Submitted:
30 April 2026
Posted:
05 May 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Related Work
2.1. Quantization in Medical Image Segmentation
2.2. Anatomical Constraints in Segmentation
2.3. Topology-Aware Deep Learning
2.4. Diversity in Training Data Using Generative AI
3. Background and Preliminaries
3.1. nnUNet Architecture and Dental Adaptation
3.2. Quantization in Deep Neural Networks
3.3. Topological Considerations in Dental Segmentation
- Incorrect tooth counts (missing or extra segments)
- Improper connections between adjacent teeth
- Spurious holes within tooth structures
4. Proposed Method: Topology-Preserving Quantization for Dental nnUNet
4.1. Tooth-Specific Topological Constraint Loss Formulation
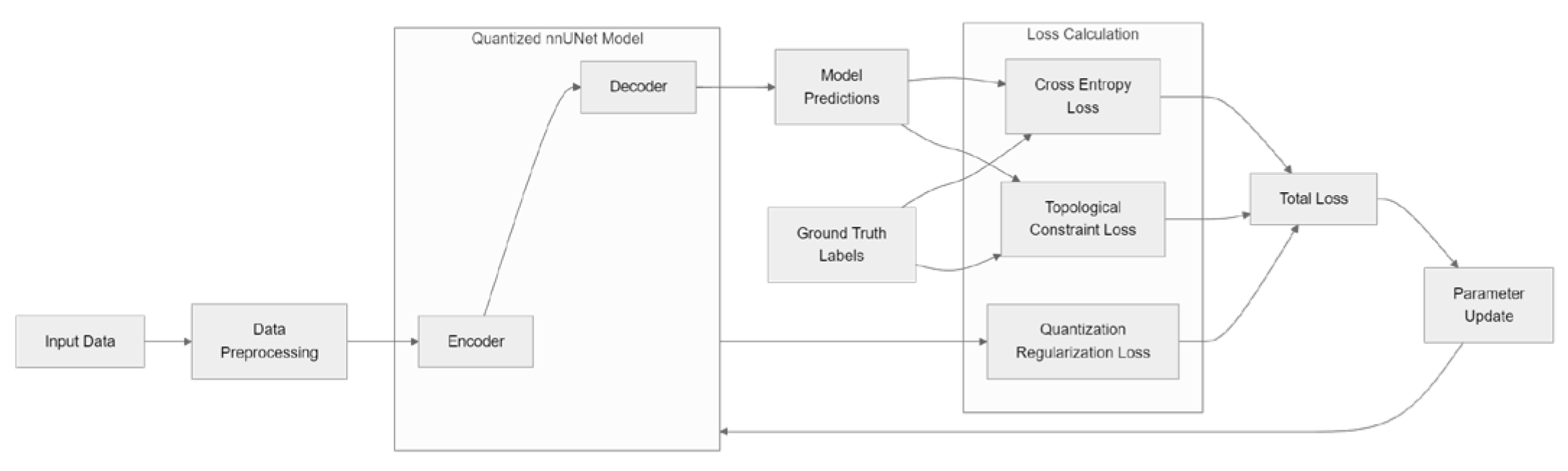
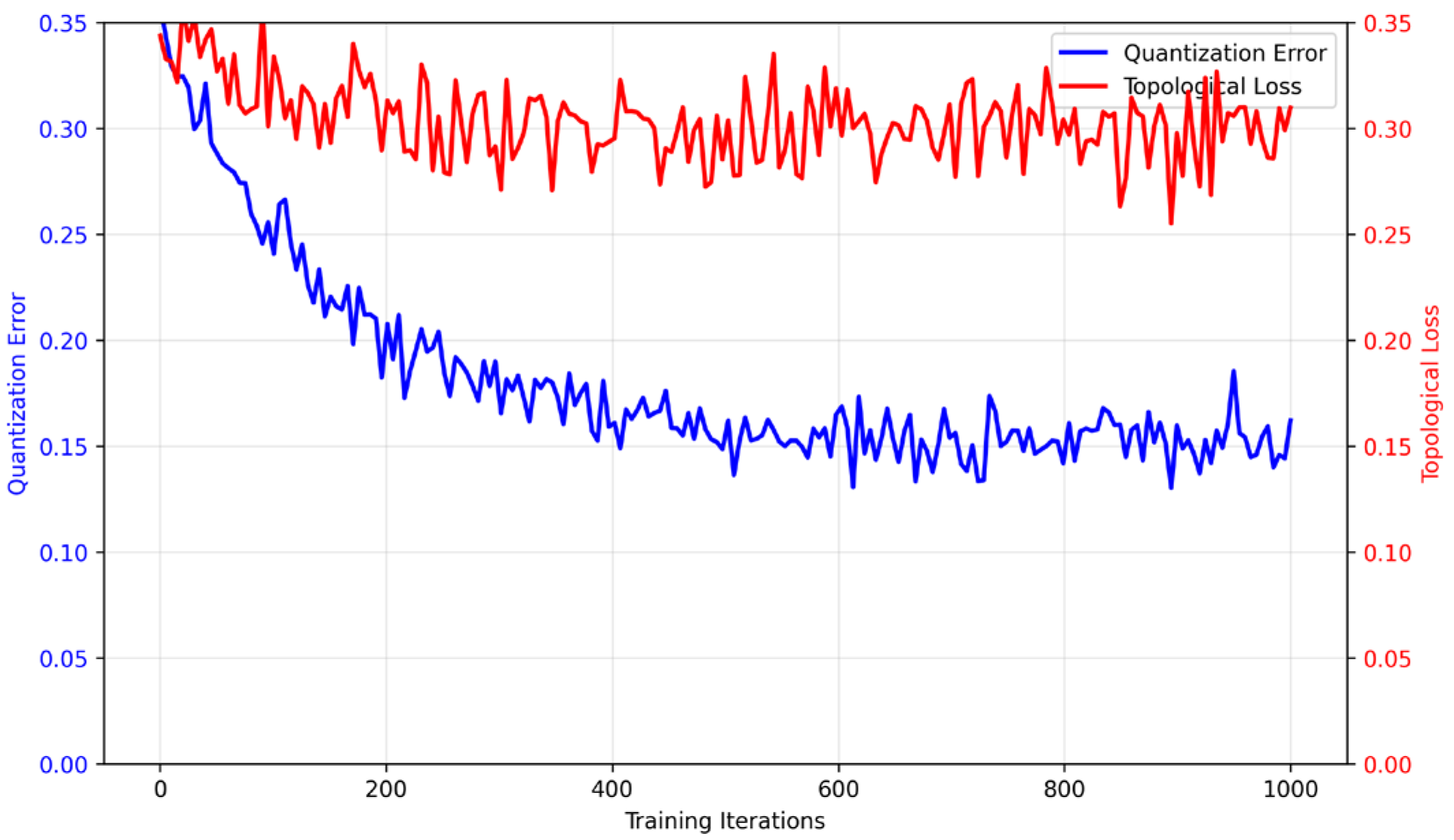
4.2. Integration of Topological Loss with Quantization-Aware Training
4.3. Differentiable Persistent Homology for Dental Data
4.4. Runtime-Efficient Inference through Implicit Encoding of Topological Constraints
5. Experimental Setup
5.1. Dataset and Preprocessing
5.2. Implementation Details
5.3. Evaluation Metrics
-
Segmentation Accuracy:
- ∘
- Dice Similarity Coefficient (DSC):
- ∘
- Intersection over Union (IoU):
- ∘
- Boundary F1 Score (BF1): Harmonic mean of precision and recall for boundary voxels
-
Topological Fidelity:
- ∘
- Tooth Count Accuracy (TCA): Percentage of scans with correct tooth instances
- ∘
- Adjacency Consistency Score (ACS): , where denotes symmetric difference
- ∘
- Cavity Error Rate (CER):
-
Computational Efficiency:
- ∘
- Model Size (MB)
- ∘
- Inference Time per Volume (seconds)
- ∘
- Multiply-Accumulate Operations (MACs)
5.4. Baseline Methods
- Full-Precision nnUNet[1]: The original floating-point implementation serving as the accuracy upper bound.
- Post-Training Quantized nnUNet[2]: Standard 8-bit quantization applied after training without fine-tuning.
- QAT-nnUNet[2]: Quantization-aware trained version without topological constraints.
- TopoNet[11]: A topology-preserving segmentation model adapted for dental data.
6. Results and Analysis
6.1. Quantitative Comparison with Baseline Methods
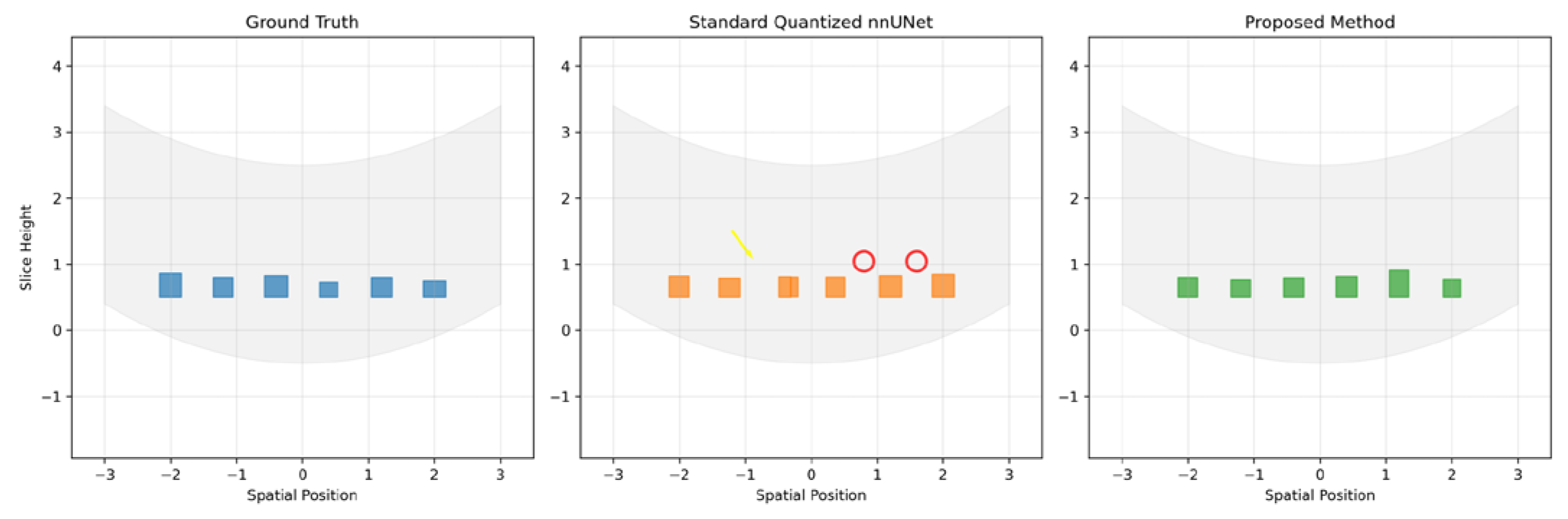
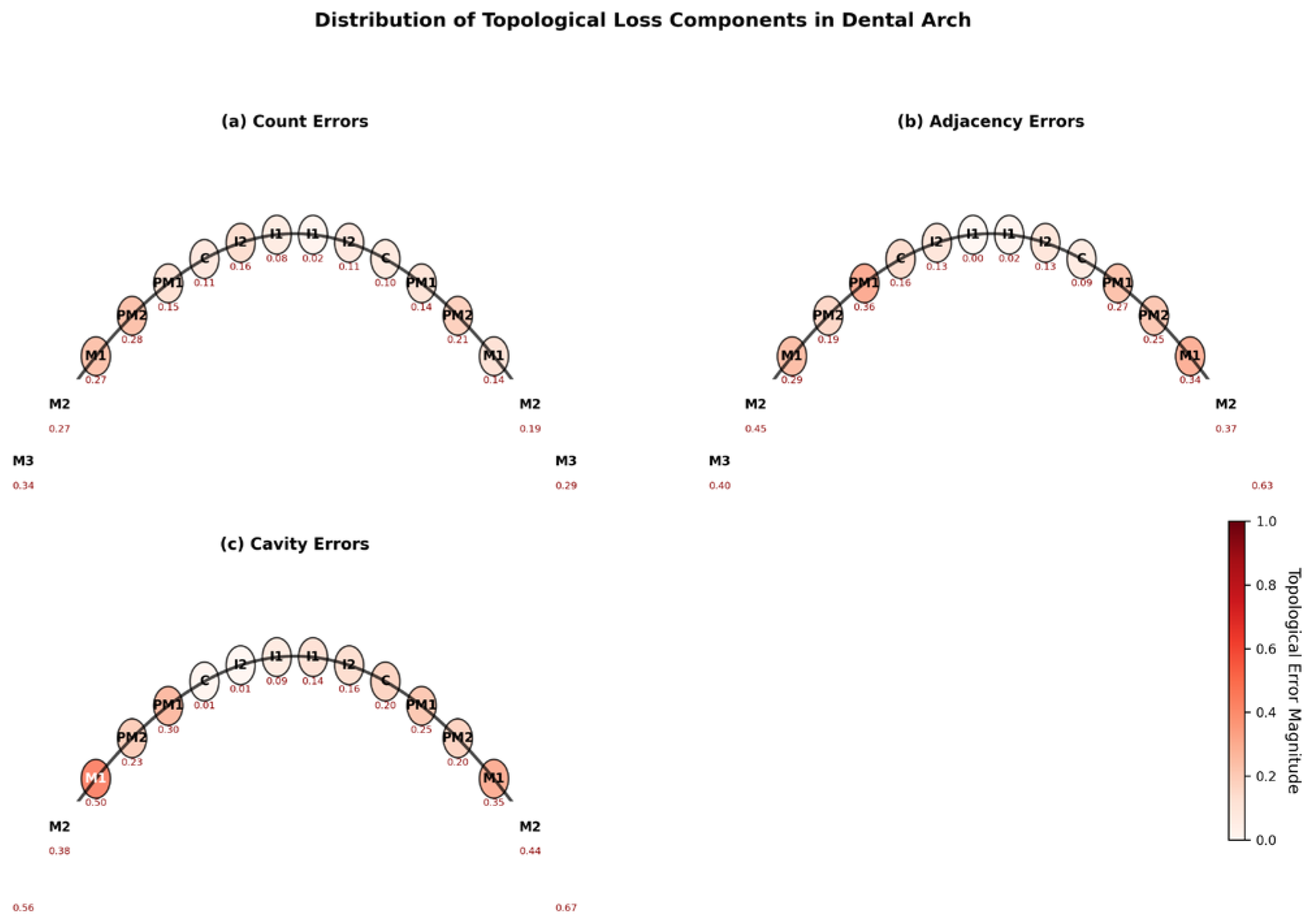
6.2. Topological Error Analysis
6.3. Ablation Study
6.4. Computational Efficiency
7. Discussion and Future Work
7.1. Limitations of the Topology-Constrained Quantized nnUNet
7.2. Potential Application Scenarios of the Proposed Method
7.3. Ethical Considerations in Dental Segmentation with the Proposed Model
7.4. Future Research Directions
8. Conclusions
References
- Isensee, F.; Jaeger, P.; Kohl, S.; Petersen, J.; et al. nnU-net: A self-configuring method for deep learning-based biomedical image segmentation. In Nature Methods; 2021. [Google Scholar]
- Jacob, B.; Kligys, S.; Chen, B.; Zhu, M.; et al. Quantization and training of neural networks for efficient integer-arithmetic-only inference. In Proceedings of the IEEE conference on computer vision and pattern recognition, 2018. [Google Scholar]
- Awari, H.; Subramani, N.; Janagaraj, A.; et al. Three-dimensional dental image segmentation and classification using deep learning with tunicate swarm algorithm. In Expert Systems; 2024. [Google Scholar]
- Bohlender, S.; Oksuz, I.; et al. A survey on shape-constraint deep learning for medical image segmentation. In Ieee Reviews in Biomedical Engineering; 2021. [Google Scholar]
- Nistelrooij, N. V.; Krämer, L.; Kempers, S.; et al. ToothSeg: Robust tooth instance segmentation and numbering in CBCT using deep learning and self-correction. IEEE J. Biomed. Health Inform. 2026. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Gu, H.; Sun, Y. Tooth segmentation on multimodal images using adapted segment anything model. Sci. Rep. 2025. [Google Scholar] [CrossRef] [PubMed]
- Lambert, Z.; Guyader, C. L.; et al. A geometrically-constrained deep network for CT image segmentation. 2021 IEEE 18th international symposium on biomedical imaging (ISBI), 2021. [Google Scholar]
- Singh, Y.; Farrelly, C.; Hathaway, Q.; Leiner, T.; et al. Topological data analysis in medical imaging: Current state of the art. In Insights into Imaging; 2023. [Google Scholar]
- Zheng, G.; Cui, X.; Song, A.; Lin, M. GFACNet: 3D dental segmentation from intraoral scans integrating geometric features and anatomical constraints. In Electronic Research Archive; 2025. [Google Scholar]
- Xi, S.; Liu, Z.; Chang, J.; Wu, H.; et al. 3D dental model segmentation with geometrical boundary preserving. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 2025. [Google Scholar]
- Demir; Massaad, E.; Kiziltan, B. Topology-aware focal loss for 3D image segmentation. In Proceedings of the IEEE conference on computer vision and pattern recognition workshops (CVPRW), 2023, 2023. [Google Scholar]
- Huang, J.; Yan, H.; Li, J.; Stewart, H.; et al. Combining anatomical constraints and deep learning for 3-d CBCT dental image multi-label segmentation. 2021 IEEE 37th international conference on data engineering, 2021. [Google Scholar]
- Wang, P. Latent anomaly detection: Masked VQ-GAN for unsupervised segmentation in medical CBCT. arXiv 2025, arXiv:2506.14209. [Google Scholar] [CrossRef]
- Ben-Hamadou; Smaoui, O.; Rekik, A.; Pujades, S.; et al. 3DTeethSeg’22: 3D teeth scan segmentation and labeling challenge. arXiv 2023, arXiv:2305.18277. [Google Scholar]
- He, Y.; Yang, D.; Roth, H.; Zhao, C.; et al. Dints: Differentiable neural network topology search for 3d medical image segmentation. In Proceedings of the IEEE conference on computer vision and pattern recognition, 2021. [Google Scholar]
- Pedersen, S.; Jain, S.; Chavez, M.; Ladehoff, V.; de Freitas, B. N.; Pauwels, R. Pano-gan: A deep generative model for panoramic dental radiographs. J. Imaging 2025, 11(2), 41. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, S.; Jain, S.; Chavez, M.; Ladehoff, V.; de Freitas, B. N.; Pauwels, R. P. G. A Deep Generative Model for Panoramic Dental Radiographs. J. Imaging 2025, 11, 41. [Google Scholar] [CrossRef] [PubMed]
- Khalil, B.; Baraka, M.; Haghighat, S.; Jain, S.; Manila, N.; Ramani, R.; Pauwels, R. Synthetic imaging in dentistry: A narrative review of deep learning techniques and applications. J. Dent. 2025, 106274. [Google Scholar] [CrossRef] [PubMed]
- Rubak, J. A. B.; Naveed, K.; Jain, S.; Esterle, L.; Iosifidis, A.; Pauwels, R. Impact of labelling inaccuracy and image noise on tooth segmentation in panoramic radiographs using federated, centralized, and local learning. Dentomaxillofacial Radiol. 2026, twag001. [Google Scholar] [CrossRef] [PubMed]
- Jain, S.; de Freitas, B. N.; Basse-OConnor, A.; Iosifidis, A.; Pauwels, R. PanoDiff-SR: synthesizing dental panoramic radiographs using diffusion and super-resolution. arXiv 2025, arXiv:2507.09227. [Google Scholar]
- Mohammad-Rahimi, H.; Jain, S.; Naveed, K.; Hosseinpour, S.; Kirkevang, L. L.; Nosrat, A.; Pauwels, R. Generative Artificial Intelligence for Computer Vision in Endodontics: A Review of Current State and Future Potential. Int. Endod. J. 2026. [Google Scholar] [CrossRef] [PubMed]
- Rubak, J. A. B.; Haghighat, S.; Jain, S.; Aldesoki, M.; Chaurasia, A.; Ehsani, S. S.; Pauwels, R. Deep Learning-based Assessment of the Relation Between the Third Molar and Mandibular Canal on Panoramic Radiographs using Local, Centralized, and Federated Learning. arXiv 2026, arXiv:2603.11850. [Google Scholar] [CrossRef]
| Method | DSC (%) | IoU (%) | BF1 (%) | TCA (%) | ACS (%) | CER (%) | Size (MB) | Time (s) |
|---|---|---|---|---|---|---|---|---|
| Full-Precision nnUNet | 92.3 | 86.1 | 89.7 | 94.2 | 91.5 | 3.1 | 1024 | 8.2 |
| Post-Training Quant | 88.7 | 80.2 | 83.4 | 82.6 | 78.3 | 12.8 | 256 | 2.1 |
| QAT-nnUNet | 90.1 | 82.3 | 86.2 | 85.4 | 83.7 | 9.5 | 256 | 2.3 |
| TopoNet | 91.8 | 85.2 | 88.9 | 93.1 | 90.2 | 4.3 | 896 | 7.5 |
| Proposed | 91.5 | 84.9 | 88.6 | 93.8 | 91.0 | 3.9 | 256 | 2.4 |
| Configuration | DSC (%) | TCA (%) | ACS (%) | CER (%) |
|---|---|---|---|---|
| QAT-only | 90.1 | 85.4 | 83.7 | 9.5 |
| + Count Loss | 90.3 | 89.2 | 85.1 | 8.7 |
| + Adjacency Loss | 90.8 | 91.6 | 89.3 | 6.4 |
| + Cavity Loss | 90.6 | 90.8 | 88.5 | 5.1 |
| Full Topo Loss | 91.5 | 93.8 | 91.0 | 3.9 |
| Method | A100 GPU | Xeon CPU | Jetson AGX |
|---|---|---|---|
| Full-Precision | 1.2 | 8.2 | 14.7 |
| Post-Training | 0.4 | 2.1 | 3.8 |
| QAT-nnUNet | 0.5 | 2.3 | 4.1 |
| Proposed | 0.5 | 2.4 | 4.2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




