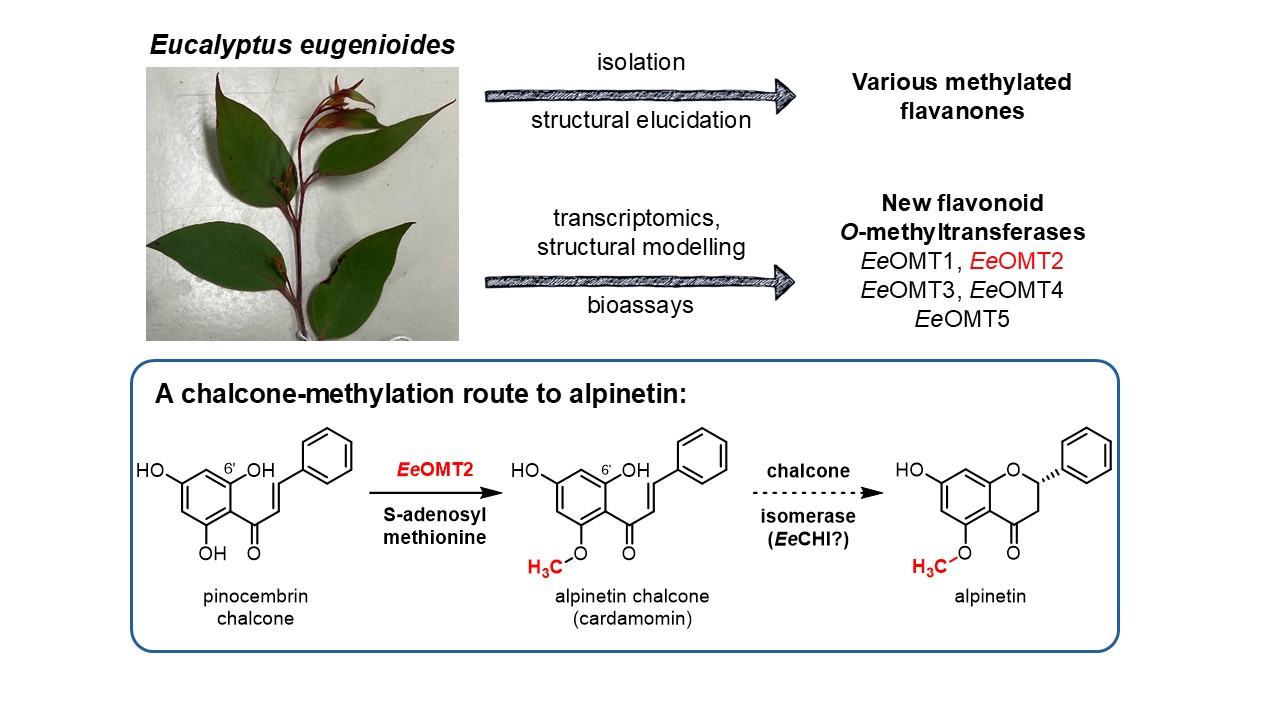
Methylated flavonoids are abundant phytochemicals in Eucalyptus and are of interest be-cause methylation can alter flavonoid diversity, bioactivity and stability. The enzymes re-sponsible for flavonoid methylation in eucalypts are largely uncharacterised. We used comparative leaf transcriptomics of two species with contrasting flavanone profiles, together with protein-structure-guided candidate selection, to identify prospective O-methyltransferases (OMTs) involved in methylated flavonoid biosynthesis. Five candidate OMTs from E. eugenioides were cloned, heterologously expressed and assayed against a panel of flavonoids and a chalcone precursor. The enzymes showed distinct substrate preferences and regioselectivities. EeOMT1 acted as a broad 7-O-methyltransferase, whereas EeOMT3–EeOMT5 preferentially methylated B and C-ring hydroxyl groups, with differing capacities for sequential methylations at different sites. EeOMT2 was of particu-lar interest because it effectively methylated pinocembrin chalcone to alpinetin chalcone, while only weakly converting pinocembrin to alpinetin. Expression–metabolite analyses across E. eugenioides genotypes supported roles relating to in planta accumulation of 5-O- and 7-O-methylated flavanones, for EeOMT2 and EeOMT1, respectively. These findings support a revised model in which alpinetin biosynthesis proceeds, at least in part, through methylation of a chalcone precursor before flavanone formation. This provides a foundation for elucidating flavonoid methylation pathways and for engineering tailored methylated flavonoids for industrial applications.