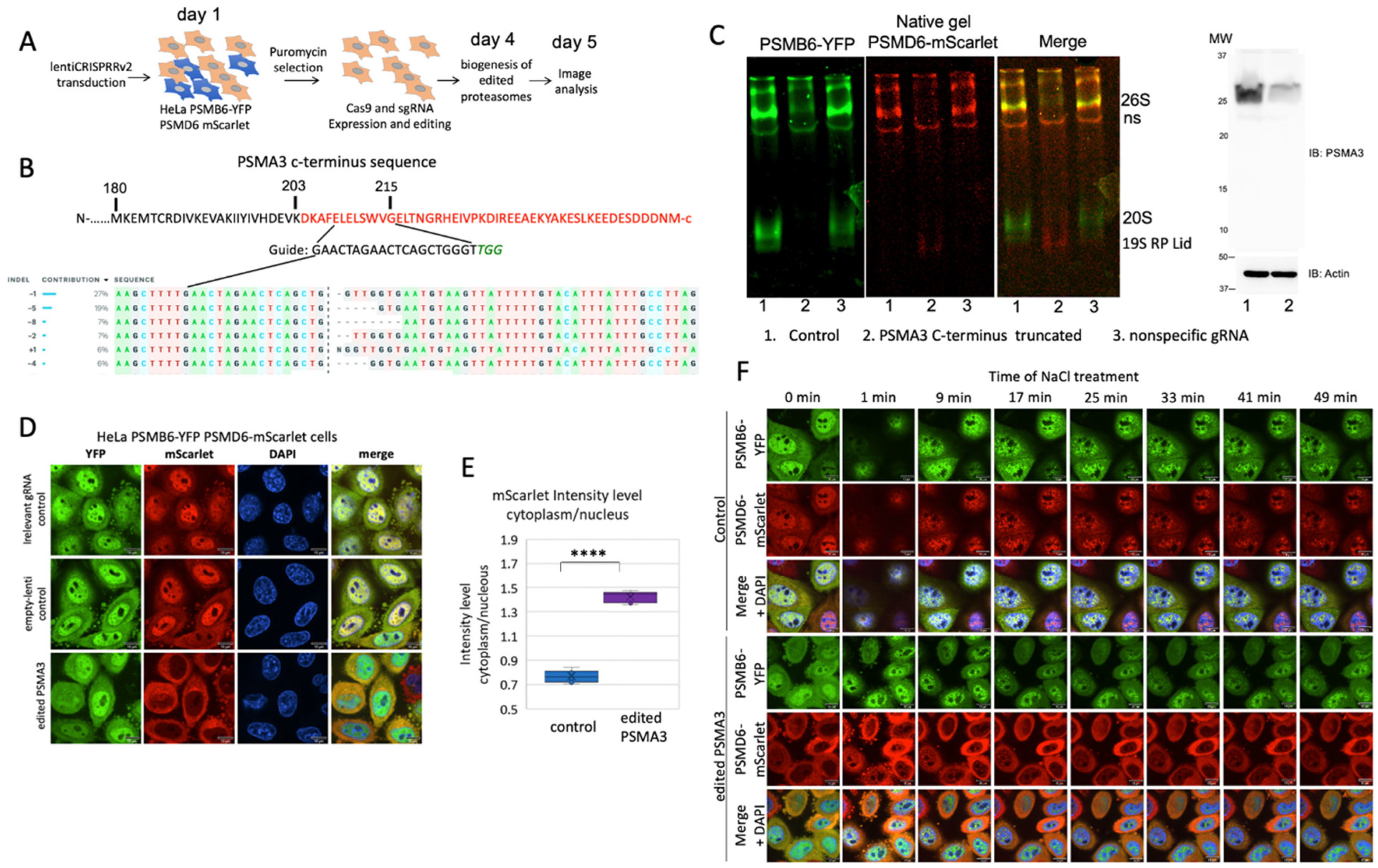
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Amzallag, E., & Hornstein, E. (2022). Crosstalk between Biomolecular Condensates and Proteostasis. Cells, 11(15). https://doi.org/10.3390/cells11152415. [CrossRef]
- Asher, G., Reuven, N., & Shaul, Y. (2006). 20S proteasomes and protein degradation “by default”. Bioessays: News and Reviews in Molecular, Cellular and Developmental Biology, 28(8), 844–849. https://doi.org/10.1002/bies.20447. [CrossRef]
- Biran, A., Myers, N., Steinberger, S., Adler, J., Riutin, M., Broennimann, K., Reuven, N., & Shaul, Y. (2022). The C-Terminus of the PSMA3 Proteasome Subunit Preferentially Traps Intrinsically Disordered Proteins for Degradation. Cells, 11(20). https://doi.org/10.3390/cells11203231. [CrossRef]
- Bose, S., Stratford, F. L. L., Broadfoot, K. I., Mason, G. G. F., & Rivett, A. J. (2004). Phosphorylation of 20S proteasome alpha subunit C8 (alpha7) stabilizes the 26S proteasome and plays a role in the regulation of proteasome complexes by gamma-interferon. The Biochemical Journal, 378(Pt 1), 177–184. https://doi.org/10.1042/BJ20031122. [CrossRef]
- Chen, X., & Walters, K. J. (2022). Nuclear destruction: A suicide mission by AKIRIN2 brings intact proteasomes into the nucleus. Molecular Cell, 82(1), 13–14. https://doi.org/10.1016/j.molcel.2021.11.020. [CrossRef]
- Coffino, P., & Cheng, Y. (2022). Allostery Modulates Interactions between Proteasome Core Particles and Regulatory Particles. Biomolecules, 12(6). https://doi.org/10.3390/biom12060764. [CrossRef]
- Cohen, D., Adamovich, Y., Reuven, N., & Shaul, Y. (2010). Hepatitis B virus activates deoxynucleotide synthesis in nondividing hepatocytes by targeting the R2 gene. Hepatology, 51(5), 1538–1546. https://doi.org/10.1002/hep.23519. [CrossRef]
- Coleman, R. A., & Trader, D. J. (2018). Development and application of a sensitive peptide reporter to discover 20S proteasome stimulators. ACS Combinatorial Science, 20(5), 269–276. https://doi.org/10.1021/acscombsci.7b00193. [CrossRef]
- de Almeida, M., Hinterndorfer, M., Brunner, H., Grishkovskaya, I., Singh, K., Schleiffer, A., Jude, J., Deswal, S., Kalis, R., Vunjak, M., Lendl, T., Imre, R., Roitinger, E., Neumann, T., Kandolf, S., Schutzbier, M., Mechtler, K., Versteeg, G. A., Haselbach, D., & Zuber, J. (2021). AKIRIN2 controls the nuclear import of proteasomes in vertebrates. Nature, 599(7885), 491–496. https://doi.org/10.1038/s41586-021-04035-8. [CrossRef]
- Dong, Y., Zhang, S., Wu, Z., Li, X., Wang, W. L., Zhu, Y., Stoilova-McPhie, S., Lu, Y., Finley, D., & Mao, Y. (2019). Cryo-EM structures and dynamics of substrate-engaged human 26S proteasome. Nature, 565(7737), 49–55. https://doi.org/10.1038/s41586-018-0736-4. [CrossRef]
- Enenkel, C., Kang, R. W., Wilfling, F., & Ernst, O. P. (2022). Intracellular localization of the proteasome in response to stress conditions. The Journal of Biological Chemistry, 298(7), 102083. https://doi.org/10.1016/j.jbc.2022.102083. [CrossRef]
- Fabre, B., Lambour, T., Garrigues, L., Ducoux-Petit, M., Amalric, F., Monsarrat, B., Burlet-Schiltz, O., & Bousquet-Dubouch, M.-P. (2014). Label-free quantitative proteomics reveals the dynamics of proteasome complexes composition and stoichiometry in a wide range of human cell lines. Journal of Proteome Research, 13(6), 3027–3037. https://doi.org/10.1021/pr500193k. [CrossRef]
- Fu, A., Cohen-Kaplan, V., Avni, N., Livneh, I., & Ciechanover, A. (2021). p62-containing, proteolytically active nuclear condensates, increase the efficiency of the ubiquitin-proteasome system. Proceedings of the National Academy of Sciences of the United States of America, 118(33). https://doi.org/10.1073/pnas.2107321118. [CrossRef]
- Gaczynska, M., Osmulski, P. A., Gao, Y., Post, M. J., & Simons, M. (2003). Proline- and arginine-rich peptides constitute a novel class of allosteric inhibitors of proteasome activity. Biochemistry, 42(29), 8663–8670. https://doi.org/10.1021/bi034784f. [CrossRef]
- Gan, J., Leestemaker, Y., Sapmaz, A., & Ovaa, H. (2019). Highlighting the proteasome: using fluorescence to visualize proteasome activity and distribution. Frontiers in Molecular Biosciences, 6, 14. https://doi.org/10.3389/fmolb.2019.00014. [CrossRef]
- Glickman, M. H., Rubin, D. M., Coux, O., Wefes, I., Pfeifer, G., Cjeka, Z., Baumeister, W., Fried, V. A., & Finley, D. (1998). A subcomplex of the proteasome regulatory particle required for ubiquitin-conjugate degradation and related to the COP9-signalosome and eIF3. Cell, 94(5), 615–623. https://doi.org/10.1016/s0092-8674(00)81603-7. [CrossRef]
- Glickman, M. H., Rubin, D. M., Fried, V. A., & Finley, D. (1998). The regulatory particle of the Saccharomyces cerevisiae proteasome. Molecular and Cellular Biology, 18(6), 3149–3162. https://doi.org/10.1128/MCB.18.6.3149. [CrossRef]
- Goldberg, A. L., Kim, H. T., Lee, D., & Collins, G. A. (2021). Mechanisms that activate 26S proteasomes and enhance protein degradation. Biomolecules, 11(6). https://doi.org/10.3390/biom11060779. [CrossRef]
- Greene, E. R., Dong, K. C., & Martin, A. (2020). Understanding the 26S proteasome molecular machine from a structural and conformational dynamics perspective. Current Opinion in Structural Biology, 61, 33–41. https://doi.org/10.1016/j.sbi.2019.10.004. [CrossRef]
- Groll, M., Bajorek, M., Köhler, A., Moroder, L., Rubin, D. M., Huber, R., Glickman, M. H., & Finley, D. (2000). A gated channel into the proteasome core particle. Nature Structural Biology, 7(11), 1062–1067. https://doi.org/10.1038/80992. [CrossRef]
- Groll, M., Ditzel, L., Löwe, J., Stock, D., Bochtler, M., Bartunik, H. D., & Huber, R. (1997). Structure of 20S proteasome from yeast at 2.4 A resolution. Nature, 386(6624), 463–471. https://doi.org/10.1038/386463a0. [CrossRef]
- Guo, X. (2022). Localized proteasomal degradation: from the nucleus to cell periphery. Biomolecules, 12(2). https://doi.org/10.3390/biom12020229. [CrossRef]
- Gu, Z. C., & Enenkel, C. (2014). Proteasome assembly. Cellular and Molecular Life Sciences, 71(24), 4729–4745. https://doi.org/10.1007/s00018-014-1699-8. [CrossRef]
- Hobbs, S., Jitrapakdee, S., & Wallace, J. C. (1998). Development of a bicistronic vector driven by the human polypeptide chain elongation factor 1alpha promoter for creation of stable mammalian cell lines that express very high levels of recombinant proteins. Biochemical and Biophysical Research Communications, 252(2), 368–372. https://doi.org/10.1006/bbrc.1998.9646. [CrossRef]
- Kumar Deshmukh, F., Yaffe, D., Olshina, M. A., Ben-Nissan, G., & Sharon, M. (2019). The contribution of the 20S proteasome to proteostasis. Biomolecules, 9(5). https://doi.org/10.3390/biom9050190. [CrossRef]
- Levy, D., Adamovich, Y., Reuven, N., & Shaul, Y. (2007). The Yes-associated protein 1 stabilizes p73 by preventing Itch-mediated ubiquitination of p73. Cell Death and Differentiation, 14(4), 743–751. https://doi.org/10.1038/sj.cdd.4402063. [CrossRef]
- Myers, N., Olender, T., Savidor, A., Levin, Y., Reuven, N., & Shaul, Y. (2018). The Disordered Landscape of the 20S Proteasome Substrates Reveals Tight Association with Phase Separated Granules. Proteomics, 18(21–22), e1800076. https://doi.org/10.1002/pmic.201800076. [CrossRef]
- Njomen, E., Osmulski, P. A., Jones, C. L., Gaczynska, M., & Tepe, J. J. (2018). Small molecule modulation of proteasome assembly. Biochemistry, 57(28), 4214–4224. https://doi.org/10.1021/acs.biochem.8b00579. [CrossRef]
- Religa, T. L., Sprangers, R., & Kay, L. E. (2010). Dynamic regulation of archaeal proteasome gate opening as studied by TROSY NMR. Science, 328(5974), 98–102. https://doi.org/10.1126/science.1184991. [CrossRef]
- Reuven, N., Adler, J., Broennimann, K., Myers, N., & Shaul, Y. (2019). Recruitment of DNA Repair MRN Complex by Intrinsically Disordered Protein Domain Fused to Cas9 Improves Efficiency of CRISPR-Mediated Genome Editing. Biomolecules, 9(10). https://doi.org/10.3390/biom9100584. [CrossRef]
- Reuven, N., Adler, J., Myers, N., & Shaul, Y. (2021). CRISPR Co-Editing Strategy for Scarless Homology-Directed Genome Editing. International Journal of Molecular Sciences, 22(7). https://doi.org/10.3390/ijms22073741. [CrossRef]
- Saeed, I. A., & Ashraf, S. S. (2009). Denaturation studies reveal significant differences between GFP and blue fluorescent protein. International Journal of Biological Macromolecules, 45(3), 236–241. https://doi.org/10.1016/j.ijbiomac.2009.05.010. [CrossRef]
- Sahu, I., Mali, S. M., Sulkshane, P., Xu, C., Rozenberg, A., Morag, R., Sahoo, M. P., Singh, S. K., Ding, Z., Wang, Y., Day, S., Cong, Y., Kleifeld, O., Brik, A., & Glickman, M. H. (2021). The 20S as a stand-alone proteasome in cells can degrade the ubiquitin tag. Nature Communications, 12(1), 6173. https://doi.org/10.1038/s41467-021-26427-0. [CrossRef]
- Schindelin, J., Arganda-Carreras, I., Frise, E., Kaynig, V., Longair, M., Pietzsch, T., Preibisch, S., Rueden, C., Saalfeld, S., Schmid, B., Tinevez, J.-Y., White, D. J., Hartenstein, V., Eliceiri, K., Tomancak, P., & Cardona, A. (2012). Fiji: an open-source platform for biological-image analysis. Nature Methods, 9(7), 676–682. https://doi.org/10.1038/nmeth.2019. [CrossRef]
- Stadtmueller, B. M., & Hill, C. P. (2011). Proteasome activators. Molecular Cell, 41(1), 8–19. https://doi.org/10.1016/j.molcel.2010.12.020. [CrossRef]
- Tanahashi, N., Murakami, Y., Minami, Y., Shimbara, N., Hendil, K. B., & Tanaka, K. (2000). Hybrid proteasomes. Induction by interferon-gamma and contribution to ATP-dependent proteolysis. The Journal of Biological Chemistry, 275(19), 14336–14345. https://doi.org/10.1074/jbc.275.19.14336. [CrossRef]
- Tsvetkov, P., Adler, J., Myers, N., Biran, A., Reuven, N., & Shaul, Y. (2018). Oncogenic addiction to high 26S proteasome level. Cell Death & Disease, 9(7), 773. https://doi.org/10.1038/s41419-018-0806-4. [CrossRef]
- Tsvetkov, P., Myers, N., Adler, J., & Shaul, Y. (2020). Degradation of intrinsically disordered proteins by the NADH 26S proteasome. Biomolecules, 10(12). https://doi.org/10.3390/biom10121642. [CrossRef]
- Tsvetkov, P., Myers, N., Eliav, R., Adamovich, Y., Hagai, T., Adler, J., Navon, A., & Shaul, Y. (2014). NADH binds and stabilizes the 26S proteasomes independent of ATP. The Journal of Biological Chemistry, 289(16), 11272–11281. https://doi.org/10.1074/jbc.M113.537175. [CrossRef]
- Uriarte, M., Sen Nkwe, N., Tremblay, R., Ahmed, O., Messmer, C., Mashtalir, N., Barbour, H., Masclef, L., Voide, M., Viallard, C., Daou, S., Abdelhadi, D., Ronato, D., Paydar, M., Darracq, A., Boulay, K., Desjardins-Lecavalier, N., Sapieha, P., Masson, J.-Y., … Affar, E. B. (2021). Starvation-induced proteasome assemblies in the nucleus link amino acid supply to apoptosis. Nature Communications, 12(1), 6984. https://doi.org/10.1038/s41467-021-27306-4. [CrossRef]
- Wang, X., Chemmama, I. E., Yu, C., Huszagh, A., Xu, Y., Viner, R., Block, S. A., Cimermancic, P., Rychnovsky, S. D., Ye, Y., Sali, A., & Huang, L. (2017). The proteasome-interacting Ecm29 protein disassembles the 26S proteasome in response to oxidative stress. The Journal of Biological Chemistry, 292(39), 16310–16320. https://doi.org/10.1074/jbc.M117.803619. [CrossRef]
- Wani, P. S., Suppahia, A., Capalla, X., Ondracek, A., & Roelofs, J. (2016). Phosphorylation of the C-terminal tail of proteasome subunit α7 is required for binding of the proteasome quality control factor Ecm29. Scientific Reports, 6, 27873. https://doi.org/10.1038/srep27873. [CrossRef]
- Welk, V., Coux, O., Kleene, V., Abeza, C., Trümbach, D., Eickelberg, O., & Meiners, S. (2016). Inhibition of proteasome activity induces formation of alternative proteasome complexes. The Journal of Biological Chemistry, 291(25), 13147–13159. https://doi.org/10.1074/jbc.M116.717652. [CrossRef]
- Wójcik, C., & DeMartino, G. N. (2003). Intracellular localization of proteasomes. The International Journal of Biochemistry & Cell Biology, 35(5), 579–589. https://doi.org/10.1016/s1357-2725(02)00380-1. [CrossRef]
- Yasuda, S., Tsuchiya, H., Kaiho, A., Guo, Q., Ikeuchi, K., Endo, A., Arai, N., Ohtake, F., Murata, S., Inada, T., Baumeister, W., Fernández-Busnadiego, R., Tanaka, K., & Saeki, Y. (2020). Stress- and ubiquitylation-dependent phase separation of the proteasome. Nature, 578(7794), 296–300. https://doi.org/10.1038/s41586-020-1982-9. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



