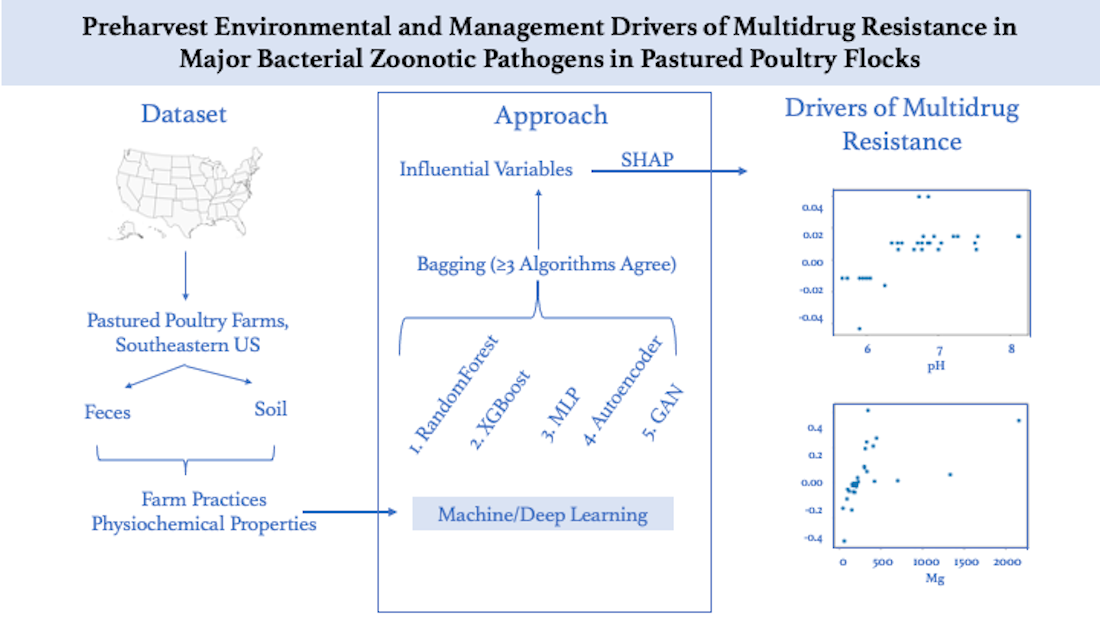
Due to nutritional benefits and perceived humane ways of treating the animals, the demand for antibiotic-free pastured poultry chicken has continued to be on a steady rise. However, despite non-usage of antibiotics in pastured poultry broiler production, antibiotic resistance (AR) is reported in zoonotic poultry pathogens. However, actors that drive multidrug resistance (MDR) in pastured poultry are not known. In this study, we used machine learning and deep learning approaches to predict farm management practices, and physicochemical properties of feces and soil that drive MDR in zoonotic poultry pathogens. Antibiotic use in agroecosystems is known to contribute to resistance. Evaluation of the development of resistance in environments that are free of antibiotics such as the all-natural antibiotic-free, pastured poultry production systems described here is critical to understand the background AR. We analyzed 1,635 preharvest (feces and soil) samples collected from forty-two pastured poultry flocks and eleven farms in the Southeastern United States. CDC National Antimicrobial Resistance Monitoring System guidelines were used to determine antimicrobial/multidrug resistance profiles of Salmonella, Listeria and Campylobacter. A combination of two traditional machine learning (RandomForest and XGBoost) and three deep learning (Multi-layer Perceptron, Generative Adversarial Network, and Auto-Encoder) approaches, identified critical farm/environmental variables that drive multidrug resistance in poultry pathogens, in broiler production systems that represents background resistance. This study enumerates management practices that contribute to AR and recommendations to potentially mitigate multidrug resistance and prevalence of Salmonella and Listeria in pastured poultry.