Submitted:
29 April 2026
Posted:
05 May 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
3. Results and Discussion
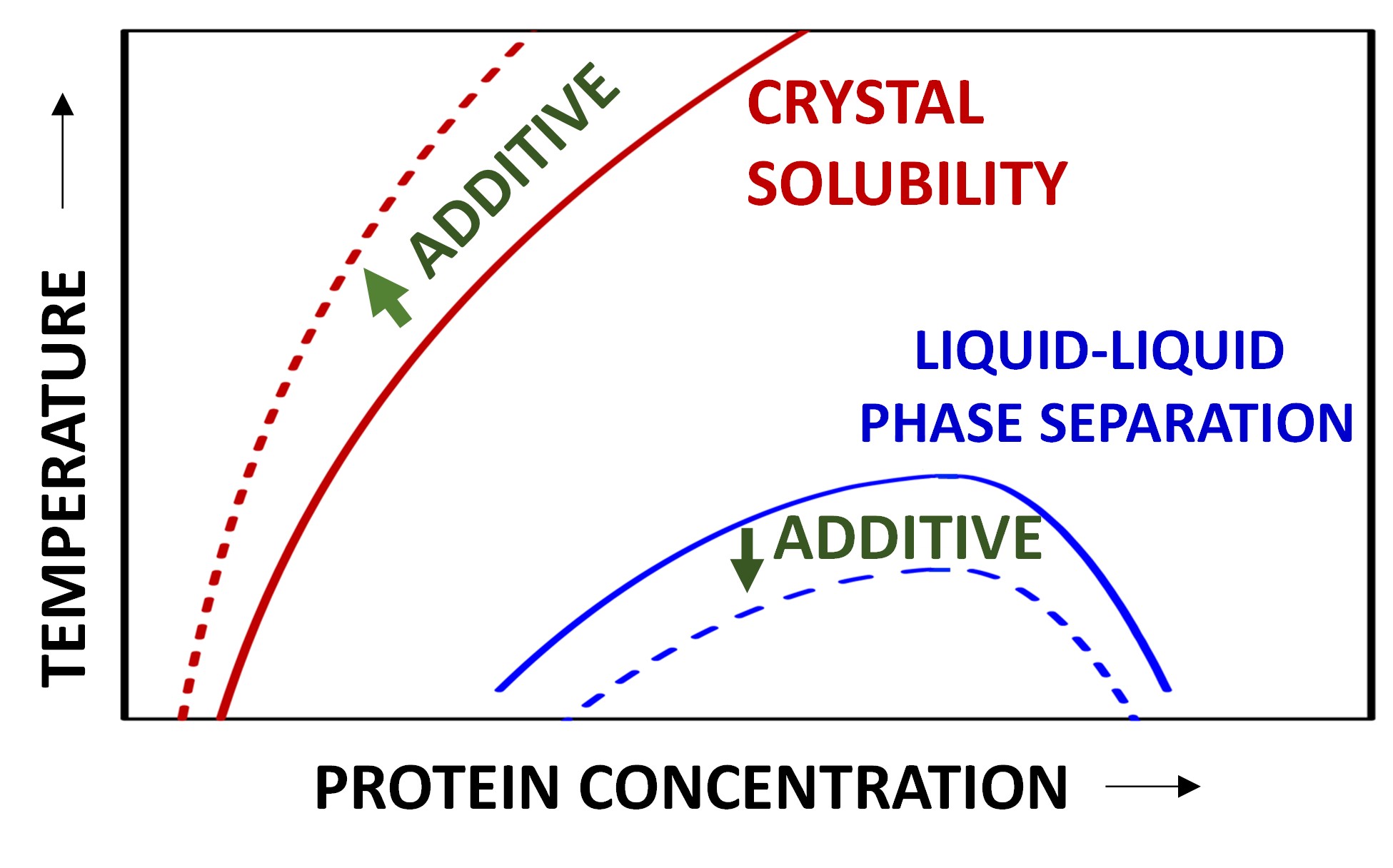
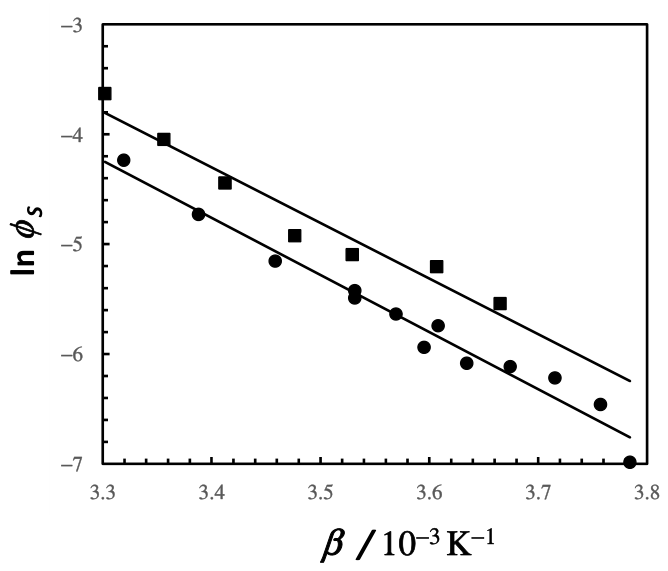
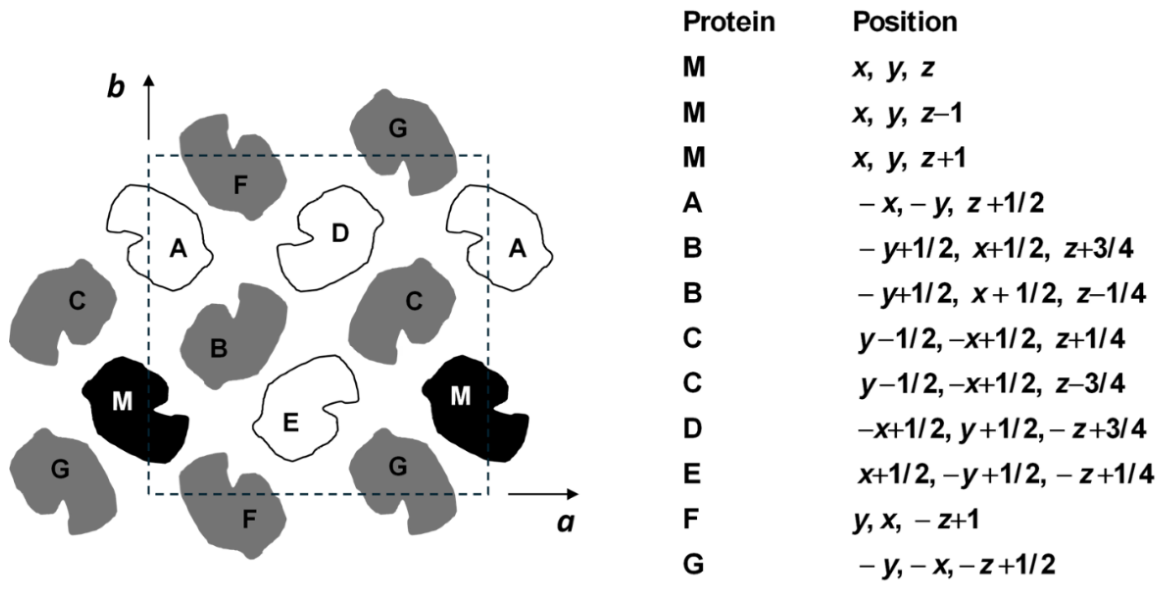
3.1. Crystal Solubility and Thermodynamics of Protein Crystal
3.2. Thermodynamic Model for the Fluid Phase
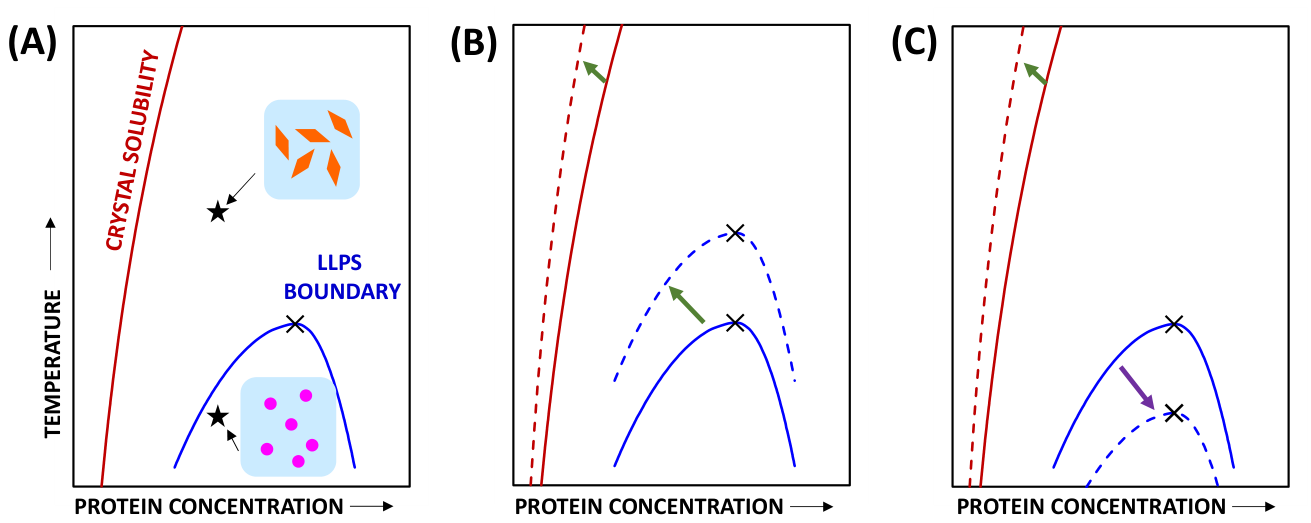
3.3. Effect of HEPES on Lysozyme LLPS Boundary
4. Conclusions
Supplementary Materials
Acknowledgments
References
- Patra, S.; Sharma, B.; George, S. Programmable Coacervate Droplets via Reaction-Coupled Liquid-Liquid Phase Separation (LLPS) and Competitive Inhibition. J. Am. Chem. Soc. 2025, 147(19), 16027–16037. [Google Scholar] [CrossRef]
- Nakashima, K.; van Haren, M.; André, A.; Robu, I.; Spruijt, E. Active coacervate droplets are protocells that grow and resist Ostwald ripening. Nat. Commun. 2021, 12(1). [Google Scholar] [CrossRef]
- Shin, Y.; Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science 2017, 357(6357). [Google Scholar] [CrossRef]
- Sudhakar, S.; Manohar, A.; Mani, E. Liquid-Liquid Phase Separation (LLPS)-Driven Fibrilization of Amyloid-β Protein. ACS Chem. Neurosci. 2023, 14(19), 3655–3664. [Google Scholar] [CrossRef]
- Carey, J.; Guo, L. Liquid-Liquid Phase Separation of TDP-43 and FUS in Physiology and Pathology of Neurodegenerative Diseases. Front. Mol. Biosci. 2022, 9. [Google Scholar] [CrossRef]
- Rowe, J. B.; Cancel, R. A.; Evangelous, T. D.; Flynn, R. P.; Pechenov, S.; Subramony, J. A.; Zhang, J. F.; Wang, Y. Metastability Gap in the Phase Diagram of Monoclonal IgG Antibody. Biophys. J. 2017, 113(8), 1750–1756. [Google Scholar] [CrossRef]
- Raut, A. S.; Kalonia, D. S. Pharmaceutical Perspective on Opalescence and Liquid-Liquid Phase Separation in Protein Solutions. Mol. Pharm. 2016, 13(5), 1431–1444. [Google Scholar] [CrossRef]
- Liu, J.; Spruijt, E.; Miserez, A.; Langer, R. Peptide-based liquid droplets as emerging delivery vehicles. Nat. Rev. Mater. 2023, 8(3), 139–141. [Google Scholar] [CrossRef]
- Ng, T.; Hoare, M.; Maristany, M.; Wilde, E.; Sneideris, T.; Huertas, J.; Agbetiameh, B.; Furukawa, M.; Joseph, J.; Knowles, T.; et al. Tandem-repeat proteins introduce tuneable properties to engineered biomolecular condensates. Chem. Sci. 2025, 16(23), 10532–10548. [Google Scholar] [CrossRef]
- Xu, S.; Zhang, H.; Qiao, B.; Wang, Y. Review of Liquid-Liquid Phase Separation in Crystallization: From Fundamentals to Application. Cryst. GROWTH Des. 2021, 21(12), 7306–7325. [Google Scholar] [CrossRef]
- McPherson, A. Introduction to the Crystallization of Biological Macromolecules. Membr. Protein Cryst. 2009, 63, 5–23. [Google Scholar] [CrossRef]
- Chen, J.; Sarma, B.; Evans, J. M. B.; Myerson, A. S. Pharmaceutical Crystallization. Cryst. Growth Des. 2011, 11(4), 887–895. [Google Scholar] [CrossRef]
- McPherson, A. Crystallization of Biological Macromolecules; Cold Spring Harbor Lab. Press, 1999. [Google Scholar]
- Hekmat, D.; Breitschwerdt, P.; Weuster-Botz, D. Purification of proteins from solutions containing residual host cell proteins via preparative crystallization. Biotechnol. Lett. 2015, 37(9), 1791–1801. [Google Scholar] [CrossRef]
- Hubbuch, J.; Kind, M.; Nirschl, H. Preparative Protein Crystallization. Chem. Eng. Technol. 2019, 42(11), 2275–2281. [Google Scholar] [CrossRef]
- Broide, M. L.; Berland, C. R.; Pande, J.; Ogun, O. O.; Benedek, G. B. BINARY-LIQUID PHASE-SEPARATION OF LENS PROTEIN SOLUTIONS. Proc. Natl. Acad. Sci. U. S. A. 1991, 88(13), 5660–5664. [Google Scholar] [CrossRef]
- Muschol, M.; Rosenberger, F. Liquid-liquid phase separation in supersaturated lysozyme solutions and associated precipitate formation/crystallization. J. Chem. Phys. 1997, 107(6), 1953–1962. [Google Scholar] [CrossRef]
- Annunziata, O.; Asherie, N.; Lomakin, A.; Pande, J.; Ogun, O.; Benedek, G. B. Effect of polyethylene glycol on the liquid-liquid phase transition in aqueous protein solutions. Proc. Natl. Acad. Sci. U. S. A. 2002, 99(22), 14165–14170, Article. [Google Scholar] [CrossRef]
- Galkin, O.; Vekilov, P. G. Control of protein crystal nucleation around the metastable liquid-liquid phase boundary. Proc. Natl. Acad. Sci. U. S. A. 2000, 97(12), 6277–6281. [Google Scholar] [CrossRef] [PubMed]
- Pantuso, E.; Mastropietro, T.; Briuglia, M.; Gerard, C.; Curcio, E.; ter Horst, J.; Nicoletta, F.; Di Profio, G. On the Aggregation and Nucleation Mechanism of the Monoclonal Antibody Anti-CD20 Near Liquid-Liquid Phase Separation (LLPS). Sci. Rep. 2020, 10(1). [Google Scholar] [CrossRef] [PubMed]
- Galkin, O.; Chen, K.; Nagel, R.; Hirsch, R.; Vekilov, P. Liquid-liquid separation in solutions of normal and sickle cell hemoglobin. Proc. Natl. Acad. Sci. U. S. A. 2002, 99(13), 8479–8483. [Google Scholar] [CrossRef]
- Babinchak, W.; Surewicz, W. Liquid-Liquid Phase Separation and Its Mechanistic Role in Pathological Protein Aggregation. J. Mol. Biol. 2020, 432(7), 1910–1925. [Google Scholar] [CrossRef]
- Liu, C. W.; Asherie, N.; Lomakin, A.; Pande, J.; Ogun, O.; Benedek, G. B. Phase separation in aqueous solutions of lens gamma-crystallins: Special role of gamma(s). Proc. Natl. Acad. Sci. U. S. A. 1996, 93(1), 377–382. [Google Scholar] [CrossRef] [PubMed]
- Annunziata, O.; Pande, A.; Pande, J.; Ogun, O.; Lubsen, N. H.; Benedek, G. B. Oligomerization and phase transitions in aqueous solutions of native and truncated human beta B1-crystalline. Biochemistry 2005, 44(4), 1316–1328. [Google Scholar] [CrossRef]
- Taratuta, V. G.; Holschbach, A.; Thurston, G. M.; Blankschtein, D.; Benedek, G. B. LIQUID LIQUID-PHASE SEPARATION OF AQUEOUS LYSOZYME SOLUTIONS - EFFECTS OF PH AND SALT IDENTITY. J. Phys. Chem. 1990, 94(5), 2140–2144. [Google Scholar] [CrossRef]
- Jim, A. I.; Goh, L. T.; Oh, S. K. W. Crystallization of IgG(1) by mapping its liquid-liquid phase separation curves. Biotechnol. Bioeng. 2006, 95(5), 911–918. [Google Scholar] [CrossRef]
- Maier, R.; Zocher, G.; Sauter, A.; Da Vela, S.; Matsarskaia, O.; Schweins, R.; Sztucki, M.; Zhang, F. J.; Stehle, T.; Schreiber, F. Protein Crystallization in the Presence of a Metastable Liquid-Liquid Phase Separation. Cryst. Growth Des. 2020, 20(12), 7951–7962. [Google Scholar] [CrossRef]
- Thomas, S.; Dougay, J.; Annunziata, O. Yield of Protein Crystallization from Metastable Liquid-Liquid Phase Separation. MOLECULES 2025, 30(11). [Google Scholar] [CrossRef] [PubMed]
- McPherson, A. Introduction to protein crystallization. Methods 2004, 34(3), 254–265. [Google Scholar] [CrossRef]
- Saridakis, E.; Chayen, N. E. Towards a 'universal' nucleant for protein crystallization. Trends Biotechnol. 2009, 27(2), 99–106. [Google Scholar] [CrossRef]
- Lomakin, A.; Asherie, N.; Benedek, G. B. Monte Carlo study of phase separation in aqueous protein solutions. J. Chem. Phys. 1996, 104(4), 1646–1656. [Google Scholar] [CrossRef]
- Hribar-Lee, B.; Luksic, M. Biophysical Principles Emerging from Experiments on Protein-Protein Association and Aggregation. Annu. Rev. Biophys. 2024, 53, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Pellicane, G. Colloidal Model of Lysozyme Aqueous Solutions: A Computer Simulation and Theoretical Study. J. Phys. Chem. B 2012, 116(7), 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Kuehner, D.; Heyer, C.; Ramsch, C.; Fornefeld, U.; Blanch, H.; Prausnitz, J. Interactions of lysozyme in concentrated electrolyte solutions from dynamic light-scattering measurements. Biophys. J. 1997, 73(6), 3211–3224. [Google Scholar] [CrossRef]
- Timasheff, S. N. Protein-solvent preferential interactions, protein hydration, and the modulation of biochemical reactions by solvent components. Proceedings of the National Academy of Sciences of the United States of America 2002, 99(15), 9721–9726. [Google Scholar] [CrossRef]
- Record, M. T.; Anderson, C. F. Interpretation of preferential interaction coefficients of nonelectrolytes and of electrolyte ions in terms of a two-domain model. Biophys. J. 1995, 68(3), 786–794. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Annunziata, O. Comparison between protein-polyethylene glycol (PEG) interactions and the effect of PEG on protein-protein interactions using the liquid-liquid phase transition. J. Phys. Chem. B 2007, 111(5), 1222–1230. [Google Scholar] [CrossRef]
- Vergara, A.; Capuano, F.; Paduano, L.; Sartorio, R. Lysozyme mutual diffusion in solutions crowded by poly(ethylene glycol). Macromolecules 2006, 39(13), 4500–4506. [Google Scholar] [CrossRef]
- Bhat, R.; Timasheff, S. N. STERIC EXCLUSION IS THE PRINCIPAL SOURCE OF THE PREFERENTIAL HYDRATION OF PROTEINS IN THE PRESENCE OF POLYETHYLENE GLYCOLS. Protein Sci. 1992, 1(9), 1133–1143. [Google Scholar] [CrossRef]
- Vivares, D.; Belloni, L.; Tardieu, A.; Bonnete, F. Catching the PEG-induced attractive interaction between proteins. Eur. Phys. J. E 2002, 9(1), 15–25. [Google Scholar] [CrossRef]
- Bloustine, J.; Virmani, T.; Thurston, G. M.; Fraden, S. Light scattering and phase behavior of lysozyme-poly(ethylene glycol) mixtures. Phys. Rev. Lett. 2006, 96(8). [Google Scholar] [CrossRef]
- Asherie, N.; Lomakin, A.; Benedek, G. B. Phase diagram of colloidal solutions. Phys. Rev. Lett. 1996, 77(23), 4832–4835. [Google Scholar] [CrossRef]
- Platten, F.; Valadez-Pérez, N.; Castañeda-Priego, R.; Egelhaaf, S. Extended law of corresponding states for protein solutions. J. Chem. Phys. 2015, 142(17). [Google Scholar] [CrossRef]
- Kastelic, M.; Kalyuzhnyi, Y. V.; Hribar-Lee, B.; Dill, K. A.; Vlachy, V. Protein aggregation in salt solutions. Proc. Natl. Acad. Sci. U. S. A. 2015, 112(21), 6766–6770. [Google Scholar] [CrossRef] [PubMed]
- Asherie, N. Protein crystallization and phase diagrams. Methods 2004, 34(3), 266–272. [Google Scholar] [CrossRef]
- Arakawa, T.; Timasheff, S. N. Preferential interactions of proteins with salts in concentrated solutions. Biochemistry 1982, 21(25), 6545–6552. [Google Scholar] [CrossRef]
- Arakawa, T.; Timasheff, S. N. THEORY OF PROTEIN SOLUBILITY. Methods Enzymol. 1985, 114, 49–77. [Google Scholar]
- Jungwirth, P.; Cremer, P. S. Beyond Hofmeister. Nat. Chem. 2014, 6(4), 261–263. [Google Scholar] [CrossRef] [PubMed]
- Annunziata, O.; Payne, A.; Wang, Y. Solubility of lysozyme in the presence of aqueous chloride salts: Common-ion effect and its role on solubility and crystal thermodynamics. J. Am. Chem. Soc. 2008, 130(40), 13347–13352, Article. [Google Scholar] [CrossRef] [PubMed]
- Retailleau, P.; RiesKautt, M.; Ducruix, A. No salting-in of lysozyme chloride observed at how ionic strength over a large range of pH. Biophys. J. 1997, 73(4), 2156–2163. [Google Scholar] [CrossRef]
- Hansen, J.; Platten, F.; Wagner, D.; Egelhaaf, S. Tuning protein-protein interactions using cosolvents: specific effects of ionic and non-ionic additives on protein phase behavior. Phys. Chem. Chem. Phys. 2016, 18(15), 10270–10280. [Google Scholar] [CrossRef]
- Fahim, A.; Annunziata, O. Effect of a Good buffer on the fate of metastable protein-rich droplets near physiological composition. Int. J. Biol. Macromol. 2021, 186, 519–527. [Google Scholar] [CrossRef]
- Fahim, A.; Pham, J.; Thomas, S.; Annunziata, O. Boosting protein crystallization from liquid-liquid phase separation by increasing metastability gap. J. Mol. Liq. 2024, 398, 124164. [Google Scholar] [CrossRef]
- Camara-Artigas, A.; Salinas-Garcia, M. C.; Plaza-Garrido, M. Crystal structure of Lysozyme in complex with Hepes. [CrossRef]
- McPherson, A.; Cudney, B. Searching for silver bullets: An alternative strategy for crystallizing macromolecules. J. Struct. Biol. 2006, 156(3), 387–406. [Google Scholar] [CrossRef]
- Albright, J. G.; Annunziata, O.; Miller, D. G.; Paduano, L.; Pearlstein, A. J. Precision measurements of binary and multicomponent diffusion coefficients in protein solutions relevant to crystal growth: Lysozyme chloride in water and aqueous NaCl at pH 4.5 and 25 degrees C-perpendicular to. In Journal of the American Chemical Society; Proceedings Paper, 1999; Volume 121, 14, pp. 3256–3266. [Google Scholar] [CrossRef]
- CARNAHAN, N.; STARLING, K. EQUATION OF STATE FOR NONATTRACTING RIGID SPHERES. J. Chem. Phys. 1969, 51(2), 635–&. [Google Scholar] [CrossRef]
- BARKER, J.; HENDERSON, D. PERTURBATION THEORY AND EQUATION OF STATE FOR FLUIDS - SQUARE-WELL POTENTIAL. J. Chem. Phys. 1967, 47(8), 2856–+. [Google Scholar] [CrossRef]
- Sear, R. Phase behavior of a simple model of globular proteins. J. Chem. Phys. 1999, 111(10), 4800–4806. [Google Scholar] [CrossRef]
- Vaney, M.; Maignan, S.; RiesKautt, M.; Ducruix, A. High-resolution structure (1.33 angstrom) of a HEW lysozyme tetragonal crystal grown in the APCF apparatus. Data and structural comparison with a crystal grown under microgravity from SpaceHab-01 mission. ACTA Crystallogr. Sect. D.-Struct. Biol. 1996, 52, 505–517. [Google Scholar] [CrossRef]
- Nadarajah, A.; Pusey, M. Growth mechanism and morphology of tetragonal lysozyme crystals. ACTA Crystallogr. Sect. D.-Biol. Crystallogr. 1996, 52, 983–996. [Google Scholar] [CrossRef]
- Weiss, M.; Palm, G.; Hilgenfeld, R. Crystallization, structure solution and refinement of hen egg-white lysozyme at pH 8.0 in the presence of MPD. ACTA Crystallogr. Sect. D.-Struct. Biol. 2000, 56, 952–958. [Google Scholar] [CrossRef]
- Lomakin, A.; Asherie, N.; Benedek, G. Aeolotopic interactions of globular proteins. Proc. Natl. Acad. Sci. U. S. A. 1999, 96(17), 9465–9468. [Google Scholar] [CrossRef]
- Fine, B. M.; Lomakin, A.; Ogun, O. O.; Benedek, G. B. Static structure factor and collective diffusion of globular proteins in concentrated aqueous solution. J. Chem. Phys. 1996, 104(1), 326–335. [Google Scholar] [CrossRef]
- HENDERSON, D.; SCALISE, O.; SMITH, W. MONTE-CARLO CALCULATIONS OF THE EQUATION OF STATE OF THE SQUARE-WELL FLUID AS A FUNCTION OF WELL WIDTH. J. Chem. Phys. 1980, 72(4), 2431–2438. [Google Scholar] [CrossRef]
- VEGA, L.; DEMIGUEL, E.; RULL, L.; JACKSON, G.; MCLURE, I. PHASE-EQUILIBRIA AND CRITICAL-BEHAVIOR OF SQUARE-WELL FLUIDS OF VARIABLE WIDTH BY GIBBS ENSEMBLE MONTE-CARLO SIMULATION. J. Chem. Phys. 1992, 96(3), 2296–2305. [Google Scholar] [CrossRef]
- LOMBA, E.; ALMARZA, N. ROLE OF THE INTERACTION RANGE IN THE SHAPING OF PHASE-DIAGRAMS IN SIMPLE FLUIDS - THE HARD-SPHERE YUKAWA FLUID AS A CASE-STUDY. J. Chem. Phys. 1994, 100(11), 8367–8372. [Google Scholar] [CrossRef]
- Kern, N.; Frenkel, D. Fluid-fluid coexistence in colloidal systems with short-ranged strongly directional attraction. J. Chem. Phys. 2003, 118(21), 9882–9889. [Google Scholar] [CrossRef]
- Bianchi, E.; Largo, J.; Tartaglia, P.; Zaccarelli, E.; Sciortino, F. Phase diagram of patchy colloids: Towards empty liquids. Phys. Rev. Lett. 2006, 97(16). [Google Scholar] [CrossRef]
- WERTHEIM, M. FLUIDS WITH HIGHLY DIRECTIONAL ATTRACTIVE FORCES.2. THERMODYNAMIC PERTURBATION-THEORY AND INTEGRAL-EQUATIONS. J. Stat. Phys. 1984, 35(1-2), 35–47. [Google Scholar] [CrossRef]
- GilVillegas, A.; Galindo, A.; Whitehead, P.; Mills, S.; Jackson, G.; Burgess, A. Statistical associating fluid theory for chain molecules with attractive potentials of variable range. J. Chem. Phys. 1997, 106(10), 4168–4186. [Google Scholar] [CrossRef]
- López-Picón, J.; Escamilla-Herrera, L.; Torres-Arenas, J. The square-well fluid: A thermodynamic geometric view. J. Mol. Liq. 2022, 368. [Google Scholar] [CrossRef]
- Brudar, S.; Hribar-Lee, B. Effect of Buffer on Protein Stability in Aqueous Solutions: A Simple Protein Aggregation Model. J. Phys. Chem. B 2021, 125(10), 2504–2512. [Google Scholar] [CrossRef]
- Patel, B.; Docherty, H.; Varga, S.; Galindo, A.; Maitland, G. Generalized equation of state for square-well potentials of variable range. Mol. Phys. 2005, 103(1), 129–139. [Google Scholar] [CrossRef]
- GilVillegas, A.; delRio, F.; Benavides, A. Deviations from corresponding-states behavior in the vapor-liquid equilibrium of the square-well fluid. FLUID PHASE Equilib. 1996, 119(1-2), 97–112. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



