Submitted:
29 April 2026
Posted:
30 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Global Colorectal Cancer Burden
1.2. Hypoxia-Associated Biological Framework
1.3. Study Rationale
1.4. Biological Basis of the Hypoxia-Associated Multimarker Panel
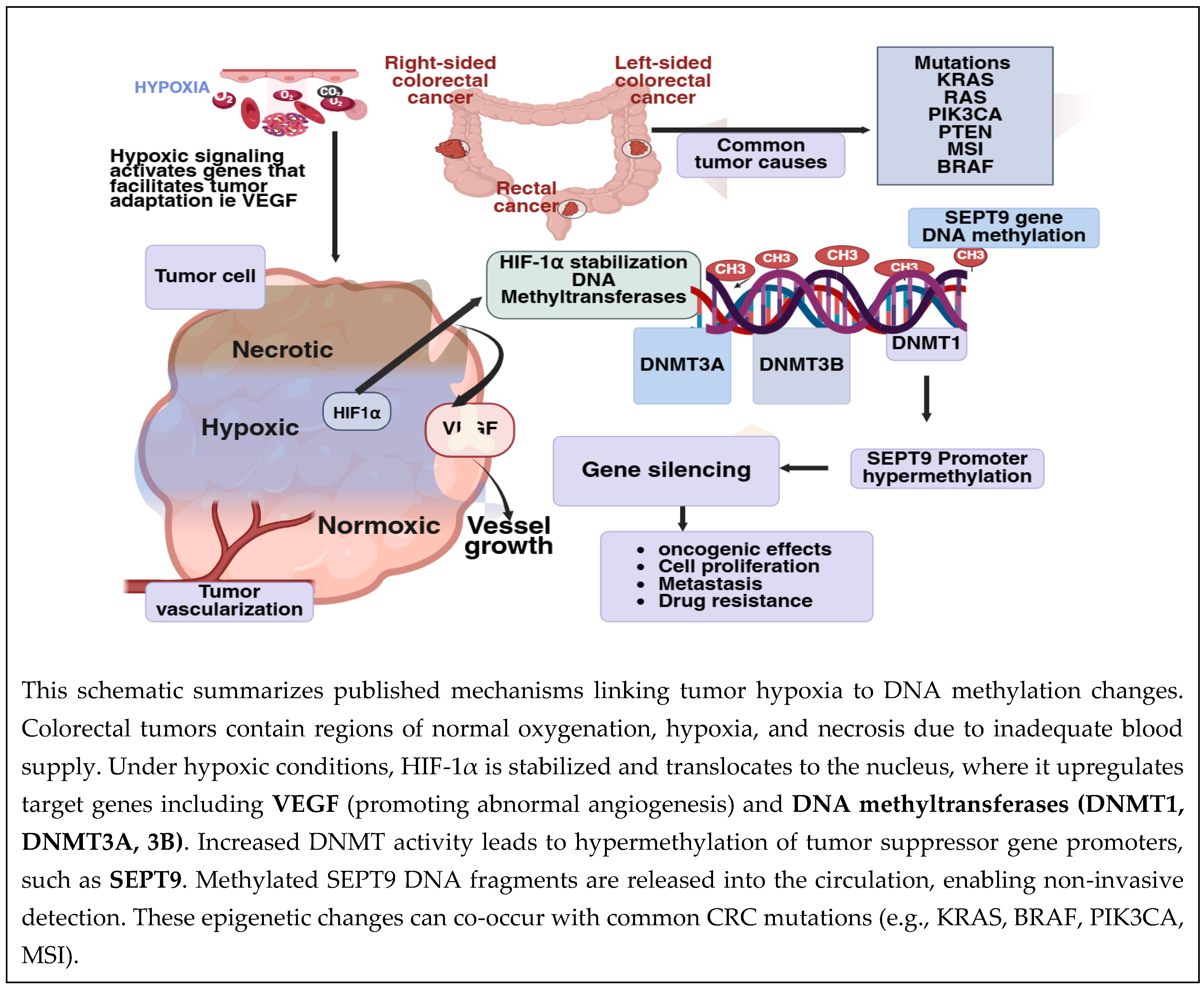
1.5. mSEPT9-Epigenetic Changes and Hypoxic Microenvironment in CRC Development
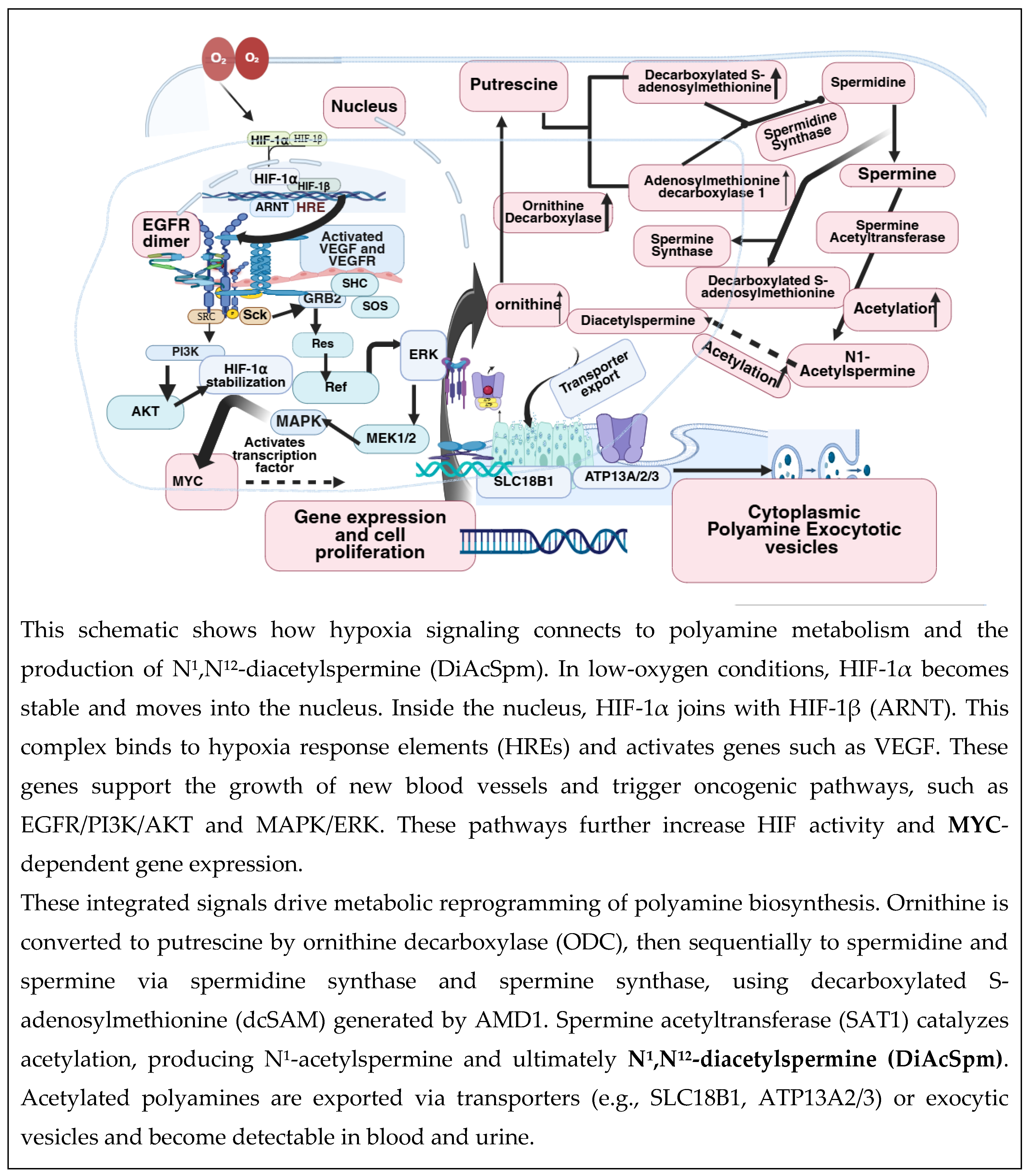
1.6. The Role of the Hypoxic Tumor Microenvironment in Enhancing DiAcSpm-Associated Metabolic Reprogramming
1.7. Neutrophil-to-Lymphocyte Ratio (NLR), Platelet-to-Lymphocyte Ratio (PLR), and Lymphocyte-to-Monocyte Ratio (LMR) Reflected In Tumor-Hypoxia Micorenvironment
Classical Serum Tumor Markers
2. Materials and Methods

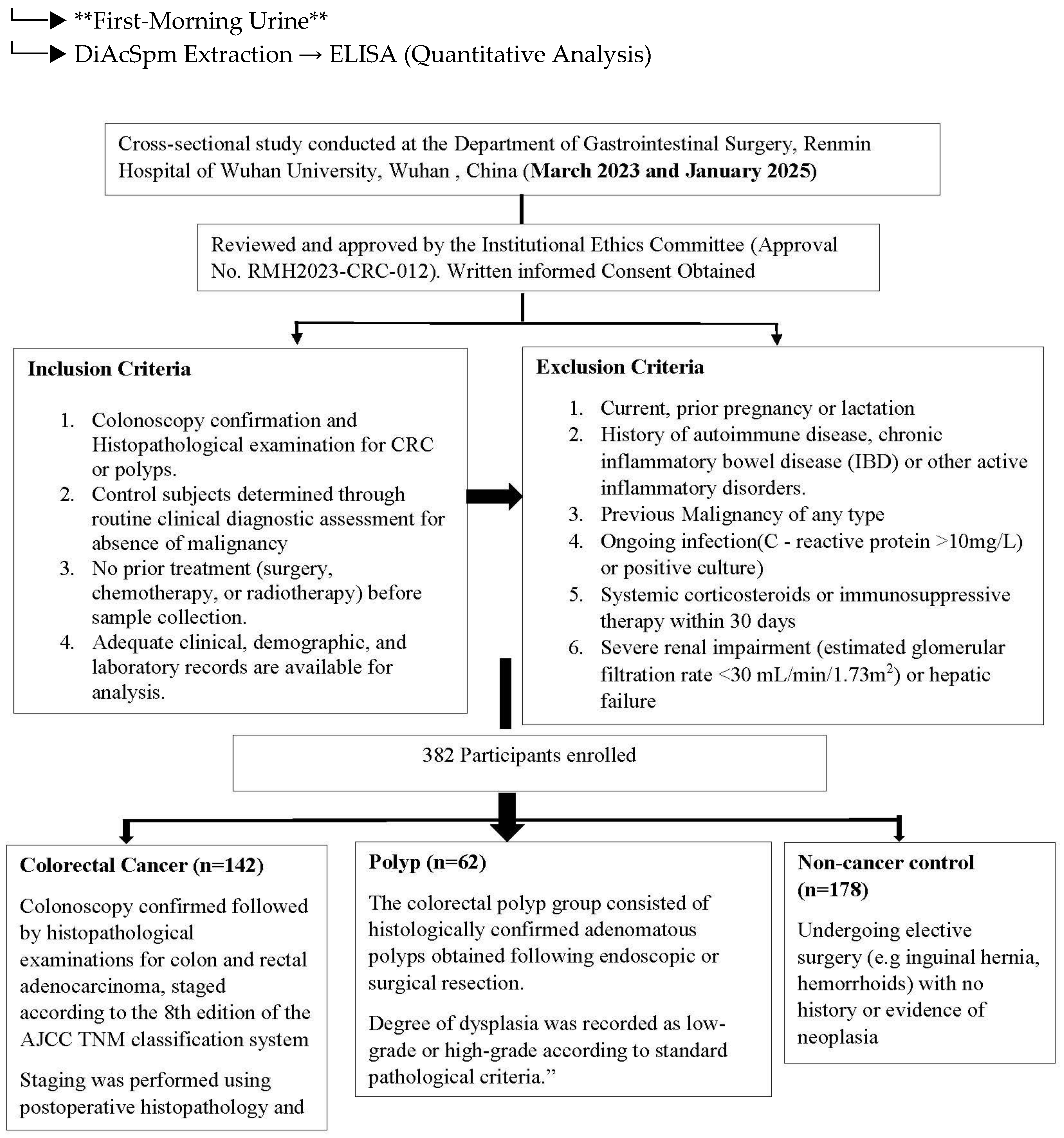
2.1. Study Design and Participants
2.2. Sample Processing and Storage
2.3. Detection and Quantification of Plasma Methylated SEPT9
2.4. Quantification of Urinary DiAcSpm by Competitive ELISA
2.5. Peripheral Inflammatory Blood Indices
2.6. Serum Tumor Marker Measurement
2.7. Determination of Diagnostic Cutoff Values
2.8. Statistical Analysis
3. Experimental Results and Analysis
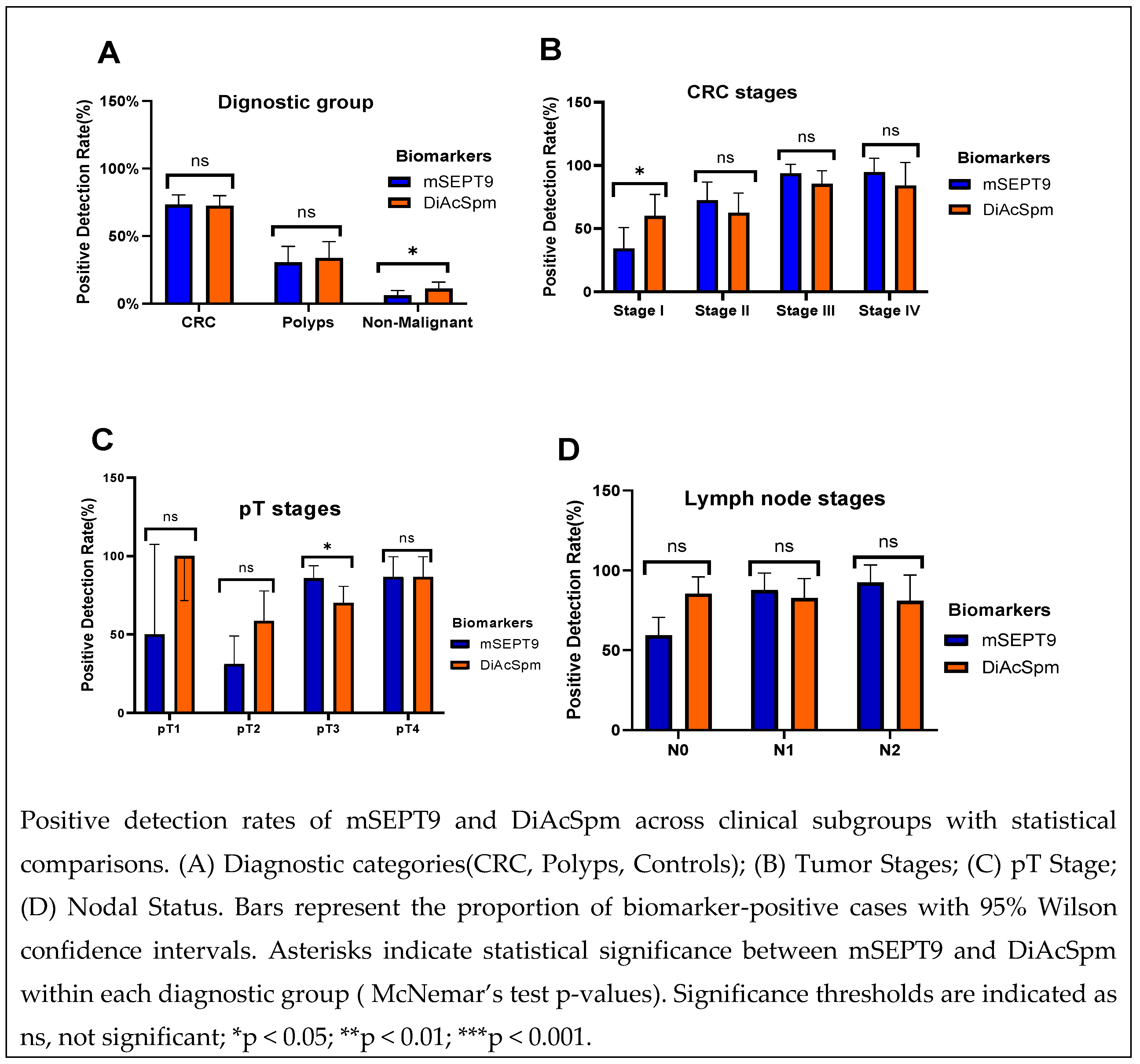
3.1. Diagnostic Yield Across Clinical Subgroups between mSEPT9 and DiAcSpm Biomarkers
3.2. Comparative Analysis of mSEPT9 and DiAcSpm Biomarker Positivity and Negativity Status: Clinicopathological Correlations
3.2. McNemar’s Test Comparison for Significance Between (mSEPT9 and DiAcSpm) Biomarkers in Different Clinical Subgroups
3.3. Comparison of mSEPT9 and DiAcSpm Biomarker Positivity and Negativity Status Correlating with Inflammatory Indices
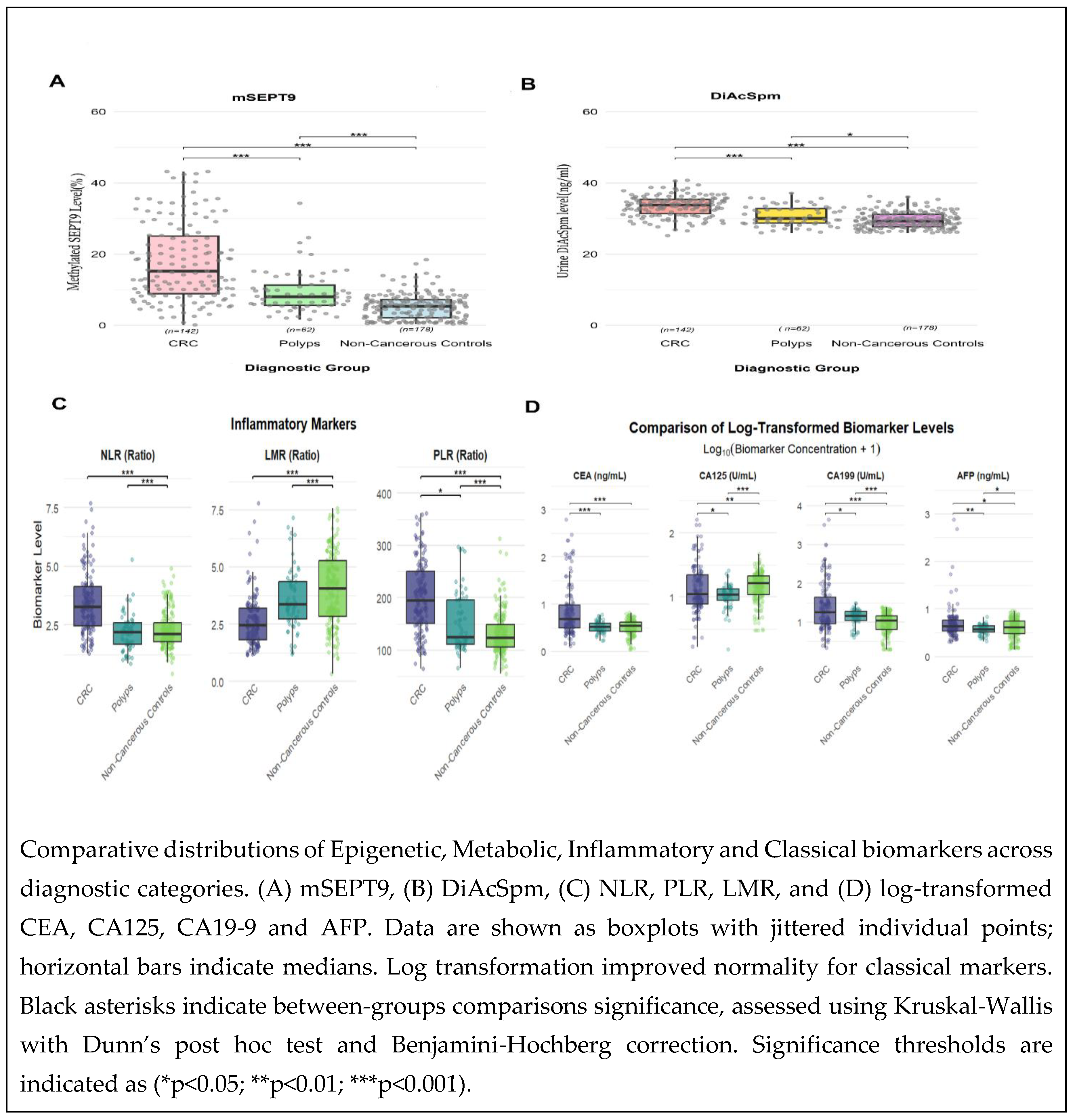
3.4. Comparative Distributions of mSEPT9, DiAcSpm, Inflammatory, and Classical Biomarkers Across Diagnostic Categories
3.5. Comparison of Classical Tumor Markers and Inflammatory Indices According to Clinico-Pathological Characteristics
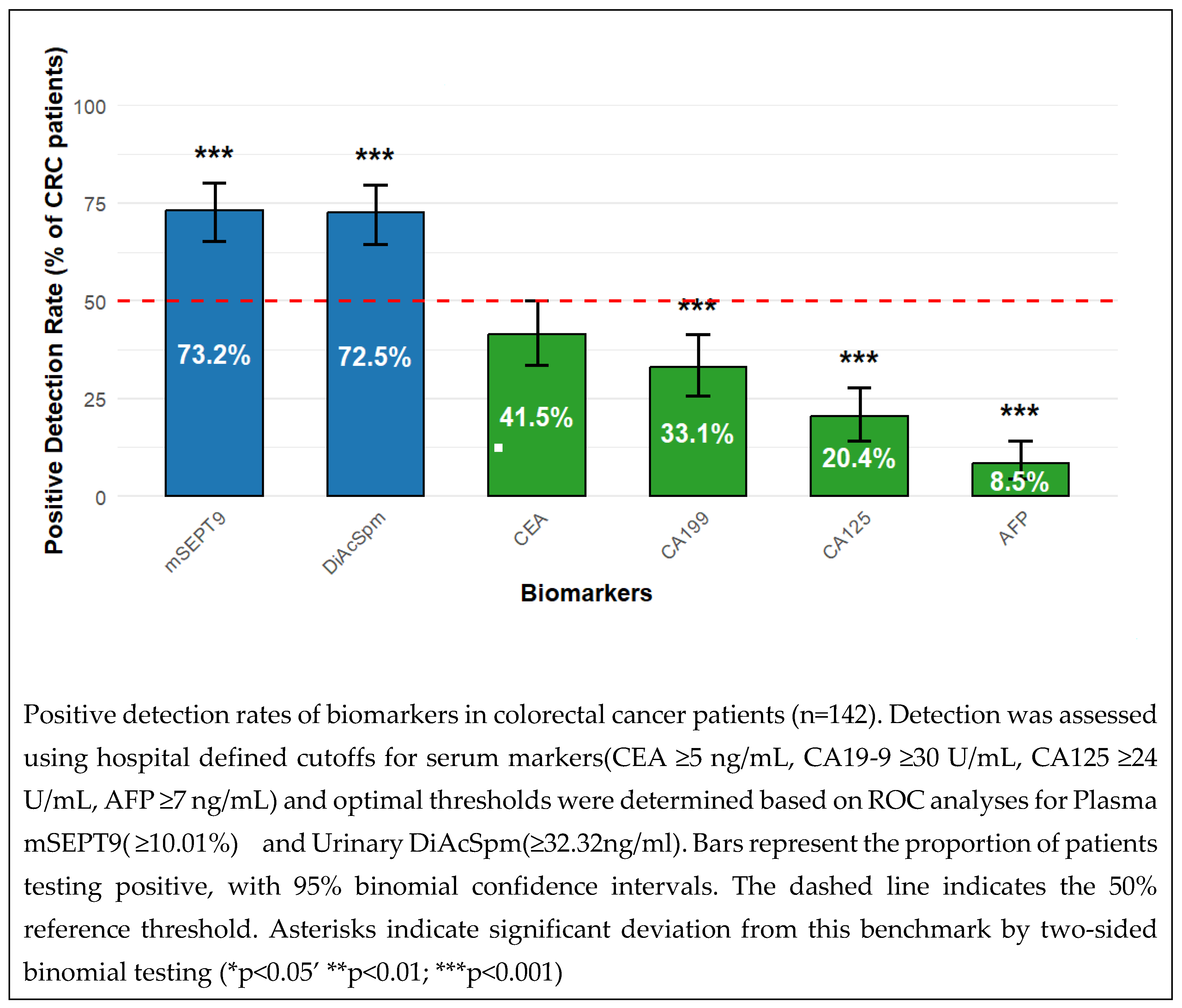
3.6. Positive Detection Rates Across Conventional Biomarkers, mSEPT9, and DiAcSpm
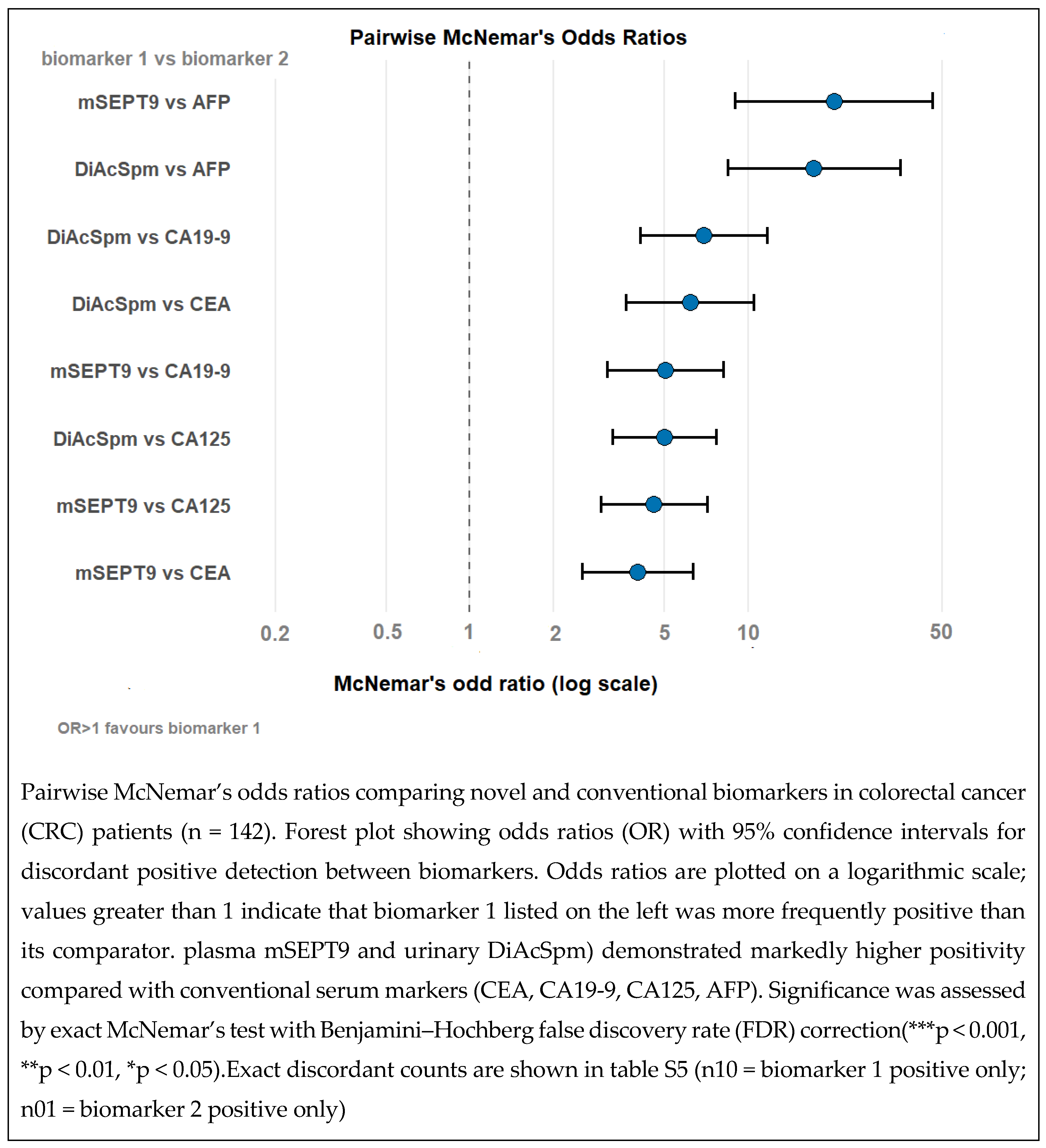
3.7. Pairwise Comparison of Biomarkers for Assessing Detection Performance for Colorectal Cancer Using McNemar’s Test
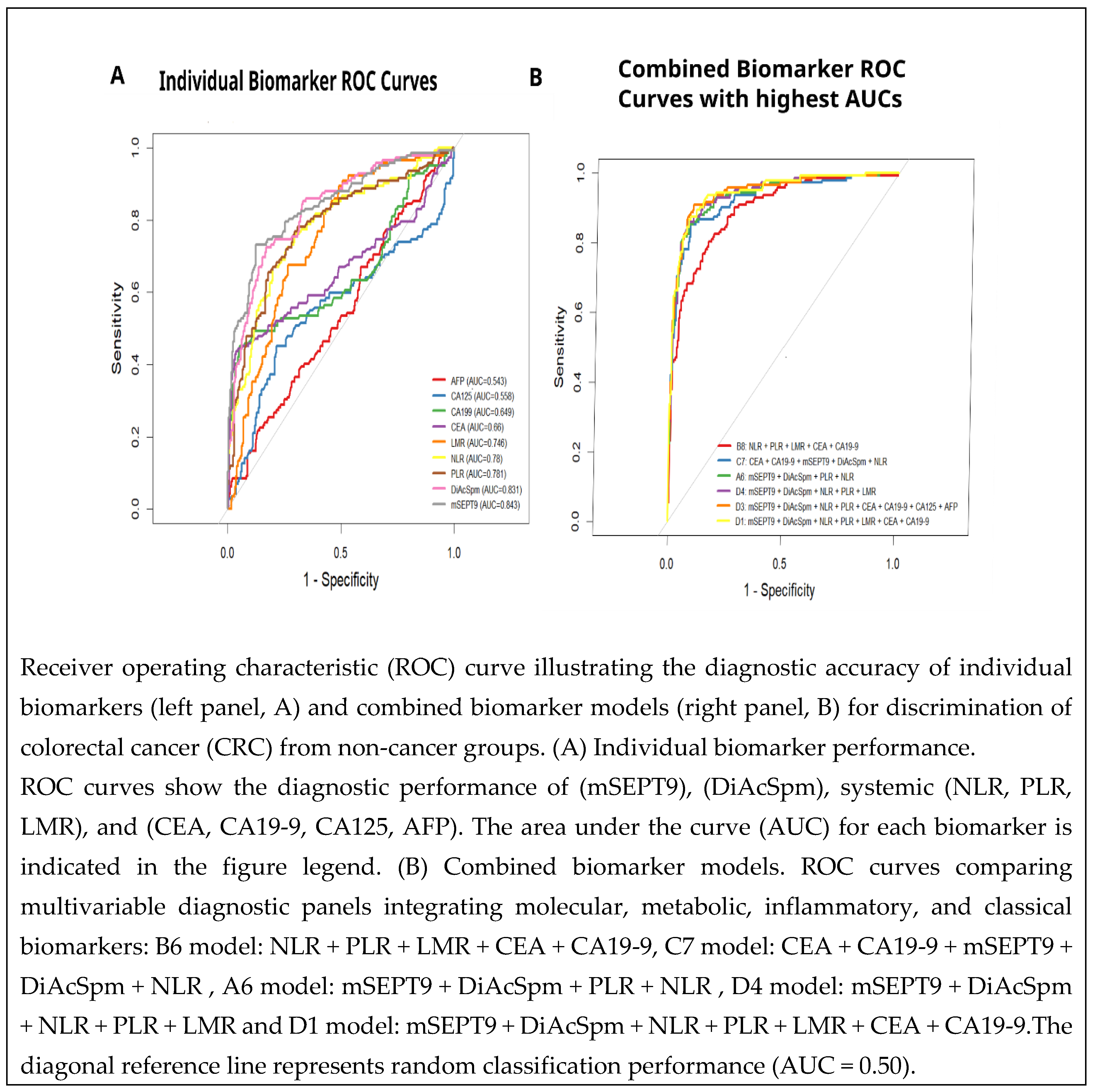
3.8. ROC Analysis and Diagnostic Performance of Individual and Multimarker Panels
3.9. Performance of Individual Biomarkers
3.10. Performance of Multimarker Models
3.11. Logistic Regression Analysis
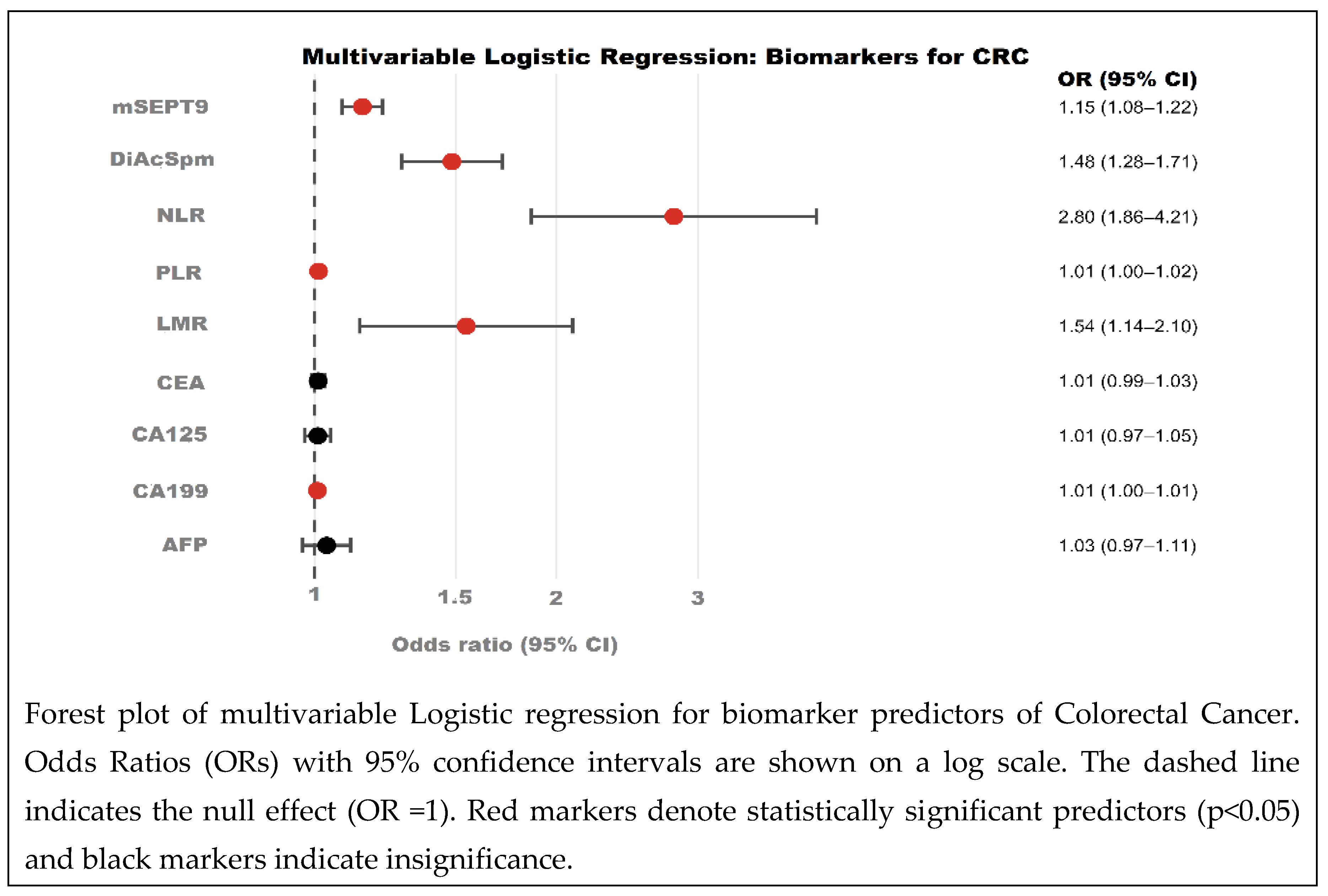
3.12. Multivariable Risk Association Analysis for Colorectal Cancer
3.13. Internal Validation of the Integrated Diagnostic Model
3.14. Delong’s test and McNemar’s test for Comparison of Optimized Multimarker Panels
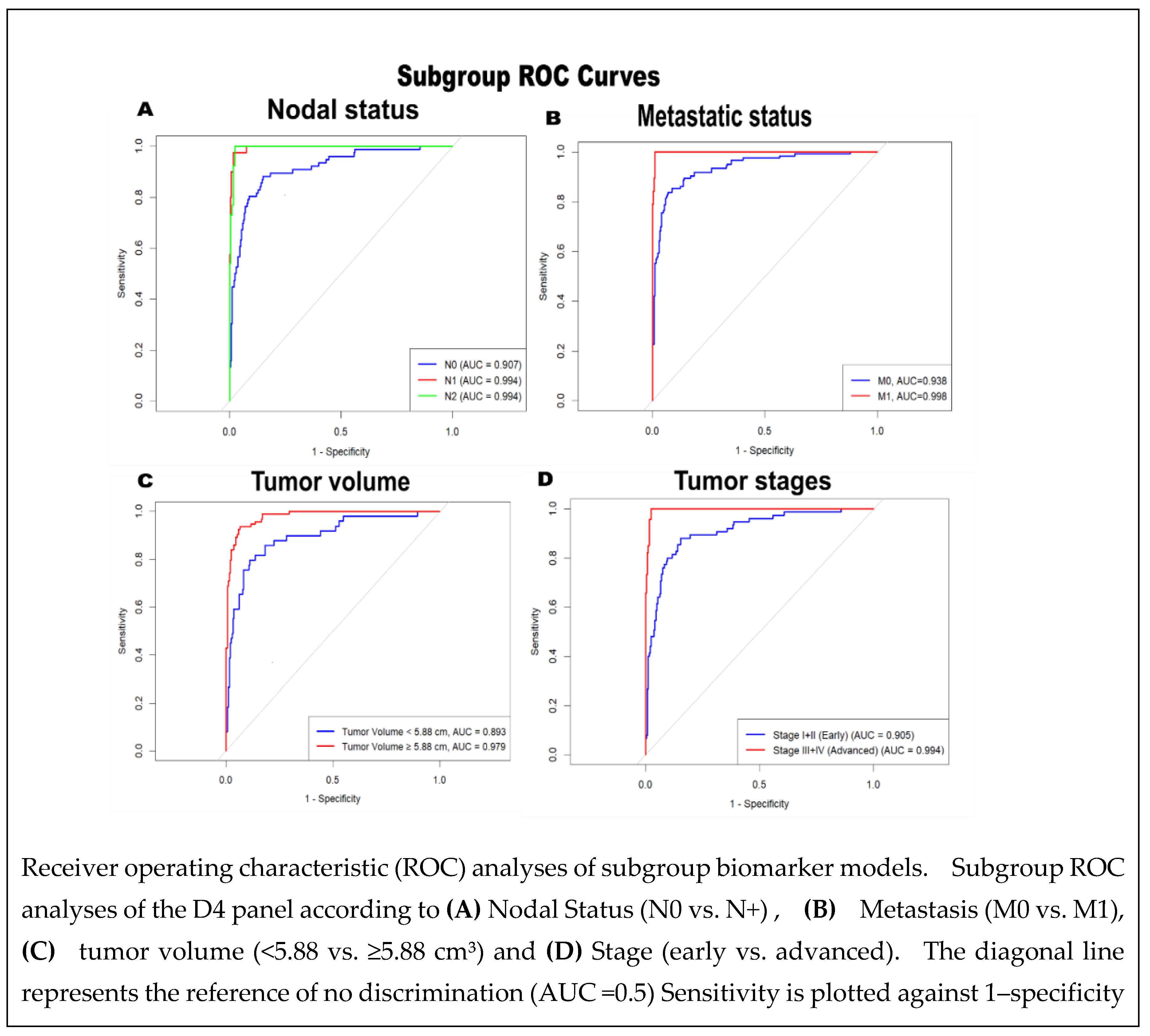
3.15. Subgroup Performance of the Optimized D4 Multimarker Model to determine robustness
4. Discussion
4.1. Study Limitations
4.2. Clinical Implications
4.3. Future Directions
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Use of Artificial Intelligence
Conflicts of Interest
Abbreviations and Full Meaning
| mSEPT9 | Methylated septin9 gene |
| DiAcSpm | N¹, N¹²-diacetylspermine |
| CEA | Carcinoembryonic antigen |
| CA19-9 | Carbohydrate antigen 19 − 9 |
| CA125 | Carbohydrate antigen 125 |
| PLT | Platelet |
| NLR | Neutrophil-lymphocyte ratio |
| PLR | Platelet-lymphocyte ratio |
| ROC | Receiver operating characteristic |
| AUC | Area under the ROC curve |
| PCR | polymerase chain reaction |
| LC–MS/MS | liquid chromatography–tandem mass spectrometry |
| ELISA | enzyme-linked immunosorbent assay |
| DNMTs | DNA methyltransferases |
| ODC | ornithine decarboxylase |
| SAT1 | spermine N¹-acetyltransferas |
| VEGF | vascular endothelial growth factor |
| EGFR | epidermal growth factor |
| dcSAM | decarboxylated S-adenosylmethionine |
| AMD1 | adenosylmethionine decarboxylase 1 |
| HRP | Horseradish peroxidase |
| TMB | (3, 3’, 5, 5’-Tetramethylbenzidine) |
| HIF | Hypoxia-Inducible Factor |
| C-MYC | A transcription factor that regulates genes involved in cell growth |
| FIT | Fecal Immunochemical Test |
| HCT | Hematocrit |
References
- Morgan, E.; Arnold, M.; Gini, A.; Lorenzoni, V.; Cabasag, C.J.; Laversanne, M.; et al. Global burden of colorectal cancer in 2020 and 2040: incidence and mortality estimates from GLOBOCAN. Gut 2023, 72(2), 338–44. [Google Scholar] [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA A Cancer J. Clin. 2021, 71(3), 209–49. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.Z.; Lengyel, C.G. Challenges in the management of colorectal cancer in low-and middle-income countries. Cancer Treat. Res. Commun. 2023, 35, 100705. [Google Scholar] [CrossRef]
- Abreu Lopez, B.A.; Pinto-Colmenarez, R.; Caliwag, F.M.C.; Ponce-Lujan, L.; Fermin, M.D.; Granillo Cortés, A.V.; et al. Colorectal cancer screening and management in low-and middle-income countries and high-income countries: a narrative review. Cureus 2024, 16(10), e70933. [Google Scholar] [CrossRef] [PubMed]
- Schliemann, D.; Ramanathan, K.; Matovu, N.; O’Neill, C.; Kee, F.; Su, T.T.; et al. The implementation of colorectal cancer screening interventions in low-and middle-income countries: a scoping review. Bmc Cancer 2021, 21(1), 1125. [Google Scholar] [CrossRef]
- Zhang, T.; Guo, Y.; Qiu, B.; Dai, X.; Wang, Y.; Cao, X. Global, regional, and national trends in colorectal cancer burden from 1990 to 2021 and projections to 2040. Front. Oncol. 2025, 14, 1466159. [Google Scholar] [CrossRef]
- Siegel, R.L.; Wagle, N.S.; Cercek, A.; Smith, R.A.; Jemal, A. Colorectal cancer statistics, 2023. CA: a cancer journal for clinicians 2023, 73(3), 233–54. [Google Scholar] [CrossRef]
- Deo, S.; Kumar, S.; Bhoriwal, S.; Shukla, N.; Sharma, A.; Thulkar, S.; et al. Colorectal cancers in low-and middle-income countries—demographic pattern and clinical profile of 970 patients treated at a tertiary care cancer center in India. JCO Glob. Oncol. 2021, 7, 1110–5. [Google Scholar] [CrossRef]
- Chen, X.; Dong, X.; Zheng, Y.; Wang, C.; Luo, Z.; Xie, J.; et al. Global trajectories of colorectal cancer burden from 1990 to 2021 and projection to 2040. Cancer Innov. 2025, 4(4), e70020. [Google Scholar] [CrossRef] [PubMed]
- Le, D.T.P.; Pham, T.D. Colorectal cancer worldwide: epidemiological trends, economic burden, and the promise of AI-driven solutions. Explor. Med. 2025, 6, 1001371. [Google Scholar] [CrossRef]
- Peterse, E.F.; Meester, R.G.; De Jonge, L.; Omidvari, A.-H.; Alarid-Escudero, F.; Knudsen, A.B.; et al. Comparing the cost-effectiveness of innovative colorectal cancer screening tests. JNCI J. Natl. Cancer Inst. 2021, 113(2), 154–61. [Google Scholar] [CrossRef] [PubMed]
- Verguet, S.; Alkire, B.C.; Bickler, S.W.; Lauer, J.A.; Uribe-Leitz, T.; Molina, G.; et al. Timing and cost of scaling up surgical services in low-income and middle-income countries from 2012 to 2030: a modelling study. Lancet Glob. Health 2015, 3, S28–S37. [Google Scholar] [CrossRef]
- Moradoghli, F.; Aghaei, M.H.; Hakimi, M.H.; Ghadimi, S.; Ebrahimoghli, R. Uptake of Colorectal Cancer Screening in Low- and Middle-Income Countries: a Systematic Review and Meta-analysis. J. Gastrointest. Cancer 2025, 56(1), 154. [Google Scholar] [CrossRef]
- Mannucci, A.; Goel, A. Stool and blood biomarkers for colorectal cancer management: an update on screening and disease monitoring. Mol. Cancer 2024, 23(1), 259. [Google Scholar] [CrossRef]
- Li, T.; Mao, C.; Wang, X.; Shi, Y.; Tao, Y. Epigenetic crosstalk between hypoxia and tumor driven by HIF regulation. J. Exp. Clin. Cancer Res. 2020, 39(1), 224. [Google Scholar] [CrossRef]
- Silina, M.; Dzhalilova, D.S.; Makarova, O. The Role of Hypoxia and HIF Transcription Factors in the Development of Ulcerative Colitis and Associated Colorectal Cancer. Mosc. Univ. Biol. Sci. Bull. 2025, 80(2), 52–65. [Google Scholar] [CrossRef]
- Chen, Z.; Han, F.; Du, Y.; Shi, H.; Zhou, W. Hypoxic microenvironment in cancer: molecular mechanisms and therapeutic interventions. Signal Transduct. Target. Ther. 2023, 8(1), 70. [Google Scholar] [CrossRef]
- Magar, A.G.; Morya, V.K.; Kwak, M.K.; Oh, J.U.; Noh, K.C. A molecular perspective on HIF-1α and angiogenic stimulator networks and their role in solid tumors: an update. Int. J. Mol. Sci. 2024, 25(6), 3313. [Google Scholar] [CrossRef]
- Muz, B.; de la Puente, P.; Azab, F.; Kareem Azab, A. The role of hypoxia in cancer progression, angiogenesis, metastasis, and resistance to therapy. Hypoxia 2015, 3(null), 83–92. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Hu, Y.; Yang, F.; Zhang, J.; Zhang, J.; Yu, W.; et al. Interaction between AhR and HIF-1 signaling pathways mediated by ARNT/HIF-1β. BMC Pharmacol. Toxicol. 2022, 23(1), 26. [Google Scholar] [CrossRef] [PubMed]
- D’Ignazio, L.; Batie, M.; Rocha, S. Hypoxia and inflammation in cancer, focus on HIF and NF-κB. Biomedicines 2017, 5(2), 21. [Google Scholar] [CrossRef]
- Bigos, K.; Quiles, C.; Lunj, S.; Smith, D.; Krause, M.; Troost, E. Tumour response to hypoxia: understanding the hypoxic tumour microenvironment to improve treatment outcome in solid tumours. Front Oncol. 2024, 14, 1331355. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, T.; Thompson, A.J.; Ashrafian, H.; Darzi, A. The measurement and modification of hypoxia in colorectal cancer: overlooked but not forgotten. Gastroenterol. Rep. 2022, 10, goac042. [Google Scholar] [CrossRef]
- Imamura, T.; Kikuchi, H.; Herraiz, M.T.; Park, D.Y.; Mizukami, Y.; Mino-Kenduson, M.; et al. HIF-1α and HIF-2α have divergent roles in colon cancer. Int. J. Cancer 2009, 124(4), 763–71. [Google Scholar] [CrossRef] [PubMed]
- Vadde, R.; Vemula, S.; Jinka, R.; Merchant, N.; Bramhachari, P.V.; Nagaraju, G.P. Role of hypoxia-inducible factors (HIF) in the maintenance of stemness and malignancy of colorectal cancer. Crit. Rev. Oncol. 2017, 113, 22–7. [Google Scholar] [CrossRef]
- Ben Lassoued, A.; Beaufils, N.; Dales, J.-P.; Gabert, J. Hypoxia-inducible factor-1α as prognostic marker. Expert Opin. Med. Diagn. 2013, 7(1), 53–70. [Google Scholar] [CrossRef] [PubMed]
- Tatum, J.L. Hypoxia: importance in tumor biology, noninvasive measurement by imaging, and value of its measurement in the management of cancer therapy. Int. J. Radiat. Biol. 2006, 82(10), 699–757. [Google Scholar] [CrossRef]
- McSorley, S.T.; Khor, B.Y.; Tsang, K.; Colville, D.; Han, S.; Horgan, P.G.; et al. The relationship between 18F-FDG-PETCT-derived markers of tumour metabolism and systemic inflammation in patients with recurrent disease following surgery for colorectal cancer. Colorectal Dis. 2018, 20(5), 407–15. [Google Scholar] [CrossRef]
- Gonzalez-Jaramillo, V.; Portilla-Fernandez, E.; Glisic, M.; Voortman, T.; Ghanbari, M.; Bramer, W.; et al. Epigenetics and inflammatory markers: a systematic review of the current evidence. Int. J. Inflamm. 2019, 2019(1), 6273680. [Google Scholar] [CrossRef]
- Deris Zayeri, Z.; Parsi, A.; Shahrabi, S.; Kargar, M.; Davari, N.; Saki, N. Epigenetic and metabolic reprogramming in inflammatory bowel diseases: diagnostic and prognostic biomarkers in colorectal cancer. Cancer Cell. Int. 2023, 23(1), 264. [Google Scholar] [CrossRef]
- Nicolini, A.; Ferrari, P. Involvement of tumor immune microenvironment metabolic reprogramming in colorectal cancer progression, immune escape, and response to immunotherapy. Front. Immunol. 2024, 15, 1353787. [Google Scholar] [CrossRef] [PubMed]
- Loomans-Kropp, H.A.; Song, Y.; Gala, M.; Parikh, A.R.; Van Seventer, E.E.; Alvarez, R.; et al. Methylated Septin9 (m SEPT9): a promising blood-based biomarker for the detection and screening of early-onset colorectal cancer. Cancer Res. Commun. 2022, 2(2), 90–8. [Google Scholar] [CrossRef]
- Xie, L.; Jiang, X.; Li, Q.; Sun, Z.; Quan, W.; Duan, Y.; et al. Diagnostic value of methylated Septin9 for colorectal cancer detection. Front. Oncol. 2018, 8, 247. [Google Scholar] [CrossRef] [PubMed]
- Nian, J.; Sun, X.; Ming, S.; Yan, C.; Ma, Y.; Feng, Y.; et al. Diagnostic accuracy of methylated SEPT9 for blood-based colorectal cancer detection: a systematic review and meta-analysis. Clin. Transl. Gastroenterol. 2017, 8(1), e216. [Google Scholar] [CrossRef]
- Wasserkort, R.; Kalmar, A.; Valcz, G.; Spisak, S.; Krispin, M.; Toth, K.; et al. Aberrant septin 9 DNA methylation in colorectal cancer is restricted to a single CpG island. BMC Cancer 2013, 13(1), 398. [Google Scholar] [CrossRef] [PubMed]
- Shahrzad, S.; Bertrand, K.; Minhas, K.; Coomber, B. Induction of DNA hypomethylation by tumor hypoxia. Epigenetics 2007, 2(2), 119–25. [Google Scholar] [CrossRef]
- Bhandari, P.N.; Cui, Y.; Elzey, B.D.; Goergen, C.J.; Long, C.M.; Irudayaraj, J. Oxygen nanobubbles revert hypoxia by methylation programming. Sci. Rep. 2017, 7(1), 9268. [Google Scholar] [CrossRef]
- Scanlon, S.E.; Glazer, P.M. Multifaceted control of DNA repair pathways by the hypoxic tumor microenvironment. DNA repair. 2015, 32, 180–9. [Google Scholar] [CrossRef]
- Han, S.; Huang, T.; Li, W.; Liu, S.; Yang, W.; Shi, Q.; et al. Association between hypoxia-inducible factor-2α (HIF-2α) expression and colorectal cancer and its prognostic role: a systematic analysis. Cell. Physiol. Biochem. 2018, 48(2), 516–27. [Google Scholar] [CrossRef]
- Thienpont, B.; Steinbacher, J.; Zhao, H.; D’Anna, F.; Kuchnio, A.; Ploumakis, A.; et al. Tumour hypoxia causes DNA hypermethylation by reducing TET activity. Nature 2016, 537(7618), 63–8. [Google Scholar] [CrossRef]
- Huang, H.; Li, Q.; Tu, X.; Yu, D.; Zhou, Y.; Ma, L.; et al. DNA hypomethylation patterns and their impact on the tumor microenvironment in colorectal cancer. Cell. Oncol. 2024, 47(4), 1375–89. [Google Scholar] [CrossRef]
- Yang, X.; Chen, Q.; Yu, Y.; Zang, Y.; Zheng, R.; Zhang, Z.; et al. Oxidative Stress and DNA Epigenetic Modifications in Cancer: Mechanisms and Targeted Therapeutics. MedComm–Oncology 2026, 5(2), e70065. [Google Scholar] [CrossRef]
- Byrne, M.B.; Leslie, M.T.; Gaskins, H.R.; Kenis, P.J. Methods to study the tumor microenvironment under controlled oxygen conditions. Trends Biotechnol. 2014, 32(11), 556–63. [Google Scholar] [CrossRef]
- D’anna, F.; Van Dyck, L.; Xiong, J.; Zhao, H.; Berrens, R.V.; Qian, J.; et al. DNA methylation repels binding of hypoxia-inducible transcription factors to maintain tumor immunotolerance. Genome Biol. 2020, 21(1), 182. [Google Scholar] [CrossRef]
- Payne, S.R. From discovery to the clinic: the novel DNA methylation biomarker m SEPT9 for the detection of colorectal cancer in blood. Epigenomics 2010, 2(4), 575–85. [Google Scholar] [CrossRef] [PubMed]
- Devos, T.; Tetzner, R.; Model, F.; Weiss, G.; Schuster, M.; Distler, J.; et al. Circulating methylated SEPT9 DNA in plasma is a biomarker for colorectal cancer. Clin. Chem. 2009, 55(7), 1337–46. [Google Scholar] [CrossRef]
- Lin, K.W. mSEPT9 blood test (Epi proColon) for colorectal cancer screening. Am. Fam. Physician 2019, 100(1), 10–1. [Google Scholar] [PubMed]
- Wang, Y.; Chen, P.-M.; Liu, R.-B. Advance in plasma SEPT9 gene methylation assay for colorectal cancer early detection. World J. Gastrointest. Oncol. 2018, 10(1), 15. [Google Scholar] [CrossRef]
- Abi Zamer, B.; Abumustafa, W.; Hamad, M.; Maghazachi, A.A.; Muhammad, J.S. Genetic mutations and non-coding RNA-based epigenetic alterations mediating the warburg effect in colorectal carcinogenesis. Biology 2021, 10(9), 847. [Google Scholar] [CrossRef]
- Gong, L.; Zhang, H.; Liu, Y.; Wang, X.; Xia, R. Interactions between non-coding RNAs and HIF-1alpha in the context of colorectal cancer. Biomolecules 2025, 15(4), 510. [Google Scholar] [CrossRef] [PubMed]
- Nimer, R.M.; Arjah, S.; Obeidat, M.; Jaradat, S.A.; Zenati, R.A.; Bustanji, Y.; et al. Untargeted LC-MS/MS- based metabolomics profiling of colorectal cancer cell lines reveals potential hypoxia-associated biomarkers. J. Pharm. Biomed. Anal. 2025, 263, 116912. [Google Scholar] [CrossRef]
- Gong, L.; Zou, C.; Zhang, H.; Yang, F.; Qi, G.; Ma, Z. Landscape of noncoding RNA in the hypoxic tumor microenvironment. Genes. 2025, 16(2), 140. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Zou, S.; Ding, J.; Gao, C.; Wang, J.; Tang, Z. Crosstalk between dysregulated amino acid sensing and glucose and lipid metabolism in colorectal cancer. Front. Oncol. 2025, 15, 1665056. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Sun, X.-X.; Qian, D.Z.; Dai, M.-S. Molecular crosstalk between MYC and HIF in cancer. Front. Cell. Dev. Biol. 2020, 8, 590576. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Xue, M.; Chung, D.C. c-Myc is regulated by HIF-2α in chronic hypoxia and influences sensitivity to 5-FU in colon cancer. Oncotarget 2016, 7(48), 78910. [Google Scholar] [CrossRef]
- Guo, Y.; Ye, Q.; Deng, P.; Cao, Y.; He, D.; Zhou, Z.; et al. Spermine synthase and MYC cooperate to maintain colorectal cancer cell survival by repressing Bim expression. Nat. Commun. 2020, 11(1), 3243. [Google Scholar] [CrossRef]
- Mahalingam, S.S.; Pandiyan, P. Polyamines: key players in immunometabolism and immune regulation. J. Cell. Immunol. 2024, 6(5), 196–208. [Google Scholar] [CrossRef]
- Kobayashi, M.; Samejima, K.; Hiramatsu, K.; Kawakita, M. Mass spectrometric separation and determination of N1, N12-diacetylspermine in the urine of cancer patients. Biol. Pharm. Bull. 2002, 25(3), 372–4. [Google Scholar] [CrossRef]
- Erben, V.; Bhardwaj, M.; Schrotz-King, P.; Brenner, H. Metabolomics biomarkers for detection of colorectal neoplasms: a systematic review. Cancers 2018, 10(8), 246. [Google Scholar] [CrossRef]
- Castillo-Rodríguez, R.A.; Trejo-Solís, C.; Cabrera-Cano, A.; Gómez-Manzo, S.; Dávila-Borja, V.M. Hypoxia as a modulator of inflammation and immune response in cancer. Cancers 2022, 14(9), 2291. [Google Scholar] [CrossRef]
- Korbecki, J.; Kojder, K.; Kapczuk, P.; Kupnicka, P.; Gawrońska-Szklarz, B.; Gutowska, I.; et al. The effect of hypoxia on the expression of CXC chemokines and CXC chemokine receptors—a review of literature. Int. J. Mol. Sci. 2021, 22(2), 843. [Google Scholar] [CrossRef]
- Minici, R.; Siciliano, M.A.; Ammendola, M.; Santoro, R.C.; Barbieri, V.; Ranieri, G.; et al. Prognostic role of neutrophil-to-lymphocyte ratio (NLR), lymphocyte-to-monocyte ratio (LMR), platelet-to-lymphocyte ratio (PLR) and lymphocyte-to-C reactive protein ratio (LCR) in patients with hepatocellular carcinoma (HCC) undergoing chemoembolizations (TACE) of the liver: the unexplored corner linking tumor microenvironment, biomarkers and interventional radiology. Cancers 2022, 15(1), 257. [Google Scholar]
- Gawiński, C.; Michalski, W.; Mróz, A.; Wyrwicz, L. Correlation between lymphocyte-to-monocyte ratio (LMR), neutrophil-to-lymphocyte ratio (NLR), platelet-to-lymphocyte ratio (PLR) and tumor-infiltrating lymphocytes (TILs) in left-sided colorectal cancer patients. Biology 2022, 11(3), 385. [Google Scholar] [CrossRef]
- Maloney, S.; Pavlakis, N.; Itchins, M.; Arena, J.; Mittal, A.; Hudson, A.; et al. The Prognostic and Predictive Role of the Neutrophil-to-Lymphocyte Ratio (NLR), Platelet-to-Lymphocyte Ratio (PLR), and Lymphocyte-to-Monocyte Ratio (LMR) as Biomarkers in Resected Pancreatic Cancer. J. Clin. Med. [Internet] 2023, 12(5). [Google Scholar] [CrossRef]
- Xing, Y.; Qin, F.; Zhai, Y.; Yang, J.; Yan, Y.; Li, D.; et al. Association of Clinical Features of Colorectal Cancer with Circulating Tumor Cells and Systemic Inflammatory Markers. Dis. Markers 2022, 2022(1), 5105599. [Google Scholar] [CrossRef]
- Talebi, V.; Hashemi, M.G.; Ghazanfari, R.; Tabrizi, M.; Saleh, M.; Saatian, M. Association of neutrophil to lymphocyte ratio and platelet to lymphocyte ratio with tumoral differentiation in colorectal cancer. Gene Rep. 2022, 29, 101701. [Google Scholar] [CrossRef]
- Park, J.W.; Chang, H.J.; Yeo, H.Y.; Han, N.; Kim, B.C.; Kong, S.-Y.; et al. The relationships between systemic cytokine profiles and inflammatory markers in colorectal cancer and the prognostic significance of these parameters. Br. J. Cancer 2020, 123(4), 610–8. [Google Scholar] [CrossRef] [PubMed]
- Nøst, T.H.; Alcala, K.; Urbarova, I.; Byrne, K.S.; Guida, F.; Sandanger, T.M.; et al. Systemic inflammation markers and cancer incidence in the UK Biobank. Eur. J. Epidemiol. 2021, 36(8), 841–8. [Google Scholar] [CrossRef]
- Castillo-Rodríguez, R.A.; Trejo-Solís, C.; Cabrera-Cano, A.; Gómez-Manzo, S.; Dávila-Borja, V.M. Hypoxia as a Modulator of Inflammation and Immune Response in Cancer. Cancers [Internet] 2022, 14(9), 2291 p. [Google Scholar] [CrossRef] [PubMed]
- Taylor, C.T.; Doherty, G.; Fallon, P.G.; Cummins, E.P. Hypoxia-dependent regulation of inflammatory pathways in immune cells. J. Clin. Investig. 2016, 126(10), 3716–24. [Google Scholar] [CrossRef]
- Zhou, Y.; Tao, L.; Qiu, J.; Xu, J.; Yang, X.; Zhang, Y.; et al. Tumor biomarkers for diagnosis, prognosis and targeted therapy. Signal Transduct. Target. Ther. 2024, 9(1), 132. [Google Scholar] [CrossRef]
- Berger, A.C.; Sigurdson, E.R.; LeVoyer, T.; Hanlon, A.; Mayer, R.J.; Macdonald, J.S.; et al. Colon Cancer Survival Is Associated With Decreasing Ratio of Metastatic to Examined Lymph Nodes. J. Clin. Oncol. 23(34), 8706–12. [CrossRef]
- Park, H.C. Clinical Significance of the Metastatic Lymph-Node Ratio in Rectal Cancer. Ann. Coloproctol. 2013, 29(3), 89–90. [Google Scholar] [CrossRef] [PubMed]
- Semaan, A.; van Ellen, A.; Meller, S.; Bergheim, D.; Branchi, V.; Lingohr, P.; et al. SEPT9 and SHOX2 DNA methylation status and its utility in the diagnosis of colonic adenomas and colorectal adenocarcinomas. Clin. Epigenet. 2016, 8(1), 100. [Google Scholar] [CrossRef]
- Dietrich, D.; Jung, M.; Puetzer, S.; Leisse, A.; Holmes, E.E.; Meller, S.; et al. Diagnostic and Prognostic Value of SHOX2 and SEPT9 DNA Methylation and Cytology in Benign, Paramalignant and Malignant Pleural Effusions. PLoS ONE 2013, 8(12), e84225. [Google Scholar] [CrossRef]
- Gunasekara, T.D.K.S.C.; Herath, C.; De Silva, P.M.C.S.; Jayasundara, N. Exploring the Utility of Urinary Creatinine Adjustment for KIM-1, NGAL, and Cystatin C for the Assessment of Kidney Function: Insights from the C-KidnEES Cohort. Children [Internet] 2024, 11(1). [Google Scholar] [CrossRef] [PubMed]
- Waikar, S.S.; Sabbisetti, V.S.; Bonventre, J.V. Normalization of urinary biomarkers to creatinine during changes in glomerular filtration rate. Kidney Int. 2010, 78(5), 486–94. [Google Scholar] [CrossRef] [PubMed]
- Di Giovannantonio, M.; Hartley, F.; Elshenawy, B.; Barberis, A.; Hudson, D.; Shafique, H.S.; et al. Defining hypoxia in cancer: A landmark evaluation of hypoxia gene expression signatures. Cell. Genom. 2025, 5(2). [Google Scholar] [CrossRef] [PubMed]
- Moen, L.; Liu, B.; Bukirwa, P.; Chingonzoh, T.; Chokunonga, E.; Finesse, A.; et al. Trends in the incidence of colorectal cancer in sub-Saharan Africa: A population-based registry study. Int. J. Cancer 2024, 155(4), 675–82. [Google Scholar] [CrossRef]
- Qiao, Y.; Jiang, X.; Li, Y.; Wang, K.; Chen, R.; Liu, J.; et al. Identification of a hypoxia-related gene prognostic signature in colorectal cancer based on bulk and single-cell RNA-seq. Sci. Rep. 2023, 13(1), 2503. [Google Scholar] [CrossRef]
- Jun, J.C.; Rathore, A.; Younas, H.; Gilkes, D.; Polotsky, V.Y. Hypoxia-Inducible Factors and Cancer. Curr. Sleep. Med. Rep. 2017, 3(1), 1–10. [Google Scholar] [CrossRef]
- Triner, D.; Shah, Y.M. Hypoxia-inducible factors: a central link between inflammation and cancer. J. Clin. Investig. 2016, 126(10), 3689–98. [Google Scholar] [CrossRef]
- Huang, M.; Deng, S.; Li, M.; Yang, Z.; Guo, J.; Deng, Y.; et al. Clinical diagnostic value of methylated SEPT9 combined with NLR, PLR and LMR in colorectal cancer. BMC Gastroenterol. 2024, 24(1), 240. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Long, L. Diagnostic performances of methylated septin9 gene, CEA, CA19-9 and platelet-to-lymphocyte ratio in colorectal cancer. BMC Cancer 2024, 24(1), 906. [Google Scholar] [CrossRef] [PubMed]
- Xue, C.; Yang, B.; Fu, L.; Hou, H.; Qiang, J.; Zhou, C.; et al. Urine biomarkers can outperform serum biomarkers in certain diseases. URINE 2023, 5, 57–64. [Google Scholar] [CrossRef]
| Diagnosis | Description | Total | Gender | Age Groups | |||||
|---|---|---|---|---|---|---|---|---|---|
| Male | Female | <50 | 50–59 | 60–69 | ≥ 70 | Mean | |||
| CRC | Overall | 142 | 76 | 66 | 6 | 29 | 66 | 41 | 65 |
| Right Colon | 38 | 16 | 22 | 2 | 6 | 17 | 13 | 66 | |
| Left Colon | 65 | 38 | 27 | 1 | 12 | 35 | 17 | 65 | |
| Rectum | 39 | 22 | 17 | 3 | 11 | 14 | 11 | 63 | |
| Stage I | 35 | 21 | 14 | 1 | 5 | 21 | 8 | 65 | |
| Stage II | 40 | 19 | 21 | 1 | 8 | 20 | 11 | 65 | |
| Stage III | 48 | 26 | 22 | 2 | 14 | 15 | 17 | 65 | |
| Stage IV | 19 | 10 | 9 | 2 | 2 | 10 | 5 | 66 | |
| Non-cancer subgroups | |||||||||
| Adenomatous Colorectal polyps | 62 | 33 | 29 | 2 | 18 | 29 | 3 | 59 | |
| Inguinal Hernia | 64 | 39 | 25 | 16 | 31 | 16 | 1 | 54 | |
| Hemorrhoids | 114 | 55 | 59 | 53 | 36 | 24 | 1 | 50 | |
| Total | 382 | 203 | 179 | 77 | 124 | 135 | 46 | 58 | |
| Category | Variable | CRC n=142 | Polyps n= 62 | Controls n=178 | p-value |
|---|---|---|---|---|---|
| Demographics | Sex | 0.992 | |||
| Male | 76(37.4%) | 33(16.3%) | 94(46.3%) | ||
| Female | 66(36.9%) | 29(16.2%) | 84(46.9%) | ||
| Clinical parameters | Tumor Gross type | ||||
| Ulcerative | 80 (56.3%) | — | — | ||
| Polypoid | 40 (28.2%) | — | — | ||
| Unknown | 22 (15.5%) | — | — | ||
| Tumor Infiltration | |||||
| Mucosa | 4 (2.8%) | — | — | ||
| Sub mucosa | 5 (3.5%) | — | — | ||
| Muscularis propria | 51 (35.9%) | — | — | ||
| Pericolic tissue | 44 (31.0%) | — | — | ||
| Serosa | 18(27.7%) | — | — | ||
| Adjacent structures | 20(14.1%) | — | — | ||
| Lymph node ratio | — | ||||
| <0.2 (low) | 112 (78.9%) | — | — | ||
| ≥0.2 (high) | 30 (21.1%) | — | — | ||
| Tumor Budding | |||||
| Bd1 (Low) | 62 (43.7%) | — | — | ||
| Bd2 (Intermediate) | 45 (31.7%) | — | — | ||
| Bd3 (High) | 35 (24.6%) | — | |||
| Lymph Node Invasion | — | ||||
| Absent | 78 (54.9%) | — | — | ||
| Present | 64 (45.1%) | — | — | ||
| Vascular Invasion | |||||
| Absent | 70 (49.3%) | — | — | ||
| Present | 72 (50.7%) | — | — | ||
| Perineural Invasion | — | ||||
| Absent | 83 (58.5%) | — | — | ||
| Present | 59 (41.5%) | — | — | ||
| p53 Mutation Pattern | — | ||||
| Wild-type | 48 (33.8%) | — | |||
| Mutant pattern | 76 (53.5%) | — | — | ||
| Indeterminate | 18 (12.7%) | — | — | ||
| Ki-67 Index Group ( Median-IQR) | 60[50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70] | — | — | ||
| Colorectal Polyps | |||||
| Tubular adenoma | — | 23(37.1%) | — | ||
| Tubulovillous adenoma | — | 17(27.4%) | — | ||
| Villous adenoma | — | 11(17.7%) | — | ||
| Serrated lesions | — | 5(8.1%) | — | ||
| Hyperplastic polyps | — | 6(9.7%) | — | ||
| Colorectal polyps Dysplasia grade | |||||
| Low-grade dysplasia | 22(35.5% | ||||
| High-grade dysplasia | 40(64.5%) | ||||
| Lifestyle parameters | Smoking History | 0.265 | |||
| Never smoked | 54 (38.0%) | 31 (50.0%) | — | ||
| Former smoker | 68 (47.9%) | 23 (37.1%) | — | ||
| Current smoker | 20 (14.1%) | 8 (12.9%) | — | ||
| Alcohol History | 0.007* | ||||
| Never drank | 52 (36.6%) | 24 (38.7%) | — | ||
| Former drinker | 72 (50.7%) | 20 (32.3%) | — | ||
| Current drinker | 18 (12.7%) | 18 (29.0%) | — | ||
| Family History of Cancer | |||||
| No | 55 (38.7%) | 21 (34.4%) | — | 0.811 | |
| Yes | 24 (16.9%) | 12 (19.7%) | — | ||
| Unknown | 63 (44.4%) | 28 (45.9%) | — |
| McNemar’s P-value for Diagnostic groups | McNemar’s P-value for Stages | McNemar’s P-value for pT stage | McNemar’s p-value for Lymph node stage |
|---|---|---|---|
| CRC (p=0.894) | Stage I(p=0.050*) | pT1 (p=0.250) | N0(p=0.596) |
| Polyps (p=0.570) | Stage II(p=0.450) | pT2 (p=0.061) | N1(p=724) |
| Non-Malignant (p=0.031*) | Stage III(p=0.340) | pT3 (p=0.0197*) | N2(p=0.450) |
| - | Stage IV(p=0.620 | pT4(p=1.00) | - |
| Marker | mSEPT9-positive | mSEPT9-negative | DiAcSpm-positive | DiAcSpm-negative | P-value |
|---|---|---|---|---|---|
| NLR | 3.05(2.23–3.86) | 2.20(1.78–2.23) | 2.83(2.12–3.69) | 2.24(1.81–2.96) | <0.001** |
| PLR | 181.25(137.79–228.96) | 126.19(107.38–166.02) | 174.26(123.51–232.65) | 128.19(108.06–168.49) | <0.001** |
| LMR | 2.49(1.92–3.28) | 3.50(2.66–4.89) | 2.65(1.93–3.42) | 3.43(2.60–4.88) | <0.001** |
| Model Group | Biomarker Combination | AUC (95% CI) | Sensitivity (%) | Specificity (%) | p-value |
|---|---|---|---|---|---|
| A6 | mSEPT9 + DiAcSpm + PLR + NLR | 0.940 (0.918–0.967) | 85.2 | 92.5 | <0.001** |
| B8 | NLR + PLR + LMR + CEA + CA19-9 | 0.899 (0.867–0.932) | 93.2 | 80.3 | <0.001** |
| C7 | CEA + CA19-9 + mSEPT9 + DiAcSpm + NLR | 0.932 (0.904–0.959) | 85.2 | 91.7 | <0.001** |
| D1 | mSEPT9 + DiAcSpm + NLR + PLR + LMR + CEA + CA19-9 | 0.950 (0.928–0.973) | 86.6 | 93.5 | <0.001** |
| D3 | mSEPT9 + DiAcSpm + NLR + PLR + CEA + CA19-9 + CA125 + AFP | 0.950 (0.927–0.973) | 90.8 | 90.4 | <0.001** |
| D4 | mSEPT9 + DiAcSpm + NLR + PLR + LMR | 0.947 (0.924–0.970) | 85.9 | 92.9 | <0.001** |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




