Submitted:
03 January 2023
Posted:
06 January 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
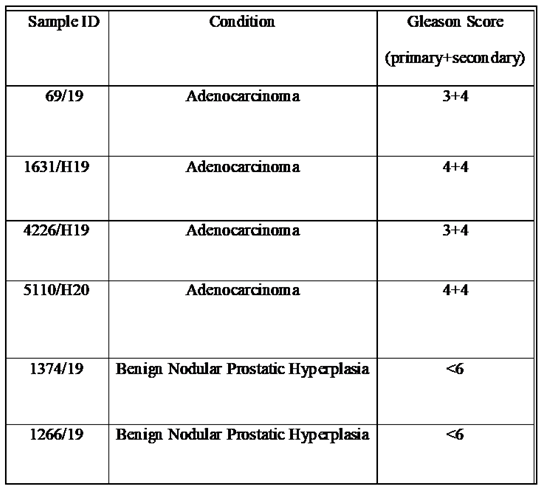
2.1. Patients, clinical samples and criteria
2.2. Tissue preparation and RNA sequencing
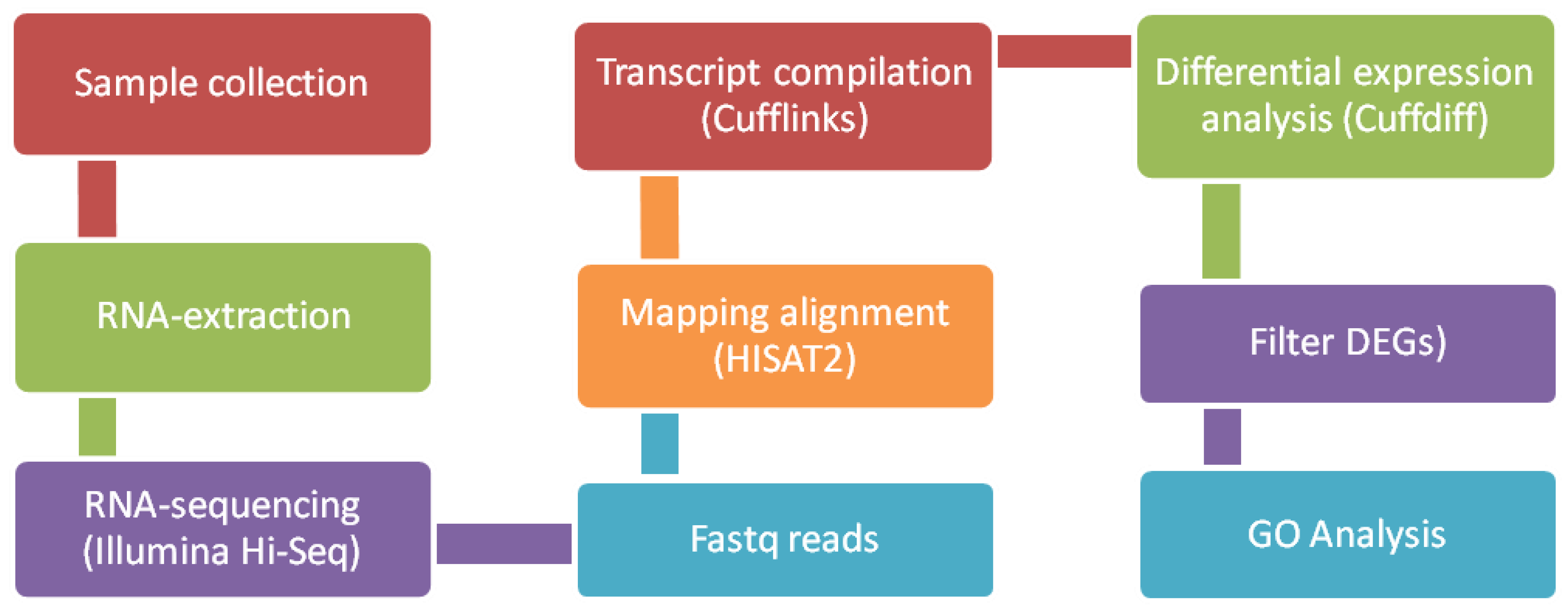
2.3. Bioinformatics and downstream RNA-Sequencing analysis
2.4. Interaction networks, statistical analysis, gene ontology and cbioportal analyses
3. Results
3.1. Distinct DEGs were obtained
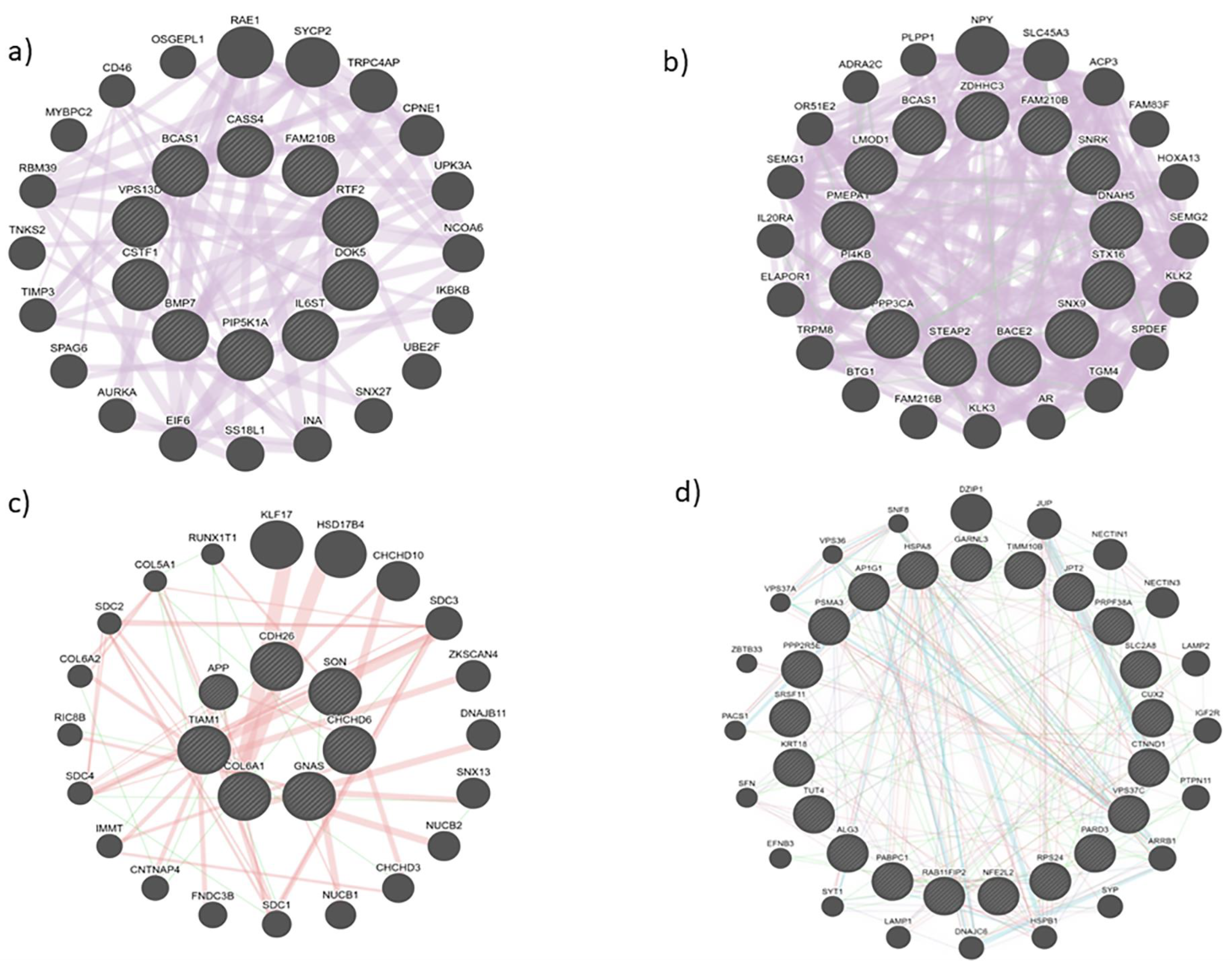
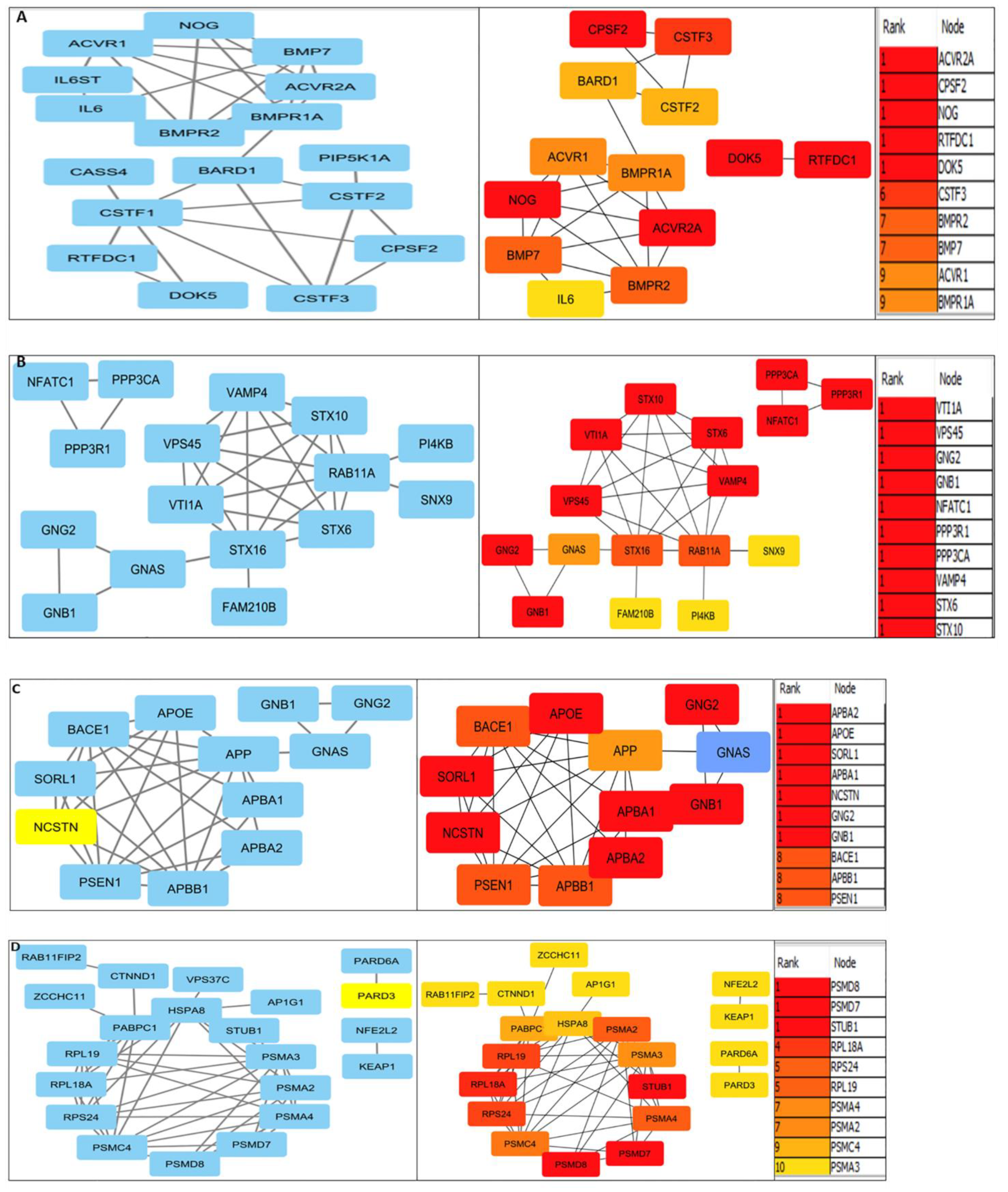
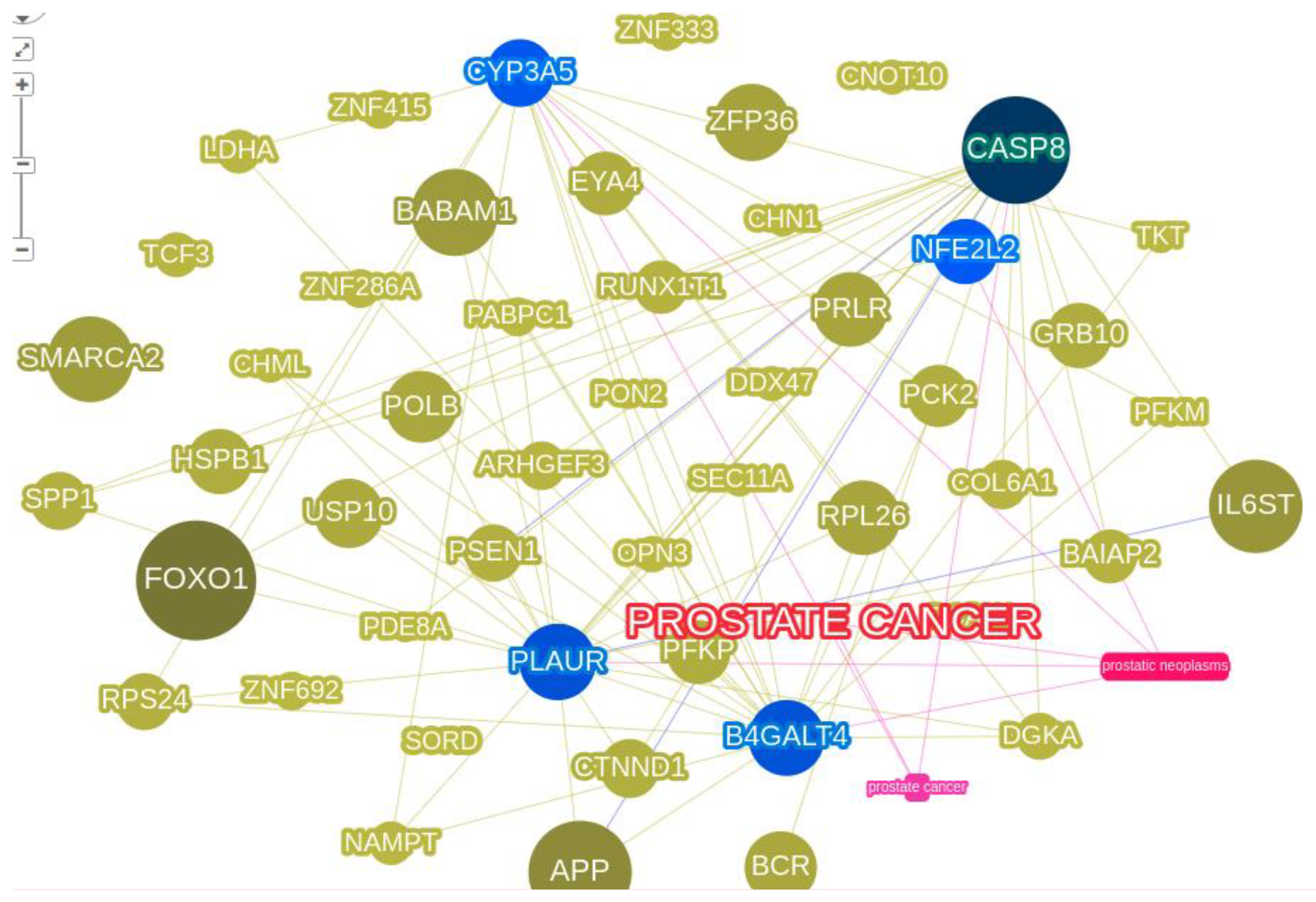
3.2. Protein interactions yielded innate pathways responsible for PCa
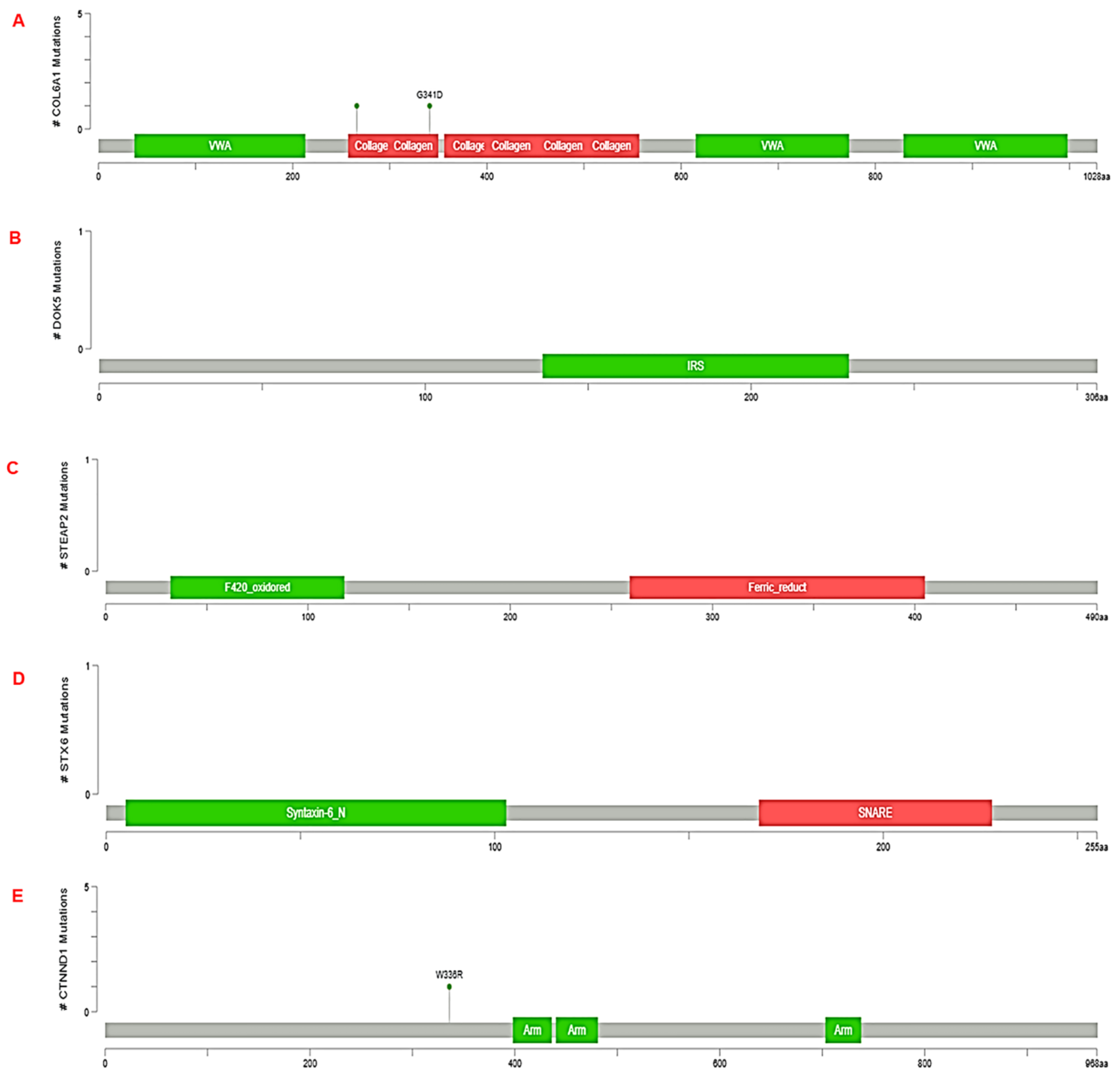
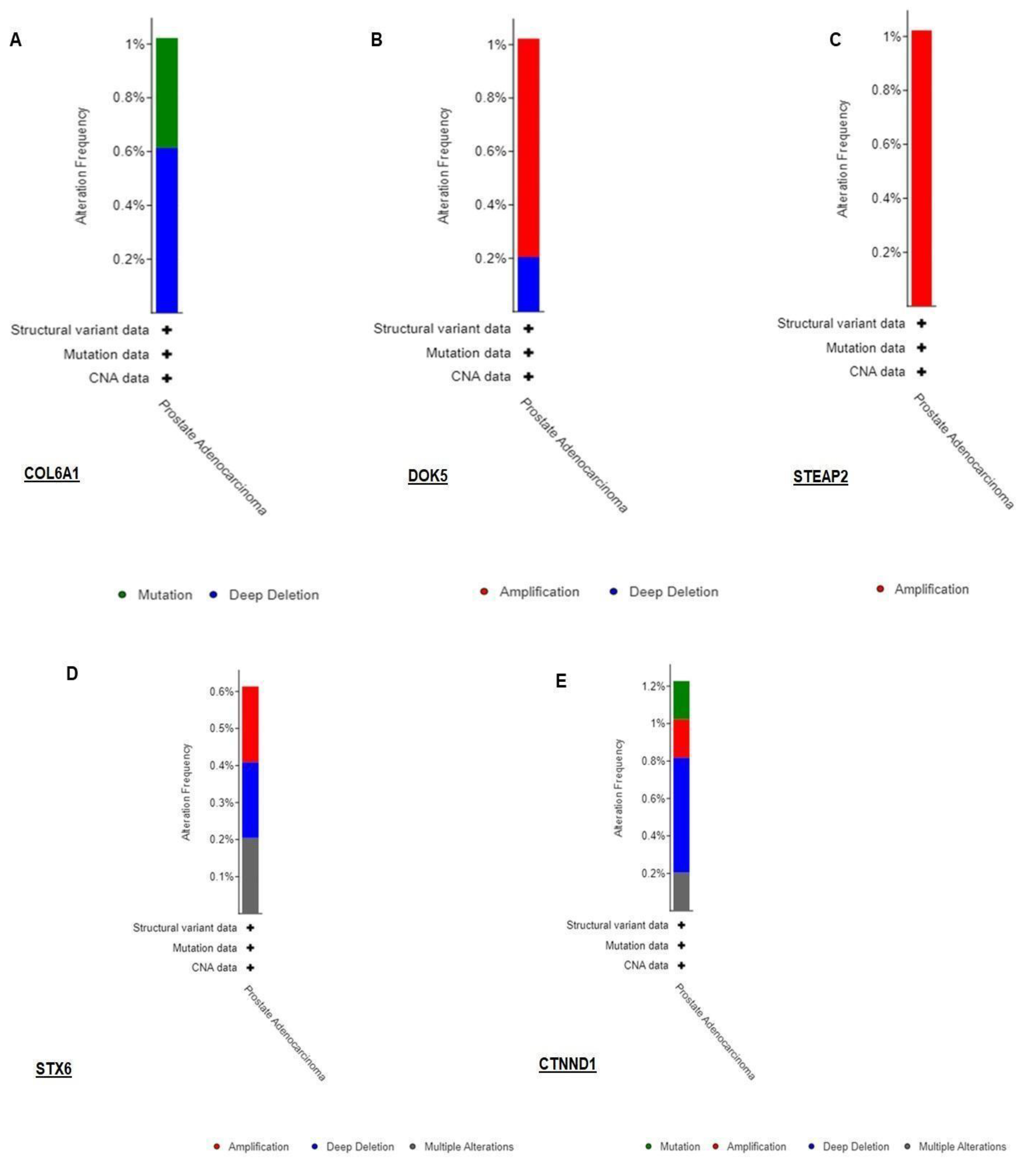
3.3. Validation of RNA-seq result using TCGA dataset by cbioportal
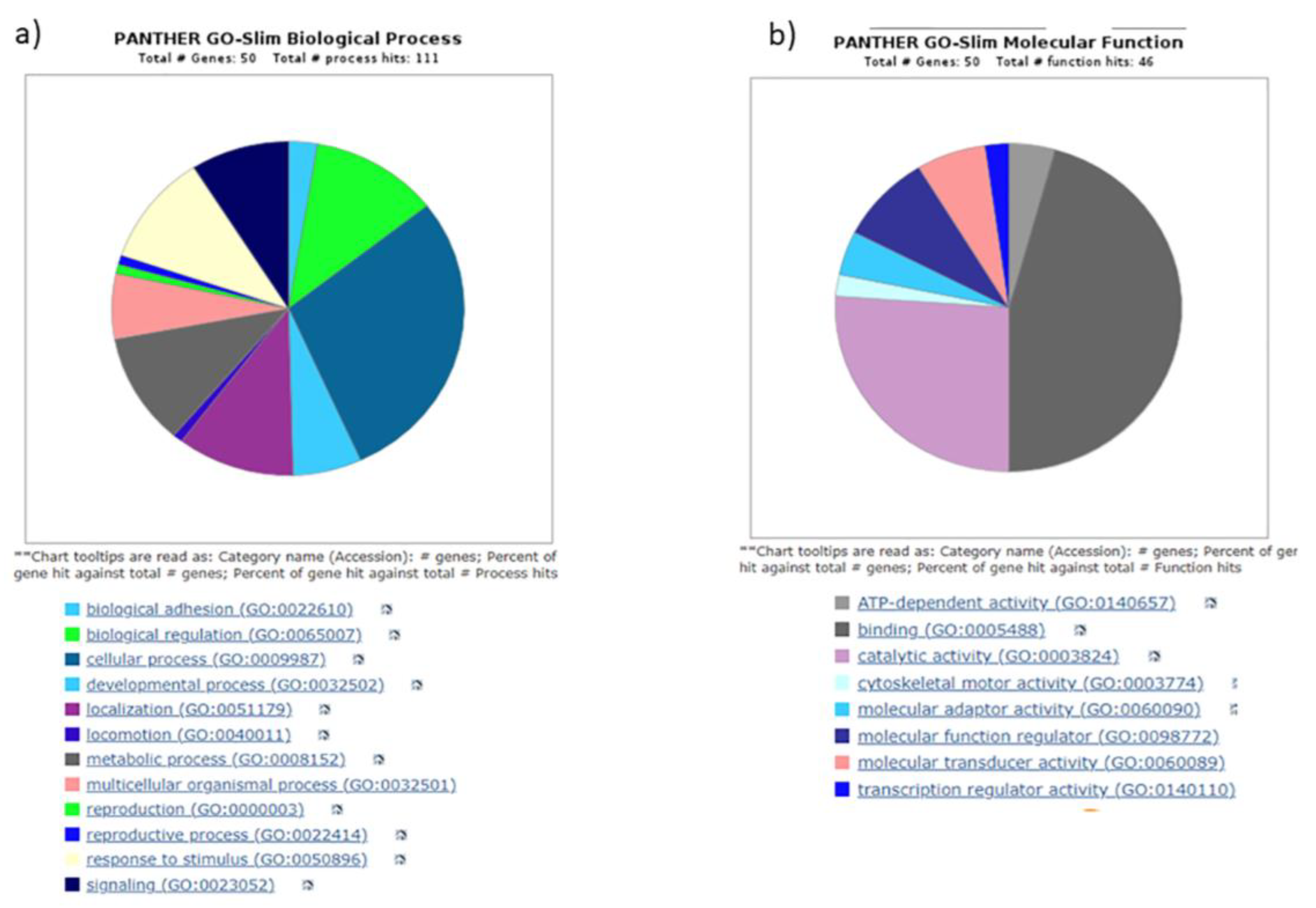
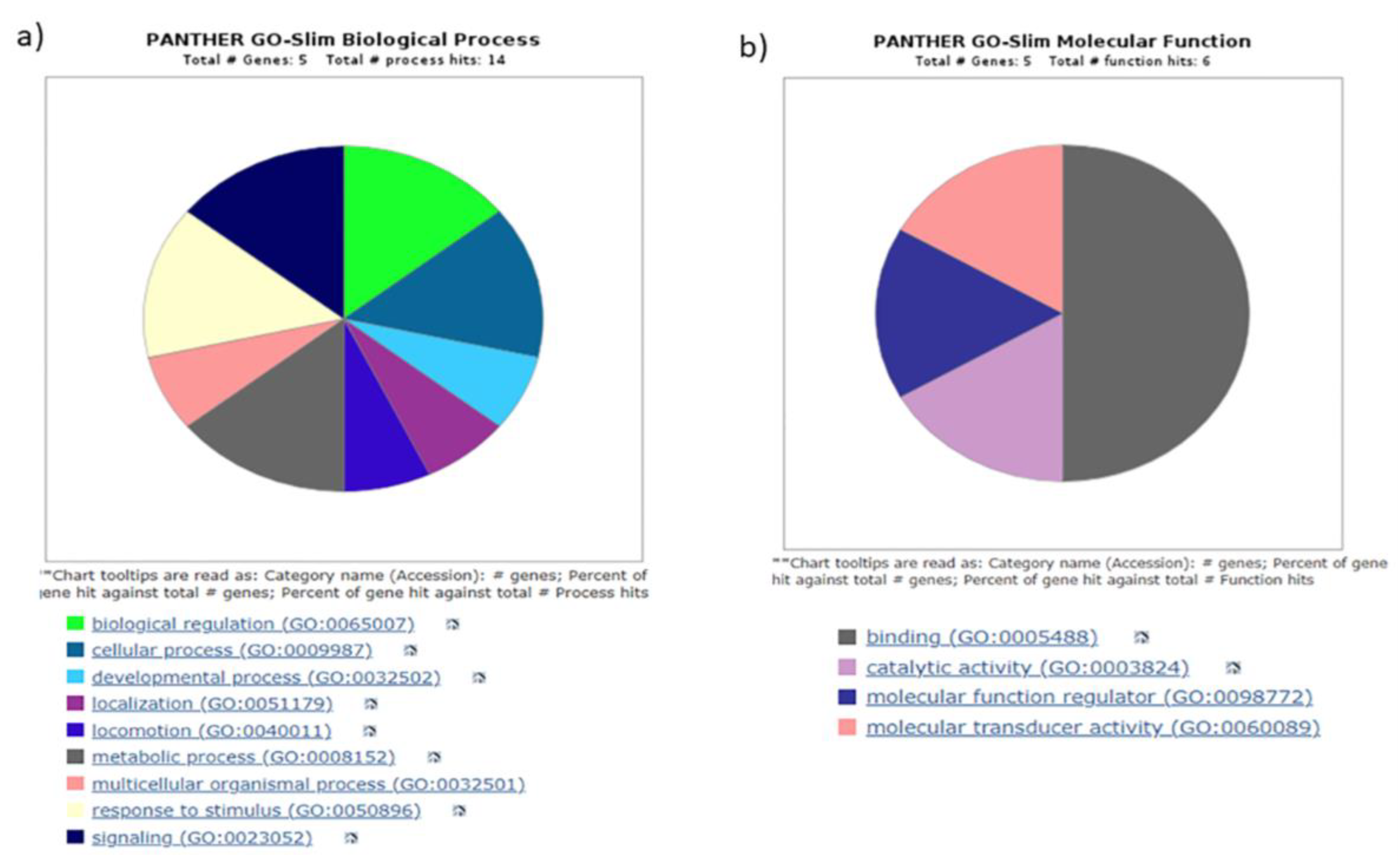
3.4. Gene ontology yielded distinct pathways regulating biogenesis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global Cancer Statistics. CA. Cancer J. Clin., 2011.
- Sim, H.G.; Cheng, C.W.S. Changing Demography of Prostate Cancer in Asia. Eur. J. Cancer, 2005.
- Jain, S.; Saxena, S.; Kumar, A. Epidemiology of Prostate Cancer in India. Meta Gene, 2014, 2, 596–605.
- NCRP. Three-Year Report of the Population Based Cancer Registries- 2009-2011; 2013.
- Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009, 1;25(9):1105-11.
- Guan YF, Li GR, Wang RJ, Yi YT, Yang L, Jiang D, Zhang XP, Peng Y. Application of next-generation sequencing in clinical oncology to advance personalized treatment of cancer. Chinese journal of cancer. 2012, 31(10):463.
- Gupta, A.; Shukla, N.; Nehra, M.; Gupta, S.; Malik, B.; Mishra, A.K.; Vijay, M.; Batra, J.; Lohiya, N.K.; Sharma, D.; Suravajhala, P. A Pilot Study on the Whole Exome Sequencing of Prostate Cancer in the Indian Phenotype Reveals Distinct Polymorphisms. Front. Genet., 2020, 11.
- Stark, R.; Grzelak, M.; Hadfield, J. RNA Sequencing: The Teenage Years. Nature Reviews Genetics, 2019.
- Marco-Puche, G.; Lois, S.; Benítez, J.; Trivino, J.C. RNA-Seq Perspectives to Improve Clinical Diagnosis. Frontiers in Genetics, 2019.
- Stupnikov A, McInerney CE, Savage KI, McIntosh SA, Emmert-Streib F, Kennedy R, Salto-Tellez M, Prise KM, McArt DG. Robustness of differential gene expression analysis of RNA-seq. Computational and structural biotechnology journal. 2021, 1;19:3470-81.
- Kukurba, K.R.; Montgomery, S.B. RNA Sequencing and Analysis. Cold Spring Harb. Protoc., 2015.
- Xi, X.; Li, T.; Huang, Y.; Sun, J.; Zhu, Y.; Yang, Y.; Lu, Z.J. RNA Biomarkers: Frontier of Precision Medicine for Cancer. Non-coding RNA, 2017.
- Chatterjee, S.K.; Zetter, B.R. Cancer Biomarkers: Knowing the Present and Predicting the Future. Future Oncology, 2005.
- Chen, L.L. The Biogenesis and Emerging Roles of Circular RNAs. Nature Reviews Molecular Cell Biology, 2016.
- Lopez, J.P.; Diallo, A.; Cruceanu, C.; Fiori, L.M.; Laboissiere, S.; Guillet, I.; Fontaine, J.; Ragoussis, J.; Benes, V.; Turecki, G.; Ernst, C. Biomarker Discovery: Quantification of MicroRNAs and Other Small Non-Coding RNAs Using next Generation Sequencing. BMC Med. Genomics, 2015.
- Krishnan P, Damaraju S. The challenges and opportunities in the clinical application of noncoding RNAs: the road map for miRNAs and piRNAs in cancer diagnostics and prognostics. International journal of genomics. 2018, 30;2018.
- Grixti, J.M.; Ayers, D. Long Noncoding RNAs and Their Link to Cancer. Non-coding RNA Research, 2020.
- Evans, J.R.; Feng, F.Y.; Chinnaiyan, A.M. The Bright Side of Dark Matter: LncRNAs in Cancer. Journal of Clinical Investigation, 2016.
- Arun, G.; Diermeier, S.D.; Spector, D.L. Therapeutic Targeting of Long Non-Coding RNAs in Cancer. Trends in Molecular Medicine, 2018.
- Gao, N.; Li, Y.; Li, J.; Gao, Z.; Yang, Z.; Li, Y.; Liu, H.; Fan, T. Long Non-Coding RNAs: The Regulatory Mechanisms, Research Strategies, and Future Directions in Cancers. Frontiers in Oncology, 2020.
- Winkle, M.; El-Daly, S.M.; Fabbri, M.; Calin, G.A. Noncoding RNA Therapeutics — Challenges and Potential Solutions. Nature Reviews Drug Discovery, 2021.
- Kumar P, Chakraborty J, Sukumar G, Dalgard C, Chatterjee R, Biswas R. Comparative RNA-seq analysis reveals dys-regulation of major canonical pathways in ERG-inducible LNCaP cell progression model of prostate cancer. Oncotarget. 2019 Jul 2;10(42):4290-4306. doi: 10.18632/oncotarget.27019. PMID: 31303963; PMCID: PMC6611515.
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential Gene and Transcript Expression Analysis of RNA-Seq Experiments with TopHat and Cufflinks. Nat. Protoc., 2012.
- Gupta, A.; Gupta, S.; Jatawa, S.K.; Kumar, A.; Suravajhala, P. A Simplest Bioinformatics Pipeline for Whole Transcriptome Sequencing: Overview of the Processing and Steps from Raw Data to Downstream Analysis. bioRxiv, 2019.
- Ferragina P, Manzini G. Proc. 41st Annual Symposium on Foundations of Computer Science. IEEE Comput. Soc.; 2000. pp. 390–398.
- Trapnell, C.; Hendrickson, D.G.; Sauvageau, M.; Goff, L.; Rinn, J.L.; Pachter, L. Differential Analysis of Gene Regulation at Transcript Resolution with RNA-Seq. Nat. Biotechnol., 2013.
- Liu, X.; Zhao, J.; Xue, L.; Zhao, T.; Ding, W.; Han, Y.; Ye, H. A Comparison of Transcriptome Analysis Methods with Reference Genome. BMC Genomics, 2022.
- Mostafavi, S.; Morris, Q. Combining Many Interaction Networks to Predict Gene Function and Analyze Gene Lists. Proteomics, 2012.
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; Jensen, L.J.; von Mering, C. The STRING Database in 2021: Customizable Protein-Protein Networks, and Functional Characterization of User-Uploaded Gene/Measurement Sets. Nucleic Acids Res., 2021.
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res., 2003.
- Saito R, Smoot ME, Ono K, Ruscheinski J, Wang PL, Lotia S, Pico AR, Bader GD, Ideker T. A travel guide to Cytoscape plugins. Nat Methods. 2012 Nov;9(11):1069-76. doi: 10.1038/nmeth.2212. Epub 2012 Nov 6. PMID: 23132118; PMCID: PMC3649846.
- Chin, C.H.; Chen, S.H.; Wu, H.H.; Ho, C.W.; Ko, M.T.; Lin, C.Y. CytoHubba: Identifying Hub Objects and Sub-Networks from Complex Interactome. BMC Syst. Biol., 2014.
- Mi, H.; Muruganujan, A.; Casagrande, J.T.; Thomas, P.D. Large-Scale Gene Function Analysis with the Panther Classification System. Nat. Protoc., 2013.
- Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y, Reva B, Goldberg AP, Sander C, Schultz N. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012 May;2(5):401-4. doi: 10.1158/2159-8290.CD-12-0095. Erratum in: Cancer Discov. 2012 Oct;2(10):960. PMID: 22588877; PMCID: PMC3956037.
- Liu, J.; Lichtenberg, T.; Hoadley, K.A.; Poisson, L.M.; Lazar, A.J.; Cherniack, A.D.; Kovatich, A.J.; Benz, C.C.; Levine, D.A.; Lee, A. V.; Omberg, L.; Wolf, D.M.; Shriver, C.D.; Thorsson, V.; Caesar-Johnson, S.J.; Demchok, J.A.; Felau, I.; Kasapi, M.; Ferguson, M.L.; Hutter, C.M.; Sofia, H.J.; Tarnuzzer, R.; Wang, Z.; Yang, L.; Zenklusen, J.C.; Zhang, J. (Julia); Chudamani, S.; Liu, J.; Lolla, L.; Naresh, R.; Pihl, T.; Sun, Q.; Wan, Y.; Wu, Y.; Cho, J.; DeFreitas, T.; Frazer, S.; Gehlenborg, N.; Getz, G.; Heiman, D.I.; Kim, J.; Lawrence, M.S.; Lin, P.; Meier, S.; Noble, M.S.; Saksena, G.; Voet, D.; Zhang, H.; Bernard, B.; Chambwe, N.; Dhankani, V.; Knijnenburg, T.; Kramer, R.; Leinonen, K.; Liu, Y.; Miller, M.; Reynolds, S.; Shmulevich, I.; Thorsson, V.; Zhang, W.; Akbani, R.; Broom, B.M.; Hegde, A.M.; Ju, Z.; Kanchi, R.S.; Korkut, A.; Li, J.; Liang, H.; Ling, S.; Liu, W.; Lu, Y.; Mills, G.B.; Ng, K.S.; Rao, A.; Ryan, M.; Wang, J.; Weinstein, J.N.; Zhang, J.; Abeshouse, A.; Armenia, J.; Chakravarty, D.; Chatila, W.K.; de Bruijn, I.; Gao, J.; Gross, B.E.; Heins, Z.J.; Kundra, R.; La, K.; Ladanyi, M.; Luna, A.; Nissan, M.G.; Ochoa, A.; Phillips, S.M.; Reznik, E.; Sanchez-Vega, F.; Sander, C.; Schultz, N.; Sheridan, R.; Sumer, S.O.; Sun, Y.; Taylor, B.S.; Wang, J.; Zhang, H.; Anur, P.; Peto, M.; Spellman, P.; Benz, C.; Stuart, J.M.; Wong, C.K.; Yau, C.; Hayes, D.N.; Parker, J.S.; Wilkerson, M.D.; Ally, A.; Balasundaram, M.; Bowlby, R.; Brooks, D.; Carlsen, R.; Chuah, E.; Dhalla, N.; Holt, R.; Jones, S.J.M.; Kasaian, K.; Lee, D.; Ma, Y.; Marra, M.A.; Mayo, M.; Moore, R.A.; Mungall, A.J.; Mungall, K.; Robertson, A.G.; Sadeghi, S.; Schein, J.E.; Sipahimalani, P.; Tam, A.; Thiessen, N.; Tse, K.; Wong, T.; Berger, A.C.; Beroukhim, R.; Cherniack, A.D.; Cibulskis, C.; Gabriel, S.B.; Gao, G.F.; Ha, G.; Meyerson, M.; Schumacher, S.E.; Shih, J.; Kucherlapati, M.H.; Kucherlapati, R.S.; Baylin, S.; Cope, L.; Danilova, L.; Bootwalla, M.S.; Lai, P.H.; Maglinte, D.T.; Van Den Berg, D.J.; Weisenberger, D.J.; Auman, J.T.; Balu, S.; Bodenheimer, T.; Fan, C.; Hoadley, K.A.; Hoyle, A.P.; Jefferys, S.R.; Jones, C.D.; Meng, S.; Mieczkowski, P.A.; Mose, L.E.; Perou, A.H.; Perou, C.M.; Roach, J.; Shi, Y.; Simons, J. V.; Skelly, T.; Soloway, M.G.; Tan, D.; Veluvolu, U.; Fan, H.; Hinoue, T.; Laird, P.W.; Shen, H.; Zhou, W.; Bellair, M.; Chang, K.; Covington, K.; Creighton, C.J.; Dinh, H.; Doddapaneni, H.V.; Donehower, L.A.; Drummond, J.; Gibbs, R.A.; Glenn, R.; Hale, W.; Han, Y.; Hu, J.; Korchina, V.; Lee, S.; Lewis, L.; Li, W.; Liu, X.; Morgan, M.; Morton, D.; Muzny, D.; Santibanez, J.; Sheth, M.; Shinbro, E.; Wang, L.; Wang, M.; Wheeler, D.A.; Xi, L.; Zhao, F.; Hess, J.; Appelbaum, E.L.; Bailey, M.; Cordes, M.G.; Ding, L.; Fronick, C.C.; Fulton, L.A.; Fulton, R.S.; Kandoth, C.; Mardis, E.R.; McLellan, M.D.; Miller, C.A.; Schmidt, H.K.; Wilson, R.K.; Crain, D.; Curley, E.; Gardner, J.; Lau, K.; Mallery, D.; Morris, S.; Paulauskis, J.; Penny, R.; Shelton, C.; Shelton, T.; Sherman, M.; Thompson, E.; Yena, P.; Bowen, J.; Gastier-Foster, J.M.; Gerken, M.; Leraas, K.M.; Lichtenberg, T.M.; Ramirez, N.C.; Wise, L.; Zmuda, E.; Corcoran, N.; Costello, T.; Hovens, C.; Carvalho, A.L.; de Carvalho, A.C.; Fregnani, J.H.; Longatto-Filho, A.; Reis, R.M.; Scapulatempo-Neto, C.; Silveira, H.C.S.; Vidal, D.O.; Burnette, A.; Eschbacher, J.; Hermes, B.; Noss, A.; Singh, R.; Anderson, M.L.; Castro, P.D.; Ittmann, M.; Huntsman, D.; Kohl, B.; Le, X.; Thorp, R.; Andry, C.; Duffy, E.R.; Lyadov, V.; Paklina, O.; Setdikova, G.; Shabunin, A.; Tavobilov, M.; McPherson, C.; Warnick, R.; Berkowitz, R.; Cramer, D.; Feltmate, C.; Horowitz, N.; Kibel, A.; Muto, M.; Raut, C.P.; Malykh, A.; Barnholtz-Sloan, J.S.; Barrett, W.; Devine, K.; Fulop, J.; Ostrom, Q.T.; Shimmel, K.; Wolinsky, Y.; Sloan, A.E.; De Rose, A.; Giuliante, F.; Goodman, M.; Karlan, B.Y.; Hagedorn, C.H.; Eckman, J.; Harr, J.; Myers, J.; Tucker, K.; Zach, L.A.; Deyarmin, B.; Hu, H.; Kvecher, L.; Larson, C.; Mural, R.J.; Somiari, S.; Vicha, A.; Zelinka, T.; Bennett, J.; Iacocca, M.; Rabeno, B.; Swanson, P.; Latour, M.; Lacombe, L.; Têtu, B.; Bergeron, A.; McGraw, M.; taugaitis, S.M.; Chabot, J.; Hibshoosh, H.; Sepulveda, A.; Su, T.; Wang, T.; Potapova, O.; Voronina, O.; Desjardins, L.; Mariani, O.; Roman-Roman, S.; Sastre, X.; Stern, M.H.; Cheng, F.; Signoretti, S.; Berchuck, A.; Bigner, D.; Lipp, E.; Marks, J.; McCall, S.; McLendon, R.; Secord, A.; Sharp, A.; Behera, M.; Brat, D.J.; Chen, A.; Delman, K.; Force, S.; Khuri, F.; Magliocca, K.; Maithel, S.; Olson, J.J.; Owonikoko, T.; Pickens, A.; Ramalingam, S.; Shin, D.M.; Sica, G.; Van Meir, E.G.; Zhang, H.; Eijckenboom, W.; Gillis, A.; Korpershoek, E.; Looijenga, L.; Oosterhuis, W.; Stoop, H.; van Kessel, K.E.; Zwarthoff, E.C.; Calatozzolo, C.; Cuppini, L.; Cuzzubbo, S.; DiMeco, F.; Finocchiaro, G.; Mattei, L.; Perin, A.; Pollo, B.; Chen, C.; Houck, J.; Lohavanichbutr, P.; Hartmann, A.; Stoehr, C.; Stoehr, R.; Taubert, H.; Wach, S.; Wullich, B.; Kycler, W.; Murawa, D.; Wiznerowicz, M.; Chung, K.; Edenfield, W.J.; Martin, J.; Baudin, E.; Bubley, G.; Bueno, R.; De Rienzo, A.; Richards, W.G.; Kalkanis, S.; Mikkelsen, T.; Noushmehr, H.; Scarpace, L.; Girard, N.; Aymerich, M.; Campo, E.; Giné, E.; Guillermo, A.L.; Van Bang, N.; Hanh, P.T.; Phu, B.D.; Tang, Y.; Colman, H.; Evason, K.; Dottino, P.R.; Martignetti, J.A.; Gabra, H.; Juhl, H.; Akeredolu, T.; Stepa, S.; Hoon, D.; Ahn, K.; Kang, K.J.; Beuschlein, F.; Breggia, A.; Birrer, M.; Bell, D.; Borad, M.; Bryce, A.H.; Castle, E.; Chandan, V.; Cheville, J.; Copland, J.A.; Farnell, M.; Flotte, T.; Giama, N.; Ho, T.; Kendrick, M.; Kocher, J.P.; Kopp, K.; Moser, C.; Nagorney, D.; O’Brien, D.; O’Neill, B.P.; Patel, T.; Petersen, G.; Que, F.; Rivera, M.; Roberts, L.; Smallridge, R.; Smyrk, T.; Stanton, M.; Thompson, R.H.; Torbenson, M.; Yang, J.D.; Zhang, L.; Brimo, F.; Ajani, J.A.; Angulo Gonzalez, A.M.; Behrens, C.; Bondaruk, J.; Broaddus, R.; Czerniak, B.; Esmaeli, B.; Fujimoto, J.; Gershenwald, J.; Guo, C.; Logothetis, C.; Meric-Bernstam, F.; Moran, C.; Ramondetta, L.; Rice, D.; Sood, A.; Tamboli, P.; Thompson, T.; Troncoso, P.; Tsao, A.; Wistuba, I.; Carter, C.; Haydu, L.; Hersey, P.; Jakrot, V.; Kakavand, H.; Kefford, R.; Lee, K.; Long, G.; Mann, G.; Quinn, M.; Saw, R.; Scolyer, R.; Shannon, K.; Spillane, A.; Stretch, J.; Synott, M.; Thompson, J.; Wilmott, J.; Al-Ahmadie, H.; Chan, T.A.; Ghossein, R.; Gopalan, A.; Levine, D.A.; Reuter, V.; Singer, S.; Singh, B.; Tien, N.V.; Broudy, T.; Mirsaidi, C.; Nair, P.; Drwiega, P.; Miller, J.; Smith, J.; Zaren, H.; Park, J.W.; Hung, N.P.; Kebebew, E.; Linehan, W.M.; Metwalli, A.R.; Pacak, K.; Pinto, P.A.; Schiffman, M.; Schmidt, L.S.; Vocke, C.D.; Wentzensen, N.; Worrell, R.; Yang, H.; Moncrieff, M.; Goparaju, C.; Melamed, J.; Pass, H.; Botnariuc, N.; Caraman, I.; Cernat, M.; Chemencedji, I.; Clipca, A.; Doruc, S.; Gorincioi, G.; Mura, S.; Pirtac, M.; Stancul, I.; Tcaciuc, D.; Albert, M.; Alexopoulou, I.; Arnaout, A.; Bartlett, J.; Engel, J.; Gilbert, S.; Parfitt, J.; Sekhon, H.; Thomas, G.; Rassl, D.M.; Rintoul, R.C.; Bifulco, C.; Tamakawa, R.; Urba, W.; Hayward, N.; Timmers, H.; Antenucci, A.; Facciolo, F.; Grazi, G.; Marino, M.; Merola, R.; de Krijger, R.; Gimenez-Roqueplo, A.P.; Piché, A.; Chevalier, S.; McKercher, G.; Birsoy, K.; Barnett, G.; Brewer, C.; Farver, C.; Naska, T.; Pennell, N.A.; Raymond, D.; Schilero, C.; Smolenski, K.; Williams, F.; Morrison, C.; Borgia, J.A.; Liptay, M.J.; Pool, M.; Seder, C.W.; Junker, K.; Omberg, L.; Dinkin, M.; Manikhas, G.; Alvaro, D.; Bragazzi, M.C.; Cardinale, V.; Carpino, G.; Gaudio, E.; Chesla, D.; Cottingham, S.; Dubina, M.; Moiseenko, F.; Dhanasekaran, R.; Becker, K.F.; Janssen, K.P.; Slotta-Huspenina, J.; Abdel-Rahman, M.H.; Aziz, D.; Bell, S.; Cebulla, C.M.; Davis, A.; Duell, R.; Elder, J.B.; Hilty, J.; Kumar, B.; Lang, J.; Lehman, N.L.; Mandt, R.; Nguyen, P.; Pilarski, R.; Rai, K.; Schoenfield, L.; Senecal, K.; Wakely, P.; Hansen, P.; Lechan, R.; Powers, J.; Tischler, A.; Grizzle, W.E.; Sexton, K.C.; Kastl, A.; Henderson, J.; Porten, S.; Waldmann, J.; Fassnacht, M.; Asa, S.L.; Schadendorf, D.; Couce, M.; Graefen, M.; Huland, H.; Sauter, G.; Schlomm, T.; Simon, R.; Tennstedt, P.; Olabode, O.; Nelson, M.; Bathe, O.; Carroll, P.R.; Chan, J.M.; Disaia, P.; Glenn, P.; Kelley, R.K.; Landen, C.N.; Phillips, J.; Prados, M.; Simko, J.; Smith-McCune, K.; VandenBerg, S.; Roggin, K.; Fehrenbach, A.; Kendler, A.; Sifri, S.; Steele, R.; Jimeno, A.; Carey, F.; Forgie, I.; Mannelli, M.; Carney, M.; Hernandez, B.; Campos, B.; Herold-Mende, C.; Jungk, C.; Unterberg, A.; von Deimling, A.; Bossler, A.; Galbraith, J.; Jacobus, L.; Knudson, M.; Knutson, T.; Ma, D.; Milhem, M.; Sigmund, R.; Godwin, A.K.; Madan, R.; Rosenthal, H.G.; Adebamowo, C.; Adebamowo, S.N.; Boussioutas, A.; Beer, D.; Giordano, T.; Mes-Masson, A.M.; Saad, F.; Bocklage, T.; Landrum, L.; Mannel, R.; Moore, K.; Moxley, K.; Postier, R.; Walker, J.; Zuna, R.; Feldman, M.; Valdivieso, F.; Dhir, R.; Luketich, J.; Mora Pinero, E.M.; Quintero-Aguilo, M.; Carlotti, C.G.; Dos Santos, J.S.; Kemp, R.; Sankarankuty, A.; Tirapelli, D.; Catto, J.; Agnew, K.; Swisher, E.; Creaney, J.; Robinson, B.; Shelley, C.S.; Godwin, E.M.; Kendall, S.; Shipman, C.; Bradford, C.; Carey, T.; Haddad, A.; Moyer, J.; Peterson, L.; Prince, M.; Rozek, L.; Wolf, G.; Bowman, R.; Fong, K.M.; Yang, I.; Korst, R.; Rathmell, W.K.; Fantacone-Campbell, J.L.; Hooke, J.A.; Kovatich, A.J.; Shriver, C.D.; DiPersio, J.; Drake, B.; Govindan, R.; Heath, S.; Ley, T.; Van Tine, B.; Westervelt, P.; Rubin, M.A.; Lee, J. Il; Aredes, N.D.; Mariamidze, A.; Hu, H. An Integrated TCGA Pan-Cancer Clinical Data Resource to Drive High-Quality Survival Outcome Analytics. Cell, 2018
- Warde-Farley, D.; Donaldson, S.L.; Comes, O.; Zuberi, K.; Badrawi, R.; Chao, P.; Franz, M.; Grouios, C.; Kazi, F.; Lopes, CT.; Maitland, A. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic acids research, 2010.
- Shukla, N., Siva, N., Sivakumar, M. B., Parveen, R., Mishra, A. K., Shah, A., Medicherla, K. M. and Suravajhala, P. (2021). Extraction of DNA and RNA from Formalin-fixed Paraffin-embedded Tissue Specimens. Bio-101: e4095. DOI: 10.21769/BioProtoc.4095.Shukla, N., Siva, N., Sivakumar, M. B., Parveen, R., Mishra, A. K., Shah, A., Medicherla, K. M. and Suravajhala, P. (2021). Extraction of DNA and RNA from Formalin-fixed Paraffin-embedded Tissue Specimens. Bio-101: e4095. [CrossRef]
- Assenov, Y.; Ramírez, F.; Schelhorn, S.E.; Lengauer, T.; Albrecht, M. Computing topological parameters of biological networks. Bioinformatics, 2008.
- Sanchez-Vega, F.; Mina, M.; Armenia, J.; Chatila, W.K.; Luna, A.; La, K.C.; Dimitriadoy, S.; Liu, D.L.; Kantheti, H.S.; Saghafinia, S.; Chakravarty, D.; Daian, F.; Gao, Q.; Bailey, M.H.; Liang, W.W.; Foltz, S.M.; Shmulevich, I.; Ding, L.; Heins, Z.; Ochoa, A.; Gross, B.; Gao, J.; Zhang, H.; Kundra, R.; Kandoth, C.; Bahceci, I.; Dervishi, L.; Dogrusoz, U.; Zhou, W.; Shen, H.; Laird, P.W.; Way, G.P.; Greene, C.S.; Liang, H.; Xiao, Y.; Wang, C.; Iavarone, A.; Berger, A.H.; Bivona, T.G.; Lazar, A.J.; Hammer, G.D.; Giordano, T.; Kwong, L.N.; McArthur, G.; Huang, C.; Tward, A.D.; Frederick, M.J.; McCormick, F.; Meyerson, M.; Caesar-Johnson, S.J.; Demchok, J.A.; Felau, I.; Kasapi, M.; Ferguson, M.L.; Hutter, C.M.; Sofia, H.J.; Tarnuzzer, R.; Wang, Z.; Yang, L.; Zenklusen, J.C.; Zhang, J. (Julia); Chudamani, S.; Liu, J.; Lolla, L.; Naresh, R.; Pihl, T.; Sun, Q.; Wan, Y.; Wu, Y.; Cho, J.; DeFreitas, T.; Frazer, S.; Gehlenborg, N.; Getz, G.; Heiman, D.I.; Kim, J.; Lawrence, M.S.; Lin, P.; Meier, S.; Noble, M.S.; Saksena, G.; Voet, D.; Zhang, H.; Bernard, B.; Chambwe, N.; Dhankani, V.; Knijnenburg, T.; Kramer, R.; Leinonen, K.; Liu, Y.; Miller, M.; Reynolds, S.; Shmulevich, I.; Thorsson, V.; Zhang, W.; Akbani, R.; Broom, B.M.; Hegde, A.M.; Ju, Z.; Kanchi, R.S.; Korkut, A.; Li, J.; Liang, H.; Ling, S.; Liu, W.; Lu, Y.; Mills, G.B.; Ng, K.S.; Rao, A.; Ryan, M.; Wang, J.; Weinstein, J.N.; Zhang, J.; Abeshouse, A.; Armenia, J.; Chakravarty, D.; Chatila, W.K.; de Bruijn, I.; Gross, B.E.; Heins, Z.J.; Kundra, R.; La, K.; Ladanyi, M.; Luna, A.; Nissan, M.G.; Ochoa, A.; Phillips, S.M.; Reznik, E.; Sanchez-Vega, F.; Sander, C.; Schultz, N.; Sheridan, R.; Sumer, S.O.; Sun, Y.; Taylor, B.S.; Wang, J.; Zhang, H.; Anur, P.; Peto, M.; Spellman, P.; Benz, C.; Stuart, J.M.; Wong, C.K.; Yau, C.; Hayes, D.N.; Parker, J.S.; Wilkerson, M.D.; Ally, A.; Balasundaram, M.; Bowlby, R.; Brooks, D.; Carlsen, R.; Chuah, E.; Dhalla, N.; Holt, R.; Jones, S.J.M.; Kasaian, K.; Lee, D.; Ma, Y.; Marra, M.A.; Mayo, M.; Moore, R.A.; Mungall, A.J.; Mungall, K.; Robertson, A.G.; Sadeghi, S.; Schein, J.E.; Sipahimalani, P.; Tam, A.; Thiessen, N.; Tse, K.; Wong, T.; Berger, A.C.; Beroukhim, R.; Cherniack, A.D.; Cibulskis, C.; Gabriel, S.B.; Gao, G.F.; Ha, G.; Meyerson, M.; Schumacher, S.E.; Shih, J.; Kucherlapati, M.H.; Kucherlapati, R.S.; Baylin, S.; Cope, L.; Danilova, L.; Bootwalla, M.S.; Lai, P.H.; Maglinte, D.T.; Van Den Berg, D.J.; Weisenberger, D.J.; Auman, J.T.; Balu, S.; Bodenheimer, T.; Fan, C.; Hoadley, K.A.; Hoyle, A.P.; Jefferys, S.R.; Jones, C.D.; Meng, S.; Mieczkowski, P.A.; Mose, L.E.; Perou, A.H.; Perou, C.M.; Roach, J.; Shi, Y.; Simons, J. V.; Skelly, T.; Soloway, M.G.; Tan, D.; Veluvolu, U.; Fan, H.; Hinoue, T.; Laird, P.W.; Shen, H.; Zhou, W.; Bellair, M.; Chang, K.; Covington, K.; Creighton, C.J.; Dinh, H.; Doddapaneni, H.V.; Donehower, L.A.; Drummond, J.; Gibbs, R.A.; Glenn, R.; Hale, W.; Han, Y.; Hu, J.; Korchina, V.; Lee, S.; Lewis, L.; Li, W.; Liu, X.; Morgan, M.; Morton, D.; Muzny, D.; Santibanez, J.; Sheth, M.; Shinbrot, E.; Wang, L.; Wang, M.; Wheeler, D.A.; Xi, L.; Zhao, F.; Hess, J.; Appelbaum, E.L.; Bailey, M.; Cordes, M.G.; Ding, L.; Fronick, C.C.; Fulton, L.A.; Fulton, R.S.; Kandoth, C.; Mardis, E.R.; McLellan, M.D.; Miller, C.A.; Schmidt, H.K.; Wilson, R.K.; Crain, D.; Curley, E.; Gardner, J.; Lau, K.; Mallery, D.; Morris, S.; Paulauskis, J.; Penny, R.; Shelton, C.; Shelton, T.; Sherman, M.; Thompson, E.; Yena, P.; Bowen, J.; Gastier-Foster, J.M.; Gerken, M.; Leraas, K.M.; Lichtenberg, T.M.; Ramirez, N.C.; Wise, L.; Zmuda, E.; Corcoran, N.; Costello, T.; Hovens, C.; Carvalho, A.L.; de Carvalho, A.C.; Fregnani, J.H.; Longatto-Filho, A.; Reis, R.M.; Scapulatempo-Neto, C.; Silveira, H.C.S.; Vidal, D.O.; Burnette, A.; Eschbacher, J.; Hermes, B.; Noss, A.; Singh, R.; Anderson, M.L.; Castro, P.D.; Ittmann, M.; Huntsman, D.; Kohl, B.; Le, X.; Thorp, R.; Andry, C.; Duffy, E.R.; Lyadov, V.; Paklina, O.; Setdikova, G.; Shabunin, A.; Tavobilov, M.; McPherson, C.; Warnick, R.; Berkowitz, R.; Cramer, D.; Feltmate, C.; Horowitz, N.; Kibel, A.; Muto, M.; Raut, C.P.; Malykh, A.; Barnholtz-Sloan, J.S.; Barrett, W.; Devine, K.; Fulop, J.; Ostrom, Q.T.; Shimmel, K.; Wolinsky, Y.; Sloan, A.E.; De Rose, A.; Giuliante, F.; Goodman, M.; Karlan, B.Y.; Hagedorn, C.H.; Eckman, J.; Harr, J.; Myers, J.; Tucker, K.; Zach, L.A.; Deyarmin, B.; Hu, H.; Kvecher, L.; Larson, C.; Mural, R.J.; Somiari, S.; Vicha, A.; Zelinka, T.; Bennett, J.; Iacocca, M.; Rabeno, B.; Swanson, P.; Latour, M.; Lacombe, L.; Têtu, B.; Bergeron, A.; McGraw, M.; Staugaitis, S.M.; Chabot, J.; Hibshoosh, H.; Sepulveda, A.; Su, T.; Wang, T.; Potapova, O.; Voronina, O.; Desjardins, L.; Mariani, O.; Roman-Roman, S.; Sastre, X.; Stern, M.H.; Cheng, F.; Signoretti, S.; Berchuck, A.; Bigner, D.; Lipp, E.; Marks, J.; McCall, S.; McLendon, R.; Secord, A.; Sharp, A.; Behera, M.; Brat, D.J.; Chen, A.; Delman, K.; Force, S.; Khuri, F.; Magliocca, K.; Maithel, S.; Olson, J.J.; Owonikoko, T.; Pickens, A.; Ramalingam, S.; Shin, D.M.; Sica, G.; Van Meir, E.G.; Zhang, H.; Eijckenboom, W.; Gillis, A.; Korpershoek, E.; Looijenga, L.; Oosterhuis, W.; Stoop, H.; van Kessel, K.E.; Zwarthoff, E.C.; Calatozzolo, C.; Cuppini, L.; Cuzzubbo, S.; DiMeco, F.; Finocchiaro, G.; Mattei, L.; Perin, A.; Pollo, B.; Chen, C.; Houck, J.; Lohavanichbutr, P.; Hartmann, A.; Stoehr, C.; Stoehr, R.; Taubert, H.; Wach, S.; Wullich, B.; Kycler, W.; Murawa, D.; Wiznerowicz, M.; Chung, K.; Edenfield, W.J.; Martin, J.; Baudin, E.; Bubley, G.; Bueno, R.; De Rienzo, A.; Richards, W.G.; Kalkanis, S.; Mikkelsen, T.; Noushmehr, H.; Scarpace, L.; Girard, N.; Aymerich, M.; Campo, E.; Giné, E.; Guillermo, A.L.; Van Bang, N.; Hanh, P.T.; Phu, B.D.; Tang, Y.; Colman, H.; Evason, K.; Dottino, P.R.; Martignetti, J.A.; Gabra, H.; Juhl, H.; Akeredolu, T.; Stepa, S.; Hoon, D.; Ahn, K.; Kang, K.J.; Beuschlein, F.; Breggia, A.; Birrer, M.; Bell, D.; Borad, M.; Bryce, A.H.; Castle, E.; Chandan, V.; Cheville, J.; Copland, J.A.; Farnell, M.; Flotte, T.; Giama, N.; Ho, T.; Kendrick, M.; Kocher, J.P.; Kopp, K.; Moser, C.; Nagorney, D.; O’Brien, D.; O’Neill, B.P.; Patel, T.; Petersen, G.; Que, F.; Rivera, M.; Roberts, L.; Smallridge, R.; Smyrk, T.; Stanton, M.; Thompson, R.H.; Torbenson, M.; Yang, J.D.; Zhang, L.; Brimo, F.; Ajani, J.A.; Gonzalez, A.M.A.; Behrens, C.; Bondaruk, J.; Broaddus, R.; Czerniak, B.; Esmaeli, B.; Fujimoto, J.; Gershenwald, J.; Guo, C.; Logothetis, C.; Meric-Bernstam, F.; Moran, C.; Ramondetta, L.; Rice, D.; Sood, A.; Tamboli, P.; Thompson, T.; Troncoso, P.; Tsao, A.; Wistuba, I.; Carter, C.; Haydu, L.; Hersey, P.; Jakrot, V.; Kakavand, H.; Kefford, R.; Lee, K.; Long, G.; Mann, G.; Quinn, M.; Saw, R.; Scolyer, R.; Shannon, K.; Spillane, A.; Stretch, J.; Synott, M.; Thompson, J.; Wilmott, J.; Al-Ahmadie, H.; Chan, T.A.; Ghossein, R.; Gopalan, A.; Levine, D.A.; Reuter, V.; Singer, S.; Singh, B.; Tien, N.V.; Broudy, T.; Mirsaidi, C.; Nair, P.; Drwiega, P.; Miller, J.; Smith, J.; Zaren, H.; Park, J.W.; Hung, N.P.; Kebebew, E.; Linehan, W.M.; Metwalli, A.R.; Pacak, K.; Pinto, P.A.; Schiffman, M.; Schmidt, L.S.; Vocke, C.D.; Wentzensen, N.; Worrell, R.; Yang, H.; Moncrieff, M.; Goparaju, C.; Melamed, J.; Pass, H.; Botnariuc, N.; Caraman, I.; Cernat, M.; Chemencedji, I.; Clipca, A.; Doruc, S.; Gorincioi, G.; Mura, S.; Pirtac, M.; Stancul, I.; Tcaciuc, D.; Albert, M.; Alexopoulou, I.; Arnaout, A.; Bartlett, J.; Engel, J.; Gilbert, S.; Parfitt, J.; Sekhon, H.; Thomas, G.; Rassl, D.M.; Rintoul, R.C.; Bifulco, C.; Tamakawa, R.; Urba, W.; Hayward, N.; Timmers, H.; Antenucci, A.; Facciolo, F.; Grazi, G.; Marino, M.; Merola, R.; de Krijger, R.; Gimenez-Roqueplo, A.P.; Piché, A.; Chevalier, S.; McKercher, G.; Birsoy, K.; Barnett, G.; Brewer, C.; Farver, C.; Naska, T.; Pennell, N.A.; Raymond, D.; Schilero, C.; Smolenski, K.; Williams, F.; Morrison, C.; Borgia, J.A.; Liptay, M.J.; Pool, M.; Seder, C.W.; Junker, K.; Omberg, L.; Dinkin, M.; Manikhas, G.; Alvaro, D.; Bragazzi, M.C.; Cardinale, V.; Carpino, G.; Gaudio, E.; Chesla, D.; Cottingham, S.; Dubina, M.; Moiseenko, F.; Dhanasekaran, R.; Becker, K.F.; Janssen, K.P.; Slotta-Huspenina, J.; Abdel-Rahman, M.H.; Aziz, D.; Bell, S.; Cebulla, C.M.; Davis, A.; Duell, R.; Elder, J.B.; Hilty, J.; Kumar, B.; Lang, J.; Lehman, N.L.; Mandt, R.; Nguyen, P.; Pilarski, R.; Rai, K.; Schoenfield, L.; Senecal, K.; Wakely, P.; Hansen, P.; Lechan, R.; Powers, J.; Tischler, A.; Grizzle, W.E.; Sexton, K.C.; Kastl, A.; Henderson, J.; Porten, S.; Waldmann, J.; Fassnacht, M.; Asa, S.L.; Schadendorf, D.; Couce, M.; Graefen, M.; Huland, H.; Sauter, G.; Schlomm, T.; Simon, R.; Tennstedt, P.; Olabode, O.; Nelson, M.; Bathe, O.; Carroll, P.R.; Chan, J.M.; Disaia, P.; Glenn, P.; Kelley, R.K.; Landen, C.N.; Phillips, J.; Prados, M.; Simko, J.; Smith-McCune, K.; VandenBerg, S.; Roggin, K.; Fehrenbach, A.; Kendler, A.; Sifri, S.; Steele, R.; Jimeno, A.; Carey, F.; Forgie, I.; Mannelli, M.; Carney, M.; Hernandez, B.; Campos, B.; Herold-Mende, C.; Jungk, C.; Unterberg, A.; von Deimling, A.; Bossler, A.; Galbraith, J.; Jacobus, L.; Knudson, M.; Knutson, T.; Ma, D.; Milhem, M.; Sigmund, R.; Godwin, A.K.; Madan, R.; Rosenthal, H.G.; Adebamowo, C.; Adebamowo, S.N.; Boussioutas, A.; Beer, D.; Giordano, T.; Mes-Masson, A.M.; Saad, F.; Bocklage, T.; Landrum, L.; Mannel, R.; Moore, K.; Moxley, K.; Postier, R.; Walker, J.; Zuna, R.; Feldman, M.; Valdivieso, F.; Dhir, R.; Luketich, J.; Pinero, E.M.M.; Quintero-Aguilo, M.; Carlotti, C.G.; Dos Santos, J.S.; Kemp, R.; Sankarankuty, A.; Tirapelli, D.; Catto, J.; Agnew, K.; Swisher, E.; Creaney, J.; Robinson, B.; Shelley, C.S.; Godwin, E.M.; Kendall, S.; Shipman, C.; Bradford, C.; Carey, T.; Haddad, A.; Moyer, J.; Peterson, L.; Prince, M.; Rozek, L.; Wolf, G.; Bowman, R.; Fong, K.M.; Yang, I. ; Korst, R.; Rathmell, W.K.; Fantacone-Campbell, J.L.; Hooke, J.A.; Kovatich, A.J.; Shriver, C.D.; DiPersio, J.; Drake, B.; Govindan, R.; Heath, S.; Ley, T.; Van Tine, B.; Westervelt, P.; Rubin, M.A.; Lee, J. Il; Aredes, N.D.; Mariamidze, A.; Van Allen, E.M.; Cherniack, A.D.; Ciriello, G.; Sander, C.; Schultz, N. Oncogenic Signaling Pathways in The Cancer Genome Atlas. Cell, 2018
- Lu, Q. δ-Catenin Dysregulation in Cancer: Interactions with E-Cadherin and Beyond. Journal of Pathology, 2010.
- Kantidze, O.L.; Kamalyukova, I.M.; Razin, S. V. Association of the Mammalian Transcriptional Regulator Kaiso with Centrosomes and the Midbody. Cell Cycle, 2009.
- Schackmann, R.C.J.; Tenhagen, M.; van de Ven, R.A.H.; Derksen, P.W.B. P120-Catenin in Cancer - Mechanisms, Models and Opportunities for Intervention. J. Cell Sci., 2013.
- Shukla, N.; Prasad, A.; Kanga, U.; Suravajhala, R.; Nigam, V.K.; Kishor, P.B.K.; Polavarapu, R.; Chaubey, G.; Singh, K.K.; Suravajhala, P. SARS-CoV-2 Transgressing LncRNAs Uncovers the Known Unknowns. Physiol. Genomics, 2021.
- Burnell, S.E.A.; Spencer-Harty, S.; Howarth, S.; Bodger, O.; Kynaston, H.; Morgan, C.; Doak, S.H. STEAP2 Knockdown Reduces the Invasive Potential of Prostate Cancer Cells. Sci. Rep., 2018.
- Stamey, T.A.; Yang, N.; Hay, A.R.; McNeal, J.E.; Freiha, F.S.; Redwine, E. Prostate-Specific Antigen as a Serum Marker for Adenocarcinoma of the Prostate. N. Engl. J. Med., 1987.
- Ahn, J.; Berndt, S.I.; Wacholder, S.; Kraft, P.; Kibel, A.S.; Yeager, M.; Albanes, D.; Giovannucci, E.; Stampfer, M.J.; Virtamo, J.; Thun, M.J.; Feigelson, H.S.; Cancel-Tassin, G.; Cussenot, O.; Thomas, G.; Hunter, D.J.; Fraumeni, J.F.; Hoover, R.N.; Chanock, S.J.; Hayes, R.B. Variation in KLK Genes, Prostate-Specific Antigen and Risk of Prostate Cancer. Nature Genetics, 2008.
- Shang, Z.; Niu, Y.; Cai, Q.; Chen, J.; Tian, J.; Yeh, S.; Lai, K.P.; Chang, C. Human Kallikrein 2 (KLK2) Promotes Prostate Cancer Cell Growth via Function as a Modulator to Promote the ARA70-Enhanced Androgen Receptor Transactivation. Tumor Biol., 2014.
- McDevitt MR, Thorek DL, Hashimoto T, Gondo T, Veach DR, Sharma SK, Kalidindi TM, Abou DS, Watson PA, Beattie BJ, Timmermand OV. Feed-forward alpha particle radiotherapy ablates androgen receptor-addicted prostate cancer. Nature communications. 2018, 24;9(1):1-1.
- Grimm, J.; Sachs, M.; Britsch, S.; Di Cesare, S.; Schwarz-Romond, T.; Alitalo, K.; Birchmeier, W. Novel P62dok Family Members, Dok-4 and Dok-5, Are Substrates of the c-Ret Receptor Tyrosine Kinase and Mediate Neuronal Differentiation. J. Cell Biol., 2001.
- Luo, F.; Wang, Z.; Chen, S.; Luo, Z.; Wang, G.; Yang, H.; Tang, L. DOK5 as a Prognostic Biomarker of Gastric Cancer Immunoinvasion: A Bioinformatics Analysis. Biomed Res. Int., 2022.
- Meng, J.; Wang, J. Role of SNARE Proteins in Tumourigenesis and Their Potential as Targets for Novel Anti-Cancer Therapeutics. Biochimica et Biophysica Acta - Reviews on Cancer, 2015.
- Han, J.; Pluhackova, K.; Böckmann, R.A. The Multifaceted Role of SNARE Proteins in Membrane Fusion. Frontiers in Physiology, 2017.
- Yan, R. Physiological Functions of the β-Site Amyloid Precursor Protein Cleaving Enzyme 1 and 2. Frontiers in Molecular Neuroscience, 2017.
- Farris, F.; Matafora, V.; Bachi, A. The Emerging Role of β-Secretases in Cancer. Journal of Experimental and Clinical Cancer Research, 2021.
- Amaral, P.P.; Neyt, C.; Wilkins, S.J.; Askarian-Amiri, M.E.; Sunkin, S.M.; Perkins, A.C.; Mattick, J.S. Complex Architecture and Regulated Expression of the Sox2ot Locus during Vertebrate Development. RNA, 2009.
- Li, Y.; Du, M.; Wang, S.; Zha, J.; Lei, P.; Wang, X.; Wu, D.; Zhang, J.; Chen, D.; Huang, D.; Lu, J.; Li, H.; Sun, M. Clinicopathological Implication of Long Non-Coding RNAs SOX2 Overlapping Transcript and Its Potential Target Gene Network in Various Cancers. Frontiers in Genetics, 2020.
- Li, S.; Zhang, Q.; Liu, W.; Zhao, C. Silencing of FTX Suppresses Pancreatic Cancer Cell Proliferation and Invasion by Upregulating MiR-513b-5p. BMC Cancer, 2021.
- Huo, X.; Wang, H.; Huo, B.; Wang, L.; Yang, K.; Wang, J.; Wang, L.; Wang, H. FTX Contributes to Cell Proliferation and Migration in Lung Adenocarcinoma via Targeting MiR-335-5p/NUCB2 Axis. Cancer Cell Int., 2020.
- Kong, D.; Zhao, Q.; Liu, W.; Wang, F. Identification of Crucial MiRNAs and IncRNAs for Ossification of Ligamentum Flavum. Mol. Med. Rep., 2019.
- Ren, S., Peng, Z., Mao, JH. et al. RNA-seq analysis of prostate cancer in the Chinese population identifies recurrent gene fusions, cancer-associated long noncoding RNAs and aberrant alternative splicings. Cell Res 22, 806–821 (2012). [CrossRef]
- Rajagopal, S.; Sharma, A.; Simlot, A.; Mathur, P.; Mehta, S.; Mehta, S.; Naravula, J.; Medicherla, K.M.; Kumar, A.; Kanga, U.; Suravajhala, R.; Bhandari, R.K.; Nair, B.G.; Kishor, P.B.K.; Suravajhala, P. Inferring bona fide Differentially Expressed Genes and Their Variants Associated with Vitamin K Deficiency Using a Systems Genetics Approach. Genes 2022, 13, 2078. 2022, 13, 2078. [CrossRef] [PubMed]
- Guo H, Ahmed M, Zhang F, Yao CQ, Li S, Liang Y, Hua J, Soares F, Sun Y, Langstein J, Li Y, Poon C, Bailey SD, Desai K, Fei T, Li Q, Sendorek DH, Fraser M, Prensner JR, Pugh TJ, Pomerantz M, Bristow RG, Lupien M, Feng FY, Boutros PC, Freedman ML, Walsh MJ, He HH. Modulation of long noncoding RNAs by risk SNPs underlying genetic predispositions to prostate cancer. Nat Genet. 2016 Oct;48(10):1142-50. doi: 10.1038/ng.3637. Epub 2016 Aug 15. PMID: 27526323.
- Ji P, Diederichs S, Wang W, Böing S, Metzger R, Schneider PM, Tidow N, Brandt B, Buerger H, Bulk E, Thomas M, Berdel WE, Serve H, Müller-Tidow C. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003 Sep 11;22(39):8031-41. doi: 10.1038/sj.onc.1206928. PMID: 12970751.
- Ren S, Liu Y, Xu W, Sun Y, Lu J, Wang F, Wei M, Shen J, Hou J, Gao X, Xu C, Huang J, Zhao Y, Sun Y. Long noncoding RNA MALAT-1 is a new potential therapeutic target for castration resistant prostate cancer. J Urol. 2013 Dec;190(6):2278-87. doi: 10.1016/j.juro.2013.07.001. Epub 2013 Jul 8. PMID: 23845456.
- Crea F, Watahiki A, Quagliata L, Xue H, Pikor L, Parolia A, Wang Y, Lin D, Lam WL, Farrar WL, Isogai T, Morant R, Castori-Eppenberger S, Chi KN, Wang Y, Helgason CD. Identification of a long non-coding RNA as a novel biomarker and potential therapeutic target for metastatic prostate cancer. Oncotarget. 2014 Feb 15;5(3):764-74. doi: 10.18632/oncotarget.1769. PMID: 24519926; PMCID: PMC3996663.
- Prensner JR, Zhao S, Erho N, Schipper M, Iyer MK, Dhanasekaran SM, Magi-Galluzzi C, Mehra R, Sahu A, Siddiqui J, Davicioni E, Den RB, Dicker AP, Karnes RJ, Wei JT, Klein EA, Jenkins RB, Chinnaiyan AM, Feng FY. RNA biomarkers associated with metastatic progression in prostate cancer: a multi-institutional high-throughput analysis of SChLAP1. Lancet Oncol. 2014 Dec;15(13):1469-1480. doi: 10.1016/S1470-2045(14)71113-1. Epub 2014 Nov 17. PMID: 25456366; PMCID: PMC4559342.
 |
| gene id | locus | log2 fold | p-value | gene name |
|---|---|---|---|---|
| CUFF.13238 | chr10:78033882-78040677 | 6.18617 | 0.0252 | RPS24 |
| CUFF.101498 | chr8:100713839-100714204 | 3.23209 | 0.03765 | PABPC1 |
| CUFF.17462 | chr11:57782675-57782947 | 4.93401 | 0.0003 | CTNND1 |
| CUFF.134059 | chr5:55937823-55938153 | 4.64641 | 0.0334 | IL6ST |
| CUFF.157342 | chr7:90232992-90235254 | 3.248 | 0.03545 | STEAP2 |
| CUFF.102981 | chr21:46003384-46005044 | 7.4652 | 0.0004 | COL6A1 |
| CUFF.119448 | chr3:181441034-181441358 | 3.356307 | 0.0131 | SOX2OT |
| CUFF.101277 | chr21:25880415-25881777 | 5.6321 | 0.03945 | APP |
| CUFF.41953 | chr17:48060789-48061102 | 4.18946 | 0.04285 | NFE2L2 |
| CUFF.110085 | chrX:74232840-74233182 | 3.23919 | 0.03765 | FTX |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





