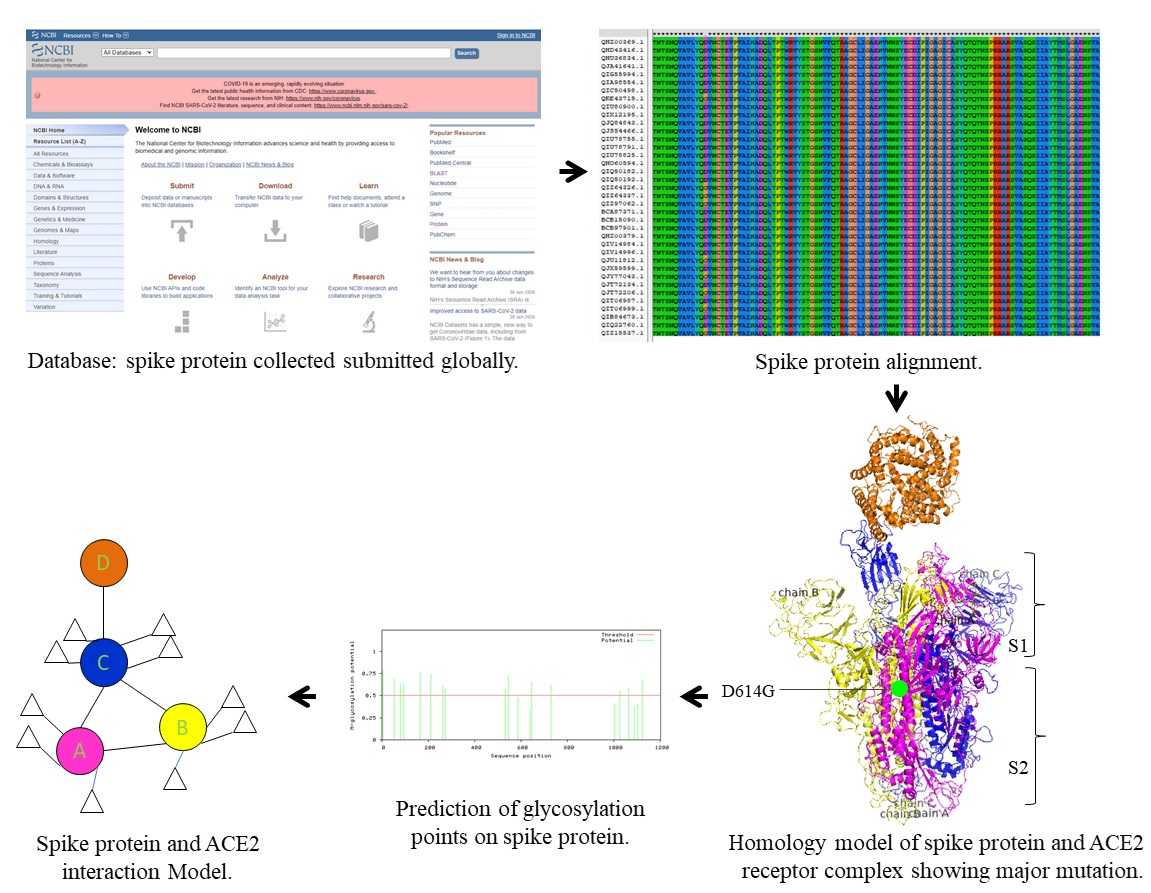
Analysis of SARS-CoV-2 spike protein sequences of over 19 countries from biological databases submitted around the globe was carried out with help of bioinformatics tools and structure prediction databases. Initial data analysis showed entry of virus into different geographic regions started in the month of January 2020. Meanwhile, alignment of spike protein sequences of SARS-CoV-2 isolates from China and other countries revealed a critical mutation of D614G. Surprisingly, mutation D614G was not seen in early samples submitted in the month of January but gradually it started appearing globally from the month of March 2020. However, the mutations of amino acids in the spike protein other than D614G exhibiting similar pI and altered polarity were found to be specific to geographical regions. Besides, prediction of homology model for interaction of spike protein showed predominant role of chain C of trimeric spike protein in adhering receptor binding domain (RBD) of human ACE2 receptor. Furthermore, the prediction of glycosylation points has revealed that there are about 20 N-glycosylation potential sites on spike protein. We believe that the information present here would not only help in thorough understanding of infectivity but also enhance the knowledge of the scientific community in developing prophylactics and/or therapeutics for SARS-CoV19-2 virus.