Submitted:
10 May 2023
Posted:
11 May 2023
Read the latest preprint version here
Abstract
Keywords:
- How do QKE and VQC algorithms compare to classical machine learning methods such as XGBoost, Ridge, Lasso, LightGBM, CatBoost, and MLP regarding accuracy and efficiency on simulated quantum circuits?
- To what extent can randomized search make the performance of quantum algorithms comparable to classical approaches?
- What are the limitations and challenges associated with the current state of quantum machine learning, and how can future research address these challenges to unlock the full potential of quantum computing in machine learning applications?
1. Related Work
2. Methodology
3. Supervised Machine Learning
3.1. Classical Supervised Machine Learning Techniques
-
Lasso and Ridge Regression/Classification: Lasso (Least Absolute Shrinkage and Selection Operator) and Ridge Regression are linear regression techniques that incorporate regularization to prevent overfitting and improve model generalization [10,28]. Lasso uses L1 regularization, which tends to produce sparse solutions, while Ridge Regression uses L2 regularization, which prevents coefficients from becoming too large.Both of these regression algorithms can also be used for classification tasks.
- Multilayer Perceptron (MLP): MLP is a type of feedforward artificial neural network with multiple layers of neurons, including input, hidden, and output layers [14]. MLPs are capable of modeling complex non-linear relationships and can be trained using backpropagation.
- Support Vector Machines (SVM): SVMs are supervised learning models used for classification and regression tasks [29]. They work by finding the optimal hyperplane that separates the data into different classes, maximizing the margin between the classes.
- Gradient Boosting Machines: Gradient boosting machines are an ensemble learning method that builds a series of weak learners, typically decision trees, to form a strong learner [30]. The weak learners are combined by iteratively adding them to the model while minimizing a loss function. Notable gradient boosting machines for classification tasks include XGBoost [9], CatBoost [13], and LightGBM [12]. These three algorithms have introduced various improvements and optimizations to the original gradient boosting framework, such as efficient tree learning algorithms, handling categorical features, and reducing memory usage.
3.2. Quantum Machine Learning
- Variational Quantum Classifier (VQC): VQC is a hybrid quantum-classical algorithm that can be viewed as a quantum analog of classical neural networks, specifically the Multilayer Perceptron (MLP) [4]. VQC employs a parametrized quantum circuit, which is trained using classical optimization techniques to find the optimal parameters for classification tasks. The learned quantum circuit can then be used to classify new data points.
- Quantum Kernel Estimator (QKE): QKE is a technique that leverages the quantum computation of kernel functions to enhance the performance of classical kernel methods, such as Support Vector Machines (SVM) [32]. By computing the kernel matrix using quantum circuits, QKE can capture complex data relationships that may be challenging for classical kernel methods to exploit.
3.3. Qiskit Machine Learning
3.4. Accuracy Score for Classification
3.5. Data Sets
- Iris Data Set: A widely known data set consisting of 150 samples of iris flowers, each with four features (sepal length, sepal width, petal length, and petal width) and one of three species labels (Iris Setosa, Iris Versicolor, or Iris Virginica). This data set is included in the Scikit-learn library [15].
- Wine Data Set: A popular data set for wine classification, which consists of 178 samples of wine, each with 13 features (such as alcohol content, color intensity, and hue) and one of three class labels (class 1, class 2, or class 3). This data set is also available in the Scikit-learn library [15].
- Indian Liver Patient Dataset (LPD): This data set contains 583 records, with 416 liver patient records and 167 non-liver patient records [33]. The data set includes ten variables: age, gender, total bilirubin, direct bilirubin, total proteins, albumin, A/G ratio, SGPT, SGOT, and Alkphos. The primary task is to classify patients into liver or non-liver patient groups.
- Breast Cancer Coimbra Dataset: This data set consists of 10 quantitative predictors and a binary dependent variable, indicating the presence or absence of breast cancer [34?]. The predictors are anthropometric data and parameters obtainable from routine blood analysis. Accurate prediction models based on these predictors can potentially serve as a biomarker for breast cancer.
- Teaching Assistant Evaluation Dataset:This data set includes 151 instances of teaching assistant (TA) assignments from the Statistics Department at the University of Wisconsin-Madison, with evaluations of their teaching performance over three regular semesters and two summer semesters [35,36]. The class variable is divided into three roughly equal-sized categories ("low", "medium", and "high"). There are six attributes, including whether the TA is a native English speaker, the course instructor, the course, the semester type (summer or regular), and the class size.
- Impedance Spectrum of Breast Tissue Dataset: This data set contains impedance measurements of freshly excised breast tissue at the following frequencies: 15.625, 31.25, 62.5, 125, 250, 500, and 1000 KHz [37,38]. The primary task is to predict the classification of either the original six classes or four classes by merging the fibro-adenoma, mastopathy, and glandular classes whose discrimination is not crucial.
4. Experimental Design
4.1. Artificially Generated Data Sets
4.2. Benchmark Data Sets and Hyperparameter Optimization
5. Results
5.1. Performance on Artificially Generated Data Sets
| Algorithm/Parametrization | Size 50 | Size 100 | Size 250 | Size 500 | Size 1000 | Size 1500 | Size 2000 |
|---|---|---|---|---|---|---|---|
| CatBoost, OutOfTheBox | 1.0 | 1.0 | 0.98 | 0.97 | 0.925 | 0.93 | 0.9425 |
| CatBoost, RandomizedSearchCV | 1.0 | 1.0 | 0.96 | 0.95 | 0.93 | 0.94 | 0.9425 |
| QKE, PauliFeatureMap, statevector-simulator, 1000.0 | 1.0 | 1.0 | 0.96 | 0.93 | 0.93 | 0.93 | 0.925 |
| XGBoost, RandomizedSearchCV | 1.0 | 0.95 | 0.94 | 0.95 | 0.935 | 0.936667 | 0.95 |
| SVM, RandomizedSearchCV | 1.0 | 1.0 | 0.92 | 0.96 | 0.94 | 0.9 | 0.94 |
| XGBoost, OutOfTheBox | 1.0 | 0.95 | 0.94 | 0.96 | 0.91 | 0.936667 | 0.95 |
| SVM, OutOfTheBox | 1.0 | 1.0 | 0.92 | 0.92 | 0.94 | 0.933333 | 0.93 |
| QKE, ZZFeatureMap, statevector-simulator, 177.82794100389228 | 1.0 | 1.0 | 0.94 | 0.93 | 0.915 | 0.926667 | 0.9225 |
| QKE, ZFeatureMap, statevector-simulator, 5.623413251903491 | 1.0 | 1.0 | 0.92 | 0.91 | 0.925 | 0.93 | 0.9375 |
| MLP, OutOfTheBox | 1.0 | 1.0 | 0.94 | 0.89 | 0.905 | 0.916667 | 0.9275 |
| MLP, RandomizedSearchCV | 1.0 | 0.95 | 0.96 | 0.88 | 0.9 | 0.933333 | 0.94 |
| Ridge, OutOfTheBox | 1.0 | 1.0 | 0.94 | 0.88 | 0.9 | 0.896667 | 0.9025 |
| Ridge, RandomizedSearchCV | 1.0 | 1.0 | 0.94 | 0.88 | 0.88 | 0.893333 | 0.9025 |
| QKE, ZFeatureMap, qasm-simulator, 5.623413251903491 | 1.0 | 1.0 | 0.94 | 0.82 | 0.91 | 0.92 | 0.9025 |
| Lasso, RandomizedSearchCV | 1.0 | 1.0 | 0.94 | 0.86 | 0.895 | 0.9 | 0.8875 |
| QKE, ZZFeatureMap, statevector-simulator, 31.622776601683793 | 1.0 | 0.95 | 0.92 | 0.88 | 0.88 | 0.926667 | 0.9175 |
| QKE, PauliFeatureMap, statevector-simulator, 5.623413251903491 | 1.0 | 0.95 | 0.92 | 0.85 | 0.895 | 0.93 | 0.92 |
| QKE, ZFeatureMap, statevector-simulator, 0.1778279410038923 | 1.0 | 0.95 | 0.9 | 0.88 | 0.9 | 0.92 | 0.9125 |
| QKE, ZFeatureMap, aer-simulator, 0.1778279410038923 | 1.0 | 0.95 | 0.9 | 0.87 | 0.905 | 0.92 | 0.9125 |
| QKE, ZZFeatureMap, qasm-simulator, 5.623413251903491 | 1.0 | 0.95 | 0.92 | 0.86 | 0.89 | 0.91 | 0.9175 |
| QKE, PauliFeatureMap, qasm-simulator, 5.623413251903491 | 1.0 | 0.95 | 0.92 | 0.86 | 0.89 | 0.91 | 0.9175 |
| VQC, ZFeatureMap, EfficientSU2, COBYLA, statevector-simulator | 1.0 | 0.95 | 0.9 | 0.9 | 0.92 | 0.893333 | 0.88 |
| VQC, ZFeatureMap, EfficientSU2, COBYLA, qasm-simulator | 1.0 | 0.95 | 0.9 | 0.88 | 0.92 | 0.91 | 0.845 |
| QKE, PauliFeatureMap, aer-simulator, 1.0 | 0.9 | 0.95 | 0.92 | 0.89 | 0.89 | 0.93 | 0.91 |
| VQC, ZFeatureMap, EfficientSU2, SPSA, qasm-simulator | 1.0 | 0.95 | 0.9 | 0.86 | 0.925 | 0.91 | 0.845 |
| VQC, ZFeatureMap, EfficientSU2, COBYLA, aer-simulator | 1.0 | 0.95 | 0.92 | 0.88 | 0.9 | 0.906667 | 0.8275 |
| VQC, ZFeatureMap, EfficientSU2, SPSA, statevector-simulator | 1.0 | 0.95 | 0.92 | 0.87 | 0.89 | 0.89 | 0.835 |
| VQC, ZFeatureMap, RealAmplitudes, COBYLA, aer-simulator | 1.0 | 0.95 | 0.9 | 0.86 | 0.905 | 0.85 | 0.865 |
| LightGBM, RandomizedSearchCV | 0.4 | 1.0 | 0.96 | 0.95 | 0.935 | 0.933333 | 0.95 |
| LightGBM, OutOfTheBox | 0.4 | 1.0 | 0.96 | 0.94 | 0.925 | 0.936667 | 0.9375 |
| VQC, PauliFeatureMap, EfficientSU2, SPSA, qasm-simulator | 0.9 | 0.75 | 0.9 | 0.84 | 0.89 | 0.86 | 0.8675 |
| VQC, ZFeatureMap, EfficientSU2, NFT, statevector-simulator | 1.0 | 0.95 | 0.86 | 0.72 | 0.9 | 0.776667 | 0.77 |
| QKE, PauliFeatureMap, aer-simulator, 31.622776601683793 | 1.0 | 0.85 | 0.96 | 0.7 | 0.875 | 0.826667 | 0.735 |
| QKE, ZFeatureMap, aer-simulator, 31.622776601683793 | 1.0 | 1.0 | 0.88 | 0.62 | 0.835 | 0.736667 | 0.7475 |
| QKE, PauliFeatureMap, aer-simulator, 1000.0 | 1.0 | 0.85 | 0.96 | 0.58 | 0.87 | 0.826667 | 0.665 |
| VQC, PauliFeatureMap, EfficientSU2, SPSA, aer-simulator | 0.8 | 0.75 | 0.9 | 0.73 | 0.845 | 0.86 | 0.8525 |
| VQC, PauliFeatureMap, EfficientSU2, NFT, statevector-simulator | 0.8 | 0.65 | 0.9 | 0.8 | 0.84 | 0.783333 | 0.8475 |
| QKE, ZFeatureMap, qasm-simulator, 177.82794100389228 | 0.9 | 1.0 | 0.88 | 0.57 | 0.875 | 0.73 | 0.6375 |
| VQC, ZZFeatureMap, EfficientSU2, COBYLA, aer-simulator | 0.7 | 0.7 | 0.9 | 0.71 | 0.82 | 0.826667 | 0.835 |
| VQC, ZZFeatureMap, RealAmplitudes, COBYLA, qasm-simulator | 0.8 | 0.7 | 0.9 | 0.62 | 0.775 | 0.816667 | 0.785 |
| VQC, ZZFeatureMap, RealAmplitudes, NFT, qasm-simulator | 0.7 | 0.7 | 0.9 | 0.86 | 0.775 | 0.786667 | 0.535 |
| VQC, PauliFeatureMap, RealAmplitudes, NFT, qasm-simulator | 0.6 | 0.7 | 0.9 | 0.49 | 0.8 | 0.763333 | 0.78 |
| VQC, ZZFeatureMap, RealAmplitudes, COBYLA, aer-simulator | 0.5 | 0.65 | 0.84 | 0.73 | 0.83 | 0.83 | 0.575 |
| QKE, PauliFeatureMap, aer-simulator, 0.03162277660168379 | 0.4 | 0.35 | 0.9 | 0.65 | 0.86 | 0.923333 | 0.8275 |
| QKE, PauliFeatureMap, aer-simulator, 0.005623413251903491 | 0.4 | 0.35 | 0.9 | 0.49 | 0.75 | 0.766667 | 0.8275 |
| QKE, PauliFeatureMap, qasm-simulator, 0.005623413251903491 | 0.4 | 0.35 | 0.9 | 0.49 | 0.75 | 0.766667 | 0.8275 |
| QKE, ZFeatureMap, statevector-simulator, 0.005623413251903491 | 0.4 | 0.35 | 0.84 | 0.49 | 0.63 | 0.85 | 0.83 |
| VQC, ZFeatureMap, TwoLocal, SPSA, statevector-simulator | 0.7 | 0.65 | 0.52 | 0.51 | 0.52 | 0.493333 | 0.58 |
| QKE, PauliFeatureMap, qasm-simulator, 0.001 | 0.4 | 0.35 | 0.9 | 0.49 | 0.48 | 0.753333 | 0.4975 |
| Lasso, OutOfTheBox | 0.4 | 0.35 | 0.5 | 0.49 | 0.48 | 0.506667 | 0.4975 |
| VQC, ZZFeatureMap, TwoLocal, COBYLA, qasm-simulator | 0.2 | 0.35 | 0.28 | 0.35 | 0.225 | 0.216667 | 0.3975 |
| VQC, PauliFeatureMap, TwoLocal, SPSA, qasm-simulator | 0.2 | 0.35 | 0.26 | 0.38 | 0.185 | 0.223333 | 0.4 |
| VQC, PauliFeatureMap, TwoLocal, COBYLA, statevector-simulator | 0.2 | 0.35 | 0.28 | 0.36 | 0.19 | 0.223333 | 0.39 |
| VQC, PauliFeatureMap, TwoLocal, SPSA, statevector-simulator | 0.2 | 0.35 | 0.28 | 0.36 | 0.19 | 0.223333 | 0.39 |
| Algorithm/Parametrization | Size 50 | Size 100 | Size 250 | Size 500 | Size 1000 | Size 1500 | Size 2000 |
|---|---|---|---|---|---|---|---|
| Lasso, OutOfTheBox | 0.002163 | 0.000836 | 0.000638 | 0.000596 | 0.000571 | 0.000552 | 0.000575 |
| Ridge, OutOfTheBox | 0.007653 | 0.001171 | 0.00121 | 0.001453 | 0.001333 | 0.00142 | 0.001452 |
| SVM, OutOfTheBox | 0.001014 | 0.000671 | 0.001049 | 0.002419 | 0.003195 | 0.005537 | 0.012677 |
| XGBoost, OutOfTheBox | 0.018497 | 0.008215 | 0.009782 | 0.021166 | 0.029465 | 0.032339 | 0.07877 |
| LightGBM, OutOfTheBox | 0.013812 | 0.012791 | 0.01488 | 0.027696 | 0.054648 | 0.04419 | 0.054521 |
| MLP, OutOfTheBox | 0.086815 | 0.090133 | 0.150594 | 0.231203 | 0.437073 | 0.647381 | 0.853657 |
| SVM, RandomizedSearchCV | 1.993112 | 0.410673 | 0.468471 | 0.4341 | 0.658507 | 0.945457 | 0.896971 |
| XGBoost, RandomizedSearchCV | 1.34873 | 0.358433 | 0.426441 | 0.519127 | 0.790996 | 1.049813 | 1.439898 |
| Ridge, RandomizedSearchCV | 2.357919 | 0.298155 | 0.469647 | 0.706193 | 0.662371 | 0.774265 | 0.835577 |
| Lasso, RandomizedSearchCV | 3.401839 | 0.482512 | 0.422395 | 0.473103 | 0.486651 | 0.438773 | 0.480838 |
| CatBoost, OutOfTheBox | 1.045732 | 1.243495 | 0.812176 | 0.762516 | 0.865055 | 1.9962 | 1.268149 |
| LightGBM, RandomizedSearchCV | 2.67037 | 0.536059 | 0.668838 | 0.900941 | 1.628797 | 1.389068 | 1.168039 |
| VQC, ZFeatureMap, TwoLocal, SPSA, statevector-simulator | 0.502447 | 0.82391 | 1.319602 | 2.9078 | 6.75953 | 11.81601 | 18.064725 |
| VQC, PauliFeatureMap, TwoLocal, COBYLA, statevector-simulator | 0.536454 | 0.886945 | 1.757877 | 3.486975 | 8.137821 | 14.688881 | 22.79476 |
| VQC, PauliFeatureMap, TwoLocal, SPSA, statevector-simulator | 1.981785 | 0.715829 | 1.621059 | 3.488372 | 8.517624 | 15.170185 | 22.300972 |
| VQC, PauliFeatureMap, TwoLocal, SPSA, qasm-simulator | 0.750719 | 1.154406 | 2.53449 | 5.000262 | 11.265137 | 19.493945 | 29.031463 |
| VQC, ZZFeatureMap, TwoLocal, COBYLA, qasm-simulator | 0.734865 | 1.097202 | 2.514703 | 4.990832 | 11.895971 | 19.283406 | 29.318269 |
| MLP, RandomizedSearchCV | 3.568304 | 2.701256 | 3.490188 | 8.736222 | 13.817605 | 20.424873 | 38.117078 |
| QKE, ZFeatureMap, statevector-simulator, 0.1778279410038923 | 1.343983 | 0.802286 | 2.170829 | 5.965899 | 18.504546 | 36.659922 | 59.889941 |
| QKE, ZFeatureMap, statevector-simulator, 0.005623413251903491 | 0.411296 | 0.697461 | 2.154164 | 6.122564 | 19.670819 | 37.297334 | 62.1901 |
| QKE, PauliFeatureMap, statevector-simulator, 1000.0 | 0.470933 | 0.956269 | 2.721257 | 7.2817 | 21.356298 | 40.130716 | 67.422908 |
| QKE, PauliFeatureMap, statevector-simulator, 5.623413251903491 | 0.501446 | 0.922237 | 2.775664 | 7.454642 | 21.780637 | 40.426036 | 66.758927 |
| QKE, ZFeatureMap, statevector-simulator, 5.623413251903491 | 0.378018 | 0.757363 | 2.141677 | 4.962464 | 19.901565 | 41.913003 | 71.453831 |
| QKE, ZZFeatureMap, statevector-simulator, 31.622776601683793 | 0.214386 | 0.567282 | 1.650304 | 5.302437 | 20.77629 | 42.614517 | 72.871078 |
| QKE, ZZFeatureMap, statevector-simulator, 177.82794100389228 | 0.461093 | 0.943574 | 2.780804 | 7.580857 | 22.906811 | 41.955521 | 68.045553 |
| CatBoost, RandomizedSearchCV | 8.292872 | 7.778893 | 19.636858 | 43.697806 | 33.126305 | 51.559816 | 42.05208 |
| VQC, ZFeatureMap, RealAmplitudes, COBYLA, aer-simulator | 47.438183 | 63.446748 | 192.148143 | 404.233954 | 1060.291657 | 1619.397205 | 2290.222381 |
| VQC, ZZFeatureMap, RealAmplitudes, COBYLA, qasm-simulator | 43.113636 | 83.175558 | 166.040938 | 421.278374 | 1064.238564 | 1702.893006 | 2719.340939 |
| VQC, ZZFeatureMap, RealAmplitudes, COBYLA, aer-simulator | 45.909504 | 83.201411 | 152.20265 | 509.1956 | 1158.902532 | 1654.065907 | 2603.942577 |
| VQC, ZFeatureMap, EfficientSU2, COBYLA, statevector-simulator | 48.546243 | 81.030425 | 190.958188 | 402.121722 | 1044.855825 | 1807.676357 | 2751.241623 |
| VQC, ZFeatureMap, EfficientSU2, COBYLA, aer-simulator | 57.728111 | 100.590997 | 240.174666 | 507.58709 | 1253.080578 | 2139.855218 | 3196.07247 |
| VQC, ZFeatureMap, EfficientSU2, COBYLA, qasm-simulator | 59.058898 | 100.862056 | 242.285405 | 507.171731 | 1262.650143 | 2151.503499 | 3191.745568 |
| VQC, ZZFeatureMap, EfficientSU2, COBYLA, aer-simulator | 59.651649 | 105.629842 | 254.918442 | 601.245125 | 1335.017904 | 2260.354294 | 3366.65501 |
| QKE, ZFeatureMap, qasm-simulator, 177.82794100389228 | 4.589478 | 13.184805 | 82.633779 | 332.71327 | 1337.102907 | 3020.689579 | 5368.201509 |
| QKE, ZZFeatureMap, qasm-simulator, 5.623413251903491 | 4.352785 | 15.921249 | 97.165028 | 390.472092 | 1573.103197 | 3549.629798 | 6282.670251 |
| QKE, PauliFeatureMap, aer-simulator, 0.03162277660168379 | 3.549125 | 15.094144 | 98.970568 | 393.496921 | 1581.662241 | 3554.962927 | 6317.355669 |
| QKE, PauliFeatureMap, aer-simulator, 0.005623413251903491 | 3.373257 | 15.311538 | 99.2351 | 390.52131 | 1574.108371 | 3555.3048 | 6339.026443 |
| QKE, PauliFeatureMap, qasm-simulator, 0.005623413251903491 | 3.812115 | 19.479307 | 101.289711 | 404.432384 | 1636.24686 | 3642.937393 | 6307.605039 |
| QKE, PauliFeatureMap, aer-simulator, 31.622776601683793 | 3.848578 | 17.062982 | 101.387533 | 408.69903 | 1635.863136 | 3674.976257 | 6555.811507 |
| VQC, ZFeatureMap, EfficientSU2, NFT, statevector-simulator | 98.831974 | 167.48274 | 394.378037 | 836.913451 | 2197.652135 | 3719.047116 | 5621.134708 |
| VQC, PauliFeatureMap, EfficientSU2, NFT, statevector-simulator | 103.914165 | 177.047181 | 423.423603 | 1014.963511 | 2338.078356 | 3953.861723 | 5905.433094 |
| VQC, ZZFeatureMap, RealAmplitudes, NFT, qasm-simulator | 105.987181 | 183.918751 | 427.016702 | 1036.605473 | 2366.463152 | 4052.521035 | 6042.538015 |
| VQC, PauliFeatureMap, RealAmplitudes, NFT, qasm-simulator | 103.625823 | 180.306618 | 425.488049 | 1041.160999 | 2371.366715 | 4044.856475 | 6048.573929 |
| VQC, ZFeatureMap, EfficientSU2, SPSA, statevector-simulator | 119.513477 | 200.101417 | 474.113288 | 1008.932874 | 2601.731917 | 4505.306268 | 6781.089745 |
| VQC, ZFeatureMap, EfficientSU2, SPSA, qasm-simulator | 145.295744 | 256.711762 | 609.791229 | 1272.675059 | 3150.527537 | 5366.116602 | 8009.649075 |
| VQC, PauliFeatureMap, EfficientSU2, SPSA, aer-simulator | 144.280811 | 259.102175 | 625.193096 | 1502.476923 | 3356.340799 | 5689.827615 | 8454.144295 |
| VQC, PauliFeatureMap, EfficientSU2, SPSA, qasm-simulator | 152.666649 | 269.680847 | 642.400747 | 1505.762521 | 3388.662998 | 5709.505826 | 8438.957709 |
| QKE, ZFeatureMap, aer-simulator, 31.622776601683793 | 5.993241 | 25.852654 | 166.703792 | 669.201309 | 2934.169598 | 6729.31411 | 12037.430687 |
| QKE, PauliFeatureMap, qasm-simulator, 5.623413251903491 | 8.384715 | 32.795287 | 206.595473 | 890.414904 | 3753.488868 | 8537.768589 | 15232.745542 |
| QKE, PauliFeatureMap, qasm-simulator, 0.001 | 7.792093 | 32.566225 | 207.832614 | 896.042249 | 3778.324351 | 8610.335147 | 15348.810142 |
| QKE, ZFeatureMap, aer-simulator, 0.1778279410038923 | 10.511296 | 43.335078 | 276.810734 | 1111.545614 | 4799.032996 | 10979.135601 | 19768.073574 |
| QKE, ZFeatureMap, qasm-simulator, 5.623413251903491 | 11.573929 | 43.186982 | 277.291314 | 1113.664313 | 4842.587094 | 10978.908476 | 19798.821156 |
| QKE, PauliFeatureMap, aer-simulator, 1000.0 | 12.596938 | 51.788837 | 332.281104 | 1434.208601 | 5986.631006 | 13592.866065 | 24280.544075 |
| QKE, PauliFeatureMap, aer-simulator, 1.0 | 12.261604 | 51.508959 | 332.561822 | 1423.111135 | 5984.902587 | 13603.956887 | 24362.83202 |
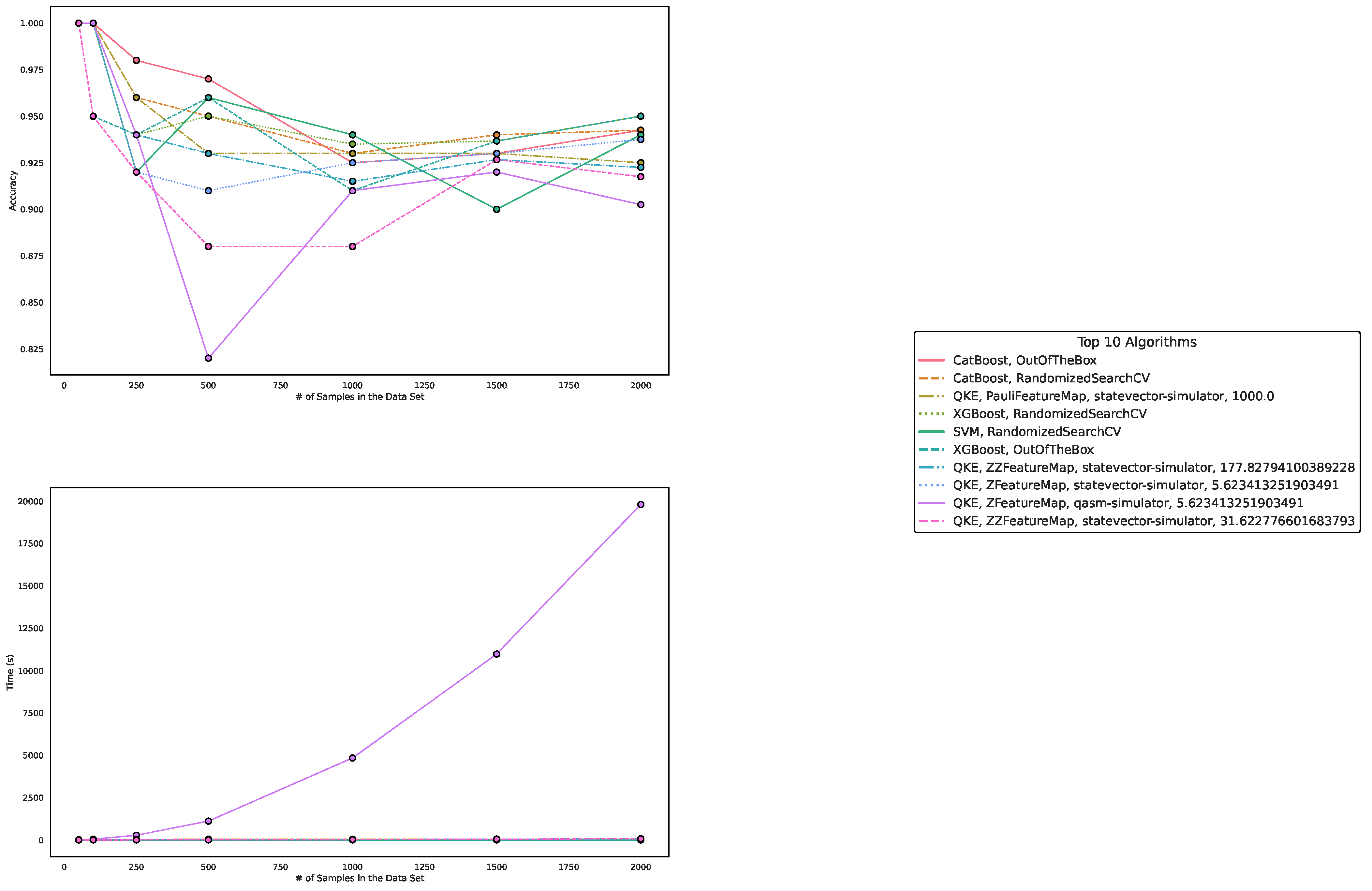
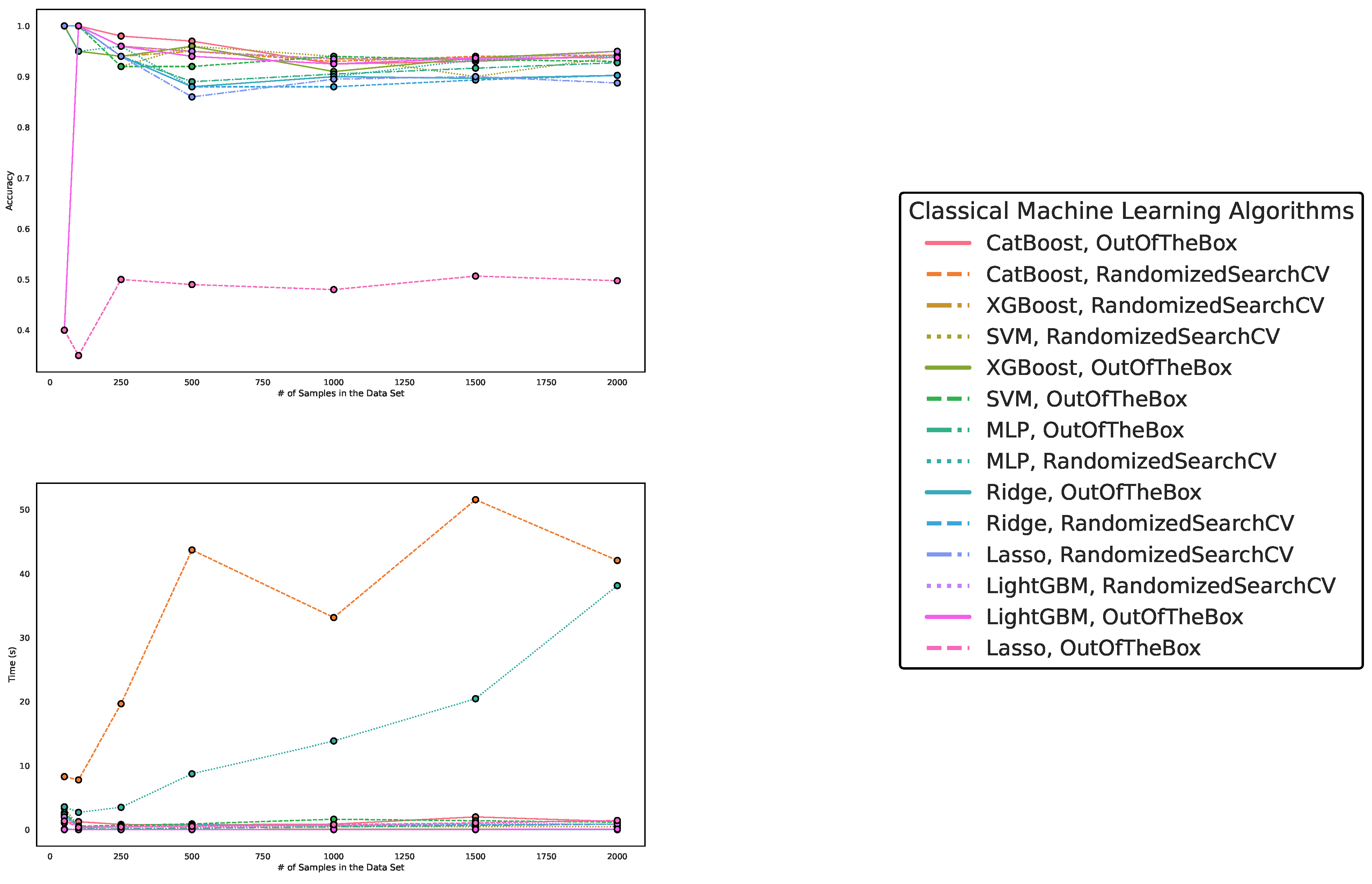
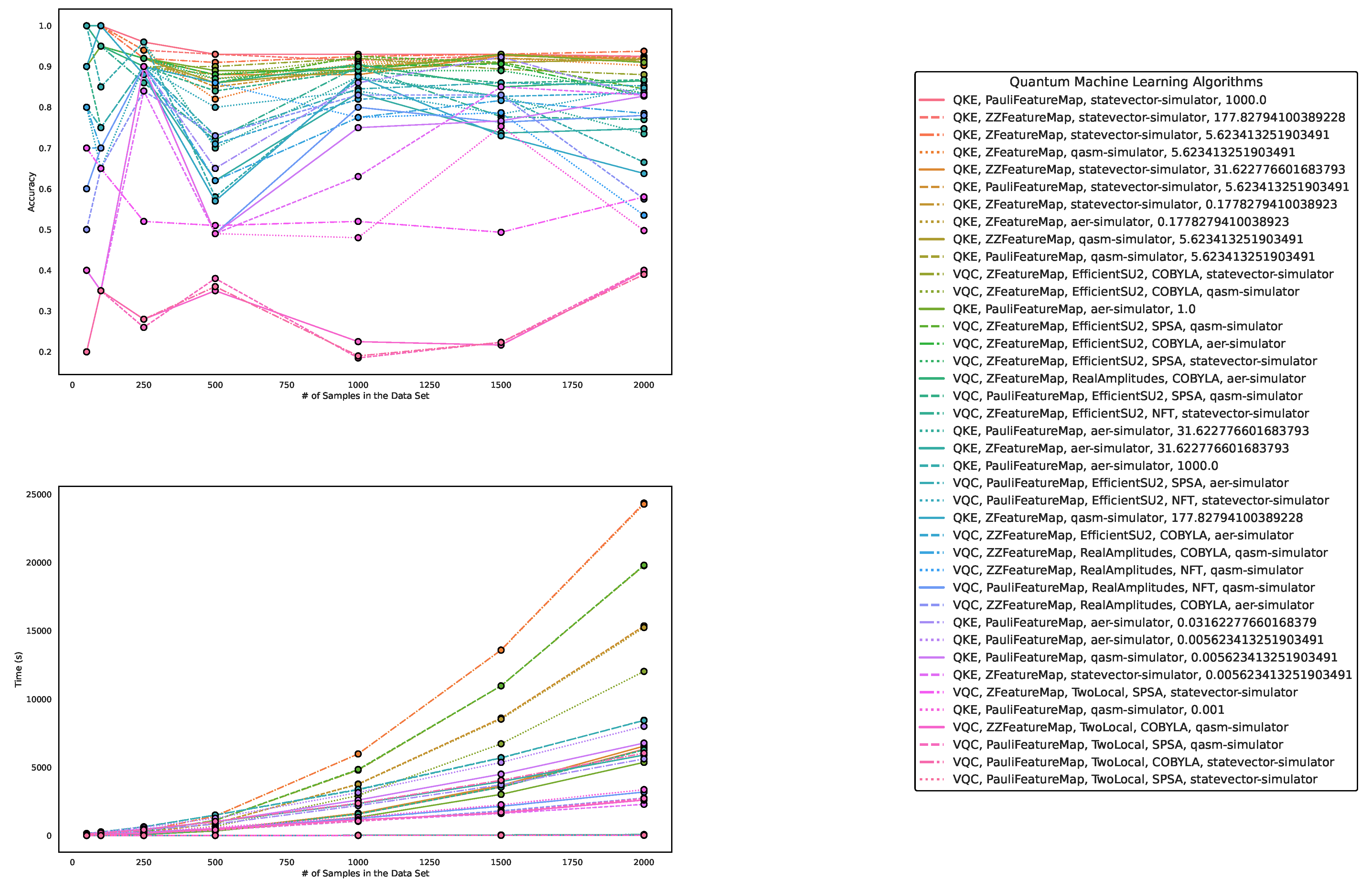
5.2. Results on Benchmark Data Sets
| Classifier\Dataset | Iris | Wine | ILPD | BC-Coimbra | TAE | Breast-Tissue |
|---|---|---|---|---|---|---|
| VQC | 0.817 | 0.817 | 0.706 | 0.599 | 0.417 | 0.339 |
| QKE | 0.908 | 0.853 | 0.706 | 0.620 | 0.483 | 0.382 |
| Ridge | 0.914 | 0.875 | 0.080 | 0.053 | 0.053 | <0.001 |
| Lasso | 0.914 | 0.870 | 0.085 | 0.004 | 0.004 | <0.001 |
| MLP | 0.975 | 0.937 | 0.712 | 0.687 | 0.425 | 0.406 |
| SVM | 0.958 | 0.759 | 0.706 | 0.630 | 0.450 | 0.382 |
| XGBoost | 0.958 | 0.986 | 0.695 | 0.656 | 0.533 | 0.441 |
| LightGBM | 0.967 | 0.986 | 0.699 | 0.666 | 0.475 | 0.393 |
| CatBoost | 0.950 | 0.979 | 0.702 | 0.688 | 0.525 | 0.440 |
| Classifier\Dataset | Iris | Wine | ILPD | BC-Coimbra | TAE | Breast-Tissue |
| VQC | 0.767 | 0.639 | 0.744 | 0.541 | 0.388 | 0.334 |
| QKE | 1.0 | 0.833 | 0.744 | 0.792 | 0.613 | 0.409 |
| Ridge | 0.947 | 0.878 | 0.115 | 0.234 | <0.001 | <0.001 |
| Lasso | 0.945 | 0.882 | 0.115 | 0.296 | <0.001 | <0.001 |
| MLP | 1.0 | 1.0 | 0.769 | 0.875 | 0.387 | 0.455 |
| SVM | 1.0 | 0.972 | 0.743 | 0.875 | 0.355 | 0.455 |
| XGBoost | 1.0 | 1.0 | 0.735 | 0.917 | 0.533 | 0.441 |
| LightGBM | 1.0 | 1.0 | 0.752 | 0.917 | 0.419 | 0.455 |
| CatBoost | 1.0 | 1.0 | 0.744 | 0.917 | 0.645 | 0.545 |
| Classifier\ Dataset |
Iris | Wine | ILPD | BC-Coimbra | TAE | Breast-Tissue |
|---|---|---|---|---|---|---|
| VQC | 3:32:16.547605 | 1 day, 13:56:59.455185 | 2 days, 23:03:26.398856 | 9:55:17.907443 | 2:46:25.921553 | 9:01:58.623806 |
| QKE | 2:03:57.921154 | 21:41:38.738255 | 7 days, 6:30:41.179676 | 5:02:26.430001 | 1:28:54.069725 | 3:37:05.655104 |
| Ridge | 0:00:00.175009 | 0:00:00.496771 | 0:00:00.399229 | 0:00:00.240857 | 0:00:00.209600 | 0:00:00.296966 |
| Lasso | 0:00:00.173051 | 0:00:00.181444 | 0:00:00.237455 | 0:00:00.192257 | 0:00:00.229508 | 0:00:00.225531 |
| MLP | 0:00:16.876288 | 0:00:10.477420 | 0:00:26.748907 | 0:00:10.951229 | 0:00:08.475263 | 0:00:13.729790 |
| SVM | 0:00:00.143353 | 0:00:00.165431 | 0:00:00.484485 | 0:00:00.180694 | 0:00:00.228508 | 0:00:00.226784 |
| XGBoost | 0:00:03.809085 | 0:00:04.030425 | 0:00:04.752627 | 0:00:02.744122 | 0:00:05.820371 | 0:00:06.864497 |
| LightGBM | 0:00:02.971164 | 0:00:03.180770 | 0:00:03.062553 | 0:00:01.462174 | 0:00:03.056615 | 0:00:04.540870 |
| CatBoost | 0:00:06.465975 | 0:00:18.511612 | 0:00:11.352944 | 0:00:07.460460 | 0:00:06.964821 | 0:00:26.639070 |
5.3. Comparison and Discussion
6. Conclusion
Acknowledgments
Appendices
- A. Parametrization
A.1. Ridge

A.2. Lasso
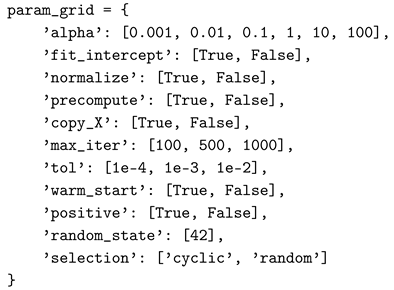

A.3. SVM

A.4. MLP

A.5. XGBoost

A.6. LightGBM

A.7. CatBoost

A.8. QKE

A.9. VQC

References
- Nielsen, M.A.; Chuang, I.L. Quantum computation and quantum information; Cambridge University Press, 2002.
- Biamonte, J.; Wittek, P.; Pancotti, N.; Rebentrost, P.; Wiebe, N.; Lloyd, S. Quantum machine learning. Nature 2017, 549, 195–202.
- Schuld, M.; Sinayskiy, I.; Petruccione, F. An introduction to quantum machine learning. Contemporary Physics 2015, 56, 172–185. [CrossRef]
- Havlíček, V.; Córcoles, A.D.; Temme, K.; Harrow, A.W.; Kandala, A.; Chow, J.M.; Gambetta, J.M. Supervised learning with quantum-enhanced feature spaces. Nature 2019, 567, 209–212.
- Farhi, E., N.H. Classification with quantum neural networks on near term processors. arXiv preprint arXiv:1802.06002 2018.
- Kuppusamy, P.; Yaswanth Kumar, N.; Dontireddy, J.; Iwendi, C. Quantum Computing and Quantum Machine Learning Classification – A Survey. 2022 IEEE 4th International Conference on Cybernetics, Cognition and Machine Learning Applications (ICCCMLA), 2022, pp. 200–204. [CrossRef]
- Blance, A.; Spannowsky, M. Quantum machine learning for particle physics using a variational quantum classifier. Journal of High Energy Physics 2021, 2021, 212. [CrossRef]
- Abohashima, Z.; Elhoseny, M.; Houssein, E.H.; Mohamed, W.M. Classification with Quantum Machine Learning: A Survey. ArXiv 2020, abs/2006.12270.
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining; ACM: New York, NY, USA, 2016; KDD ’16, pp. 785–794. [CrossRef]
- Hoerl A.E., K.R. Ridge regression: Biased estimation for nonorthogonal problems. Technometrics 1970, 12, 55–67.
- R., T. Regression shrinkage and selection via the lasso. Journal of the Royal Statistical Society: Series B (Methodological) 1996, 58, 267–288.
- Ke, G.; Meng, Q.; Finley, T.; Wang, T.; Chen, W.; Ma, W.; Ye, Q.; Liu, T.Y. LightGBM: A Highly Efficient Gradient Boosting Decision Tree. Proceedings of the 31st International Conference on Neural Information Processing Systems; Curran Associates Inc.: Red Hook, NY, USA, 2017; NIPS’17, p. 3149–3157.
- Prokhorenkova, L.; Gusev, G.; Vorobev, A.; Dorogush, A.V.; Gulin, A. CatBoost: Unbiased Boosting with Categorical Features. Proceedings of the 32nd International Conference on Neural Information Processing Systems; Curran Associates Inc.: Red Hook, NY, USA, 2018; NIPS’18, p. 6639–6649.
- Rumelhart, D.E.; Hinton, G.E.; Williams, R.J. Learning internal representations by error propagation. Technical report, California Univ San Diego La Jolla Inst for Cognitive Science, 1985.
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; others. Scikit-learn: Machine learning in Python. Journal of Machine Learning Research 2011, 12, 282–290. Accessed on April 18th, 2023.
- Raubitzek, S. Quantum_Machine_Learning, 2023. doi:not_yet_applicable.
- Zeguendry, A.; Jarir, Z.; Quafafou, M. Quantum Machine Learning: A Review and Case Studies. Entropy 2023, 25. [CrossRef]
- Schuld, M.; Killoran, N. Quantum machine learning in feature Hilbert spaces. Physical Review Letters 2019, 122, 040504.
- Mitarai, K.; Negoro, M.; Kitagawa, M.; Fujii, K. Quantum circuit learning. Physical Review A 2018, 98, 032309.
- Rebentrost, P.; Mohseni, M.; Lloyd, S. Quantum support vector machine for big data classification. Physical Review Letters 2014, 113, 130503.
- Liu, D.; Rebentrost, P. Quantum machine learning for quantum anomaly detection. Physical Review A 2019, 100, 042328. [CrossRef]
- Broughton, M.; Verdon, G.; McCourt, T.; Martinez, A.J.; Yoo, J.H.; Isakov, S.V.; King, A.D.; Smelyanskiy, V.N.; Neven, H. TensorFlow Quantum: A Software Framework for Quantum Machine Learning. arXiv preprint arXiv:2003.02989 2020.
- Qiskit contributors. Qiskit: An Open-source Framework for Quantum Computing, 2023. [CrossRef]
- Bishop, C.M. Pattern Recognition and Machine Learning (Information Science and Statistics); Springer-Verlag: Berlin, Heidelberg, 2006.
- Murphy, K.P. Machine learning : a probabilistic perspective; MIT Press: Cambridge, Mass. [u.a.], 2013.
- Kotsiantis, S.B. Supervised Machine Learning: A Review of Classification Techniques. Proceedings of the 2007 Conference on Emerging Artificial Intelligence Applications in Computer Engineering: Real Word AI Systems with Applications in EHealth, HCI, Information Retrieval and Pervasive Technologies; IOS Press: NLD, 2007; p. 3–24.
- Liu, L.; Özsu, M.T., Eds. Encyclopedia of Database Systems; Springer Reference, Springer: New York, 2009. [CrossRef]
- Tibshirani, R. Regression shrinkage and selection via the lasso. Journal of the Royal Statistical Society: Series B (Methodological) 1996, 58, 267–288.
- Cortes, C.; Vapnik, V. Support-vector networks. Machine Learning 1995, 20, 273–297. [CrossRef]
- Friedman, J.H. Greedy function approximation: A gradient boosting machine. The Annals of Statistics 2001, 29, 1189 – 1232. [CrossRef]
- Biamonte, J.; Wittek, P.; Pancotti, N.; Rebentrost, P.; Wiebe, N.; Lloyd, S. Quantum machine learning. Nature 2017, 549, 195–202. [CrossRef]
- Schuld, M.; Sinayskiy, I. Quantum Machine Learning: An Applied Approach; Cambridge University Press, 2021.
- Ramana, B.V.; Babu, M.S.P.; Venkateswarlu, N.B. LPD (Indian Liver Patient Dataset) Data Set. https://archive.ics.uci.edu/ml/datasets/ILPD+(Indian+Liver+Patient+Dataset), 2012.
- Crisóstomo, J.; Matafome, P.; Santos-Silva, D.; Gomes, A.L.; Gomes, M.; Patrício, M.; Letra, L.; Sarmento-Ribeiro, A.B.; Santos, L.; Seiça, R. Hyperresistinemia and metabolic dysregulation: a risky crosstalk in obese breast cancer. Endocrine 2016, 53, 433–442. [CrossRef]
- Loh, W.Y.; Shih, Y.S. SPLIT SELECTION METHODS FOR CLASSIFICATION TREES. Statistica Sinica 1997, 7, 815–840.
- Lim, T.S.; Loh, W.Y.; Shih, Y.S. A comparison of prediction accuracy, complexity, and training time of thirty-three old and new classification algorithms. Machine learning 2000, 40, 203–228.
- Marques de Sá, J.; Jossinet, J. Breast Tissue Impedance Data Set. https://archive.ics.uci.edu/ml/datasets/Breast+Tissue, 2002.
- Estrela da Silva, J.; Marques de Sá, J.P.; Jossinet, J. Classification of breast tissue by electrical impedance spectroscopy. Medical and Biological Engineering and Computing 2000, 38, 26–30. [CrossRef]
- Schuld, M.; Petruccione, F. Quantum ensembles of quantum classifiers. Scientific Reports 2018, 8, 2772. [CrossRef]
- Moiseyev, N. Non-Hermitian Quantum Mechanics; Cambridge University Press, 2011. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




