Submitted:
26 January 2023
Posted:
30 January 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.0. Biochemicals and other reagents
2.1. Generation of a peptide binder for CCMV
2.1.1. Experimental peptide binder generation
2.1.2. Molecular dynamics simulations of the peptide with GROMACS
2.1.3. Markov modeling to obtain the dominant equilibrium peptide structure in solution
2.2. Preparation of the affinity column with CCMV-binding peptide
2.3. Optimized protocol for the purification of CCMV
2.4. Characterization of purification steps with silver-stained SDS-PAGE
2.5. Characterization of purification steps with size exclusion chromatography (SEC)
2.6. Characterization of purification steps with MALDI-TOF-MS
2.7. Purity determination by reversed-phase HPLC
2.8. Determination of integrity and monodispersity
2.9. Quantification of CCMV by ELISA
3. Results
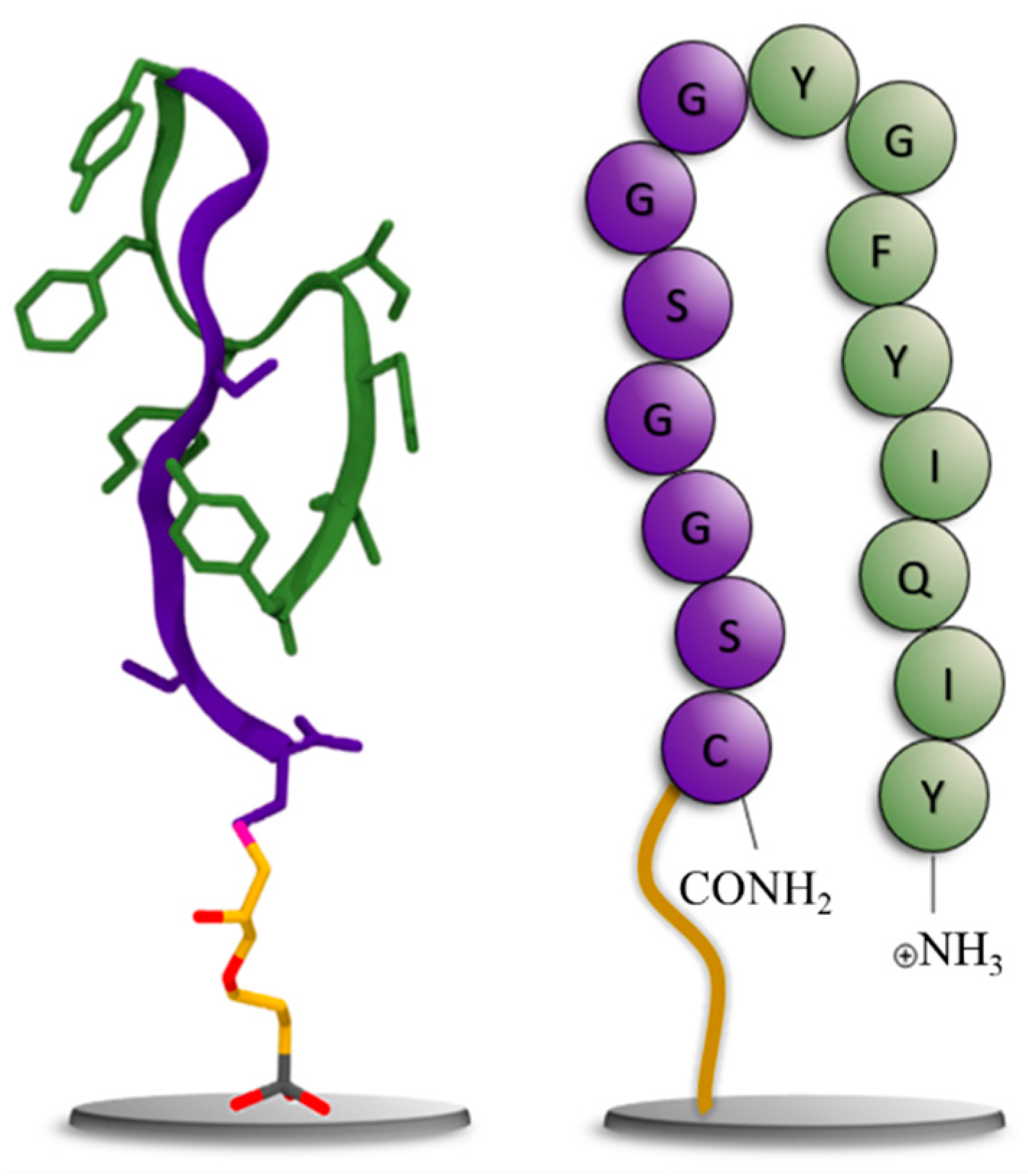
3.1. Generation of a peptide aptamer for CCMV
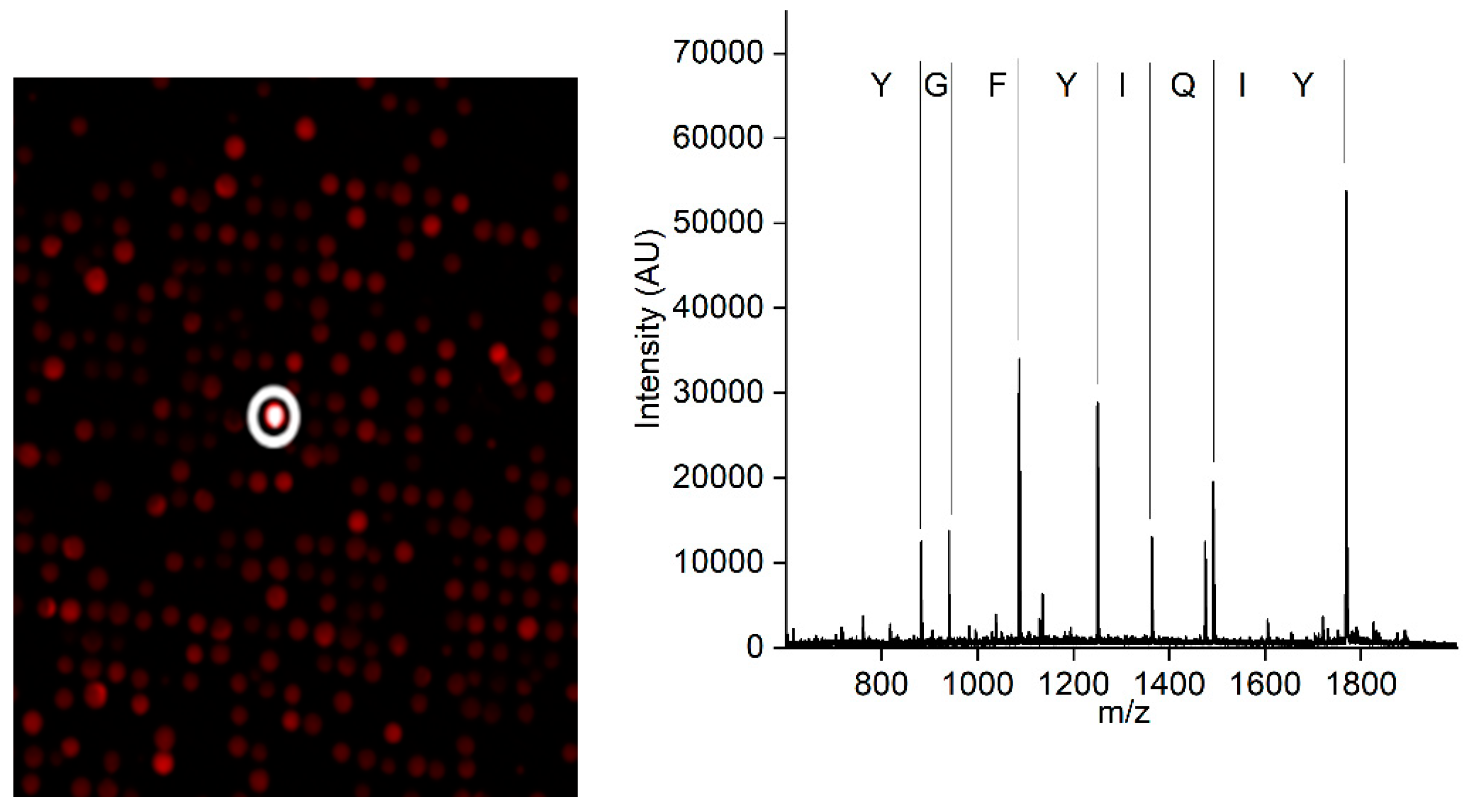
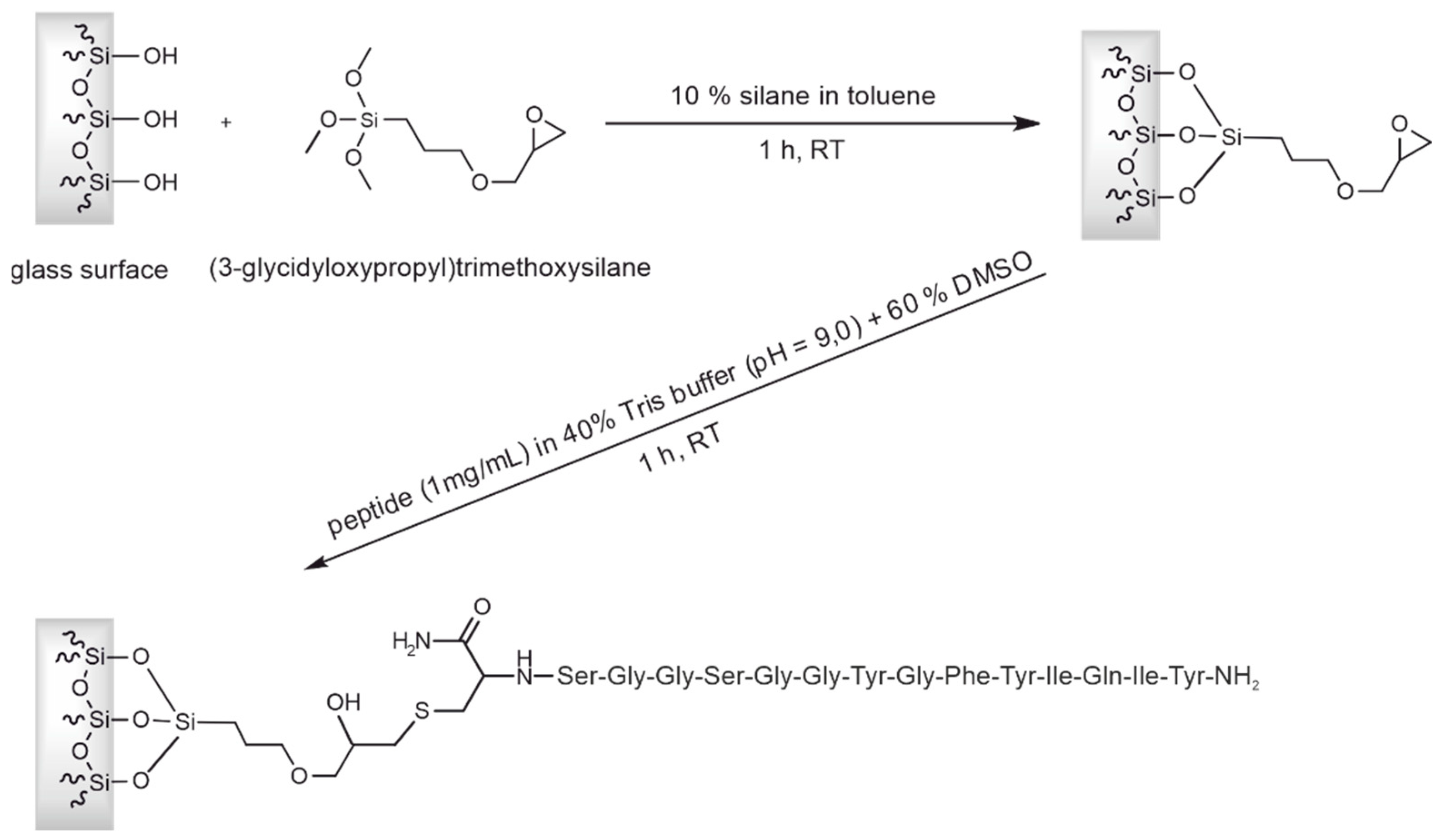
3.2. Preparation of an affinity column with the CCMV-binding peptide aptamer
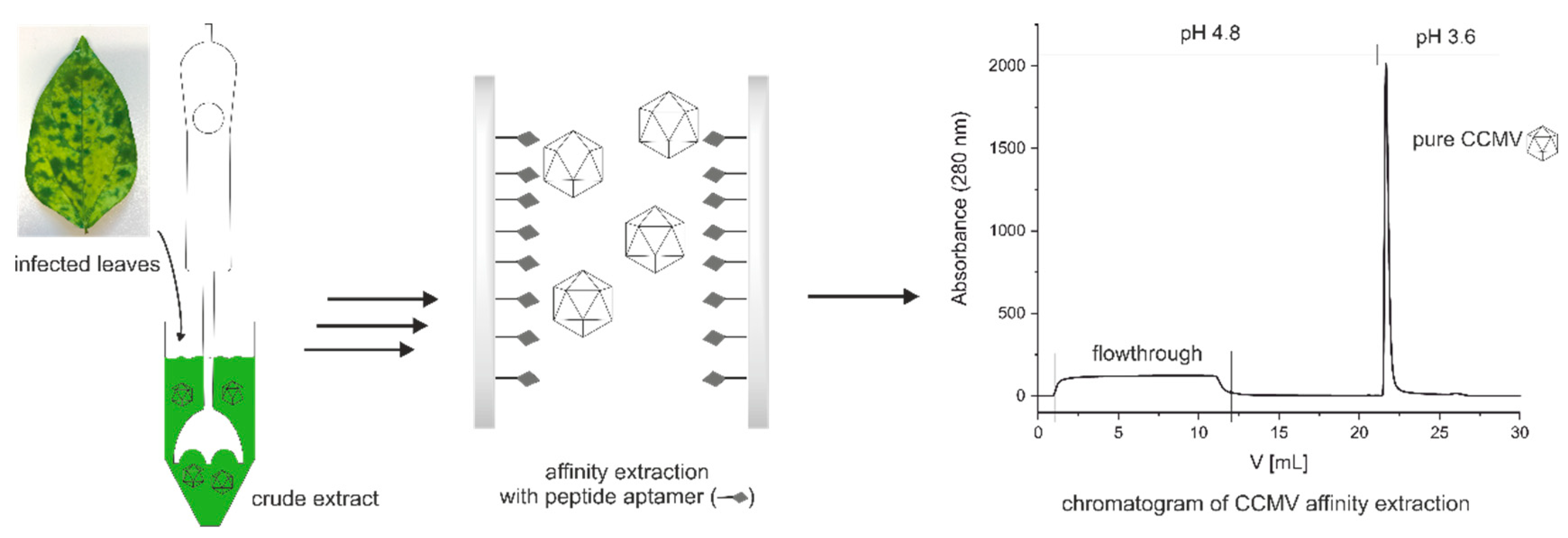
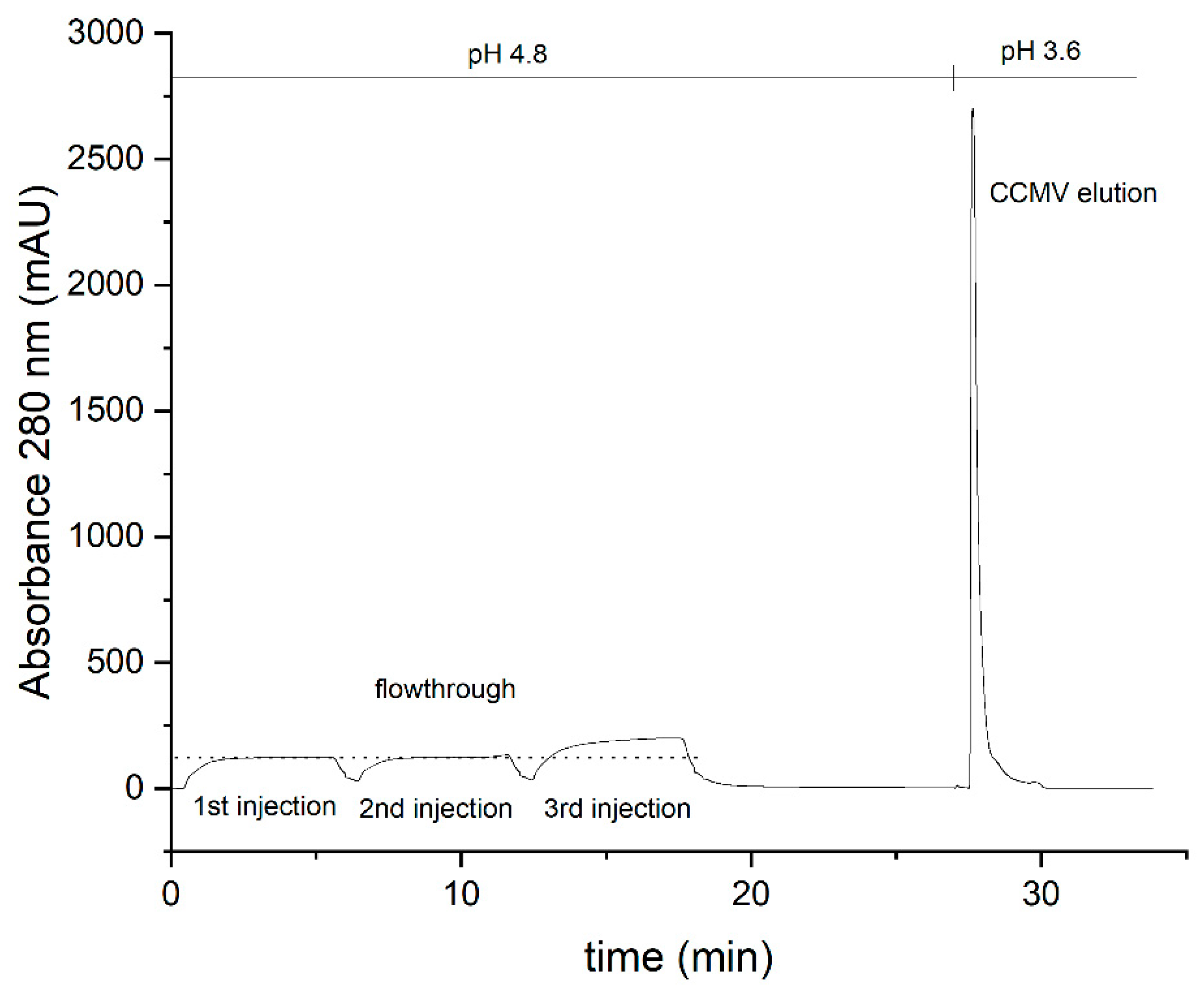
3.3. Optimized protocol for the purification of CCMV
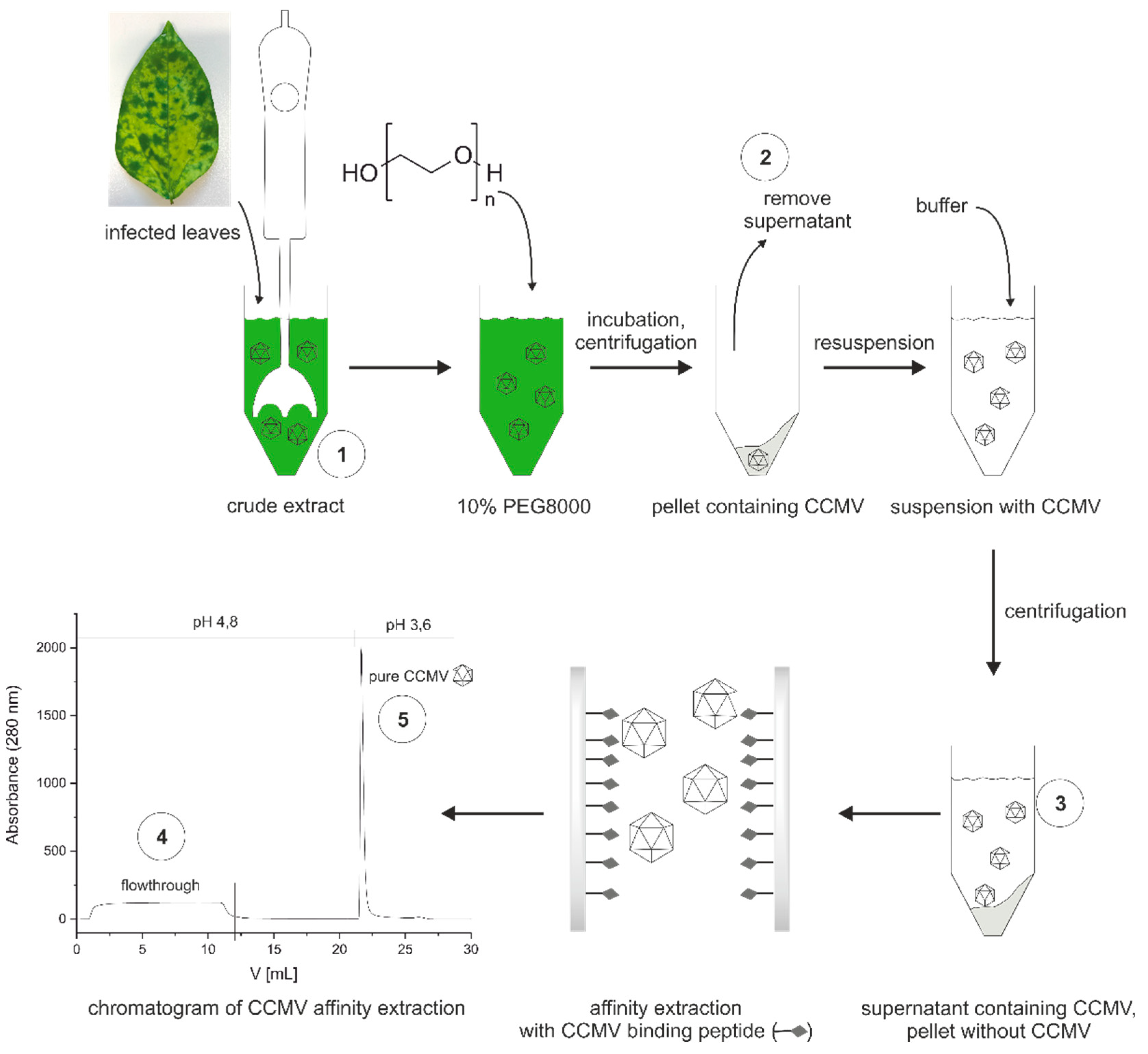
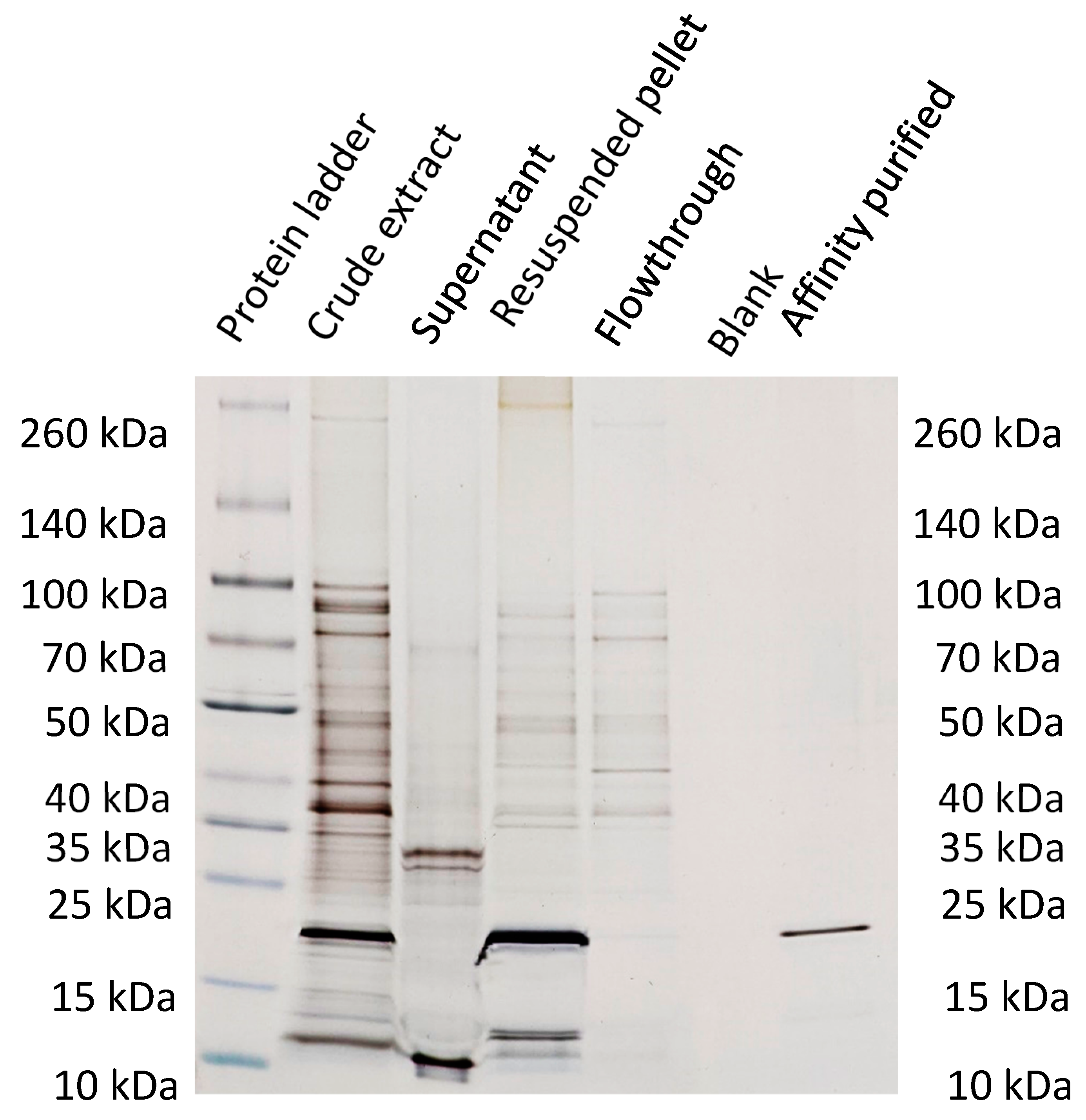
3.4. Characterization of purification steps with silver-stained SDS-PAGE
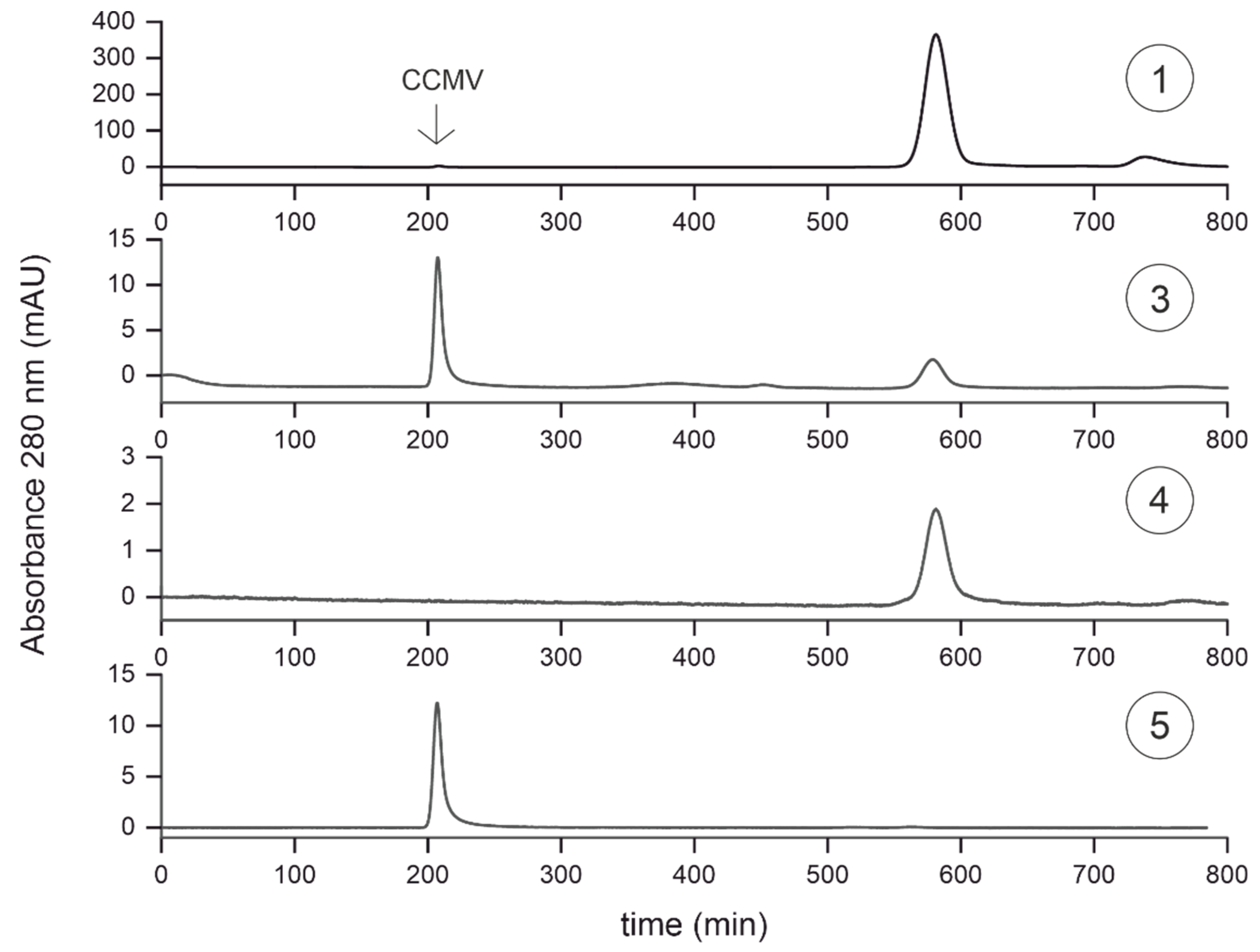
3.5. Characterization of purification steps with size exclusion chromatography
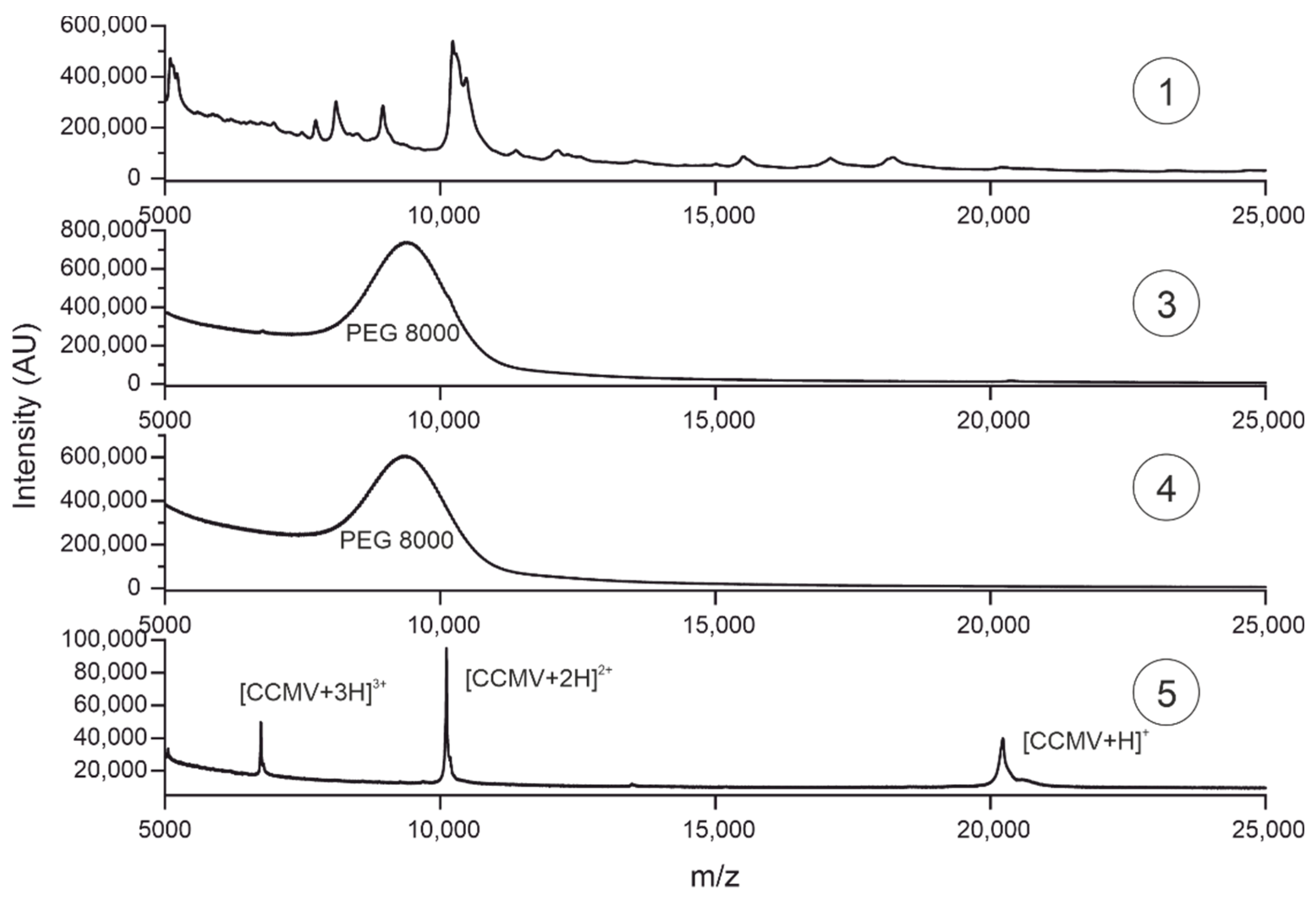
3.6. Characterization of purification steps with MALDI-TOF-MS
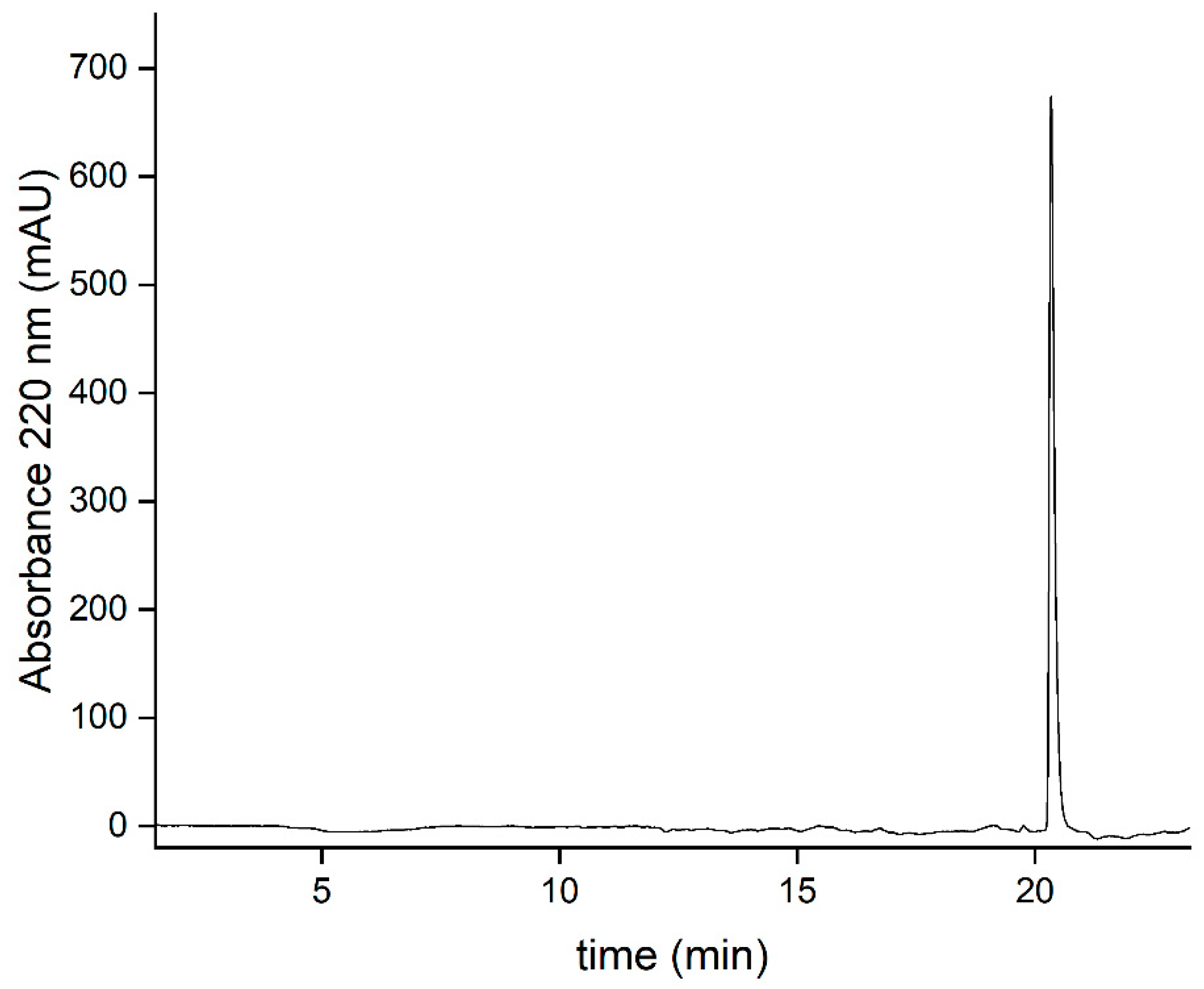
3.7. Purity determination by reversed-phase liquid chromatography (HPLC)
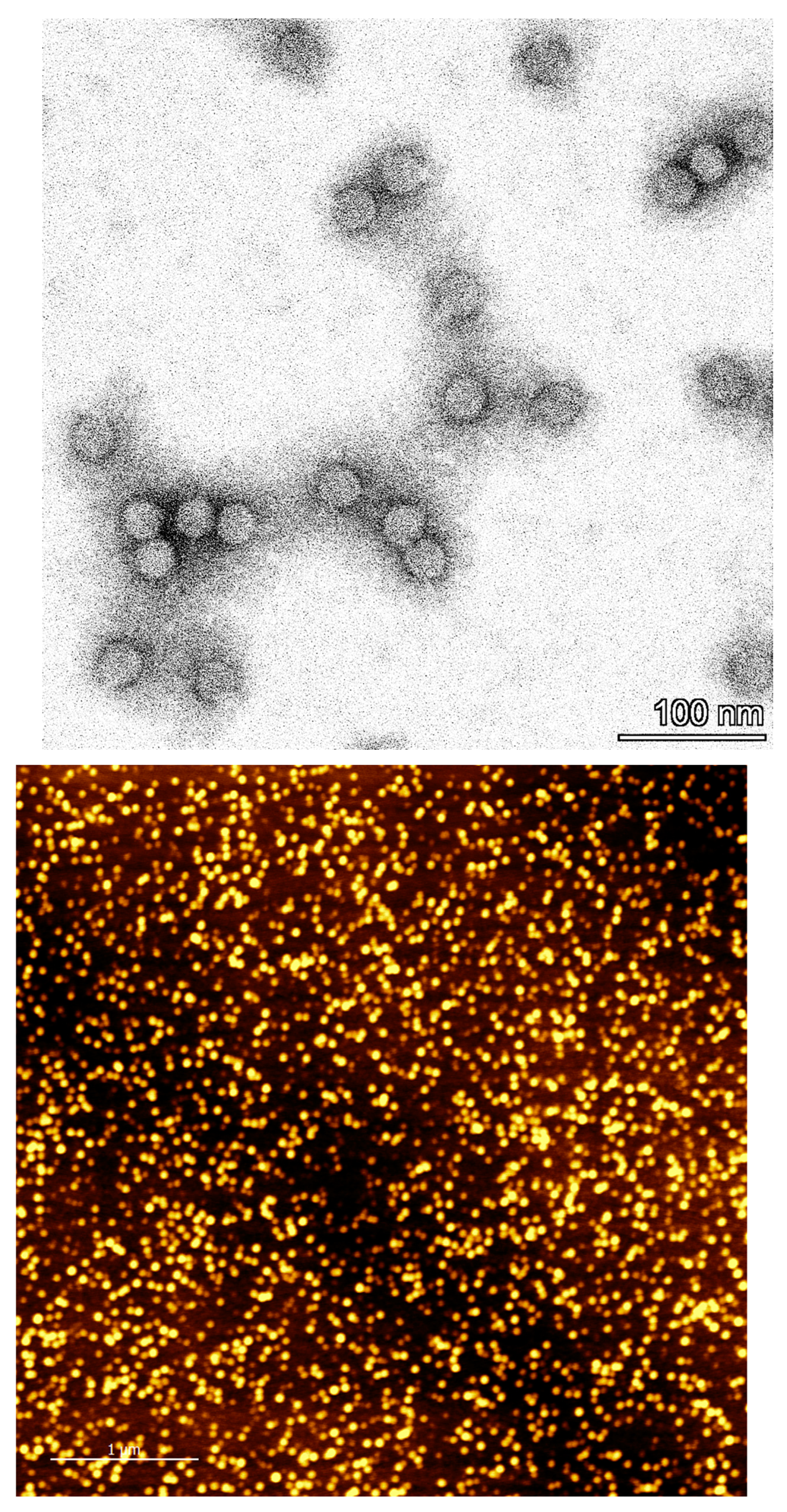
3.8. Determination of particle integrity and size distribution
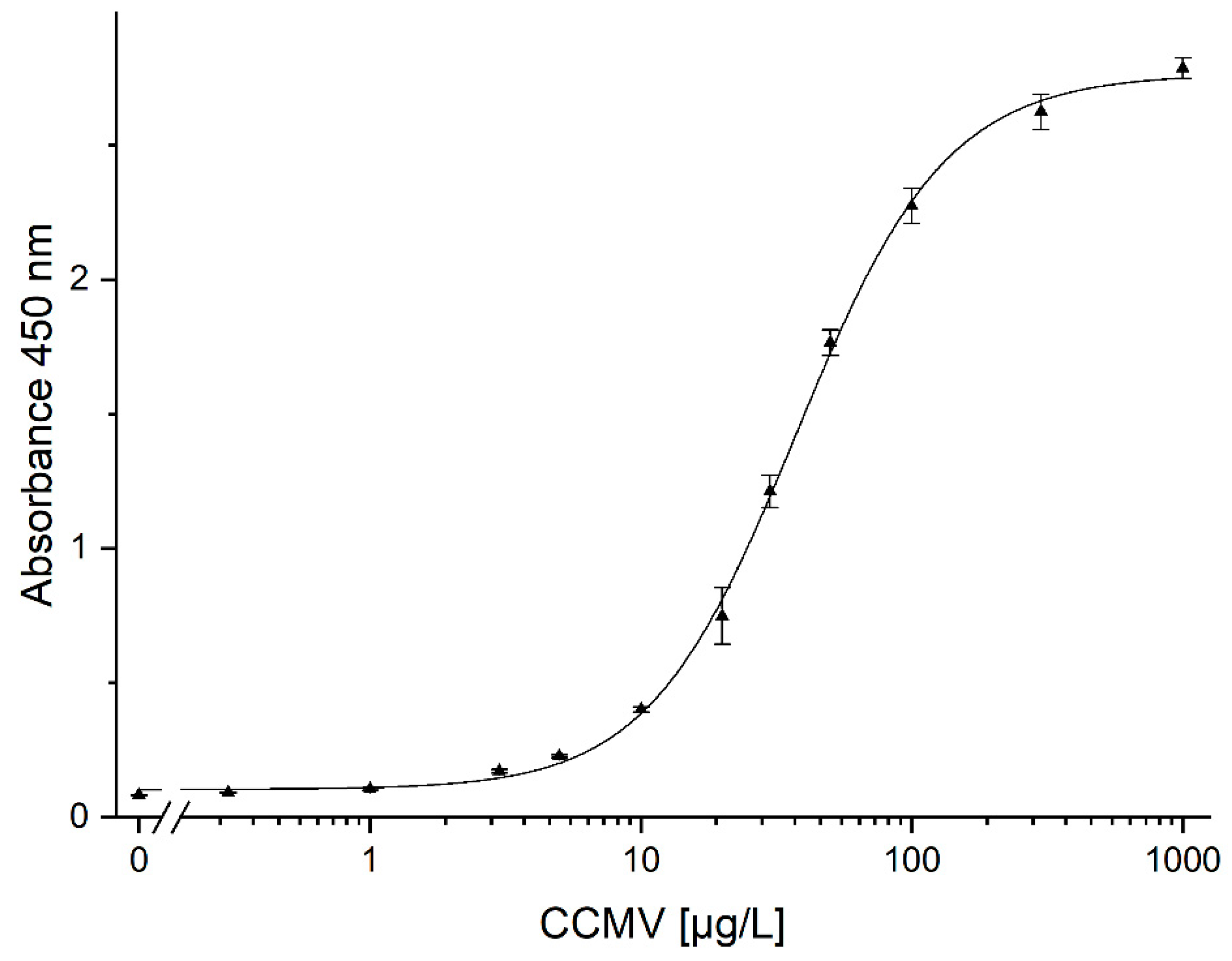
3.9. Quantification of CCMV by ELISA
4. Discussion
5. Conclusion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hespenheide, B.M.; Jacobs, D.J.; Thorpe, M.F. Structural rigidity in the capsid assembly of cowpea chlorotic mottle virus. Journal of Physics-Condensed Matter 2004, 16, S5055-S5064. [CrossRef]
- Kaiser, C.R.; Flenniken, M.L.; Gillitzer, E.; Harmsen, A.L.; Harmsen, A.G.; Jutila, M.A.; Douglas, T.; Young, M.J. Biodistribution studies of protein cage nanoparticles demonstrate broad tissue distribution and rapid clearance in vivo. International journal of nanomedicine 2007, 2, 715-733.
- Azizgolshani, O.; Garmann, R.F.; Cadena-Nava, R.; Knobler, C.M.; Gelbart, W.M. Reconstituted plant viral capsids can release genes to mammalian cells. Virology 2013, 441, 12-17. [CrossRef]
- Sanchez-Sanchez, L.; Cadena-Nava, R.D.; Palomares, L.A.; Ruiz-Garcia, J.; Koay, M.S.; Cornelissen, J.J.; Vazquez-Duhalt, R. Chemotherapy pro-drug activation by biocatalytic virus-like nanoparticles containing cytochrome P450. Enzyme and microbial technology 2014, 60, 24-31. [CrossRef]
- Pretto, C.; Tang, M.; Chen, M.; Xu, H.P.; Subrizi, A.; Urtti, A.; van Hest, J.C.M. Cowpea Chlorotic Mottle Virus-Like Particles as Potential Platform for Antisense Oligonucleotide Delivery in Posterior Segment Ocular Diseases. Macromolecular Bioscience 2021, 21, 2100095. [CrossRef]
- Villagrana-Escareño, M.V.; Reynaga-Hernández, E.; Galicia-Cruz, O.G.; Durán-Meza, A.L.; la Cruz-González, D.; Hernández-Carballo, C.Y.; Ruíz-García, J. VLPs derived from the CCMV plant virus can directly transfect and deliver heterologous genes for translation into mammalian cells. BioMed Research International 2019. [CrossRef]
- Timmermans, S.; Mesman, R.; Blezer, K.J.R.; van Niftrik, L.; van Hest, J.C.M. Cargo-loading of hybrid cowpea chlorotic mottle virus capsids via a co-expression approach. Virology 2022, 577, 99-104. [CrossRef]
- Tresset, G.; Chen, J.Z.; Chevreuil, M.; Nhiri, N.; Jacquet, E.; Lansac, Y. Two-Dimensional Phase Transition of Viral Capsid Gives Insights into Subunit Interactions. Physical Review Applied 2017, 7, 014005. [CrossRef]
- Chen, J.Z.; Lansac, Y.; Tresset, G. Interactions between the Molecular Components of the Cowpea Chlorotic Mottle Virus Investigated by Molecular Dynamics Simulations. J Phys Chem B 2018, 122, 9490-9498. [CrossRef]
- Lu, X.Y.; Thompson, J.R.; Perry, K.L. Encapsidation of DNA, a protein and a fluorophore into virus-like particles by the capsid protein of cucumber mosaic virus. Journal of General Virology 2012, 93, 1120-1126. [CrossRef]
- Gillitzer, E.; Willits, D.; Young, M.; Douglas, T. Chemical modification of a viral cage for multivalent presentation. Chem Commun (Camb) 2002, 2390-2391. [CrossRef]
- Hassani-Mehraban, A.; Creutzburg, S.; van Heereveld, L.; Kormelink, R. Feasibility of Cowpea chlorotic mottle virus-like particles as scaffold for epitope presentations. Bmc Biotechnology 2015, 15, 1-17. [CrossRef]
- Barwal, I.; Kumar, R.; Kateriya, S.; Dinda, A.K.; Yadav, S.C. Targeted delivery system for cancer cells consist of multiple ligands conjugated genetically modified CCMV capsid on doxorubicin GNPs complex. Scientific reports 2016, 6, 37096. [CrossRef]
- Vervoort, D.F.; Heiringhoff, R.; Timmermans, S.B.; Van Stevendaal, M.H.; Van Hest, J.C. Dual site-selective presentation of functional handles on protein-engineered cowpea chlorotic mottle virus-like particles. Bioconjugate chemistry 2021, 32, 958-963. [CrossRef]
- Hommersom, C.A.; Matt, B.; van der Ham, A.; Cornelissen, J.J.; Katsonis, N. Versatile post-functionalization of the external shell of cowpea chlorotic mottle virus by using click chemistry. Organic & biomolecular chemistry 2014, 12, 4065-4069. [CrossRef]
- Pomwised, R.; Intamaso, U.; Teintze, M.; Young, M.; Pincus, S.H. Coupling Peptide Antigens to Virus-Like Particles or to Protein Carriers Influences the Th1/Th2 Polarity of the Resulting Immune Response. Vaccines (Basel) 2016, 4, 15. [CrossRef]
- Eiben, S.; Koch, C.; Altintoprak, K.; Southan, A.; Tovar, G.; Laschat, S.; Weiss, I.M.; Wege, C. Plant virus-based materials for biomedical applications: Trends and prospects. Adv Drug Deliv Rev 2019, 145, 96-118. [CrossRef]
- Smith, T.J.; Chase, E.; Schmidt, T.; Perry, K.L. The structure of cucumber mosaic virus and comparison to cowpea chlorotic mottle virus. Journal of Virology 2000, 74, 7578-7586. [CrossRef]
- Tama, F.; Brooks, C.L., 3rd. The mechanism and pathway of pH induced swelling in cowpea chlorotic mottle virus. Journal of molecular biology 2002, 318, 733-747. [CrossRef]
- Speir, J.A.; Munshi, S.; Wang, G.; Baker, T.S.; Johnson, J.E. Structures of the native and swollen forms of cowpea chlorotic mottle virus determined by X-ray crystallography and cryo-electron microscopy. Structure 1995, 3, 63-78. [CrossRef]
- Sehnal, D.; Bittrich, S.; Deshpande, M.; Svobodova, R.; Berka, K.; Bazgier, V.; Velankar, S.; Burley, S.K.; Koca, J.; Rose, A.S. Mol* Viewer: modern web app for 3D visualization and analysis of large biomolecular structures. Nucleic Acids Res 2021, 49, W431-W437. [CrossRef]
- Bancroft, J.B.; Hills, G.J.; Markham, R. A study of the self-assembly process in a small spherical virus. Formation of organized structures from protein subunits in vitro. Virology 1967, 31, 354-379. [CrossRef]
- Lam, P.; Steinmetz, N.F. Delivery of siRNA therapeutics using cowpea chlorotic mottle virus-like particles. Biomaterials science 2019, 7, 3138-3142. [CrossRef]
- Hu, Y.F.; Zandi, R.; Anavitarte, A.; Knobler, C.M.; Gelbart, W.M. Packaging of a polymer by a viral capsid: The interplay between polymer length and capsid size. Biophysical Journal 2008, 94, 1428-1436. [CrossRef]
- Douglas, T.; Young, M. Host-guest encapsulation of materials by assembled virus protein cages. Nature 1998, 393, 152-155. [CrossRef]
- Aniagyei, S.E.; Kennedy, C.J.; Stein, B.; Willits, D.A.; Douglas, T.; Young, M.J.; De, M.; Rotello, V.M.; Srisathiyanarayanan, D.; Kao, C.C.; et al. Synergistic effects of mutations and nanoparticle templating in the self-assembly of cowpea chlorotic mottle virus capsids. Nano letters 2009, 9, 393-398. [CrossRef]
- Rurup, W.F.; Verbij, F.; Koay, M.S.; Blum, C.; Subramaniam, V.; Cornelissen, J.J. Predicting the loading of virus-like particles with fluorescent proteins. Biomacromolecules 2014, 15, 558-563. [CrossRef]
- Shukla, S.; Wang, C.; Beiss, V.; Cai, H.; Washington, T., 2nd; Murray, A.A.; Gong, X.; Zhao, Z.; Masarapu, H.; Zlotnick, A.; et al. The unique potency of Cowpea mosaic virus (CPMV) in situ cancer vaccine. Biomaterials science 2020, 8, 5489-5503. [CrossRef]
- Almendarez-Rodriguez, C.; Solis-Andrade, K.I.; Govea-Alonso, D.O.; Comas-Garcia, M.; Rosales-Mendoza, S. Production and characterization of chimeric SARS-CoV-2 antigens based on the capsid protein of cowpea chlorotic mottle virus. International Journal of Biological Macromolecules 2022, 213, 1007-1017. [CrossRef]
- Vajda, B.P. Concentration and purification of viruses and bacteriophages with polyethylene glycol. Folia Microbiol (Praha) 1978, 23, 88-96. [CrossRef]
- Bhat, A.I.; Rao, G.P. Characterization of plant viruses; Springer: 2020.
- Ali, A.; Roossinck, M.J. Rapid and efficient purification of Cowpea chlorotic mottle virus by sucrose cushion ultracentrifugation. Journal of virological methods 2007, 141, 84-86. [CrossRef]
- Michel, J.P.; Gingery, M.; Lavelle, L. Efficient purification of bromoviruses by ultrafiltration. Journal of virological methods 2004, 122, 195-198. [CrossRef]
- Office of Environment, H.S. Ultracentrifuges: Hazards and Precautions. Available online: https://ehs.berkeley.edu/publications/ultracentrifuges-hazards-and-precautions (accessed on 24 January 2023).
- Zhao, X.; Fox, J.M.; Olson, N.H.; Baker, T.S.; Young, M.J. In vitro assembly of cowpea chlorotic mottle virus from coat protein expressed in Escherichia coli and in vitro-transcribed viral cDNA. Virology 1995, 207, 486-494. [CrossRef]
- Cantin, G.T.; Resnick, S.; Jin, H.F.; O'Hanlon, R.; Espinosa, O.; Stevens, A.; Payne, J.; Glenn, N.R.; Rasochova, L.; Allen, J.R. Comparison of Methods for Chemical Conjugation of an Influenza Peptide to Wild-Type and Cysteine-Mutant Virus-Like Particles Expressed in Pseudomonas fluorescens. International Journal of Peptide Research and Therapeutics 2011, 17, 217-224. [CrossRef]
- Wilke, M.; Roder, B.; Paul, M.; Weller, M.G. Sintered Glass Monoliths as Supports for Affinity Columns. Separations 2021, 8, 56. [CrossRef]
- Schwaar, T.; Lettow, M.; Remmler, D.; Borner, H.G.; Weller, M.G. Efficient Screening of Combinatorial Peptide Libraries by Spatially Ordered Beads Immobilized on Conventional Glass Slides. High Throughput 2019, 8, 11. [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1, 19-25. [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Zidek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583-589. [CrossRef]
- Hornak, V.; Abel, R.; Okur, A.; Strockbine, B.; Roitberg, A.; Simmerling, C. Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins 2006, 65, 712-725. [CrossRef]
- Hockney, R.W.; Goel, S.P.; Eastwood, J.W. Quiet High-Resolution Computer Models of a Plasma. Journal of Computational Physics 1974, 14, 148-158. [CrossRef]
- Bussi, G.; Donadio, D.; Parrinello, M. Canonical sampling through velocity rescaling. The Journal of chemical physics 2007, 126, 014101. [CrossRef]
- Parrinello, M.; Rahman, A. Polymorphic Transitions in Single-Crystals - a New Molecular-Dynamics Method. Journal of Applied Physics 1981, 52, 7182-7190. [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A Smooth Particle Mesh Ewald Method. Journal of Chemical Physics 1995, 103, 8577-8593. [CrossRef]
- Scherer, M.K.; Trendelkamp-Schroer, B.; Paul, F.; Perez-Hernandez, G.; Hoffmann, M.; Plattner, N.; Wehmeyer, C.; Prinz, J.H.; Noe, F. PyEMMA 2: A Software Package for Estimation, Validation, and Analysis of Markov Models. Journal of chemical theory and computation 2015, 11, 5525-5542. [CrossRef]
- Wehmeyer, C.; Scherer, M.K.; Hempel, T.; Husic, B.E.; Olsson, S.; Noé, F. Introduction to Markov state modeling with the PyEMMA software—v1. 0. Living journal of computational molecular science 2018, 1, 10.33011. [CrossRef]
- Vitalini, F.; Mey, A.S.; Noe, F.; Keller, B.G. Dynamic properties of force fields. Journal of chemical physics 2015, 142, 084101. [CrossRef]
- Schor, M.; Mey, A.; MacPhee, C.E. Analytical methods for structural ensembles and dynamics of intrinsically disordered proteins. Biophys Rev 2016, 8, 429-439. [CrossRef]
- Perez-Hernandez, G.; Paul, F.; Giorgino, T.; De Fabritiis, G.; Noe, F. Identification of slow molecular order parameters for Markov model construction. Journal of chemical physics 2013, 139, 015102. [CrossRef]
- Röblitz, S.; Weber, M. Fuzzy spectral clustering by PCCA+: application to Markov state models and data classification. Advances in Data Analysis and Classification 2013, 7, 147-179. [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: visual molecular dynamics. Journal of molecular graphics 1996, 14, 33-38, 27-38. [CrossRef]
- Bancroft, J.B.; Hiebert, E.; Rees, M.W.; Markham, R. Properties of cowpea chlorotic mottle virus, its protein and nucleic acid. Virology 1968, 34, 224-239. [CrossRef]
- De, S.; Khan, A. Efficient synthesis of multifunctional polymers via thiol-epoxy "click" chemistry. Chemical Communications 2012, 48, 3130-3132. [CrossRef]
- Reinmuth-Selzle, K.; Tchipilov, T.; Backes, A.T.; Tscheuschner, G.; Tang, K.; Ziegler, K.; Lucas, K.; Pöschl, U.; Fröhlich-Nowoisky, J.; Weller, M.G. Determination of the protein content of complex samples by aromatic amino acid analysis, liquid chromatography-UV absorbance, and colorimetry. Analytical and Bioanalytical Chemistry 2022, 1-14.
- Tscheuschner, G.; Schwaar, T.; Weller, M.G. Fast Confirmation of Antibody Identity by MALDI-TOF MS Fingerprints. Antibodies (Basel) 2020, 9, 8. [CrossRef]
- Tscheuschner, G.; Kaiser, M.N.; Lisec, J.; Beslic, D.; Muth, T.; Kruger, M.; Mages, H.W.; Dorner, B.G.; Knospe, J.; Schenk, J.A.; et al. MALDI-TOF-MS-Based Identification of Monoclonal Murine Anti-SARS-CoV-2 Antibodies within One Hour. Antibodies 2022, 11, 27. [CrossRef]
- Chan, S.K.; Steinmetz, N.F. Isolation of Cowpea Mosaic Virus-Binding Peptides. Biomacromolecules 2021, 22, 3613-3623. [CrossRef]


| Purification step | Absolute yield [mg] | Relative yield [%] |
| 1 - Crude extract | 3.52 ± 0.27 | 100 (def.) |
| 2 - Supernatant of 1 | 0.040 ± 0.001 | (1.1) |
| 3 – Resuspended pellet | 2.91 ± 0.04 | 83 |
| 4 – Flowthrough of affinity column | < 0.001 | (0) |
| 5 – Eluate of affinity step | 1.57 ± 0.02 | 45 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




