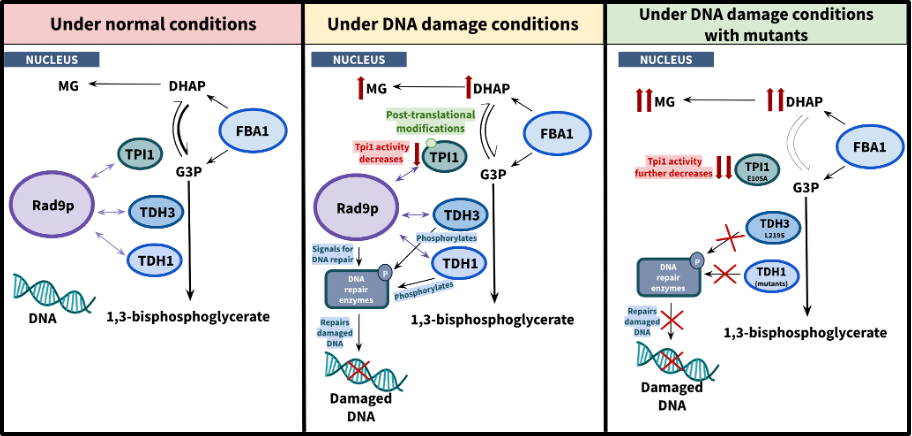
Cancer cells, like yeast, use fermentation despite the presence of oxygen, a phenomenon called aerobic glycolysis. The advantage is that it maintains most of the C-C bonds of glucose, allowing highly proliferating cells to produce the biomolecules that are necessary for cytokinesis. However, aerobic glycolysis is less energy-efficient than respiration, and it must operate at a higher frequency and produces more toxic by-products like methylglyoxal, which damages DNA. Cancer cells, like yeast cells, developed efficient systems to repair their damaged DNA. This makes cancer cells resistant to radiotherapy, which requires a combination with chemotherapy using drugs that inhibit DNA repair. However, this converts healthy cells to cancer cells, indicating that more research is required regarding the relationship between glycolysis and cancer. Using yeast as a model, we have discovered that the glycolytic enzymes TPI1 and GAPDH interact with the DNA damage-dependent checkpoint Rad9p. Furthermore, we have isolated TPI1 and GAPDH mutant strains that are unable to repair their damaged DNA. The TPI1 mutant strain has lower TPI enzymatic activity, suggesting that it accumulates methylglyoxal, while the GAPDH mutant strains have normal GAPDH enzymatic activity, confirming that GAPDH moonlights in the DNA Damage Response.