Submitted:
20 February 2023
Posted:
23 February 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Convolutional Neural Networks
3. CNN Ensemble Learning
- Data level: by splitting the dataset into different subsets;
- Feature level: by pre-processing the dataset with unique methods;
- Classifier level: by training different classifiers on the same dataset;
- Decision level: by combining the decisions of multiple models.
3.1. Image Pre-Processing
- The "Percentile" method presented in [3], also called "Baseline".
- A "Gaussian" image processing method that encodes each color channel based on the normal distribution of the grayscale intensities of the sixteen images.
- Two "Mean-based" methods focused on utilizing an average or mean of the sixteen images to reconstruct the R, G, and B values.
- The "HSVPP" method that utilizes a different color space composed of hue, saturation, and value of brightness information.
3.1.1. Percentile
- Read the sixteen images;
- Populate a matrix with the grayscale values;
- For each pixel, extract its sixteen grayscale values into a list;
- Sort the list;
- Use elements 2, 8, and 15 as RGB values for the new image;
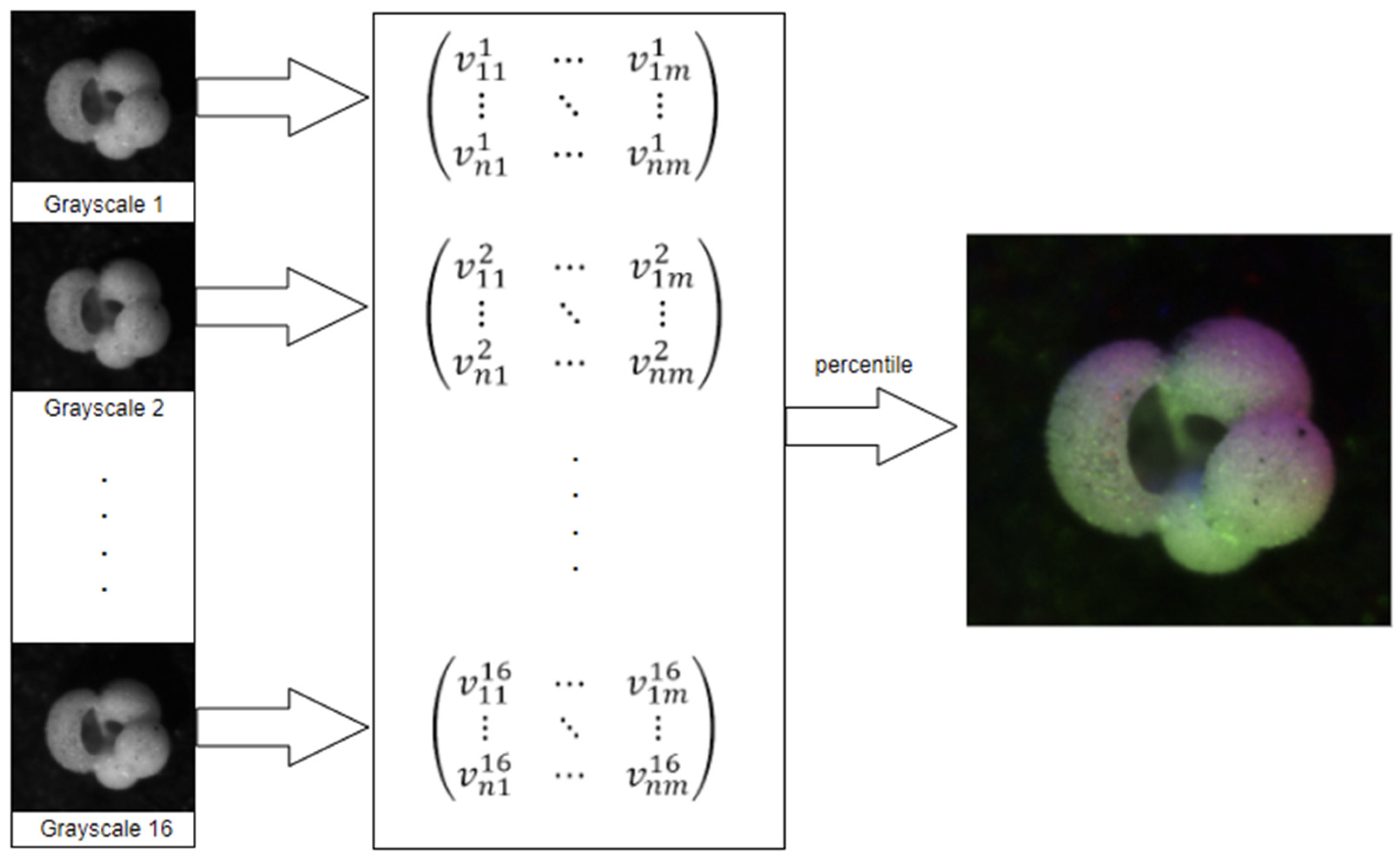
3.1.2. Gaussian
3.1.3. Mean-based
3.1.3.1. Luma Scaling
- .
3.1.3.2. Means Reconstruction
- .
3.1.3. HSVPP: Hue, Saturation, Value of brightness + Post-Processing
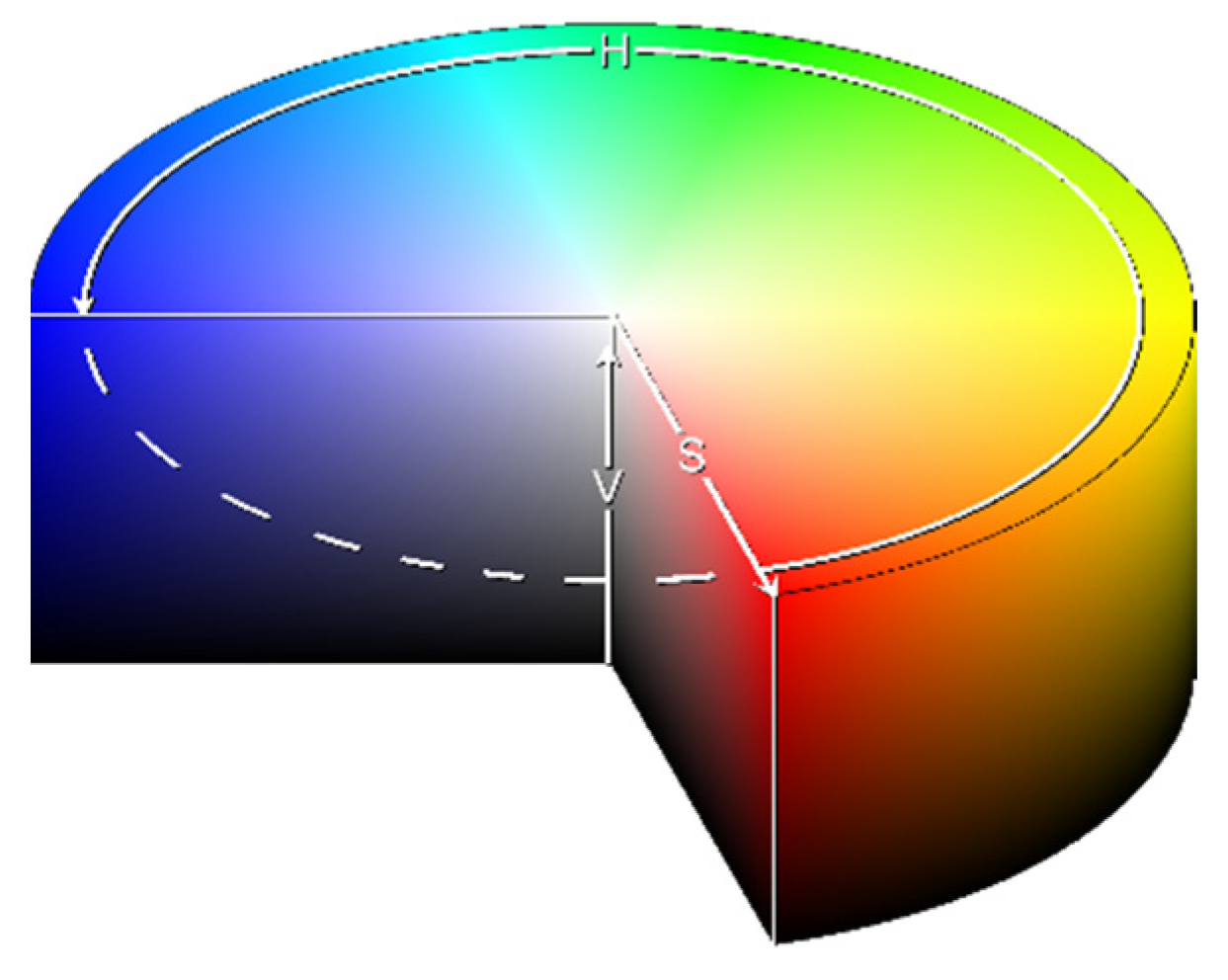
- Hue (H) encodes the angle of the color vector on the HSV space, with 0° being red, 120° being green, and 240° being blue, rescaled to [0,1] during computation;
- Saturation (S) determines how far from the center of the circumference the color is placed in the range [0,1];
- Value of brightness (V) calculates the height in the color space cylinder and measures color luminosity in the range [0,1].
- H is assigned based on the index of the image, giving each a different color hue;
- S is set to 1 by default, for maximum diversity between colors;
- V is set to the grayscale image's original intensity, i.e., its brightness.
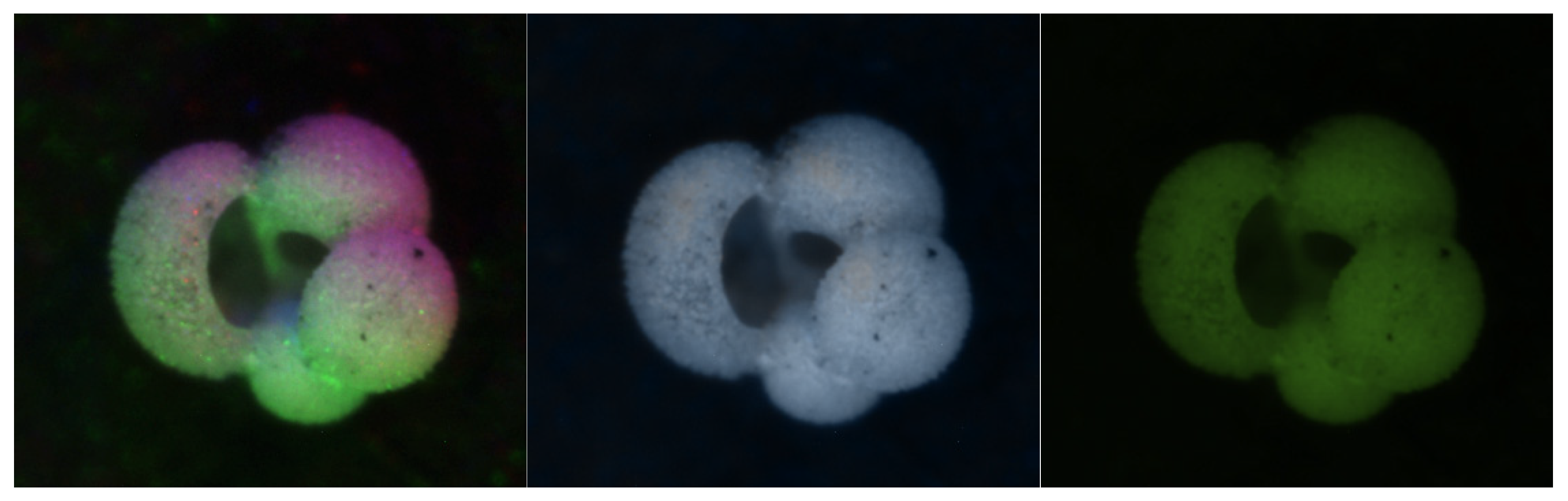
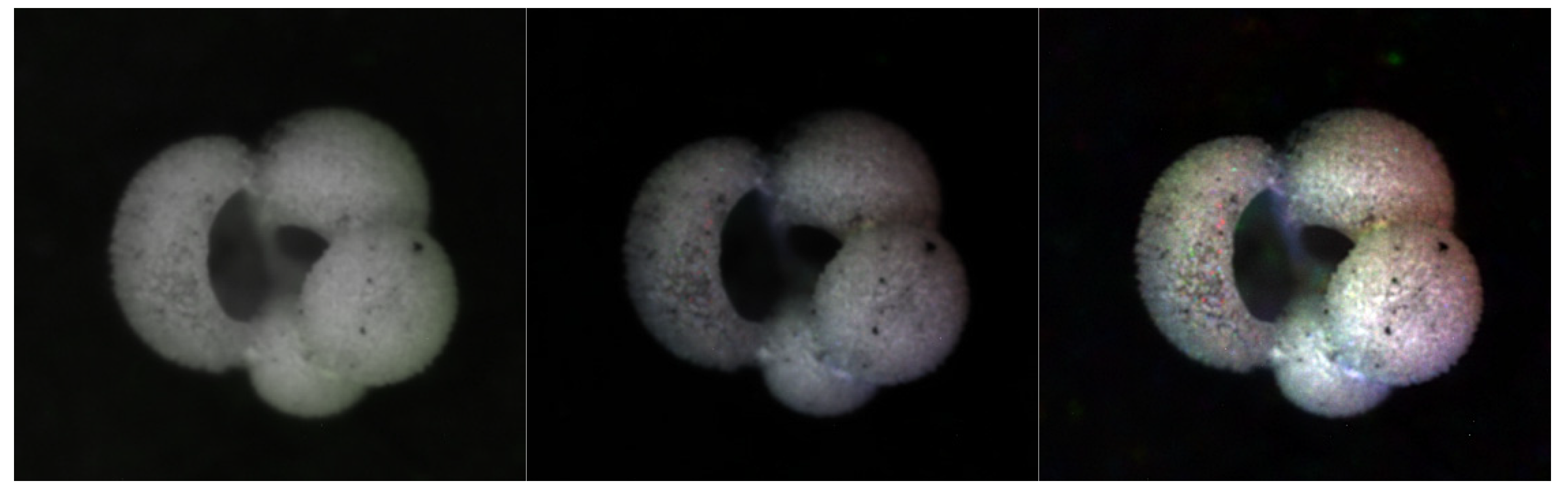
3.2. Training
- Mini Batch Size: 30
- Max Epochs: 20
- Learning Rate: 10-3
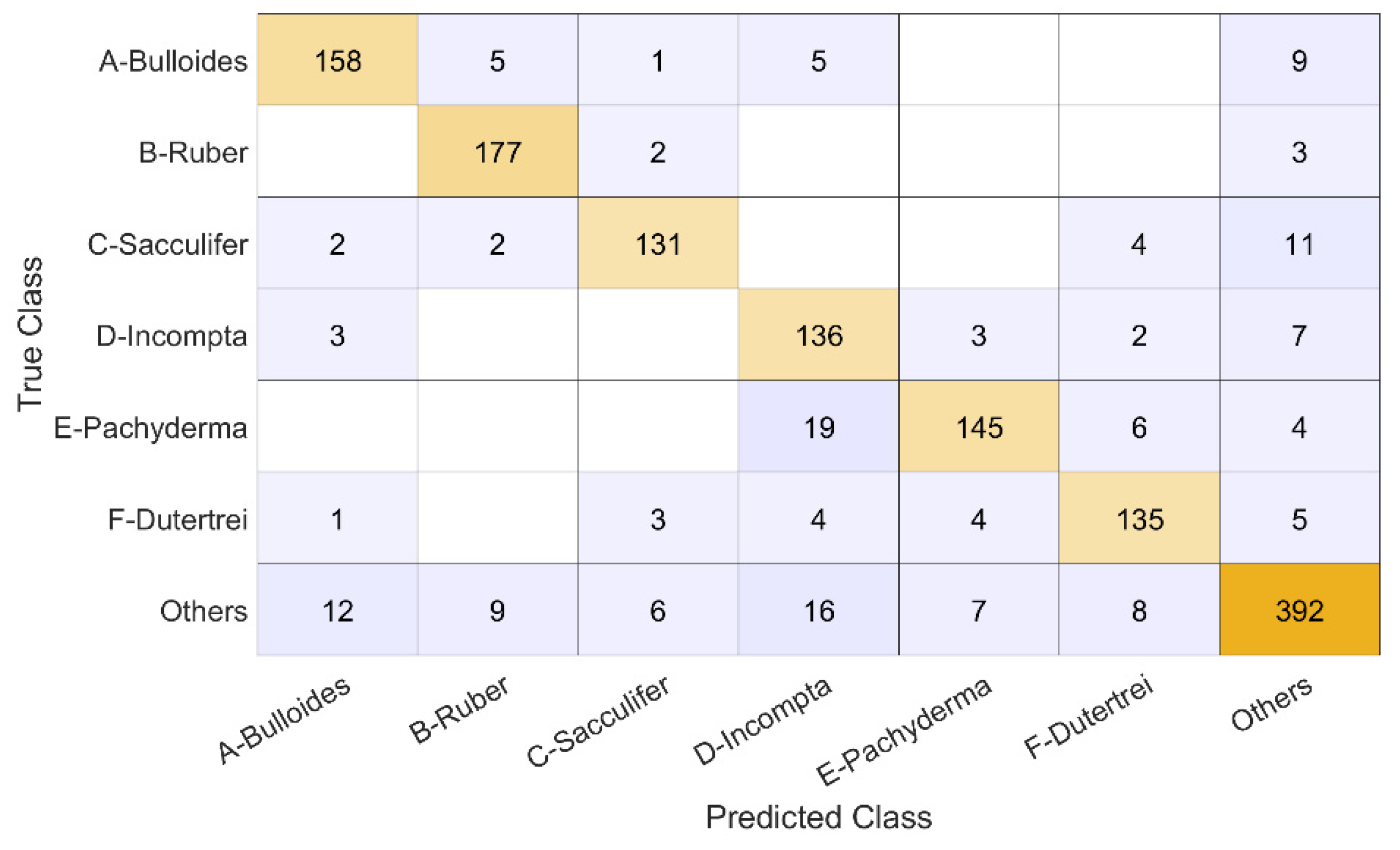
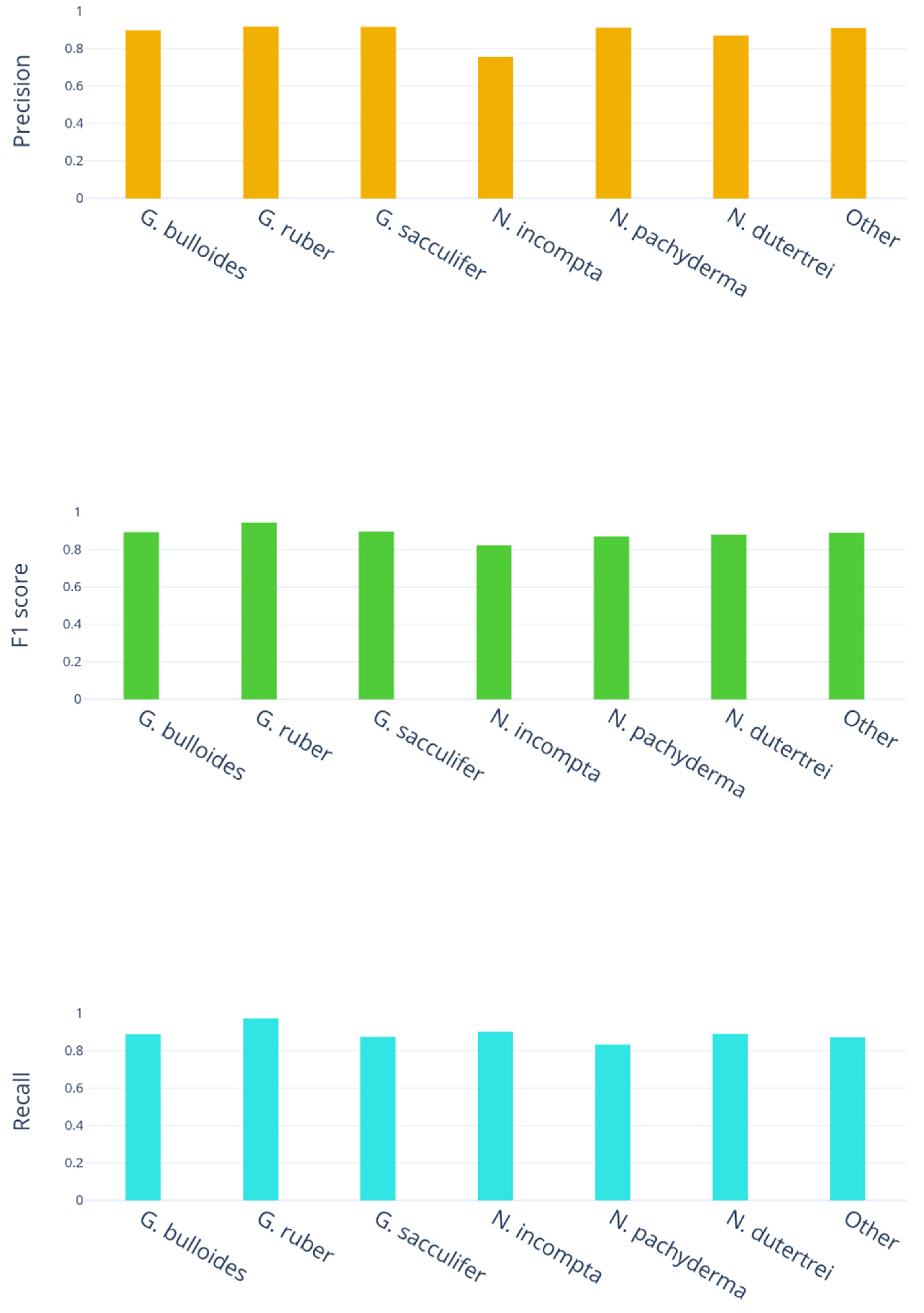
4. Results
- 178 images are G. bulloides;
- 182 images are G. ruber;
- 150 images are G. sacculifer;
- 174 images are N. incompta;
- 152 images are N. pachyderma;
- 151 images are N. dutertrei;
- 450 images are "rest of the world," i.e., they belong to other species of planktic foraminifera.
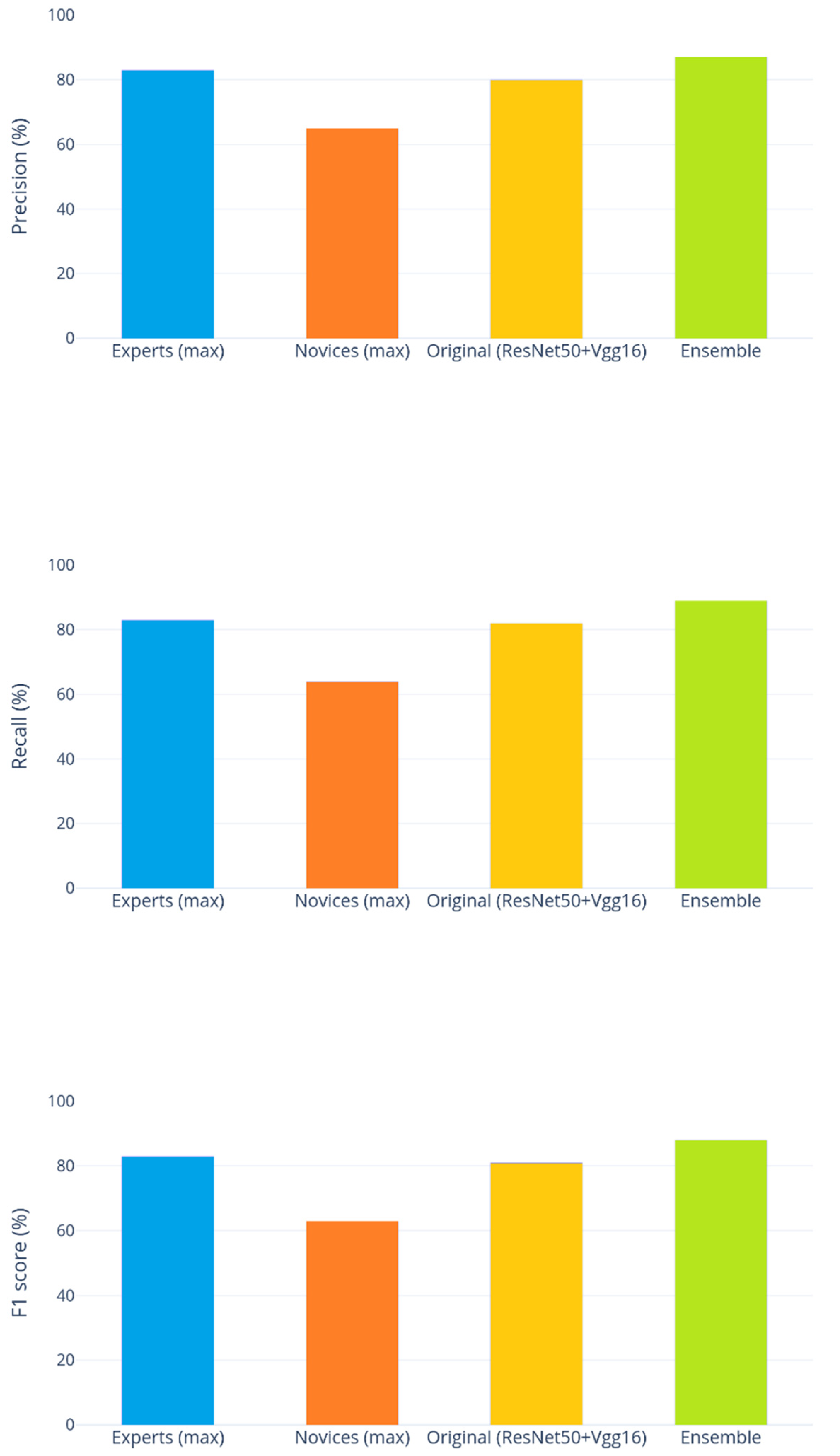
- The best-performing ensemble produces results that significantly improve those obtained by the method presented in [3] (Percentile), whose score was reported as 81%.
- It appears that, in general, increasing the diversity of the ensemble yields better results. The approaches combining multiple preprocessed images sets consistently rank higher in scores than any individual method, iterated ten times. Combining fewer iterations of all the approaches yielded the best results overall.
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zewen Li, Fan Liu, Wenjie Yang, Shouheng Peng and Jun Zhou, A Survey of Convolutional Neural Networks: Analysis, Applications, and Prospects, 2021. [CrossRef]
- Christopher Menart, Evaluating the variance in convolutional neural network behavior stemming from randomness, 2020. [CrossRef]
- R. Mitra, T.M. Marchitto, Q. Ge, B. Zhong, B. Kanakiya, M.S. Cook, J.S. Fehrenbacher, J.D. Ortiz, A. Tripati, E. Lobaton, Automated species-level identification of planktic foraminifera using convolutional neural networks, with comparison to human performance, 2019. [CrossRef]
- Luc Beaufort, Denis Dollfus, Automatic recognition of coccoliths by dynamical neural networks, 2004. [CrossRef]
- Luis F. Pedraza, Cesar A. Hernández, Danilo A. López, "A Model to Determine the Propagation Losses Based on the Integration of Hata-Okumura and Wavelet Neural Models", International Journal of Antennas and Propagation, vol. 2017, Article ID 1034673, 8 pages, 2017. [CrossRef]
- Bing Huang, Feng Yang, Mengxiao Yin, Xiaoying Mo, Cheng Zhong, "A Review of Multimodal Medical Image Fusion Techniques", Computational and Mathematical Methods in Medicine, 2020. [CrossRef]
- Akrem Sellami, Ali Ben Abbes, Vincent Barra, Imed Riadh Farah, Fused 3-D spectral-spatial deep neural networks and spectral clustering for hyperspectral image classification, Pattern Recognition Letters, Volume 138, 2020. [CrossRef]
- Xingchen Zhang, Ping Ye, Gang Xiao, VIFB: A Visible and Infrared Image Fusion Benchmark, Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) Workshops, 2020, pp. 104-105.
- Alex Pappachen James, Belur V. Dasarathy, Medical image fusion: A survey of the state of the art, Information Fusion, Volume 19, 2014. [CrossRef]
- Sarmad Maqsood, Umer Javed, Multi-modal Medical Image Fusion based on Two-scale Image Decomposition and Sparse Representation, Biomedical Signal Processing and Control, Volume 57, 2020. [CrossRef]
- Y. LeCun et al., "Backpropagation Applied to Handwritten Zip Code Recognition," in Neural Computation, vol. 1, no. 4, pp. 541-551, Dec. 1989. [CrossRef]
- Bengio, Y. & Lecun, Yann, Convolutional Networks for Images, Speech, and Time-Series, 1997.
- Y. LeCun, L. Bottou, Y. Bengio and P. Haffner, "Gradient-based learning applied to document recognition," in Proceedings of the IEEE, vol. 86, no. 11, pp. 2278-2324, Nov. 1998. [CrossRef]
- Xingchen Zhang, Ping Ye, Henry Leung, Ke Gong, Gang Xiao, Object fusion tracking based on visible and infrared images: A comprehensive review, Information Fusion, Volume 63, 2020. [CrossRef]
- Ian Goodfellow, Yoshua Bengio, Aaron Courville, Deep Learning, The MIT Press, 2016.
- Kuncheva L.I. Combining Pattern Classifiers. Methods and Algorithms, Wiley, 2nd edition, 2014.
- http://hdl.handle.net/20.500.12608/29285.
- https://it.wikipedia.org/wiki/Hue_Saturation_Brightness.
- Fuzhen Zhuang, Zhiyuan Qi, Keyu Duan, Dongbo Xi, Yongchun Zhu, Hengshu Zhu, Hui Xiong, Qing He, A Comprehensive Survey on Transfer Learning. [CrossRef]
| 4-fold cross-validation | |
|---|---|
| [3] | 0.850 |
| Percentile(1) | 0.811 |
| Percentile(10) | 0.853 |
| Luma Scaling(10) | 0.870 |
| Means Reconstruction(10) | 0.874 |
| Gaussian(10) | 0.873 |
| HSVPP(10) | 0.843 |
| Percentile(3)+Luma Scaling(3)+ Means Reconstruction(3) |
0.877 |
| Gaussian(3)+Luma Scaling(3)+ Means Reconstruction(3) |
0.879 |
| Percentile(2)+Gaussian(2)+Luma Scaling(2)+ Means Reconstruction(2)+HSVPP(2) |
0.885 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




