Submitted:
30 November 2023
Posted:
30 November 2023
Read the latest preprint version here
Abstract
Keywords:
Author summary
Introduction
Results
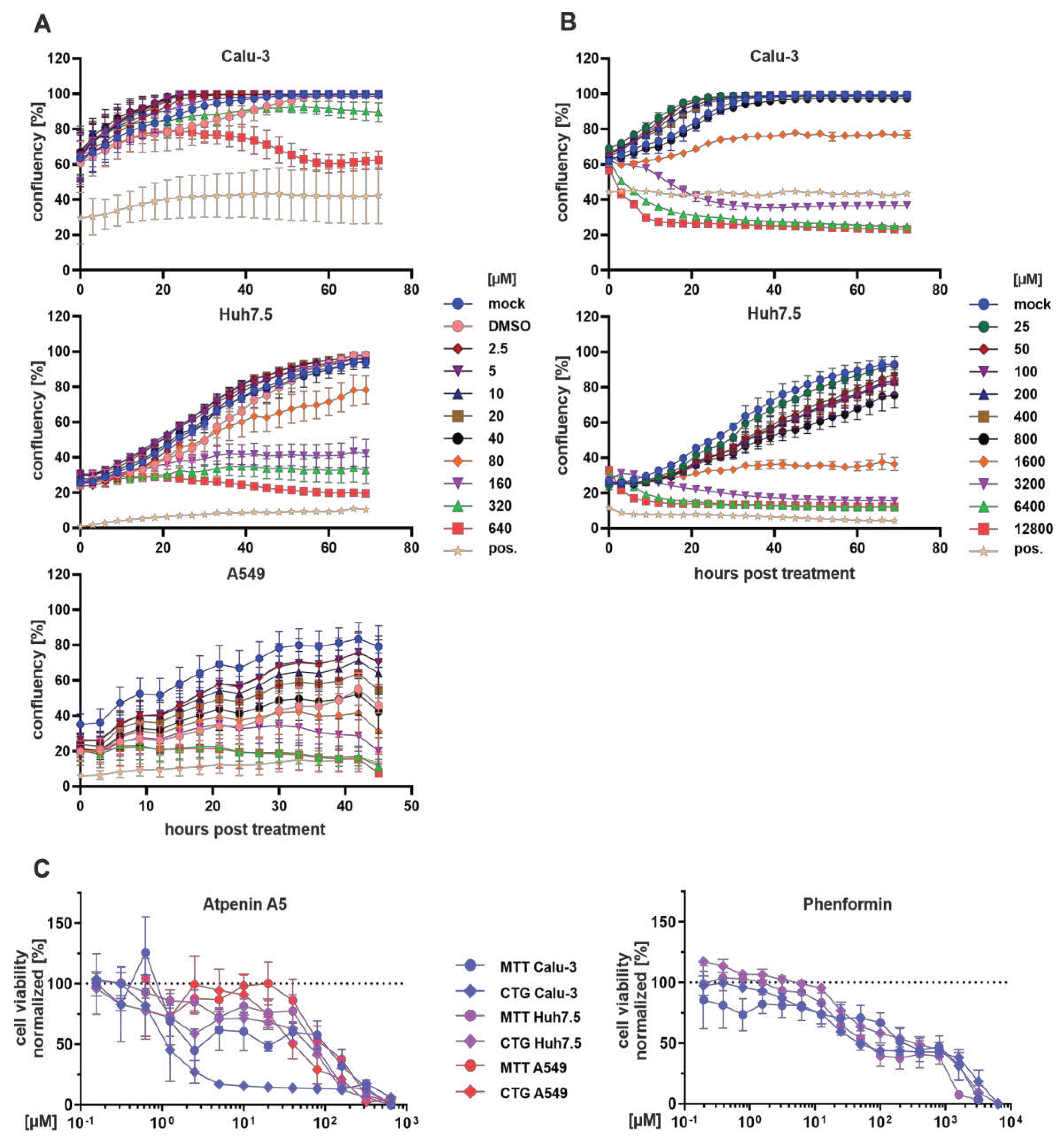
Detailed analysis of atpenin A5 and phenformin cytotoxicity and inhibition of metabolic activity in various cell lines.
Discussion
Methods
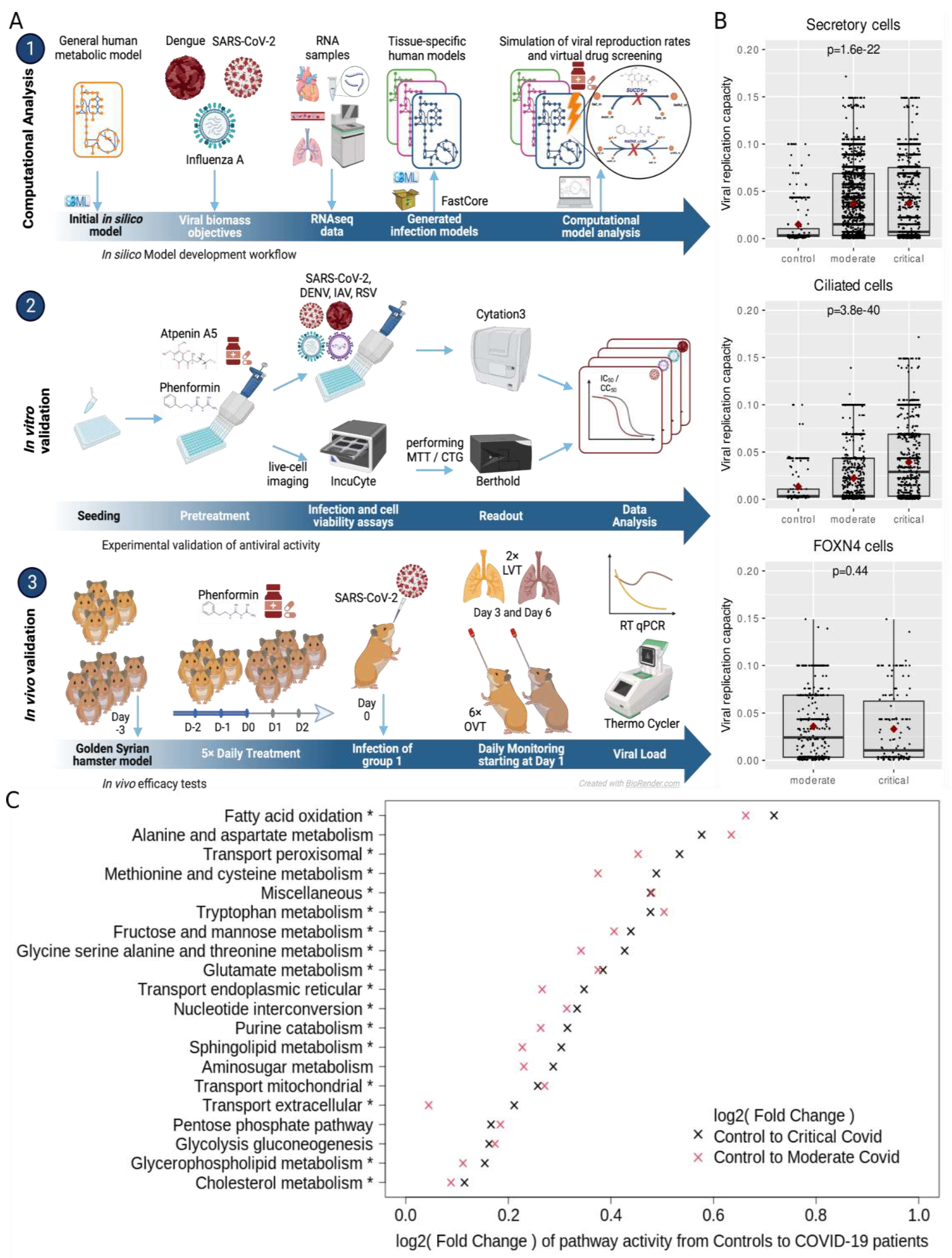
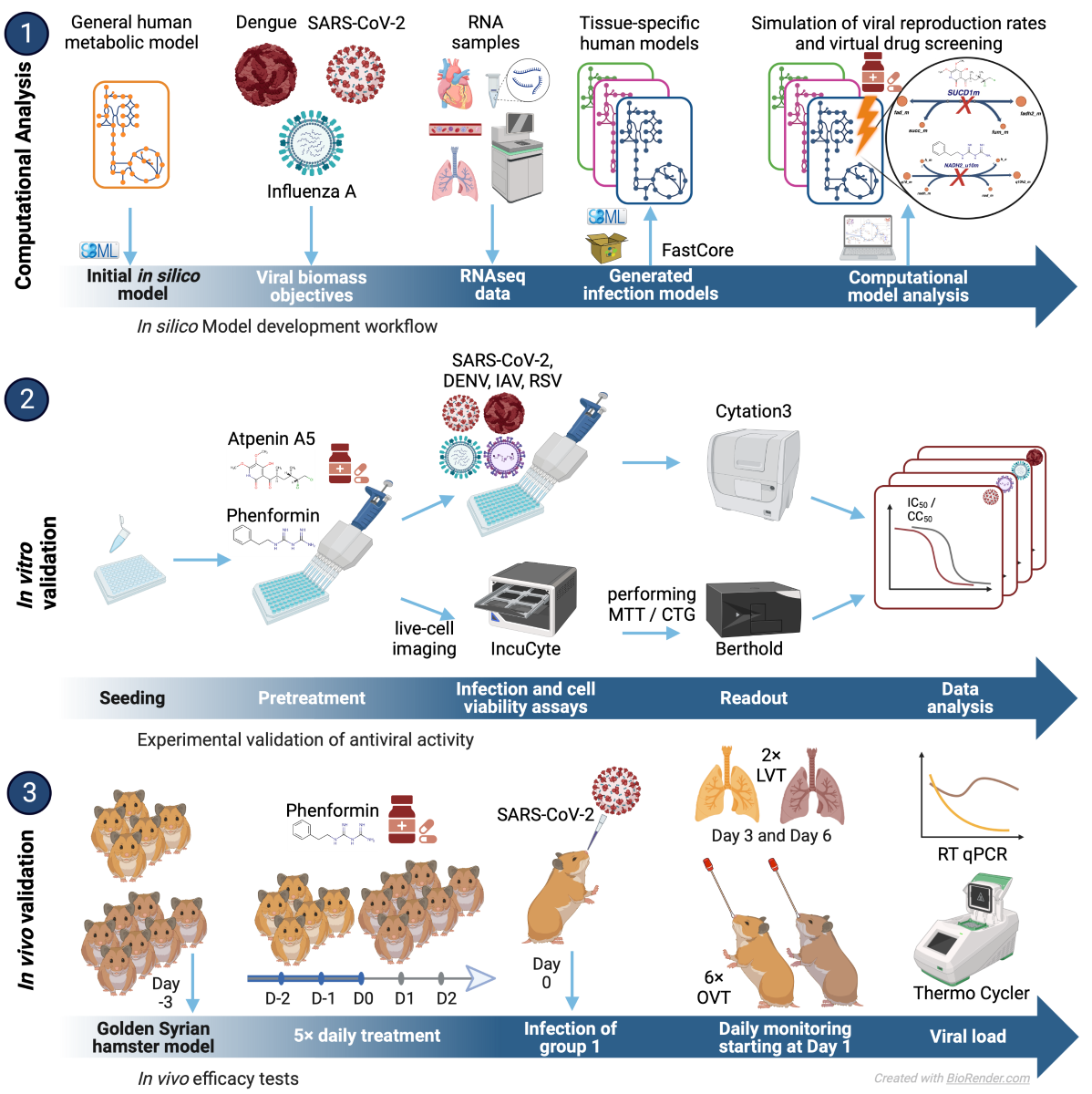
Computational analysis
Single-cell sequencing datasets
Viral replication models based on Recon 2.2
Cell identity annotation
scRNA data processing and reconstruction of context-specific metabolic models
Statistical analysis
Experimental approaches
Cell culture
Viruses
Compound information
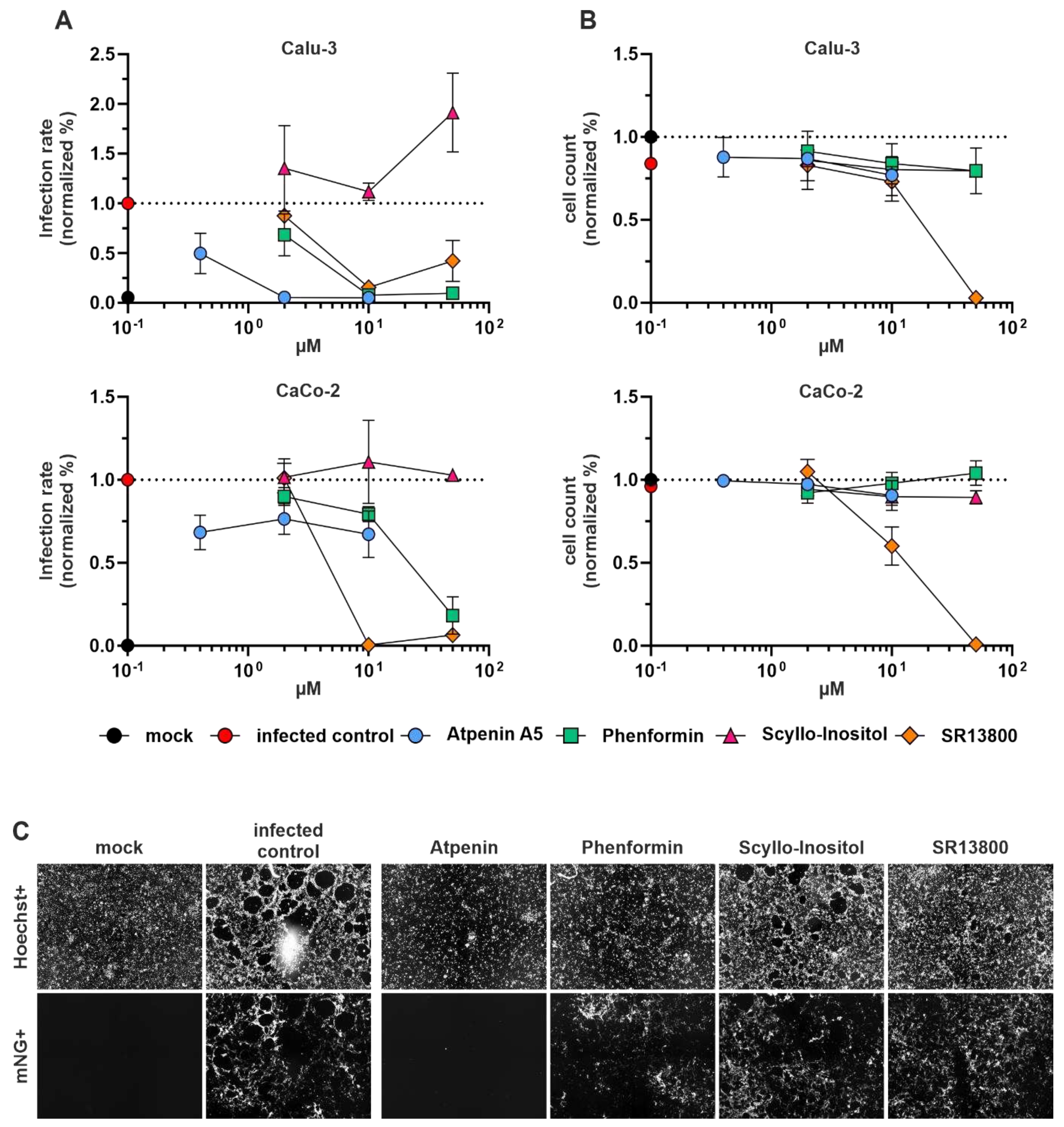
Initial screening of four drug candidates against SARS-CoV-2
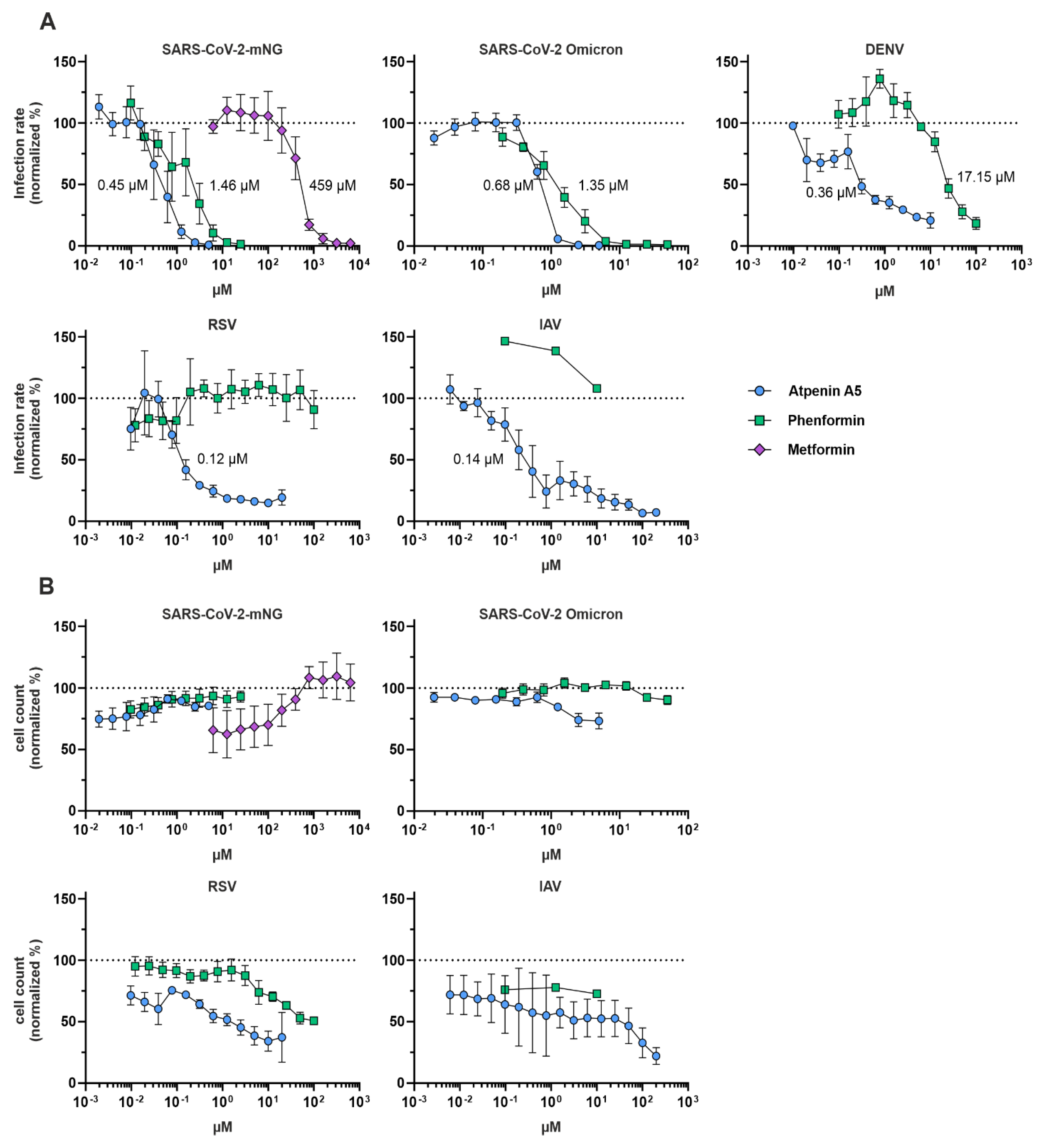
IC50 calculation of phenformin, metformin, and atpenin A5
CC50 calculation of phenformin and atpenin A5
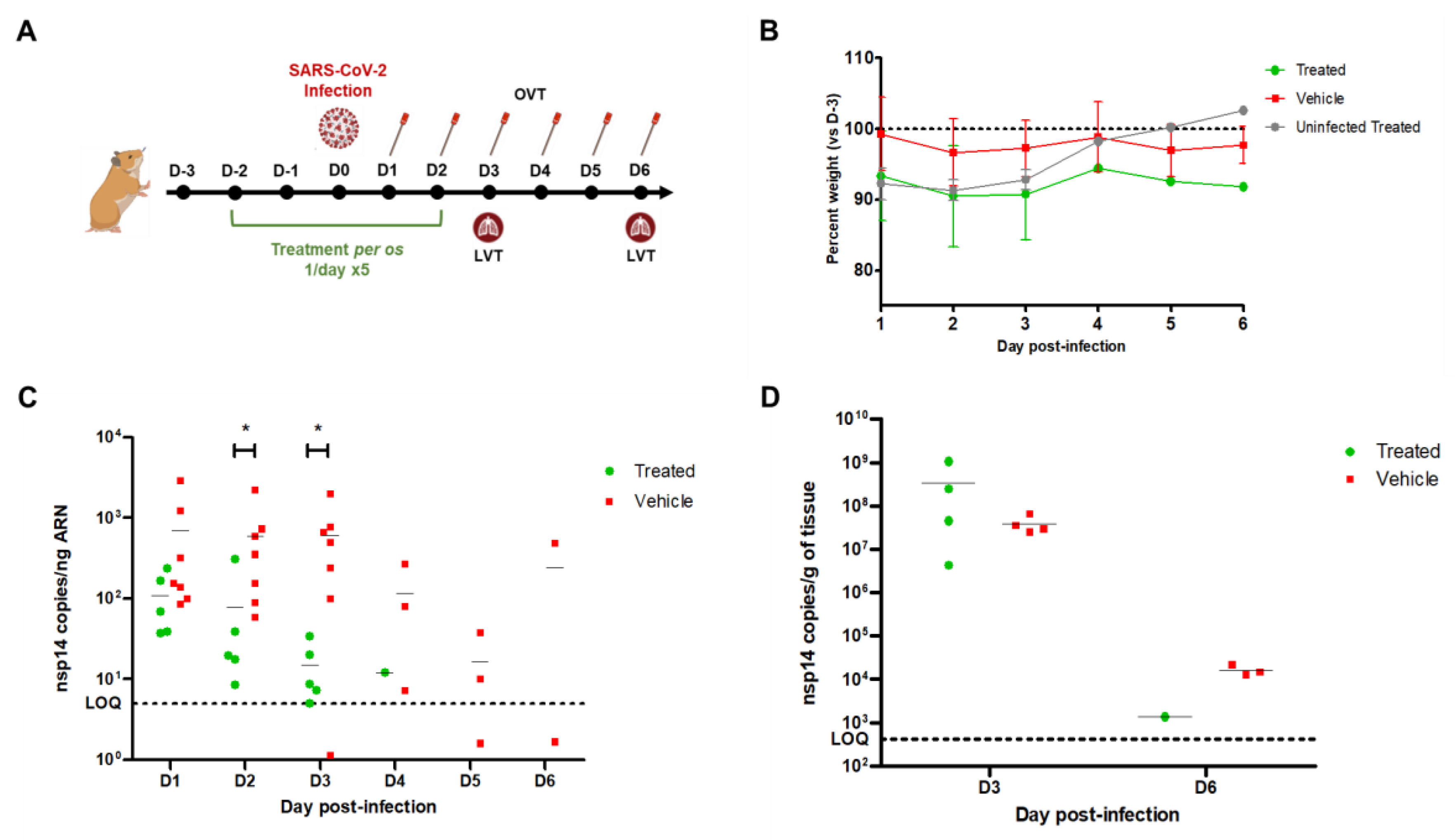
In vivo infection experiments
Supplementary Materials
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Adams TS, Schupp JC, Poli S, Ayaub EA, Neumark N, Ahangari F, Chu SG, Raby BA, DeIuliis G, Januszyk M, Duan Q, Arnett HA, Siddiqui A, Washko GR, Homer R, Yan X, Rosas IO, Kaminski N. 2020. Single-cell RNA-seq reveals ectopic and aberrant lung-resident cell populations in idiopathic pulmonary fibrosis. Sci Adv 6:eaba1983.
- Aller S, Scott A, Sarkar-Tyson M, Soyer OS. 2018. Integrated human-virus metabolic stoichiometric modelling predicts host-based antiviral targets against Chikungunya, Dengue and Zika viruses. J R Soc Interface 15. [CrossRef]
- Arrizabalaga J. 2006. The Black Death, 1346-1353: The Complete History (review). Bulletin of the History of Medicine. [CrossRef]
- Azhar EI, Hui DSC, Memish ZA, Drosten C, Zumla A. 2019. The Middle East Respiratory Syndrome (MERS). Infect Dis Clin North Am 33:891–905.
- Barbosa JAF, Sparapani S, Boulais J, Lodge R, Cohen ÉA. 2021. Human Immunodeficiency Virus Type 1 Vpr Mediates Degradation of APC1, a Scaffolding Component of the Anaphase-Promoting Complex/Cyclosome. J Virol 95:e0097120.
- Benjamini Y, Hochberg Y. 1995. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society: Series B (Methodological). [CrossRef]
- Bergmann FT, Adams R, Moodie S, Cooper J, Glont M, Golebiewski M, Hucka M, Laibe C, Miller AK, Nickerson DP, Olivier BG, Rodriguez N, Sauro HM, Scharm M, Soiland-Reyes S, Waltemath D, Yvon F, Le Novère N. 2014. COMBINE archive and OMEX format: one file to share all information to reproduce a modeling project. BMC Bioinformatics 15:369.
- Bernardes JP, Mishra N, Tran F, Bahmer T, Best L, Blase JI, Bordoni D, Franzenburg J, Geisen U, Josephs-Spaulding J, Köhler P, Künstner A, Rosati E, Aschenbrenner AC, Bacher P, Baran N, Boysen T, Brandt B, Bruse N, Dörr J, Dräger A, Elke G, Ellinghaus D, Fischer J, Forster M, Franke A, Franzenburg S, Frey N, Friedrichs A, Fuß J, Glück A, Hamm J, Hinrichsen F, Hoeppner MP, Imm S, Junker R, Kaiser S, Kan YH, Knoll R, Lange C, Laue G, Lier C, Lindner M, Marinos G, Markewitz R, Nattermann J, Noth R, Pickkers P, Rabe KF, Renz A, Röcken C, Rupp J, Schaffarzyk A, Scheffold A, Schulte-Schrepping J, Schunk D, Skowasch D, Ulas T, Wandinger K-P, Wittig M, Zimmermann J, Busch H, Hoyer BF, Kaleta C, Heyckendorf J, Kox M, Rybniker J, Schreiber S, Schultze JL, Rosenstiel P, HCA Lung Biological Network, Deutsche COVID-19 Omics Initiative (DeCOI). 2020. Longitudinal Multi-omics Analyses Identify Responses of Megakaryocytes, Erythroid Cells, and Plasmablasts as Hallmarks of Severe COVID-19. Immunity 53:1296–1314.e9.
- Bordbar A, Monk JM, King ZA, Palsson BO. 2014. Constraint-based models predict metabolic and associated cellular functions. Nat Rev Genet 15:107–120.
- Bramante CT, Buse JB, Liebovitz DM, Nicklas JM, Puskarich MA, Cohen K, Belani HK, Anderson BJ, Huling JD, Tignanelli CJ, Thompson JL, Pullen M, Wirtz EL, Siegel LK, Proper JL, Odde DJ, Klatt NR, Sherwood NE, Lindberg SM, Karger AB, Beckman KB, Erickson SM, Fenno SL, Hartman KM, Rose MR, Mehta T, Patel B, Griffiths G, Bhat NS, Murray TA, Boulware DR. 2023. Outpatient treatment of COVID-19 and incidence of post-COVID-19 condition over 10 months (COVID-OUT): a multicentre, randomised, quadruple-blind, parallel-group, phase 3 trial. Lancet Infect Dis. [CrossRef]
- Bramante CT, Huling JD, Tignanelli CJ, Buse JB, Liebovitz DM, Nicklas JM, Cohen K, Puskarich MA, Belani HK, Proper JL, Siegel LK, Klatt NR, Odde DJ, Luke DG, Anderson B, Karger AB, Ingraham NE, Hartman KM, Rao V, Hagen AA, Patel B, Fenno SL, Avula N, Reddy NV, Erickson SM, Lindberg S, Fricton R, Lee S, Zaman A, Saveraid HG, Tordsen WJ, Pullen MF, Biros M, Sherwood NE, Thompson JL, Boulware DR, Murray TA, COVID-OUT Trial Team. 2022. Randomized Trial of Metformin, Ivermectin, and Fluvoxamine for Covid-19. N Engl J Med 387:599–610.
- Butler A, Hoffman P, Smibert P, Papalexi E, Satija R. 2018. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol 36:411–420.
- Cavallari I, Scattolin G, Silic-Benussi M, Raimondi V, D’Agostino DM, Ciminale V. 2018. Mitochondrial Proteins Coded by Human Tumor Viruses. Front Microbiol 9:81.
- Chan JF-W, Zhang AJ, Yuan S, Poon VK-M, Chan CC-S, Lee AC-Y, Chan W-M, Fan Z, Tsoi H-W, Wen L, Liang R, Cao J, Chen Y, Tang K, Luo C, Cai J-P, Kok K-H, Chu H, Chan K-H, Sridhar S, Chen Z, Chen H, To KK-W, Yuen K-Y. 2020. Simulation of the Clinical and Pathological Manifestations of Coronavirus Disease 2019 (COVID-19) in a Golden Syrian Hamster Model: Implications for Disease Pathogenesis and Transmissibility. Clin Infect Dis 71:2428–2446.
- Chen X, Ji ZL, Chen YZ. 2002. TTD: Therapeutic Target Database. Nucleic Acids Res 30:412–415.
- Chua RL, Lukassen S, Trump S, Hennig BP, Wendisch D, Pott F, Debnath O, Thürmann L, Kurth F, Völker MT, Kazmierski J, Timmermann B, Twardziok S, Schneider S, Machleidt F, Müller-Redetzky H, Maier M, Krannich A, Schmidt S, Balzer F, Liebig J, Loske J, Suttorp N, Eils J, Ishaque N, Liebert UG, von Kalle C, Hocke A, Witzenrath M, Goffinet C, Drosten C, Laudi S, Lehmann I, Conrad C, Sander L-E, Eils R. 2020. COVID-19 severity correlates with airway epithelium-immune cell interactions identified by single-cell analysis. Nat Biotechnol 38:970–979.
- Clough E, Barrett T. 2016. The Gene Expression Omnibus Database. Methods in Molecular Biology. [CrossRef]
- Cotto KC, Wagner AH, Feng Y-Y, Kiwala S, Coffman AC, Spies G, Wollam A, Spies NC, Griffith OL, Griffith M. 2018. DGIdb 3.0: a redesign and expansion of the drug-gene interaction database. Nucleic Acids Res 46:D1068–D1073.
- Crouse AB, Grimes T, Li P, Might M, Ovalle F, Shalev A. 2020. Metformin Use Is Associated With Reduced Mortality in a Diverse Population With COVID-19 and Diabetes. Front Endocrinol 11:600439.
- Cunningham F, Achuthan P, Akanni W, Allen J, Amode MR, Armean IM, Bennett R, Bhai J, Billis K, Boddu S, Cummins C, Davidson C, Dodiya KJ, Gall A, Girón CG, Gil L, Grego T, Haggerty L, Haskell E, Hourlier T, Izuogu OG, Janacek SH, Juettemann T, Kay M, Laird MR, Lavidas I, Liu Z, Loveland JE, Marugán JC, Maurel T, McMahon AC, Moore B, Morales J, Mudge JM, Nuhn M, Ogeh D, Parker A, Parton A, Patricio M, Abdul Salam AI, Schmitt BM, Schuilenburg H, Sheppard D, Sparrow H, Stapleton E, Szuba M, Taylor K, Threadgold G, Thormann A, Vullo A, Walts B, Winterbottom A, Zadissa A, Chakiachvili M, Frankish A, Hunt SE, Kostadima M, Langridge N, Martin FJ, Muffato M, Perry E, Ruffier M, Staines DM, Trevanion SJ, Aken BL, Yates AD, Zerbino DR, Flicek P. 2019. Ensembl 2019. Nucleic Acids Res 47:D745–D751.
- Cutler DM, Summers LH. 2020. The COVID-19 Pandemic and the $16 Trillion Virus. JAMA 324:1495–1496.
- Daniloski Z, Jordan TX, Wessels H-H, Hoagland DA, Kasela S, Legut M, Maniatis S, Mimitou EP, Lu L, Geller E, Danziger O, Rosenberg BR, Phatnani H, Smibert P, Lappalainen T, tenOever BR, Sanjana NE. 2021. Identification of Required Host Factors for SARS-CoV-2 Infection in Human Cells. Cell 184:92–105.e16.
- Davis ZH, Verschueren E, Jang GM, Kleffman K, Johnson JR, Park J, Von Dollen J, Maher MC, Johnson T, Newton W, Jäger S, Shales M, Horner J, Hernandez RD, Krogan NJ, Glaunsinger BA. 2015. Global mapping of herpesvirus-host protein complexes reveals a transcription strategy for late genes. Mol Cell 57:349–360.
- de Chassey B, Aublin-Gex A, Ruggieri A, Meyniel-Schicklin L, Pradezynski F, Davoust N, Chantier T, Tafforeau L, Mangeot P-E, Ciancia C, Perrin-Cocon L, Bartenschlager R, André P, Lotteau V. 2013. The interactomes of influenza virus NS1 and NS2 proteins identify new host factors and provide insights for ADAR1 playing a supportive role in virus replication. PLoS Pathog 9:e1003440.
- de Kanter JK, Lijnzaad P, Candelli T, Margaritis T, Holstege FCP. 2019. CHETAH: a selective, hierarchical cell type identification method for single-cell RNA sequencing. Nucleic Acids Res 47:e95.
- Deprez M, Zaragosi L-E, Truchi M, Becavin C, Ruiz García S, Arguel M-J, Plaisant M, Magnone V, Lebrigand K, Abelanet S, Brau F, Paquet A, Pe’er D, Marquette C-H, Leroy S, Barbry P. 2020. A Single-Cell Atlas of the Human Healthy Airways. Am J Respir Crit Care Med 202:1636–1645.
- Di Magno L, Di Pastena F, Bordone R, Coni S, Canettieri G. 2022. The Mechanism of Action of Biguanides: New Answers to a Complex Question. Cancers 14. [CrossRef]
- Doherty JR, Yang C, Scott KEN, Cameron MD, Fallahi M, Li W, Hall MA, Amelio AL, Mishra JK, Li F, Tortosa M, Genau HM, Rounbehler RJ, Lu Y, Dang CV, Kumar KG, Butler AA, Bannister TD, Hooper AT, Unsal-Kacmaz K, Roush WR, Cleveland JL. 2014. Blocking lactate export by inhibiting the Myc target MCT1 Disables glycolysis and glutathione synthesis. Cancer Res 74:908–920.
- Fischl W, Bartenschlager R. 2013. High-throughput screening using dengue virus reporter genomes. Methods Mol Biol 1030:205–219.
- Garcia CK, Goldstein JL, Pathak RK, Anderson RG, Brown MS. 1994. Molecular characterization of a membrane transporter for lactate, pyruvate, and other monocarboxylates: implications for the Cori cycle. Cell 76:865–873.
- Gawron P, Ostaszewski M, Satagopam V, Gebel S, Mazein A, Kuzma M, Zorzan S, McGee F, Otjacques B, Balling R, Schneider R. 2016. MINERVA-a platform for visualization and curation of molecular interaction networks. NPJ Syst Biol Appl 2:16020.
- Geraghty RJ, Aliota MT, Bonnac LF. 2021. Broad-Spectrum Antiviral Strategies and Nucleoside Analogues. Viruses 13. [CrossRef]
- Germain M-A, Chatel-Chaix L, Gagné B, Bonneil É, Thibault P, Pradezynski F, de Chassey B, Meyniel-Schicklin L, Lotteau V, Baril M, Lamarre D. 2014. Elucidating novel hepatitis C virus-host interactions using combined mass spectrometry and functional genomics approaches. Mol Cell Proteomics 13:184–203.
- Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, O’Meara MJ, Rezelj VV, Guo JZ, Swaney DL, Tummino TA, Hüttenhain R, Kaake RM, Richards AL, Tutuncuoglu B, Foussard H, Batra J, Haas K, Modak M, Kim M, Haas P, Polacco BJ, Braberg H, Fabius JM, Eckhardt M, Soucheray M, Bennett MJ, Cakir M, McGregor MJ, Li Q, Meyer B, Roesch F, Vallet T, Mac Kain A, Miorin L, Moreno E, Naing ZZC, Zhou Y, Peng S, Shi Y, Zhang Z, Shen W, Kirby IT, Melnyk JE, Chorba JS, Lou K, Dai SA, Barrio-Hernandez I, Memon D, Hernandez-Armenta C, Lyu J, Mathy CJP, Perica T, Pilla KB, Ganesan SJ, Saltzberg DJ, Rakesh R, Liu X, Rosenthal SB, Calviello L, Venkataramanan S, Liboy-Lugo J, Lin Y, Huang X-P, Liu Y, Wankowicz SA, Bohn M, Safari M, Ugur FS, Koh C, Savar NS, Tran QD, Shengjuler D, Fletcher SJ, O’Neal MC, Cai Y, Chang JCJ, Broadhurst DJ, Klippsten S, Sharp PP, Wenzell NA, Kuzuoglu-Ozturk D, Wang H-Y, Trenker R, Young JM, Cavero DA, Hiatt J, Roth TL, Rathore U, Subramanian A, Noack J, Hubert M, Stroud RM, Frankel AD, Rosenberg OS, Verba KA, Agard DA, Ott M, Emerman M, Jura N, von Zastrow M, Verdin E, Ashworth A, Schwartz O, d’Enfert C, Mukherjee S, Jacobson M, Malik HS, Fujimori DG, Ideker T, Craik CS, Floor SN, Fraser JS, Gross JD, Sali A, Roth BL, Ruggero D, Taunton J, Kortemme T, Beltrao P, Vignuzzi M, García-Sastre A, Shokat KM, Shoichet BK, Krogan NJ. 2020. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583:459–468.
- Gu C, Kim GB, Kim WJ, Kim HU, Lee SY. 2019. Current status and applications of genome-scale metabolic models. Genome Biol 20:121.
- He J, Cai S, Feng H, Cai B, Lin L, Mai Y, Fan Y, Zhu A, Huang H, Shi J, Li D, Wei Y, Li Y, Zhao Y, Pan Y, Liu H, Mo X, He X, Cao S, Hu F, Zhao J, Wang J, Zhong N, Chen X, Deng X, Chen J. 2020. Single-cell analysis reveals bronchoalveolar epithelial dysfunction in COVID-19 patients. Protein Cell.
- Honigsbaum M. 2020. Revisiting the 1957 and 1968 influenza pandemics. Lancet 395:1824–1826.
- Horsefield R, Yankovskaya V, Sexton G, Whittingham W, Shiomi K, Omura S, Byrne B, Cecchini G, Iwata S. 2006. Structural and computational analysis of the quinone-binding site of complex II (succinate-ubiquinone oxidoreductase): a mechanism of electron transfer and proton conduction during ubiquinone reduction. J Biol Chem 281:7309–7316.
- Howe KL, Achuthan P, Allen J, Allen J, Alvarez-Jarreta J, Amode MR, Armean IM, Azov AG, Bennett R, Bhai J, Billis K, Boddu S, Charkhchi M, Cummins C, Da Rin Fioretto L, Davidson C, Dodiya K, El Houdaigui B, Fatima R, Gall A, Garcia Giron C, Grego T, Guijarro-Clarke C, Haggerty L, Hemrom A, Hourlier T, Izuogu OG, Juettemann T, Kaikala V, Kay M, Lavidas I, Le T, Lemos D, Gonzalez Martinez J, Marugán JC, Maurel T, McMahon AC, Mohanan S, Moore B, Muffato M, Oheh DN, Paraschas D, Parker A, Parton A, Prosovetskaia I, Sakthivel MP, Salam AIA, Schmitt BM, Schuilenburg H, Sheppard D, Steed E, Szpak M, Szuba M, Taylor K, Thormann A, Threadgold G, Walts B, Winterbottom A, Chakiachvili M, Chaubal A, De Silva N, Flint B, Frankish A, Hunt SE, IIsley GR, Langridge N, Loveland JE, Martin FJ, Mudge JM, Morales J, Perry E, Ruffier M, Tate J, Thybert D, Trevanion SJ, Cunningham F, Yates AD, Zerbino DR, Flicek P. 2021. Ensembl 2021. Nucleic Acids Res 49:D884–D891.
- Hu M, Bogoyevitch MA, Jans DA. 2019. Subversion of Host Cell Mitochondria by RSV to Favor Virus Production is Dependent on Inhibition of Mitochondrial Complex I and ROS Generation. Cells 8. [CrossRef]
- Jäger S, Cimermancic P, Gulbahce N, Johnson JR, McGovern KE, Clarke SC, Shales M, Mercenne G, Pache L, Li K, Hernandez H, Jang GM, Roth SL, Akiva E, Marlett J, Stephens M, D’Orso I, Fernandes J, Fahey M, Mahon C, O’Donoghue AJ, Todorovic A, Morris JH, Maltby DA, Alber T, Cagney G, Bushman FD, Young JA, Chanda SK, Sundquist WI, Kortemme T, Hernandez RD, Craik CS, Burlingame A, Sali A, Frankel AD, Krogan NJ. 2011. Global landscape of HIV-human protein complexes. Nature 481:365–370.
- Joshi CJ, Schinn S-M, Richelle A, Shamie I, O’Rourke EJ, Lewis NE. 2020. StanDep: Capturing transcriptomic variability improves context-specific metabolic models. PLoS Comput Biol 16:e1007764.
- Kandeel M, Abdelrahman AHM, Oh-Hashi K, Ibrahim A, Venugopala KN, Morsy MA, Ibrahim MAA. 2021. Repurposing of FDA-approved antivirals, antibiotics, anthelmintics, antioxidants, and cell protectives against SARS-CoV-2 papain-like protease. J Biomol Struct Dyn 39:5129–5136.
- Keating SM, Waltemath D, König M, Zhang F, Dräger A, Chaouiya C, Bergmann FT, Finney A, Gillespie CS, Helikar T, Hoops S, Malik-Sheriff RS, Moodie SL, Moraru II, Myers CJ, Naldi A, Olivier BG, Sahle S, Schaff JC, Smith LP, Swat MJ, Thieffry D, Watanabe L, Wilkinson DJ, Blinov ML, Begley K, Faeder JR, Gómez HF, Hamm TM, Inagaki Y, Liebermeister W, Lister AL, Lucio D, Mjolsness E, Proctor CJ, Raman K, Rodriguez N, Shaffer CA, Shapiro BE, Stelling J, Swainston N, Tanimura N, Wagner J, Meier-Schellersheim M, Sauro HM, Palsson B, Bolouri H, Kitano H, Funahashi A, Hermjakob H, Doyle JC, Hucka M, SBML Level 3 Community members. 2020. SBML Level 3: an extensible format for the exchange and reuse of biological models. Mol Syst Biol 16:e9110.
- Kelly B, Tannahill GM, Murphy MP, O’Neill LAJ. 2015. Metformin Inhibits the Production of Reactive Oxygen Species from NADH:Ubiquinone Oxidoreductase to Limit Induction of Interleukin-1β (IL-1β) and Boosts Interleukin-10 (IL-10) in Lipopolysaccharide (LPS)-activated Macrophages. J Biol Chem 290:20348–20359.
- Lee S, Yu Y, Trimpert J, Benthani F, Mairhofer M, Richter-Pechanska P, Wyler E, Belenki D, Kaltenbrunner S, Pammer M, Kausche L, Firsching TC, Dietert K, Schotsaert M, Martínez-Romero C, Singh G, Kunz S, Niemeyer D, Ghanem R, Salzer HJF, Paar C, Mülleder M, Uccellini M, Michaelis EG, Khan A, Lau A, Schönlein M, Habringer A, Tomasits J, Adler JM, Kimeswenger S, Gruber AD, Hoetzenecker W, Steinkellner H, Purfürst B, Motz R, Di Pierro F, Lamprecht B, Osterrieder N, Landthaler M, Drosten C, García-Sastre A, Langer R, Ralser M, Eils R, Reimann M, Fan DNY, Schmitt CA. 2021. Virus-induced senescence is a driver and therapeutic target in COVID-19. Nature 599:283–289.
- Lehrer S. 2020. Inhaled biguanides and mTOR inhibition for influenza and coronavirus (Review). World Acad Sci J 2. [CrossRef]
- Liao M, Liu Y, Yuan J, Wen Y, Xu G, Zhao J, Cheng L, Li J, Wang X, Wang F, Liu L, Amit I, Zhang S, Zhang Z. 2020. Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19. Nat Med 26:842–844.
- Licata L, Briganti L, Peluso D, Perfetto L, Iannuccelli M, Galeota E, Sacco F, Palma A, Nardozza AP, Santonico E, Castagnoli L, Cesareni G. 2012. MINT, the molecular interaction database: 2012 update. Nucleic Acids Res 40:D857–61.
- Li W, O’Neill KR, Haft DH, DiCuccio M, Chetvernin V, Badretdin A, Coulouris G, Chitsaz F, Derbyshire MK, Durkin AS, Gonzales NR, Gwadz M, Lanczycki CJ, Song JS, Thanki N, Wang J, Yamashita RA, Yang M, Zheng C, Marchler-Bauer A, Thibaud-Nissen F. 2021. RefSeq: expanding the Prokaryotic Genome Annotation Pipeline reach with protein family model curation. Nucleic Acids Res 49:D1020–D1028.
- Li Y, Liu D, Wang Y, Su W, Liu G, Dong W. 2021. The Importance of Glycans of Viral and Host Proteins in Enveloped Virus Infection. Front Immunol 12:638573.
- Magalhaes I, Yogev O, Mattsson J, Schurich A. 2019. The Metabolic Profile of Tumor and Virally Infected Cells Shapes Their Microenvironment Counteracting T Cell Immunity. Front Immunol 10:2309.
- Mahmoudabadi G, Milo R, Phillips R. 2017. Energetic cost of building a virus. Proc Natl Acad Sci U S A 114:E4324–E4333.
- Malik-Sheriff RS, Glont M, Nguyen TVN, Tiwari K, Roberts MG, Xavier A, Vu MT, Men J, Maire M, Kananathan S, Fairbanks EL, Meyer JP, Arankalle C, Varusai TM, Knight-Schrijver V, Li L, Dueñas-Roca C, Dass G, Keating SM, Park YM, Buso N, Rodriguez N, Hucka M, Hermjakob H. 2020. BioModels-15 years of sharing computational models in life science. Nucleic Acids Res 48:D407–D415.
- Marani M, Katul GG, Pan WK, Parolari AJ. 2021. Intensity and frequency of extreme novel epidemics. Proc Natl Acad Sci U S A 118. https://doi.org/10.1073/pnas.2105482118. [CrossRef]
- Martinez-Gil L, Vera-Velasco NM, Mingarro I. 2017. Exploring the Human-Nipah Virus Protein-Protein Interactome. J Virol 91. [CrossRef]
- Marty AM, Jones MK. 2020. The novel Coronavirus (SARS-CoV-2) is a one health issue. One Health. [CrossRef]
- Mayer KA, Stöckl J, Zlabinger GJ, Gualdoni GA. 2019. Hijacking the Supplies: Metabolism as a Novel Facet of Virus-Host Interaction. Front Immunol 10:1533.
- Medaglia C, Kolpakov I, Zhu Y, Constant S, Huang S, Zwygart AC-A, Cagno V, Dermitzakis ET, Stellacci F, Xenarios I, Tapparel C. n.d. A novel anti-influenza combined therapy assessed by single cell RNA-sequencing. [CrossRef]
- Michon C, Kang C-M, Karpenko S, Tanaka K, Ishikawa S, Yoshida K-I. 2020. A bacterial cell factory converting glucose into scyllo-inositol, a therapeutic agent for Alzheimer’s disease. Commun Biol 3:93.
- Miyadera H, Shiomi K, Ui H, Yamaguchi Y, Masuma R, Tomoda H, Miyoshi H, Osanai A, Kita K, Omura S. 2003. Atpenins, potent and specific inhibitors of mitochondrial complex II (succinate-ubiquinone oxidoreductase). Proc Natl Acad Sci U S A 100:473–477.
- Mora C, McKenzie T, Gaw IM, Dean JM, von Hammerstein H, Knudson TA, Setter RO, Smith CZ, Webster KM, Patz JA, Franklin EC. 2022. Over half of known human pathogenic diseases can be aggravated by climate change. Nat Clim Chang 1–7.
- Moreno-Altamirano MMB, Kolstoe SE, Sánchez-García FJ. 2019. Virus Control of Cell Metabolism for Replication and Evasion of Host Immune Responses. Front Cell Infect Microbiol 9:95.
- Morris ME, Felmlee MA. 2008. Overview of the proton-coupled MCT (SLC16A) family of transporters: characterization, function and role in the transport of the drug of abuse gamma-hydroxybutyric acid. AAPS J 10:311–321.
- Muller M, Jacob Y, Jones L, Weiss A, Brino L, Chantier T, Lotteau V, Favre M, Demeret C. 2012. Large scale genotype comparison of human papillomavirus E2-host interaction networks provides new insights for e2 molecular functions. PLoS Pathog 8:e1002761.
- Nattrass M, Sizer K, Alberti KG. 1980. Correlation of plasma phenformin concentration with metabolic effects in normal subjects. Clin Sci 58:153–155.
- Neal ML, König M, Nickerson D, Mısırlı G, Kalbasi R, Dräger A, Atalag K, Chelliah V, Cooling MT, Cook DL, Crook S, de Alba M, Friedman SH, Garny A, Gennari JH, Gleeson P, Golebiewski M, Hucka M, Juty N, Myers C, Olivier BG, Sauro HM, Scharm M, Snoep JL, Touré V, Wipat A, Wolkenhauer O, Waltemath D. 2019. Harmonizing semantic annotations for computational models in biology. Brief Bioinform 20:540–550.
- Noronha A, Daníelsdóttir AD, Gawron P, Jóhannsson F, Jónsdóttir S, Jarlsson S, Gunnarsson JP, Brynjólfsson S, Schneider R, Thiele I, Fleming RMT. 2017. ReconMap: an interactive visualization of human metabolism. Bioinformatics 33:605–607.
- Orth JD, Thiele I, Palsson BØ. 2010. What is flux balance analysis? Nat Biotechnol 28:245–248.
- Oughtred R, Stark C, Breitkreutz B-J, Rust J, Boucher L, Chang C, Kolas N, O’Donnell L, Leung G, McAdam R, Zhang F, Dolma S, Willems A, Coulombe-Huntington J, Chatr-Aryamontri A, Dolinski K, Tyers M. 2019. The BioGRID interaction database: 2019 update. Nucleic Acids Res 47:D529–D541.
- Ravindra NG, Alfajaro MM, Gasque V, Huston NC, Wan H, Szigeti-Buck K, Yasumoto Y, Greaney AM, Habet V, Chow RD, Chen JS, Wei J, Filler RB, Wang B, Wang G, Niklason LE, Montgomery RR, Eisenbarth SC, Chen S, Williams A, Iwasaki A, Horvath TL, Foxman EF, Pierce RW, Pyle AM, van Dijk D, Wilen CB. 2021. Single-cell longitudinal analysis of SARS-CoV-2 infection in human airway epithelium identifies target cells, alterations in gene expression, and cell state changes. PLoS Biol 19:e3001143.
- Renz A, Widerspick L, Dräger A. 2020. FBA reveals guanylate kinase as a potential target for antiviral therapies against SARS-CoV-2. Bioinformatics 36:i813–i821.
- Reuther P, Göpfert K, Dudek AH, Heiner M, Herold S, Schwemmle M. 2015. Generation of a variety of stable Influenza A reporter viruses by genetic engineering of the NS gene segment. Sci Rep 5:11346.
- Richelle A, Chiang AWT, Kuo C-C, Lewis NE. 2019. Increasing consensus of context-specific metabolic models by integrating data-inferred cell functions. PLoS Comput Biol 15:e1006867.
- Rozenblatt-Rosen O, Deo RC, Padi M, Adelmant G, Calderwood MA, Rolland T, Grace M, Dricot A, Askenazi M, Tavares M, Pevzner SJ, Abderazzaq F, Byrdsong D, Carvunis A-R, Chen AA, Cheng J, Correll M, Duarte M, Fan C, Feltkamp MC, Ficarro SB, Franchi R, Garg BK, Gulbahce N, Hao T, Holthaus AM, James R, Korkhin A, Litovchick L, Mar JC, Pak TR, Rabello S, Rubio R, Shen Y, Singh S, Spangle JM, Tasan M, Wanamaker S, Webber JT, Roecklein-Canfield J, Johannsen E, Barabási A-L, Beroukhim R, Kieff E, Cusick ME, Hill DE, Münger K, Marto JA, Quackenbush J, Roth FP, DeCaprio JA, Vidal M. 2012. Interpreting cancer genomes using systematic host network perturbations by tumour virus proteins. Nature 487:491–495.
- Sanchez EL, Lagunoff M. 2015. Viral activation of cellular metabolism. Virology 479-480:609–618.
- Shah PS, Link N, Jang GM, Sharp PP, Zhu T, Swaney DL, Johnson JR, Von Dollen J, Ramage HR, Satkamp L, Newton B, Hüttenhain R, Petit MJ, Baum T, Everitt A, Laufman O, Tassetto M, Shales M, Stevenson E, Iglesias GN, Shokat L, Tripathi S, Balasubramaniam V, Webb LG, Aguirre S, Willsey AJ, Garcia-Sastre A, Pollard KS, Cherry S, Gamarnik AV, Marazzi I, Taunton J, Fernandez-Sesma A, Bellen HJ, Andino R, Krogan NJ. 2018. Comparative Flavivirus-Host Protein Interaction Mapping Reveals Mechanisms of Dengue and Zika Virus Pathogenesis. Cell 175:1931–1945.e18.
- Shutters ST. 2021. Modelling long-term COVID-19 impacts on the U.S. workforce of 2029. PLoS One 16:e0260797.
- Sia SF, Yan L-M, Chin AWH, Fung K, Choy K-T, Wong AYL, Kaewpreedee P, Perera RAPM, Poon LLM, Nicholls JM, Peiris M, Yen H-L. 2020. Pathogenesis and transmission of SARS-CoV-2 in golden hamsters. Nature 583:834–838.
- Smith LP, Hucka M, Hoops S, Finney A, Ginkel M, Myers CJ, Moraru I, Liebermeister W. 2015. SBML Level 3 package: Hierarchical Model Composition, Version 1 Release 3. J Integr Bioinform 12:268.
- Sogame Y, Kitamura A, Yabuki M, Komuro S. 2011. Liver uptake of biguanides in rats. Biomed Pharmacother 65:451–455.
- Sogame Y, Kitamura A, Yabuki M, Komuro S. 2009. A comparison of uptake of metformin and phenformin mediated by hOCT1 in human hepatocytes. Biopharm Drug Dispos 30:476–484.
- Stang M, Wysowski DK, Butler-Jones D. 1999. Incidence of lactic acidosis in metformin users. Diabetes Care 22:925–927.
- Stelzer G, Rosen N, Plaschkes I, Zimmerman S, Twik M, Fishilevich S, Stein TI, Nudel R, Lieder I, Mazor Y, Kaplan S, Dahary D, Warshawsky D, Guan-Golan Y, Kohn A, Rappaport N, Safran M, Lancet D. 2016. The GeneCards Suite: From Gene Data Mining to Disease Genome Sequence Analyses. Curr Protoc Bioinformatics 54:1.30.1–1.30.33.
- Sterne J. 1963. [Report on 5-years’ experience with dimethylbiguanide (metformin, glucophage) in diabetic therapy]. Wien Med Wochenschr 113:599–602.
- Sumbria D, Berber E, Mathayan M, Rouse BT. 2020. Virus Infections and Host Metabolism-Can We Manage the Interactions? Front Immunol 11:594963.
- Swainston N, Smallbone K, Hefzi H, Dobson PD, Brewer J, Hanscho M, Zielinski DC, Ang KS, Gardiner NJ, Gutierrez JM, Kyriakopoulos S, Lakshmanan M, Li S, Liu JK, Martínez VS, Orellana CA, Quek L-E, Thomas A, Zanghellini J, Borth N, Lee D-Y, Nielsen LK, Kell DB, Lewis NE, Mendes P. 2016. Recon 2.2: from reconstruction to model of human metabolism. Metabolomics 12:109.
- Teo G, Liu G, Zhang J, Nesvizhskii AI, Gingras A-C, Choi H. 2014. SAINTexpress: improvements and additional features in Significance Analysis of INTeractome software. J Proteomics 100:37–43.
- Thaker SK, Ch’ng J, Christofk HR. 2019. Viral hijacking of cellular metabolism. BMC Biol 17:59.
- Touré V, Dräger A, Luna A, Dogrusoz U, Rougny A. 2021. The Systems Biology Graphical Notation: Current Status and Applications in Systems Medicine. Systems Medicine. [CrossRef]
- Tripathi LP, Kataoka C, Taguwa S, Moriishi K, Mori Y, Matsuura Y, Mizuguchi K. 2010. Network based analysis of hepatitis C virus core and NS4B protein interactions. Mol Biosyst 6:2539–2553.
- Venables WN, Ripley BD. 2012. Modern Applied Statistics with S. Springer.
- Villenave R, Broadbent L, Douglas I, Lyons JD, Coyle PV, Teng MN, Tripp RA, Heaney LG, Shields MD, Power UF. 2015. Induction and Antagonism of Antiviral Responses in Respiratory Syncytial Virus-Infected Pediatric Airway Epithelium. J Virol 89:12309–12318.
- Vlassis N, Pacheco MP, Sauter T. 2014. Fast reconstruction of compact context-specific metabolic network models. PLoS Comput Biol 10:e1003424.
- Wang H, Huwaimel B, Verma K, Miller J, Germain TM, Kinarivala N, Pappas D, Brookes PS, Trippier PC. 2017. Synthesis and Antineoplastic Evaluation of Mitochondrial Complex II (Succinate Dehydrogenase) Inhibitors Derived from Atpenin A5. ChemMedChem 12:1033–1044.
- Wang R, Simoneau CR, Kulsuptrakul J, Bouhaddou M, Travisano KA, Hayashi JM, Carlson-Stevermer J, Zengel JR, Richards CM, Fozouni P, Oki J, Rodriguez L, Joehnk B, Walcott K, Holden K, Sil A, Carette JE, Krogan NJ, Ott M, Puschnik AS. 2021. Genetic Screens Identify Host Factors for SARS-CoV-2 and Common Cold Coronaviruses. Cell 184:106–119.e14.
- Watanabe T, Kawakami E, Shoemaker JE, Lopes TJS, Matsuoka Y, Tomita Y, Kozuka-Hata H, Gorai T, Kuwahara T, Takeda E, Nagata A, Takano R, Kiso M, Yamashita M, Sakai-Tagawa Y, Katsura H, Nonaka N, Fujii H, Fujii K, Sugita Y, Noda T, Goto H, Fukuyama S, Watanabe S, Neumann G, Oyama M, Kitano H, Kawaoka Y. 2014. Influenza virus-host interactome screen as a platform for antiviral drug development. Cell Host Microbe 16:795–805.
- Wei J, Alfajaro MM, DeWeirdt PC, Hanna RE, Lu-Culligan WJ, Cai WL, Strine MS, Zhang S-M, Graziano VR, Schmitz CO, Chen JS, Mankowski MC, Filler RB, Ravindra NG, Gasque V, de Miguel FJ, Patil A, Chen H, Oguntuyo KY, Abriola L, Surovtseva YV, Orchard RC, Lee B, Lindenbach BD, Politi K, van Dijk D, Kadoch C, Simon MD, Yan Q, Doench JG, Wilen CB. 2021. Genome-wide CRISPR Screens Reveal Host Factors Critical for SARS-CoV-2 Infection. Cell 184:76–91.e13.
- Wishart DS, Feunang YD, Guo AC, Lo EJ, Marcu A, Grant JR, Sajed T, Johnson D, Li C, Sayeeda Z, Assempour N, Iynkkaran I, Liu Y, Maciejewski A, Gale N, Wilson A, Chin L, Cummings R, Le D, Pon A, Knox C, Wilson M. 2018. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res 46:D1074–D1082.
- Wojtovich AP, Brookes PS. 2009. The complex II inhibitor atpenin A5 protects against cardiac ischemia-reperfusion injury via activation of mitochondrial KATP channels. Basic Res Cardiol 104:121–129.
- Wyler E, Mösbauer K, Franke V, Diag A, Gottula LT, Arsiè R, Klironomos F, Koppstein D, Hönzke K, Ayoub S, Buccitelli C, Hoffmann K, Richter A, Legnini I, Ivanov A, Mari T, Del Giudice S, Papies J, Praktiknjo S, Meyer TF, Müller MA, Niemeyer D, Hocke A, Selbach M, Akalin A, Rajewsky N, Drosten C, Landthaler M. 2021. Transcriptomic profiling of SARS-CoV-2 infected human cell lines identifies HSP90 as target for COVID-19 therapy. iScience 24:102151.
- Xie X, Muruato A, Lokugamage KG, Narayanan K, Zhang X, Zou J, Liu J, Schindewolf C, Bopp NE, Aguilar PV, Plante KS, Weaver SC, Makino S, LeDuc JW, Menachery VD, Shi P-Y. 2020. An Infectious cDNA Clone of SARS-CoV-2. Cell Host Microbe 27:841–848.e3.
- Yang W, Petkova E, Shaman J. 2014. The 1918 influenza pandemic in New York City: age-specific timing, mortality, and transmission dynamics. Influenza Other Respi Viruses 8:177–188.
- Yim KHW, Borgoni S, Chahwan R. 2022. Serum extracellular vesicles profiling is associated with COVID-19 progression and immune responses. J Extracell Biol 1:e37.
- Zanini F, Robinson ML, Croote D, Sahoo MK, Sanz AM, Ortiz-Lasso E, Albornoz LL, Rosso F, Montoya JG, Goo L, Pinsky BA, Quake SR, Einav S. 2018. Virus-inclusive single-cell RNA sequencing reveals the molecular signature of progression to severe dengue. Proc Natl Acad Sci U S A 115:E12363–E12369.
- Zeileis A, Kleiber C, Jackman S. 2008. Regression Models for Count Data inR. Journal of Statistical Software. [CrossRef]
- Zhao T, Mu X, You Q. 2017. Succinate: An initiator in tumorigenesis and progression. Oncotarget 8:53819–53828.
- Zhu S-L, Wang L, Cao Z-Y, Wang J, Jing M-Z, Xia Z-C, Ao F, Ye L-B, Liu S, Zhu Y. 2016. Inducible CYP4F12 enhances Hepatitis C virus infection via association with viral nonstructural protein 5B. Biochem Biophys Res Commun 471:95–102.
- Zitzmann C, Kaderali L. 2018. Mathematical Analysis of Viral Replication Dynamics and Antiviral Treatment Strategies: From Basic Models to Age-Based Multi-Scale Modeling. Front Microbiol 9:1546.
| Atpenin A5 | IC50 | CC50 | selectivity index: CC50/IC50 |
|---|---|---|---|
|
ic-SARS-CoV-2-mNG (in Calu-3 cells) |
0.45 µM | MTT: 3.6 µM CTG: 1.0 µM IncuCyte: > 160 µM |
MTT: 8 CTG: 2.2 IncuCyte: > 355 |
|
SARS-CoV-2 Omicron (in Calu-3 cells) |
0.68 µM | MTT: 3.6 µM CTG: 1.0 µM IncuCyte: > 160 µM |
MTT: 5.3 CTG: 1.5 IncuCyte: > 235 |
|
Dengue Virus (in Huh7.5 cells) |
0.36 µM | MTT: 83.3 µM CTG: 85.4 µM IncuCyte: 115 µM |
MTT: 231.4 CTG: 258.8 IncuCyte: 319.4 |
|
Influenza A Virus (in A549 cells) |
0.14 µM (n = 2) |
MTT: 114.7 µM CTG: 49.5 µM Incucyte: 67 µM |
MTT: 819.3 CTG: 353.6 Incucyte: 478.6 |
|
RSV (in A549 cells) |
0.12 µM | MTT: 114.7 µM CTG: 49.5 µM IncuCyte: 67 µM |
MTT: 955.8 CTG: 412.5 Incucyte: 558.3 |
| Phenformin | IC50 | CC50 | selectivity index: CC50/IC50 |
|
ic-SARS-CoV-2-mNG (in Calu-3 cells) |
1.46 µM | MTT: 456 µM CTG: 26.3 µM IncuCyte: 2060 µM |
MTT: 312.3 CTG: 18.0 IncuCyte: 1411 |
|
SARS-CoV-2 Omicron (in Calu-3 cells) |
1.35 µM | MTT: 456 µM CTG: 26.3 µM IncuCyte: 2060 µM |
MTT: 337.8 CTG: 19.5 IncuCyte: 1525.9 |
|
Dengue Virus (in Huh7.5 cells) |
17.15 µM | MTT: 64.2 µM CTG: 80.2 µM IncuCyte: 1313 µM |
MTT: 3.7 CTG: 4.7 IncuCyte: 76.6 |
|
Influenza A Virus (in A549 cells) |
not active | ||
|
RSV (in A459 cells) |
not active |
| Dataset | Reference | Accession Number | Celltype annotation source |
|---|---|---|---|
| BALF1 | (Liao et al., 2020) | GSE145926 | Annotation via CHETAH version 1.8.0 with single cell lung atlas (GSE136831,(Adams et al., 2020)) |
| BALF2 | (Chua et al., 2020) | EGAS00001004481 | Original authors (https://doi.org/10.6084/m9.figshare.12436517) |
| CALU-3 | (Wyler et al., 2021) | GSE148729 | Single cell experiment |
| scH1299 | (Wyler et al., 2021) | GSE148729 | Single cell experiment |
| Influenza H1N1 | (Medaglia et al., n.d.) | GSE191176 | Manual annotation following methods of original publication |
| Dengue | (Zanini et al., 2018) | GSE116672 | Annotation via CHETAH version 1.8.0 with PBMC single cell reference 62 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).