Submitted:
28 April 2026
Posted:
30 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
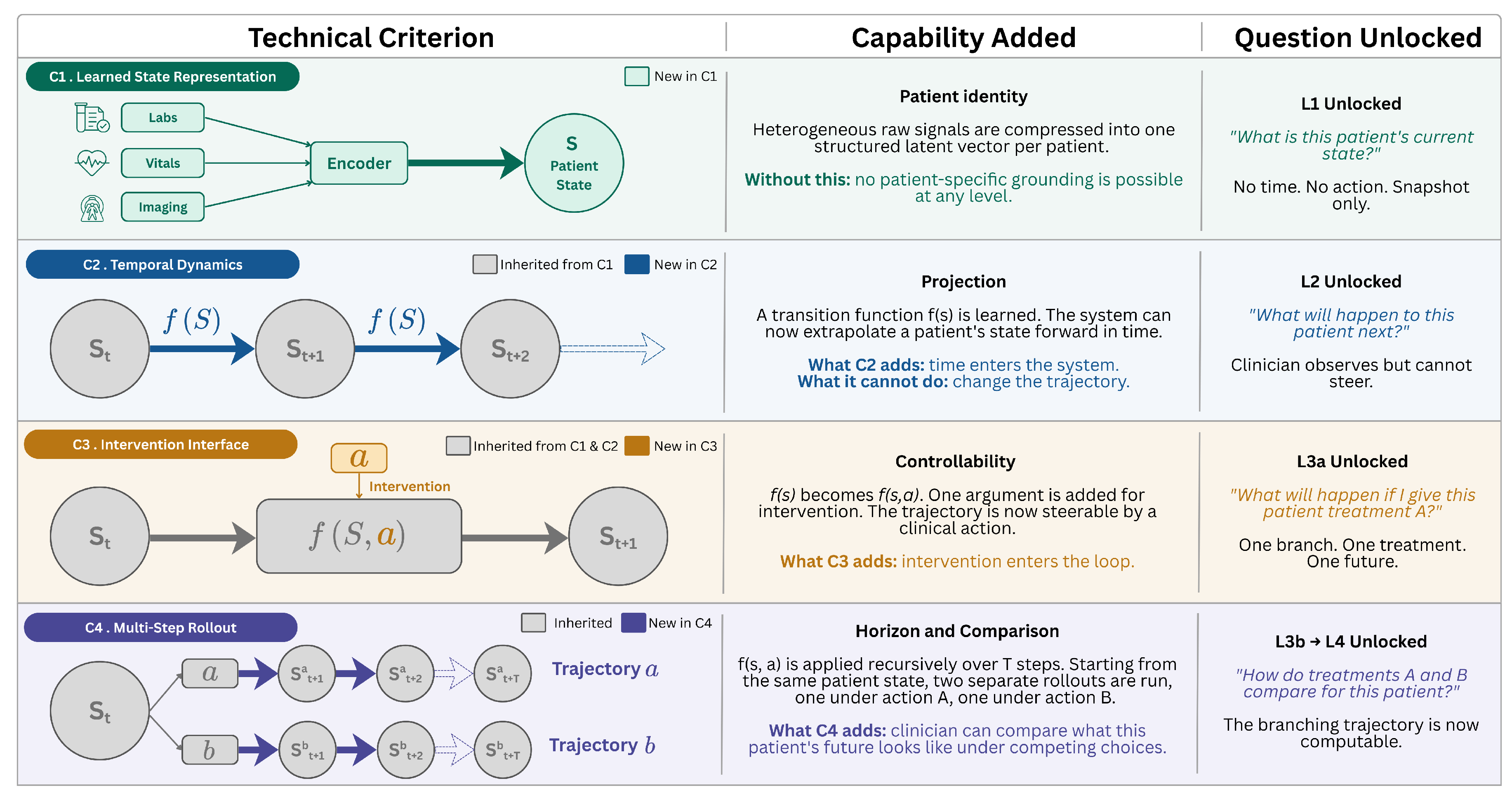
- 1.
- Learned state representation. The model learns a compressed representation of the patient, organ system, or clinical environment from data, rather than relying only on hand-engineered features.
- 2.
- Temporal dynamics (state transitions). The model learns how the patient’s state evolves over time by modeling transitions between states, rather than making isolated, single-timepoint predictions.
- 3.
- Intervention interface. The model accepts clinically meaningful actions (drugs, procedures, device settings) as inputs that causally influence state transitions, enabling simulation of alternative treatment choices rather than reflecting historical correlations alone.
- 4.
- Multi-step rollout. The model can iteratively apply its transition dynamics under specified interventions to generate forward trajectories over multiple time steps (i.e., simulations), which can be assessed for plausibility, calibration, and clinical utility.
2. A Five-Level Capability Ladder for Medical World Models
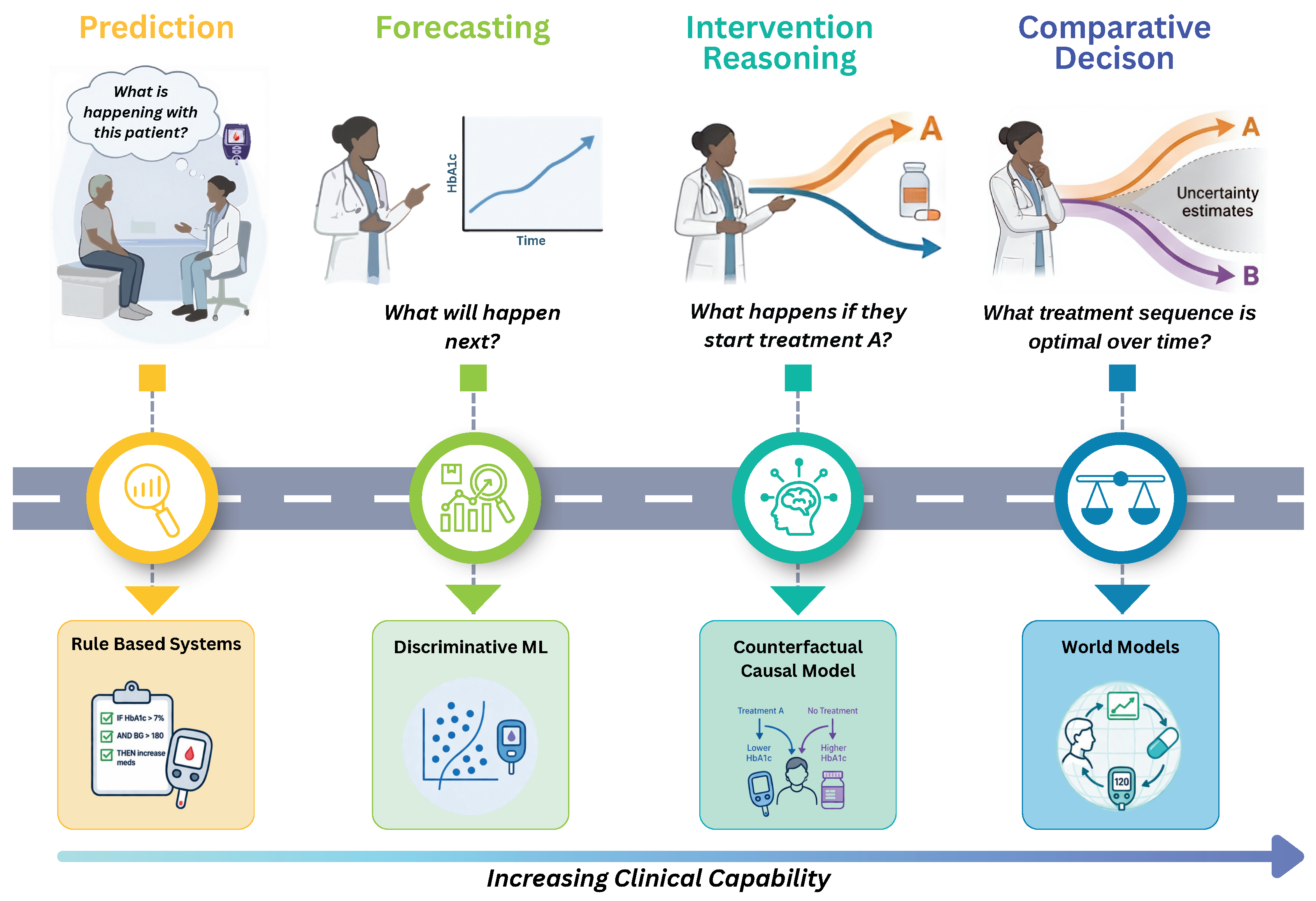
2.1. Five Levels of Capability
Representing the patient state (L1).
Forecasting without intervention (L2).
Projecting one treatment path (L3a).
Comparing alternative interventions (L3b).
Planning over time (L4).
2.2. Clinical Domains and Boundaries
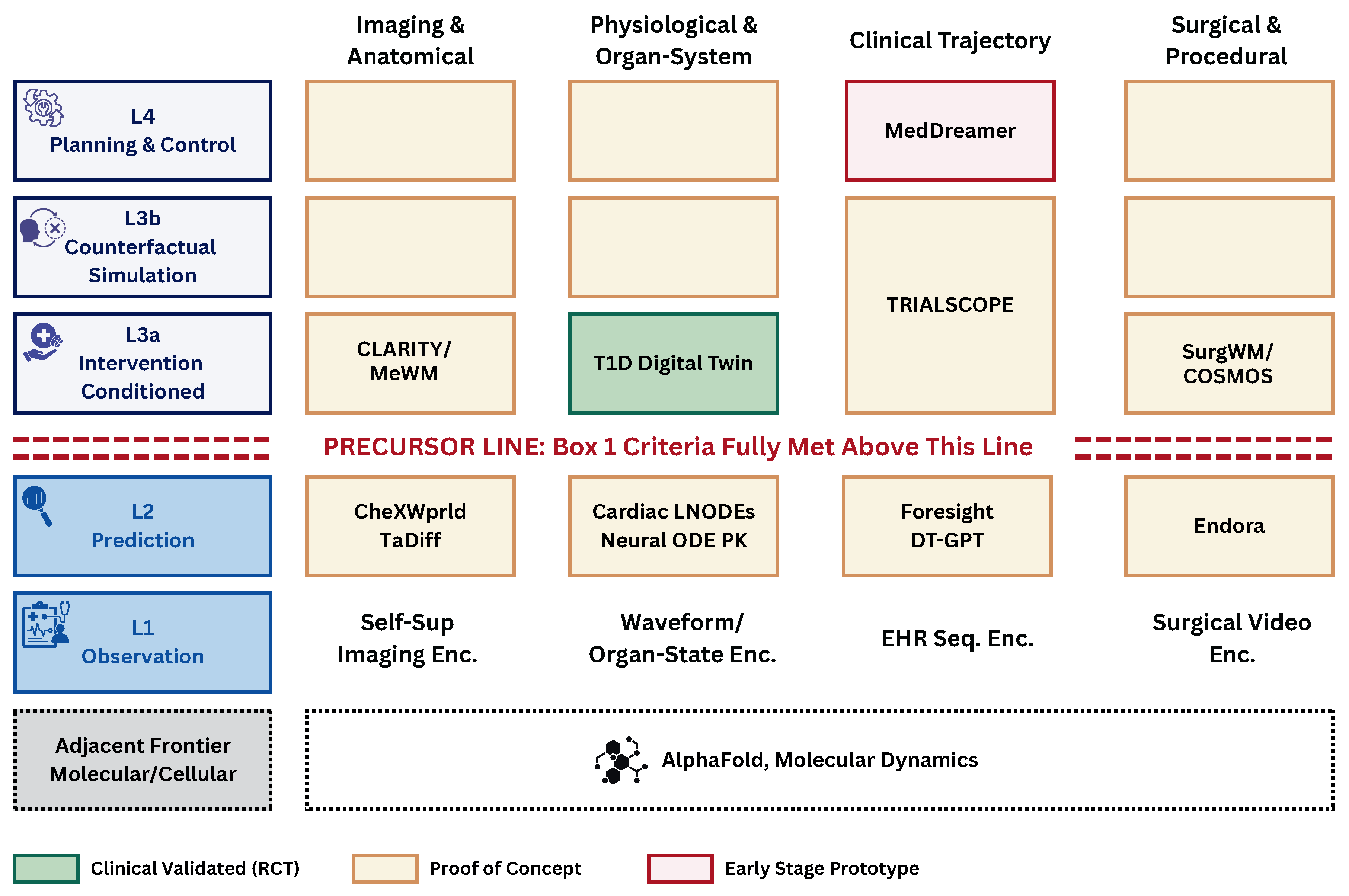
3. The Current Landscape of Medical World Models
3.1. Imaging Offers Clear Prototypes with Narrow Scope
3.2. Physiology: The Deepest Roots, the Strongest Clinical Evidence
3.3. Clinical Trajectories: Rich Data, Limited Causal Footing
3.4. Surgery: Visual Realism Without Physical Grounding
3.5. Foundation Models: Provisional Bridges, not Grounded Simulators
4. Barriers to Reliable Medical World Models
4.1. Validating Unobserved Futures
4.2. Failure Modes of Medical World Models
4.3. Data Gaps in Patient Modeling
4.4. Generalizability Across Populations and Settings
4.5. Prioritizing the Barriers
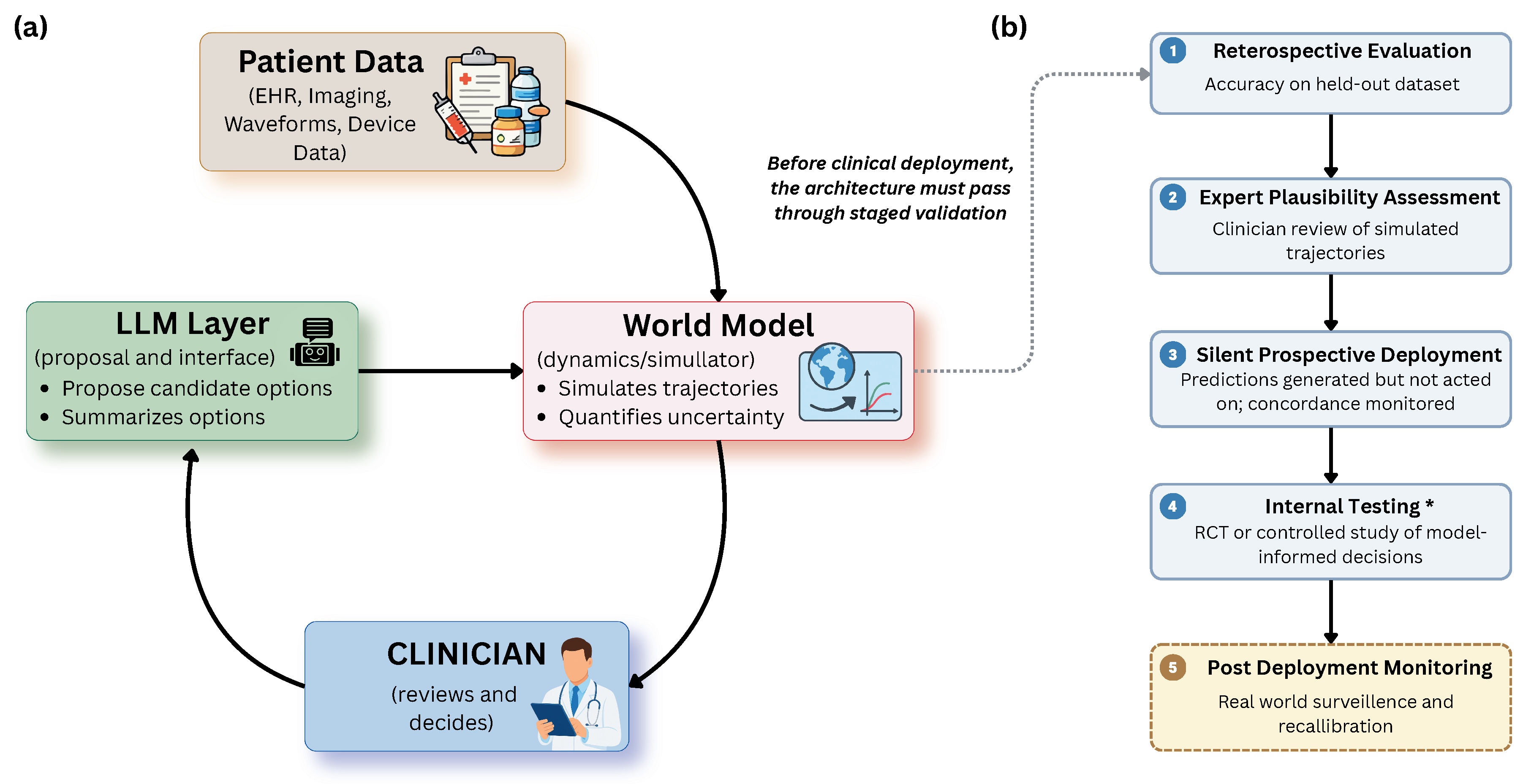
5. Translating Medical World Models to the Clinic
5.1. Integrating Simulation into Clinical Workflows
5.2. Recognizing When Simulation Is Unreliable
5.3. Near-Term Opportunities Without New Model Training
5.4. Building Composite and Hybrid Systems
5.5. Toward Causal Simulation and Closed-Loop Care
6. From Prediction to Clinical Simulation
References
- LeCun, Y. A Path Towards Autonomous Machine Intelligence. OpenReview Preprint 2022. Version 0.9.2, 2022-06-27.
- Ha, D.; Schmidhuber, J. Recurrent world models facilitate policy evolution. Adv. Neural Inf. Process. Syst. 2018, 31. [Google Scholar]
- Hafner, D.; Pasukonis, J.; Ba, J.; Lillicrap, T. Mastering diverse control tasks through world models. Nature 2025, 640, 647–653. [Google Scholar] [CrossRef]
- Brooks, T.; Peebles, B.; Holmes, C.; DePue, W.; Guo, Y.; Jing, L.; Schnurr, D.; Taylor, J.; Luhman, T.; Luhman, E.; et al. Video generation models as world simulators. OpenAI Blog 2024, 1, 1. [Google Scholar]
- Parker-Holder, J.; Ball, P.; Bruce, J.; Dasagi, V.; Holsheimer, K.; Kaplanis, C.; Moufarek, A.; Scully, G.; Shar, J.; Shi, J.; et al. Genie 2: A large-scale foundation world model. URL: https://deepmind. google/discover/blog/genie-2-a-large-scale-foundation-world-model 2024, 2.
- Topol, E.J. High-performance medicine: the convergence of human and artificial intelligence. Nat. Med. 2019, 25, 44–56. [Google Scholar] [CrossRef]
- Esteva, A.; et al. A guide to deep learning in healthcare. Nat. Med. 2019, 25, 24–29. [Google Scholar] [CrossRef] [PubMed]
- Rajkomar, A.; et al. Scalable and accurate deep learning with electronic health records. npj Digit. Med. 2018, 1, 18. [Google Scholar] [CrossRef]
- Hernán, M.A.; Robins, J.M. Causal Inference: What If; Chapman & Hall/CRC: Boca Raton, 2020. [Google Scholar]
- Sadée, C.; et al. Medical digital twins: enabling precision medicine and medical artificial intelligence. Lancet Digit. Health 2025, 7, 100864. [Google Scholar] [CrossRef]
- Kraljevic, Z.; et al. Foresight: a generative pretrained transformer for modelling of patient timelines using electronic health records: a retrospective modelling study. Lancet Digit. Health 2024, 6, e281–e290. [Google Scholar] [CrossRef]
- Makarov, N.; et al. Large language models forecast patient health trajectories enabling digital twins. npj Digit. Med. 2025, 8, 588. [Google Scholar] [CrossRef] [PubMed]
- Long, Y.; Lin, A.; Kwok, D.H.C.; Zhang, L.; Yang, Z.; Shi, K.; Song, L.; Fu, J.; Lin, H.; Wei, W.; et al. Surgical embodied intelligence for generalized task autonomy in laparoscopic robot-assisted surgery. Sci. Robot. 2025, 10, eadt3093. [Google Scholar] [CrossRef]
- Yang, Z.; et al. TransformEHR: transformer-based encoder-decoder generative model to enhance prediction of disease outcomes using electronic health records. Nat. Commun. 2023, 14, 7857. [Google Scholar] [CrossRef] [PubMed]
- Singhal, K.; et al. Large language models encode clinical knowledge. Nature 2023, 620, 172–180. [Google Scholar] [CrossRef] [PubMed]
- Qazi, M.A.; Nadeem, M.; Yaqub, M. Beyond Generative AI: World Models for Clinical Prediction, Counterfactuals, and Planning. In Proceedings of the NeurIPS 2025 Workshop on Bridging Language, Agent, and World Models for Reasoning and Planning, 2025.
- Tomašev, N.; et al. A clinically applicable approach to continuous prediction of future acute kidney injury. Nature 2019, 572, 116–119. [Google Scholar] [CrossRef]
- Pham, T.; Tran, T.; Phung, D.; Venkatesh, S. Deepcare: A deep dynamic memory model for predictive medicine. In Proceedings of the Pacific-Asia conference on knowledge discovery and data mining. Springer, 2016, pp. 30–41.
- Shalit, U.; Johansson, F.D.; Sontag, D. Estimating individual treatment effect: generalization bounds and algorithms. In Proceedings of the International conference on machine learning. PMLR, 2017, pp. 3076–3085.
- Komorowski, M.; Celi, L.A.; Badawi, O.; Gordon, A.C.; Faisal, A.A. The Artificial Intelligence Clinician learns optimal treatment strategies for sepsis in intensive care. Nat. Med. 2018, 24, 1716–1720. [Google Scholar] [CrossRef]
- Gottesman, O.; Johansson, F.; Komorowski, M.; Faisal, A.; Sontag, D.; Doshi-Velez, F.; Celi, L.A. Guidelines for reinforcement learning in healthcare. Nat. Med. 2019, 25, 16–18. [Google Scholar] [CrossRef]
- Murphy, S.A. Optimal dynamic treatment regimes. J. R. Stat. Soc. Ser. B Stat. Methodol. 2003, 65, 331–355. [Google Scholar] [CrossRef]
- Chakraborty, B.; Murphy, S.A. Dynamic Treatment Regimes. Annu. Rev. Stat. Its Appl. 2014, 1, 447–464. [Google Scholar] [CrossRef]
- Pearl, J. Causality; Cambridge university press, 2009. [Google Scholar]
- Jumper, J.; Evans, R.; Pritzel, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Hollingsworth, S.A.; Dror, R.O. Molecular Dynamics Simulation for All. Neuron 2018, 99, 1129–1143. [Google Scholar] [CrossRef]
- Cheng, X.; Li, P.; Guo, H.; Liang, Y.; Gong, J.; de Vazelhes, W.; Gou, C.; Xie, P.; Song, L.; Xing, E.P. Harnessing AI to Build Virtual Cells. bioRxiv 2026, pp. 2026–04. [Google Scholar] [CrossRef]
- Yang, Y.; Wang, Z.Y.; Liu, Q.; Sun, S.; Wang, K.; Chellappa, R.; Zhou, Z.; Yuille, A.; Zhu, L.; Zhang, Y.D.; et al. Medical world model: Generative simulation of tumor evolution for treatment planning. arXiv preprint arXiv:2506.02327 2025. [Google Scholar]
- Ding, T.; Zou, Y.; Chen, C.; Shah, M.; Tian, Y. CLARITY: Medical World Model for Guiding Treatment Decisions by Modeling Context-Aware Disease Trajectories in Latent Space. arXiv preprint arXiv:2512.08029 2025. [Google Scholar]
- Yue, Y.; Wang, Y.; Tao, C.; Liu, P.; Song, S.; Huang, G. CheXWorld: Exploring image world modeling for radiograph representation learning. In Proceedings of the Proceedings of the Computer Vision and Pattern Recognition Conference, 2025, pp. 20778–20788.
- Liu, Q.; Fuster-Garcia, E.; Hovden, I.T.; MacIntosh, B.J.; Grødem, E.O.; Brandal, P.; Lopez-Mateu, C.; Sederevičius, D.; Skogen, K.; Schellhorn, T.; et al. Treatment-aware diffusion probabilistic model for longitudinal MRI generation and diffuse glioma growth prediction. IEEE Trans. Med. Imaging 2025, 44, 2449–2462. [Google Scholar] [CrossRef]
- Corral-Acero, J.; et al. The `Digital Twin’ to enable the vision of precision cardiology. Eur. Heart J. 2020, 41, 4556–4564. [Google Scholar] [CrossRef] [PubMed]
- Kuang, K.; Dean, F.; Jedlicki, J.B.; Ouyang, D.; Philippakis, A.; Sontag, D.; Alaa, A. Med-real2sim: Non-invasive medical digital twins using physics-informed self-supervised learning. Adv. Neural Inf. Process. Syst. 2024, 37, 5757–5788. [Google Scholar]
- Builes-Montaño, C.E.; et al. A digital twin-enhanced decision support system improves time-in-range in type 1 diabetes: a randomized clinical trial. Sci. Rep. 2025, 15, 39738. [Google Scholar] [CrossRef]
- Kovatchev, B.P.; Colmegna, P.; Pavan, J.; Diaz Castañeda, J.L.; Villa-Tamayo, M.F.; Koravi, C.L.; Santini, G.; Alix, C.; Stumpf, M.; Brown, S.A. Human-machine co-adaptation to automated insulin delivery: a randomised clinical trial using digital twin technology. npj Digit. Med. 2025, 8, 253. [Google Scholar] [CrossRef] [PubMed]
- Mujahid, O.; et al. Generative deep learning for the development of a type 1 diabetes simulator. Commun. Med. 2024, 4, 51. [Google Scholar] [CrossRef]
- Salvador, M.; Strocchi, M.; Regazzoni, F.; Augustin, C.M.; Dede’, L.; Niederer, S.A.; Quarteroni, A. Whole-heart electromechanical simulations using latent neural ordinary differential equations. npj Digit. Med. 2024, 7, 90. [Google Scholar] [CrossRef]
- Qian, S.; et al. Developing cardiac digital twin populations powered by machine learning provides electrophysiological insights in conduction and repolarization. Nat. Cardiovasc. Res. 2025, 4, 624–636. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Deng, K.; Zhang, X.; Liu, G.; Guan, Y. Neural-ODE for Pharmacokinetics Modeling and Its Advantage to Alternative Machine Learning Models in Predicting New Dosing Regimens. iScience 2021, 24, 102804. [Google Scholar] [CrossRef]
- Mould, D.R.; Upton, R.N. Basic concepts in population modeling, simulation, and model-based drug development. CPT Pharmacomet. Syst. Pharmacol. 2012, 1, 1–14. [Google Scholar] [CrossRef]
- Xu, Q.; Habib, G.; Wu, F.; Perera, D.; Feng, M. MedDreamer: Model-Based Reinforcement Learning with Latent Imagination on Complex EHRs for Clinical Decision Support. arXiv preprint arXiv:2505.19785 2025. [Google Scholar]
- González, J.; Ueno, R.; Wong, C.; Gero, Z.; Bagga, J.; Chien, I.; Oravkin, E.; Kiciman, E.; Nori, A.; Weerasinghe, R.; et al. TRIALSCOPE—A framework for clinical trial simulation from real-world data. NEJM AI 2025, 2, AIoa2400859. [Google Scholar] [CrossRef]
- Koju, S.; Bastola, S.; Shrestha, P.; Amgain, S.; Shrestha, Y.R.; Poudel, R.P.; Bhattarai, B. Surgical vision world model. In Proceedings of the MICCAI Workshop on Data Engineering in Medical Imaging. Springer, 2025, pp. 1–10.
- He, Y.; Guo, P.; Xu, M.; Li, Z.; Myronenko, A.; Imans, D.; Liu, B.; Yang, D.; Gu, M.; Ji, Y.; et al. Cosmos-H-Surgical: Learning Surgical Robot Policies from Videos via World Modeling. arXiv preprint arXiv:2512.23162 2025. [Google Scholar]
- Chen, Z.; Xu, Q.; Wu, J.; Yang, B.; Zhai, Y.; Guo, G.; Zhang, J.; Ding, Y.; Navab, N.; Luo, J. How Far Are Surgeons from Surgical World Models? A Pilot Study on Zero-shot Surgical Video Generation with Expert Assessment. arXiv preprint arXiv:2511.01775 2025. [Google Scholar]
- Li, K.; Hopkins, A.K.; Bau, D.; Viégas, F.; Pfister, H.; Wattenberg, M. Emergent World Representations: Exploring a Sequence Model Trained on a Synthetic Task. ICLR 2023. [Google Scholar]
- Thirunavukarasu, A.J.; Ting, D.S.J.; Elangovan, K.; Gutierrez, L.; Tan, T.F.; Ting, D.S.W. Large language models in medicine. Nat. Med. 2023, 29, 1930–1940. [Google Scholar] [CrossRef]
- Hernán, M.A.; Robins, J.M. Using big data to emulate a target trial when a randomized trial is not available. Am. J. Epidemiol. 2016, 183, 758–764. [Google Scholar] [CrossRef]
- Li, R.; Hu, S.; Lu, M.; Utsumi, Y.; Chakraborty, P.; Sow, D.M.; Madan, P.; Li, J.; Ghalwash, M.; Shahn, Z.; et al. G-net: a recurrent network approach to g-computation for counterfactual prediction under a dynamic treatment regime. In Proceedings of the Machine Learning for Health. PMLR, 2021, pp. 282–299.
- Robins, J.M.; Hernan, M.A.; Brumback, B. Marginal structural models and causal inference in epidemiology. Epidemiology 2000, 11, 550–560. [Google Scholar] [CrossRef]
- Abdar, M.; et al. A Review of Uncertainty Quantification in Deep Learning: Techniques, Applications and Challenges. Inf. Fusion 2021, 76, 243–297. [Google Scholar] [CrossRef]
- Teo, Z.L.; et al. Generative Artificial Intelligence in Medicine. Nat. Med. 2025, 31, 3270–3282. [Google Scholar] [CrossRef]
- Rieke, N.; Hancox, J.; Li, W.; Milletari, F.; Roth, H.R.; Albarqouni, S.; Bakas, S.; Galtier, M.N.; Landman, B.A.; Maier-Hein, K.; et al. The future of digital health with federated learning. npj Digit. Med. 2020, 3, 119. [Google Scholar] [CrossRef] [PubMed]
- Obermeyer, Z.; Powers, B.; Vogeli, C.; Mullainathan, S. Dissecting Racial Bias in an Algorithm Used to Manage the Health of Populations. Science 2019, 366, 447–453. [Google Scholar] [CrossRef] [PubMed]
- Li, M.M.; Reis, B.Y.; Rodman, A.; Cai, T.; Dagan, N.; Balicer, R.D.; Loscalzo, J.; Kohane, I.S.; Zitnik, M. Scaling Medical AI across Clinical Contexts. Nat. Med. 2026, 32, 439–448. [Google Scholar] [CrossRef]
- Regulation (EU) 2017/745 of the European Parliament and of the Council on medical devices. Official Journal of the European Union, 2017.
- Regulation (EU) 2024/1689 of the European Parliament and of the Council laying down harmonised rules on artificial intelligence. Official Journal of the European Union, 2024.
- U.S. Food and Drug Administration. Marketing Submission Recommendations for a Predetermined Change Control Plan for Artificial Intelligence-Enabled Device Software Functions. FDA Guidance Document, 2024.
- Bica, I.; et al. From Real-World Patient Data to Individualized Treatment Effects Using Machine Learning: Current and Future Methods to Address Underlying Challenges. Clin. Pharmacol. Ther. 2021, 109, 87–100. [Google Scholar] [CrossRef]
| Level | Clinical question | Allowable claim | Minimum evidence | Representative system |
|---|---|---|---|---|
| L1 | What is the patient’s current state? | Patient state can be meaningfully compressed | Reconstruction fidelity on held-out data | Self-supervised imaging encoders |
| L2 | What will happen next? | Future trajectory can be forecast from history | Held-out temporal accuracy | Foresight, DT-GPT |
| L3a | What happens under treatment A? | Tr ajectory under a specified treatment is plausible | Held-out trajectory evaluation against observed outcomes in patients who received the indexed treatment | MeWM, T1D digital twin |
| L3b | How do treatments A and B compare? | Comparative treatment effect is credible | Trial emulation, causal inference, or randomized data | TRIALSCOPE (partial) |
| L4 | What treatment sequence is optimal? | Recommended strategy improves outcomes | Prospective randomized study | None yet validated |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




