Submitted:
28 April 2026
Posted:
29 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Study Cohort and Sample Availability
2.2. Baseline Associations Between Biomarkers and Clinical Variables
2.3. Clinical Trajectories After Tofersen Administration
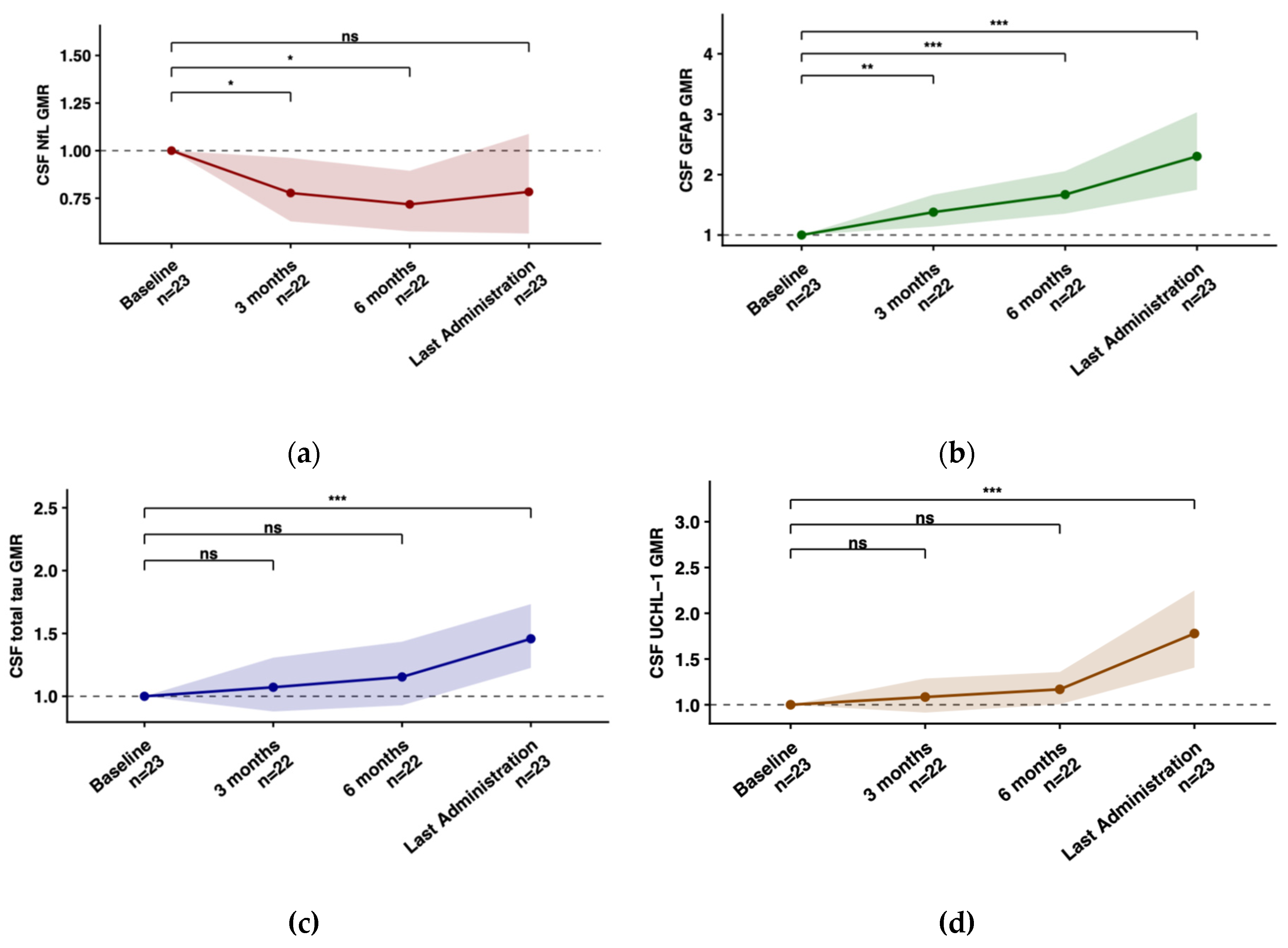
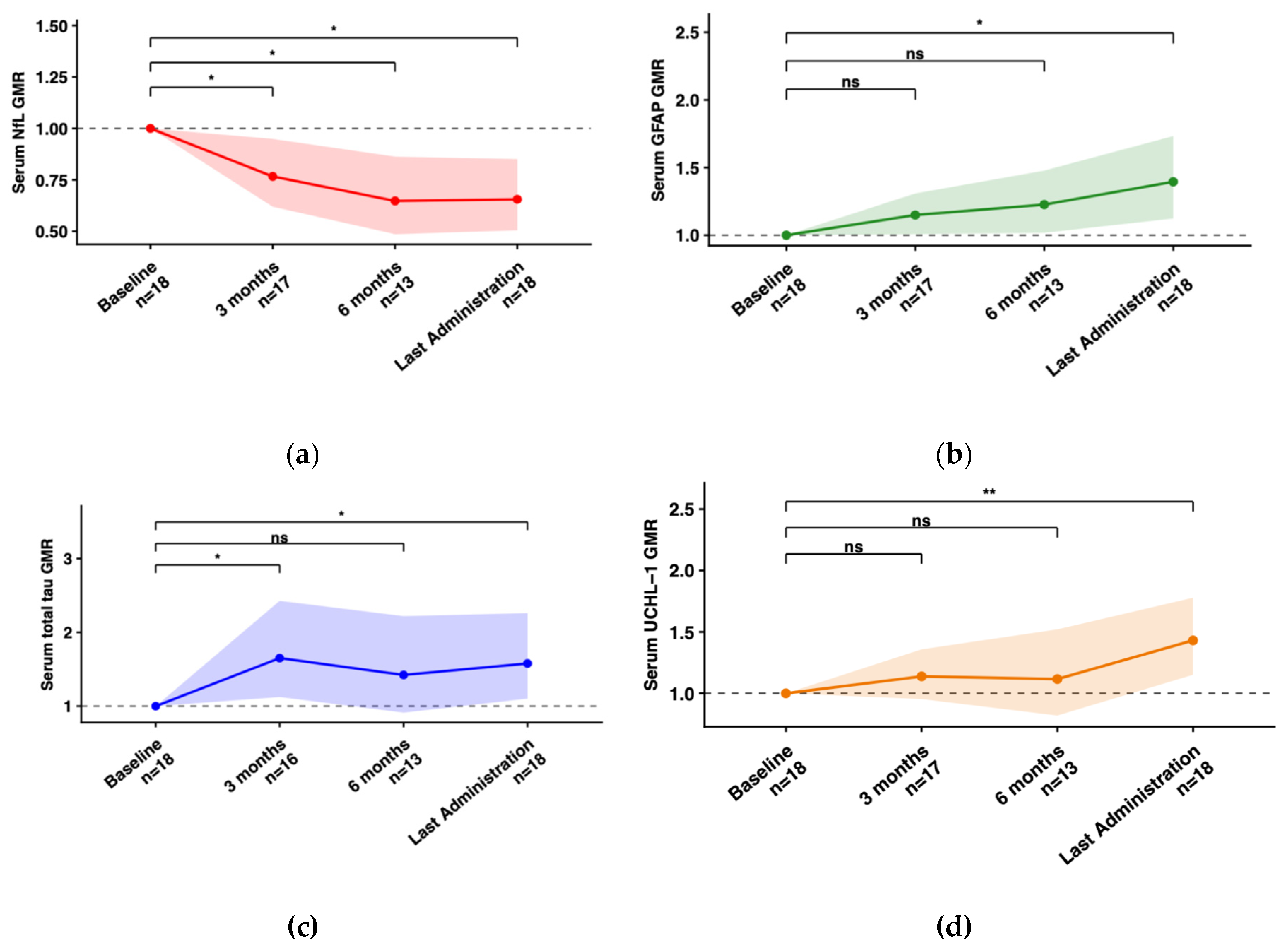
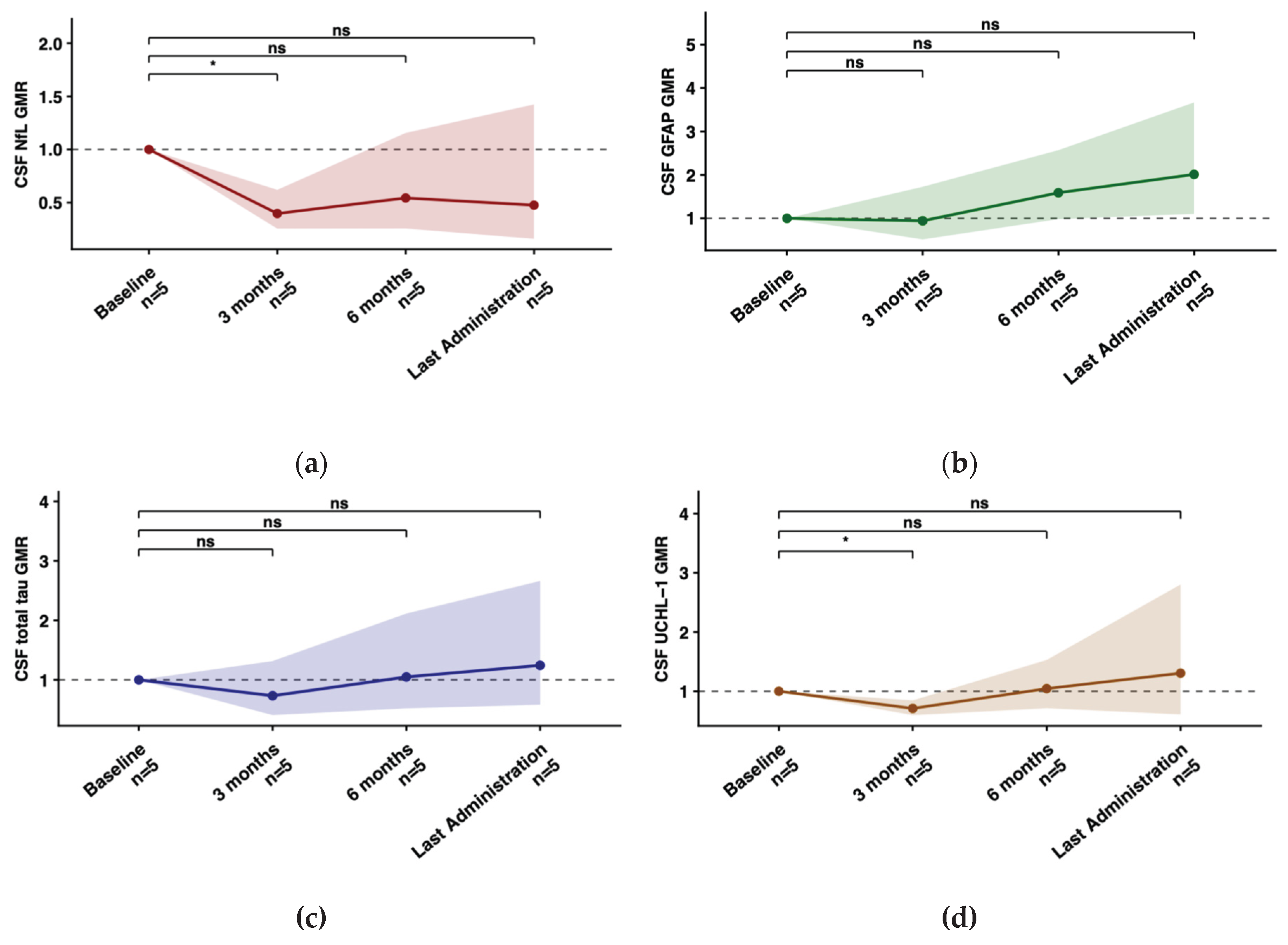
2.4. Longitudinal Biomarker Trajectories
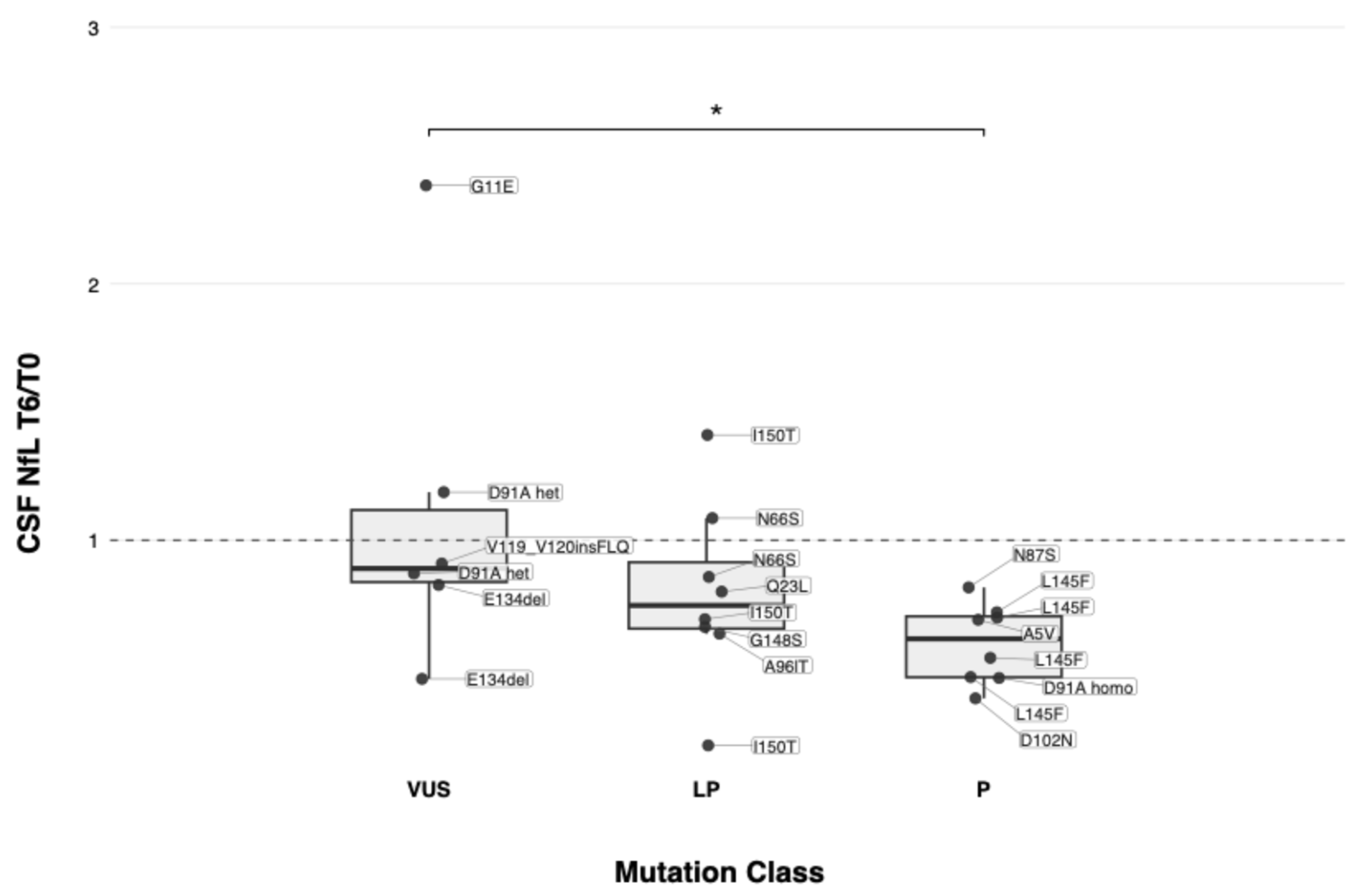
2.5. Genotype–Biomarker Analysis
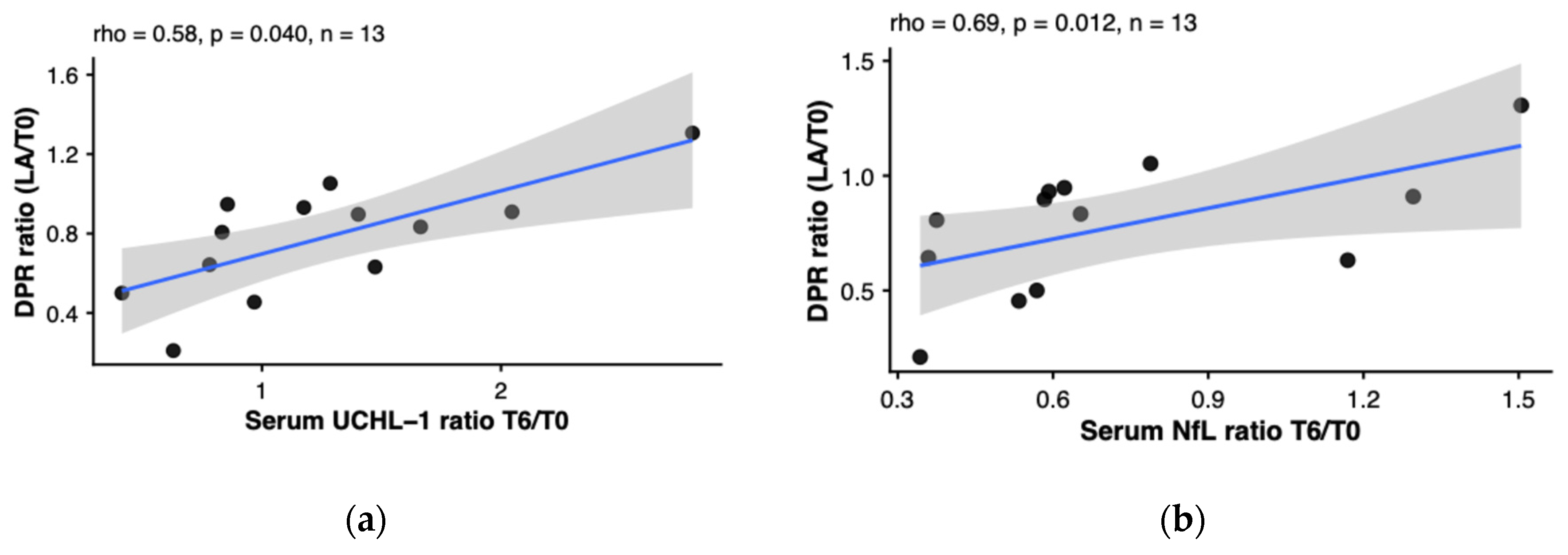
2.6. Early Biomarker Changes as Predictors of Long-Term DPR Change
2.7. Early Biomarker Responder Subgroup Analysis
3. Discussion
4. Materials and Methods
4.1. Study Design and Cohort
4.2. Clinical Variables and Outcomes
4.3. Genetic Annotation
4.4. Biospecimen Collection and Biomarker Assays
4.5. Statistical Analysis
5. Limitations
6. Conclusion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ACMG/AMP | American College of Medical Genetics and Genomics / Association for Molecular Pathology |
| ALS | Amyotrophic Lateral Sclerosis |
| ALSFRS-R | Amyotrophic Lateral Sclerosis Functional Rating Scale–Revised |
| CADD | Combined Annotation Dependent Depletion |
| CSF | Cerebrospinal Fluid |
| DPR | Disease Progression Rate |
| FVC | Forced Vital Capacity |
| GFAP | Glial Fibrillary Acidic Protein |
| NfL | Neurofilament Light Chain |
| NIV | Non-Invasive Ventilation |
| REVEL | Rare Exome Variant Ensemble Learner |
| RNase H | Ribonuclease H |
| SOD1 | Superoxide Dismutase 1 |
| TARDBP | TAR DNA-Binding Protein |
| UCHL-1 | Ubiquitin C-Terminal Hydrolase L1 |
| UTR | Untranslated Region |
References
- Miller, T.M.; Cudkowicz, M.E.; Genge, A.; Shaw, P.J.; Sobue, G.; Bucelli, R.C.; Chiò, A.; Van Damme, P.; Ludolph, A.C.; Glass, J.D.; et al. Trial of Antisense Oligonucleotide Tofersen for SOD1 ALS . New England Journal of Medicine 2022, 387, 1099–1110. [CrossRef]
- McCampbell, A.; Cole, T.; Wegener, A.J.; Tomassy, G.S.; Setnicka, A.; Farley, B.J.; Schoch, K.M.; Hoye, M.L.; Shabsovich, M.; Sun, L.; et al. Antisense Oligonucleotides Extend Survival and Reverse Decrement in Muscle Response in ALS Models. Journal of Clinical Investigation 2018, 128, 3558–3567. [CrossRef]
- Miller, T.; Cudkowicz, M.; Shaw, P.J.; Andersen, P.M.; Atassi, N.; Bucelli, R.C.; Genge, A.; Glass, J.; Ladha, S.; Ludolph, A.L.; et al. Phase 1–2 Trial of Antisense Oligonucleotide Tofersen for SOD1 ALS . New England Journal of Medicine 2020, 383, 109–119. [CrossRef]
- Miller, T.M.; Cudkowicz, M.E.; Shaw, P.J.; Genge, A.; Sobue, G.; Bucelli, R.C.; Chiò, A.; Van Damme, P.; Ludolph, A.C.; Glass, J.D.; et al. Long-Term Tofersen in SOD1 Amyotrophic Lateral Sclerosis. JAMA Neurol. 2025. [CrossRef]
- Meyer, T.; Schumann, P.; Weydt, P.; Petri, S.; Koc, Y.; Spittel, S.; Bernsen, S.; Günther, R.; Weishaupt, J.H.; Dreger, M.; et al. Neurofilament Light-Chain Response during Therapy with Antisense Oligonucleotide Tofersen in SOD1-Related ALS: Treatment Experience in Clinical Practice. Muscle Nerve 2023, 67, 515–521. [CrossRef]
- Sabatelli, M.; Cerri, F.; Zuccarino, R.; Patanella, A.K.; Bernardo, D.; Bisogni, G.; Tanel, R.; Sansone, V.; Filosto, M.; Lattante, S.; et al. Long-Term Treatment of SOD1 ALS with Tofersen: A Multicentre Experience in 17 Patients. J. Neurol. 2024, 271, 5177–5186. [CrossRef]
- Wiesenfarth, M.; Dorst, J.; Brenner, D.; Elmas, Z.; Parlak, Ö.; Uzelac, Z.; Kandler, K.; Mayer, K.; Weiland, U.; Herrmann, C.; et al. Effects of Tofersen Treatment in Patients with SOD1-ALS in a “Real-World” Setting – a 12-Month Multicenter Cohort Study from the German Early Access Program. EClinicalMedicine 2024, 69. [CrossRef]
- Khalil, M.; Teunissen, C.E.; Lehmann, S.; Otto, M.; Piehl, F.; Ziemssen, T.; Bittner, S.; Sormani, M.P.; Gattringer, T.; Abu-Rumeileh, S.; et al. Neurofilaments as Biomarkers in Neurological Disorders — towards Clinical Application. Nat. Rev. Neurol. 2024, 20, 269–287.
- Riva, N.; Domi, T.; Pozzi, L.; Lunetta, C.; Schito, P.; Spinelli, E.G.; Cabras, S.; Matteoni, E.; Consonni, M.; Bella, E.D.; et al. Update on Recent Advances in Amyotrophic Lateral Sclerosis. J. Neurol. 2024, 271, 4693–4723. [CrossRef]
- Liu, J.; Wang, F. Role of Neuroinflammation in Amyotrophic Lateral Sclerosis: Cellular Mechanisms and Therapeutic Implications. Front. Immunol. 2017, 8.
- Rizea, R.E.; Corlatescu, A.D.; Costin, H.P.; Dumitru, A.; Ciurea, A.V. Understanding Amyotrophic Lateral Sclerosis: Pathophysiology, Diagnosis, and Therapeutic Advances. Int. J. Mol. Sci. 2024, 25.
- Agnello, L.; Colletti, T.; Lo Sasso, B.; Vidali, M.; Spataro, R.; Gambino, C.M.; Giglio, R.V.; Piccoli, T.; Bivona, G.; La Bella, V.; et al. Tau Protein as a Diagnostic and Prognostic Biomarker in Amyotrophic Lateral Sclerosis. Eur. J. Neurol. 2021, 28, 1868–1875. [CrossRef]
- Falzone, Y.M.; Domi, T.; Mandelli, A.; Pozzi, L.; Schito, P.; Russo, T.; Barbieri, A.; Fazio, R.; Volontè, M.A.; Magnani, G.; et al. Integrated Evaluation of a Panel of Neurochemical Biomarkers to Optimize Diagnosis and Prognosis in Amyotrophic Lateral Sclerosis. Eur. J. Neurol. 2022, 29, 1930–1939. [CrossRef]
- Li, R.; Wang, J.; Xie, W.; Liu, J.; Wang, C. UCHL1 from Serum and CSF Is a Candidate Biomarker for Amyotrophic Lateral Sclerosis. Ann. Clin. Transl. Neurol. 2020, 7, 1420–1428. [CrossRef]
- Thapa, S.; Bhattarai, A.; Shah, S.; Chand, S.; Bagherieh, S.; Mirmosayyeb, O.; Mishra, S.K. Diagnostic Role of Tau Proteins in Amyotrophic Lateral Sclerosis: A Systematic Review and Meta-Analysis. Acta Neurol. Scand. 2023, 2023.
- Sofroniew, M. V.; Vinters, H. V. Astrocytes: Biology and Pathology. Acta Neuropathol. 2010, 119, 7–35.
- Nagy, D.; Kato, T.; Kushner, P.D. Reactive Astrocytes Are Widespread in the Cortical Gray Matter of Amyotrophic Lateral Sclerosis; 1994;
- Schiffer, D.; Cordera, S.; Cavalla, P.; Migheli, A. Reactive Astrogliosis of the Spinal Cord in Amyotrophic Lateral Sclerosis; 1996; Vol. 139;.
- Heller, C.; Foiani, M.S.; Moore, K.; Convery, R.; Bocchetta, M.; Neason, M.; Cash, D.M.; Thomas, D.; Greaves, C. V.; Woollacott, I.O.C.; et al. Plasma Glial Fibrillary Acidic Protein Is Raised in Progranulin-Associated Frontotemporal Dementia. J. Neurol. Neurosurg. Psychiatry 2020, 91. [CrossRef]
- Day, I.N.M.; Thompson, R.J. UCHL1 (PGP 9.5): Neuronal Biomarker and Ubiquitin System Protein. Prog. Neurobiol. 2010, 90, 327–362.
- Zhu, S.; Wuolikainen, A.; Wu, J.; Öhman, A.; Wingsle, G.; Moritz, T.; Andersen, P.M.; Forsgren, L.; Trupp, M. Targeted Multiple Reaction Monitoring Analysis of CSF Identifies UCHL1 and GPNMB as Candidate Biomarkers for ALS. Journal of Molecular Neuroscience 2019, 69, 643–657. [CrossRef]
- Oeckl, P.; Weydt, P.; Thal, D.R.; Weishaupt, J.H.; Ludolph, A.C.; Otto, M. Proteomics in Cerebrospinal Fluid and Spinal Cord Suggests UCHL1, MAP2 and GPNMB as Biomarkers and Underpins Importance of Transcriptional Pathways in Amyotrophic Lateral Sclerosis. Acta Neuropathol. 2020, 139, 119–134. [CrossRef]
- Brettschneider, J.; Petzold, ; A; Sü, ; S D; Ludolph, ; A C; Tumani, H. Axonal Damage Markers in Cerebrospinal Fluid Are Increased in ALS; 2006;
- Wilke, C.; Deuschle, C.; Rattay, T.W.; Maetzler, W.; Synofzik, M. Total Tau Is Increased, but Phosphorylated Tau Not Decreased, in Cerebrospinal Fluid in Amyotrophic Lateral Sclerosis. Neurobiol. Aging 2015, 36, 1072–1074. [CrossRef]
- Paladino, P.; Valentino, F.; Piccoli, T.; Piccoli, F.; La Bella, V. Cerebrospinal Fluid Tau Protein Is Not a Biological Marker in Amyotrophic Lateral Sclerosis. Eur. J. Neurol. 2009, 16, 257–261. [CrossRef]
- Scarafino, A.; D’Errico, E.; Introna, A.; Fraddosio, A.; Distaso, E.; Tempesta, I.; Morea, A.; Mastronardi, A.; Leante, R.; Ruggieri, M.; et al. Diagnostic and Prognostic Power of CSF Tau in Amyotrophic Lateral Sclerosis. J. Neurol. 2018, 265, 2353–2362. [CrossRef]
- Lanznaster, D.; Hergesheimer, R.C.; Bakkouche, S.E.; Beltran, S.; Vourc’H, P.; Andres, C.R.; Dufour-Rainfray, D.; Corcia, P.; Blasco, H. Aβ1-42 and Tau as Potential Biomarkers for Diagnosis and Prognosis of Amyotrophic Lateral Sclerosis. Int. J. Mol. Sci. 2020, 21. [CrossRef]
- Bridel, C.; Van Wieringen, W.N.; Zetterberg, H.; Tijms, B.M.; Teunissen, C.E.; Alvarez-Cermeño, J.C.; Andreasson, U.; Axelsson, M.; Bäckström, D.C.; Bartos, A.; et al. Diagnostic Value of Cerebrospinal Fluid Neurofilament Light Protein in Neurology: A Systematic Review and Meta-Analysis. JAMA Neurol. 2019, 76, 1035–1048. [CrossRef]
- Zucchi, E.; Lu, C.H.; Cho, Y.; Chang, R.; Adiutori, R.; Zubiri, I.; Ceroni, M.; Cereda, C.; Pansarasa, O.; Greensmith, L.; et al. A Motor Neuron Strategy to Save Time and Energy in Neurodegeneration: Adaptive Protein Stoichiometry. J. Neurochem. 2018, 146, 631–641. [CrossRef]
- Weishaupt, J.H.; Körtvélyessy, P.; Schumann, P.; Valkadinov, I.; Weyen, U.; Hesebeck-Brinckmann, J.; Weishaupt, K.; Endres, M.; Andersen, P.M.; Regensburger, M.; et al. Tofersen Decreases Neurofilament Levels Supporting the Pathogenesis of the SOD1 p.D91A Variant in Amyotrophic Lateral Sclerosis Patients. Communications Medicine 2024, 4. [CrossRef]
- Steffke, C.; Baskar, K.; Bachhuber, F.; Wiesenfarth, M.; Dorst, J.; Schuster, J.; Schöberl, F.; Reilich, P.; Regensburger, M.; German, A.; et al. Targeted Proteomics upon Treatment with Tofersen Identifies Novel Response Markers for Superoxide Dismutase 1-Linked Amyotrophic Lateral Sclerosis. Ann. Neurol. 2025. [CrossRef]
- Simonini, C.; Zucchi, E.; Martinelli, I.; Gianferrari, G.; Lunetta, C.; Sorarù, G.; Trojsi, F.; Pepe, R.; Piras, R.; Giacchino, M.; et al. Neurodegenerative and Neuroinflammatory Changes in SOD1-ALS Patients Receiving Tofersen. Sci. Rep. 2025, 15. [CrossRef]
- Kanaan, N.M.; Grabinski, T. Neuronal and Glial Distribution of Tau Protein in the Adult Rat and Monkey. Front. Mol. Neurosci. 2021, 14. [CrossRef]
- Kubo, A.; Misonou, H.; Matsuyama, M.; Nomori, A.; Wada-Kakuda, S.; Takashima, A.; Kawata, M.; Murayama, S.; Ihara, Y.; Miyasaka, T. Distribution of Endogenous Normal Tau in the Mouse Brain. Journal of Comparative Neurology 2019, 527, 985–998. [CrossRef]
- Chiò, A.; Calvo, A.; Moglia, C.; Mazzini, L.; Mora, G.; Mutani, R.; Balma, M.; Cammarosano, S.; Canosa, A.; Gallo, S.; et al. Phenotypic Heterogeneity of Amyotrophic Lateral Sclerosis: A Population Based Study. J. Neurol. Neurosurg. Psychiatry 2011, 82, 740–746. [CrossRef]
- Schito, P.; Russo, T.; Domi, T.; Mandelli, A.; Pozzi, L.; Del Carro, U.; Carrera, P.; Agosta, F.; Quattrini, A.; Furlan, R.; et al. Clinical Features and Biomarkers to Differentiate Primary and Amyotrophic Lateral Sclerosis in Patients With an Upper Motor Neuron Syndrome. Neurology 2023, 101, 352–356. [CrossRef]
- Schito, P.; Manera, U.; Russo, T.; Cremona, G.; Riboldi, E.; Tettamanti, A.; Agosta, F.; Quattrini, A.; Chiò, A.; Filippi, M.; et al. Use of the Combination of Spirometry, Arterial Blood Gas Analysis and Overnight Oximetry to Predict the Outcomes of Patients Affected by Motor Neuron Disease: The Milan-Torin Respiratory Score (Mi-To-RS). Eur. J. Neurol. 2024, 31. [CrossRef]
- Kimura, F.; Fujimura, C; Ishida, S; Nakajima, H; Furutama, D; Uehara, H; Shinoda, K; Sugino, M; Hanafusa, T. Progression Rate of ALSFRS-R at Time of Diagnosis Predicts Survival Time in ALS; 2006; Vol. 66;.
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and Guidelines for the Interpretation of Sequence Variants: A Joint Consensus Recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genetics in Medicine 2015, 17, 405–424. [CrossRef]
- Kircher, M.; Witten, D.M.; Jain, P.; O’roak, B.J.; Cooper, G.M.; Shendure, J. A General Framework for Estimating the Relative Pathogenicity of Human Genetic Variants. Nat. Genet. 2014, 46, 310–315. [CrossRef]
- Ioannidis, N.M.; Rothstein, J.H.; Pejaver, V.; Middha, S.; McDonnell, S.K.; Baheti, S.; Musolf, A.; Li, Q.; Holzinger, E.; Karyadi, D.; et al. REVEL: An Ensemble Method for Predicting the Pathogenicity of Rare Missense Variants. Am. J. Hum. Genet. 2016, 99, 877–885. [CrossRef]
- Wilson, D.H.; Rissin, D.M.; Kan, C.W.; Fournier, D.R.; Piech, T.; Campbell, T.G.; Meyer, R.E.; Fishburn, M.W.; Cabrera, C.; Patel, P.P.; et al. The Simoa HD-1 Analyzer: A Novel Fully Automated Digital Immunoassay Analyzer with Single-Molecule Sensitivity and Multiplexing. J. Lab. Autom. 2016, 21, 533–547. [CrossRef]
- Graham, N.S.N.; Zimmerman, K.A.; Moro, F.; Heslegrave, A.; Maillard, S.A.; Bernini, A.; Miroz, J.P.; Donat, C.K.; Lopez, M.Y.; Bourke, N.; et al. Axonal Marker Neurofilament Light Predicts Long-Term Outcomes and Progressive Neurodegeneration after Traumatic Brain Injury. Sci. Transl. Med. 2021, 13. [CrossRef]
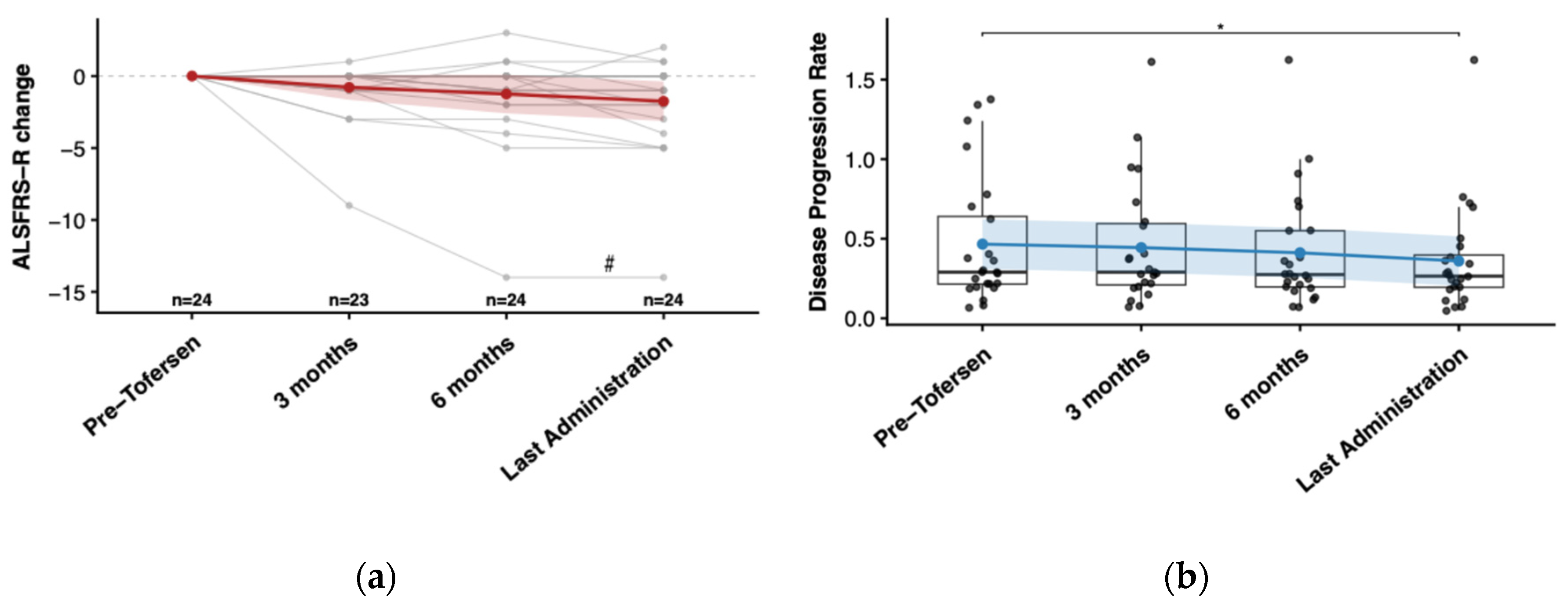
| Clinical feature | Summary |
|---|---|
| Enrolled patients | N = 24 |
| Sex | |
| Male | 11/24 (46%) |
| Female | 13/24 (54%) |
| Site of onset | N = 15 |
| Proximal UL | 5/15 (33.3%) |
| Distal LL | 8/15 (53.3%) |
| Distal UL | 2/15 (13.3%) |
| Weight at baseline (kg) | 72.71 ± 12.42 |
| Phenotype | |
| PLMN | 3/22 (13.6%) |
| Classic | 10/22 (45.5%) |
| Flail arm | 2/22 (9.1%) |
| Flail leg | 5/22 (22.7%) |
| Pyramidal | 1/22 (4.5%) |
| PUMN | 1/22 (4.5%) |
| Age at onset, years | 50.82 ± 9.38 |
| Age at diagnosis, years | 52.61 ± 9.58 |
| Age at baseline, years | 56.77 ± 10.50 |
| Months from onset to baseline, median [IQR] | 58.79 [28.13 - 103.48] |
| ALSFRS-R at baseline, median [IQR] | 29.50 [20.00 - 34.00] |
| DPR at baseline, median [IQR] | 0.29 [0.21 - 0.66] |
| FVC (% predicted) at baseline, median [IQR] | 65 [42 - 80] |
| Mutation | Exon | CADD | REVEL | ACGM | Age at onset |
Sex | Phenotype | ALSFRS-R at T0 | DPR Pre-Tofersen | DPR at LA |
|---|---|---|---|---|---|---|---|---|---|---|
| p.Gly11Glu | 5’UTR | VUS | 61.7 | F | Fail arm | 26 | Slow | Slow | ||
|
c.14C>T (p. Ala5Val) |
I | 25.7 | 0.85 | Pathogenic | 27 | F | Classic | 38 | Fast | Intermediate |
|
c.68A>T (p.Gln23Leu) |
I | 23.5 | 0.79 |
Likely pathogenic |
32.4 | F | PUMN | 22 | Slow | Slow |
|
c.197A>G (p.Asn66Ser) |
III | 26.1 | 0.94 |
Likely pathogenic |
47.1 | F | Classic | 15 | Slow | Slow |
|
c.197A>G (p.Asn66Ser) |
III | 26.1 | 0.94 |
Likely pathogenic |
56 | F | Fail leg | 16 | Slow | Slow |
| p.Ser108Leufs*15 | IV | NA | NA | Pathogenic | 64.8 | M | NA | 16 | Slow | Slow |
|
c.260A>G (p. Asn87Ser) |
IV | 22.7 | 0.85 | Pathogenic | 58.8 | F | PLMN | 45 | Slow | Slow |
|
c.272A>C (p.Asp91Ala) Heterozygous # |
IV | 9.48 | 0.55 | VUS | 54.3 | F | Pyramidal | 34 | Fast | Fast |
|
c.272A>C (p.Asp91Ala) Heterozygous |
IV | 9.48 | 0.55 | VUS | 58.3 | M | Classic | 30 | Slow | Slow |
|
c.272A>C (p.Asp91Ala) Homozygous |
IV | 9.48 | 0.55 | Pathogenic | 59.2 | M | Classic | 34 | Slow | Slow |
|
c.286G>A (p.Ala96Thr) |
IV | 22.9 | 0.78 |
Likely pathogenic |
45.3 | M | PLMN | 34 | Slow | Slow |
| c.286G>A (p.Ala96Thr) | IV | 22.9 | 0.776 |
Likely pathogenic |
60.3 | F | Classic | 27 | Slow | Slow |
| c.304G>A (p.Asp102Asn) | IV | 25.3 | 0.83 | Pathogenic | 50.4 | F | Classic | 29 | Fast | Slow |
| c.358-10 T>G | Intronic | VUS | 49 | M | Fail leg | 33 | Intermediate | Slow | ||
| p.Glu134del | V | NA | NA | VUS | 53.4 | F | Classic | 34 | Fast | Intermediate |
| p.Glu134del | V | NA | NA | VUS | 61.6 | F | Fail leg | 33 | Slow | Slow |
|
c.435G>C (p.Leu145Phe) |
V | 24 | 0.91 | Pathogenic | 41.2 | M | Classic | 22 | Slow | Slow |
|
c.435G>C (p.Leu145Phe) |
V | 24 | 0.91 | Pathogenic | 43.3 | M | Fail leg | 40 | Slow | Slow |
|
c.435G>C (p.Leu145Phe) |
V | 24 | 0.91 | Pathogenic | 45.7 | F | Classic | 20 | Slow | Slow |
|
c.435G>C (p.Leu145Phe) |
V | 24 | 0.91 | pathogenic | 49.9 | M | Classic | 18 | Intermediate | Imtermediate |
|
c.442G>A (p.Gly148Ser) |
V | 26.3 | 0.969 |
Likely pathogenic |
54.8 | M | NA | 14 | Intermediate | Intermediate |
|
c.449T>C (p.Ile150Thr) |
V | 26.6 | 0.968 |
Likely pathogenic |
40.5 | M | Fail arm | 44 | Slow | Slow |
|
c.449T>C (p.Ile150Thr) |
V | 26.6 | 0.55 |
Likely pathogenic |
50 | M | Fail leg | 20 | Slow | Slow |
|
C.449T>C (p.Ile150Thr) |
V | 26.6 | 0.96 |
Likely pathogenic |
54.6 | F | PLMN | 39 | Slow | Slow |
| Baseline | Month 3 | Month 6 | Last Administration | |
|---|---|---|---|---|
| Serum | ||||
| UCHL-1 (pg/mL) |
39.97 [24.97] (n=18) | 41.50 [27.64] (n=17) GMRΔ +13.8% |
44.66 [37.94] (n=13) GMRΔ +11.6% |
49.09 [40.30] (n=18) GMRΔ +43.1% |
| NfL (pg/mL) |
31.20 [29.23] (n=18) | 21.60 [14.28] (n=17) GMRΔ -23.3% |
18.45 [10.62] (n=13) GMRΔ -35.2% |
19.40 [14.80] (n=18) GMRΔ -34.5% |
| GFAP (pg/mL) |
137.72 [165.19] (n=18) | 147.95 [113.95] (n=17) GMRΔ +14.9% |
127.47 [154.88] (n=13) GMRΔ +22.6% |
174.80 [270.82] (n=18) GMRΔ +39.5% |
| Total Tau (pg/mL) |
0.42 [0.84] (n=18) | 0.97 [1.19] (n=16) GMRΔ +65.2% |
0.61 [1.07] (n=13) GMRΔ +42.3% |
0.76 [0.97] (n=18) GMRΔ +57.9% |
| CSF | ||||
| UCHL-1 (pg/mL) |
1828.72 [1231.43] (n=23) | 2158.38 [1235.20] (n=22) GMRΔ +8.4% |
2300.55 [1793.13] (n=22) GMRΔ +16.8% |
3105.21 [2194.22] (n=23) GMRΔ +77.8% |
| NfL (pg/mL) |
2156.56 [2816.57] (n=23) | 1986.47 [2112.00] (n=22) GMRΔ -22.3% |
1460.28 [2572.63] (n=22) GMRΔ -28.2% |
1652.33 [1621.83] (n=23) GMRΔ -21.6% |
| GFAP (pg/mL) |
5630.31 [7821.74] (n=23) | 7418.40 [12733.05] (n=22) GMRΔ +37.9% |
9932.60 [11416.80] (n=22) GMRΔ +67.0% |
15250.40 [11879.26] (n=23) GMRΔ +130.3% |
| Total Tau (pg/mL) |
58.83 [27.16] (n=23) | 69.41 [29.03] (n=22) GMRΔ +7.2% |
73.61 [26.93] (n=22) GMRΔ +15.4% |
80.47 [28.36] (n=23) GMRΔ +45.8% |
| Baseline | Month 3 | Month 6 | Last Administration | |
|---|---|---|---|---|
| Serum | ||||
| UCHL-1 (pg/mL) |
42.83 [20.79] (n=4) | 39.83 [26.51] (n=4) GMRΔ +8.1% |
48.84 [56.61] (n=3) GMRΔ +58.6% |
57.41 [43.28] (n=4) GMRΔ +67.1% |
| NfL (pg/mL) |
57.24 [75.00] (n=4) | 27.20 [55.13] (n=4) GMRΔ -19.6% |
15.85 [91.62] (n=3) GMRΔ -15.1% |
26.12 [62.07] (n=4) GMRΔ -34.0% |
| GFAP (pg/mL) |
201.88 [186.64] (n=4) | 303.92 [276.54] (n=4) GMRΔ +51.5% |
413.76 [186.75] (n=3) GMRΔ +63.9% |
445.26 [143.26] (n=4) GMRΔ +103.6% |
| Total Tau (pg/mL) |
0.60 [0.41] (n=4) | 1.25 [1.94] (n=4) GMRΔ +11.9% |
0.61 [0.78] (n=3) GMRΔ -21.1% |
0.56 [0.39] (n=4) GMRΔ -18.4% |
| CSF | ||||
| UCHL-1 (pg/mL) |
3002.72 [1307.35] (n=5) | 2138.66 [1779.03] (n=5) GMRΔ -29.0% |
3105.21 [1533.11] (n=5) GMRΔ +4.4% |
4306.88 [3609.25] (n=5) GMRΔ +30.5% |
| NfL (pg/mL) |
4623.43 [5502.97] (n=5) | 1775.23 [1765.43] (n=5) GMRΔ -60.3% |
3211.90 [2545.60] (n=5) GMRΔ -45.6% |
3170.65 [2545.60] (n=5) GMRΔ -52.4% |
| GFAP (pg/mL) |
9342.73 [17097.69] (n=5) | 11274.85 [12774.94] (n=5) GMRΔ -5.7% |
15251.59 [21494.28] (n=5) GMRΔ +58.9% |
22772.54 [21494.28] (n=5) GMRΔ +101.2% |
| Total Tau (pg/mL) |
72.62 [24.56] (n=5) | 56.20 [31.55] (n=5) GMRΔ -26.8% |
74.31 [26.77] (n=5) GMRΔ +4.9% |
95.53 [55.80] (n=5) GMRΔ +24.4% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




