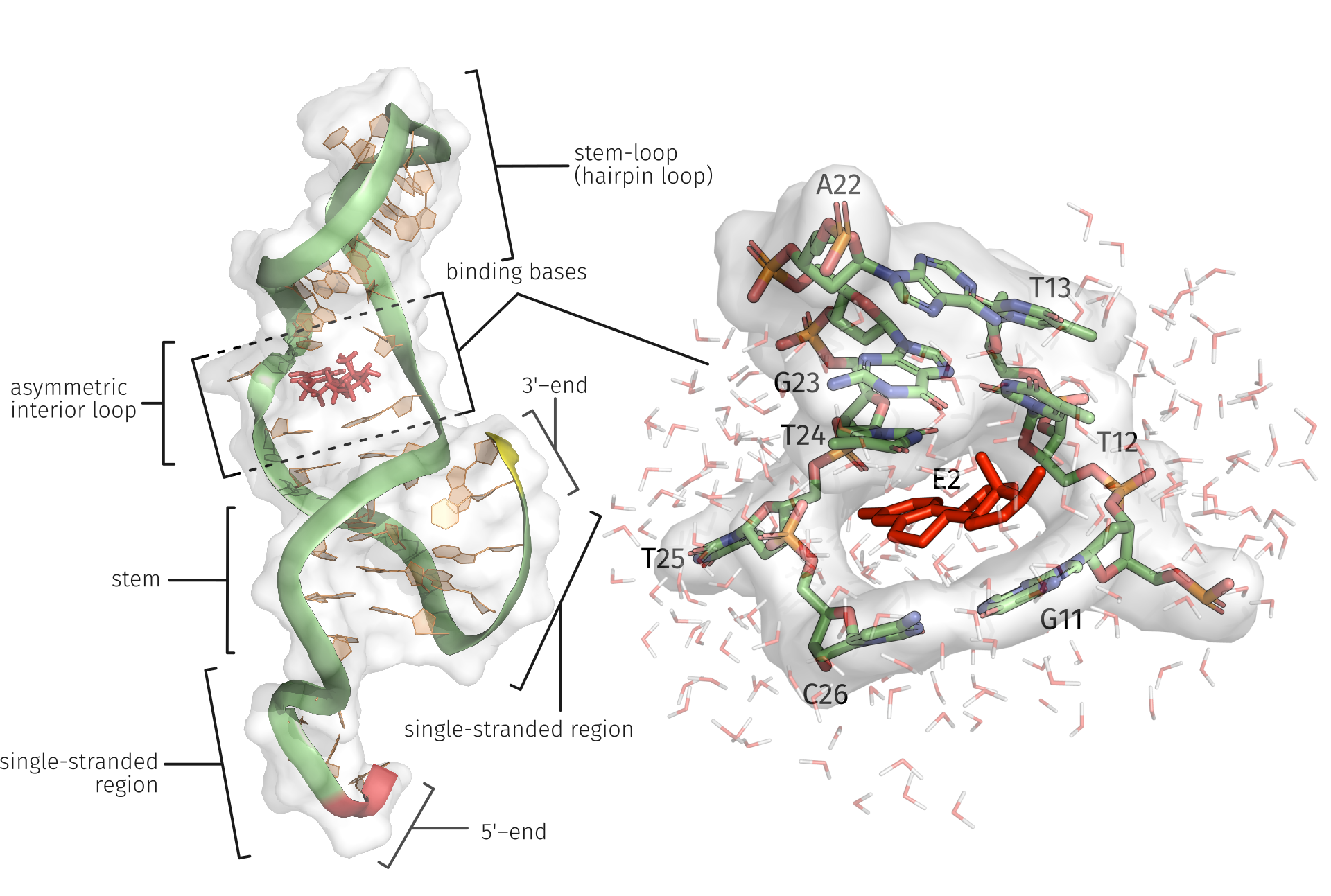
Micro pollutants such as 17β-Estradiol (E2) have been detected in low concentrations in different water resources and their negative effects on the environment and organisms are observed. In this study, a previously described 35-mer E2-specific aptamer was used to investigate the underlying binding characteristics between E2 and the aptamer through an MD simulation in an aqueous medium. There is no 3D structure information for this aptamer, so it was modeled using coarse-grained modeling method. The E2 ligand was positioned inside the potential binding area of the aptamer structure, the underlying complex was used for an 25 ns MD simulation, and the interactions were examined by an interaction profiler tool for each time step. The E2-specific bases of the aptamer had presumably dominant roles in the binding of E2 in terms of the different interaction types. We identified the specific bases within the interior loop of the aptamer and we also demonstrate the influence of oftentimes underestimated water-mediated hydrogen bonds. The study contributes to the understanding of the behavior of ligands binding through aptamer structure in an aqueous solution. The developed workflow allows to generate and examine further appealing ligand aptamer complexes.