Submitted:
28 April 2026
Posted:
29 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Result and Discussion
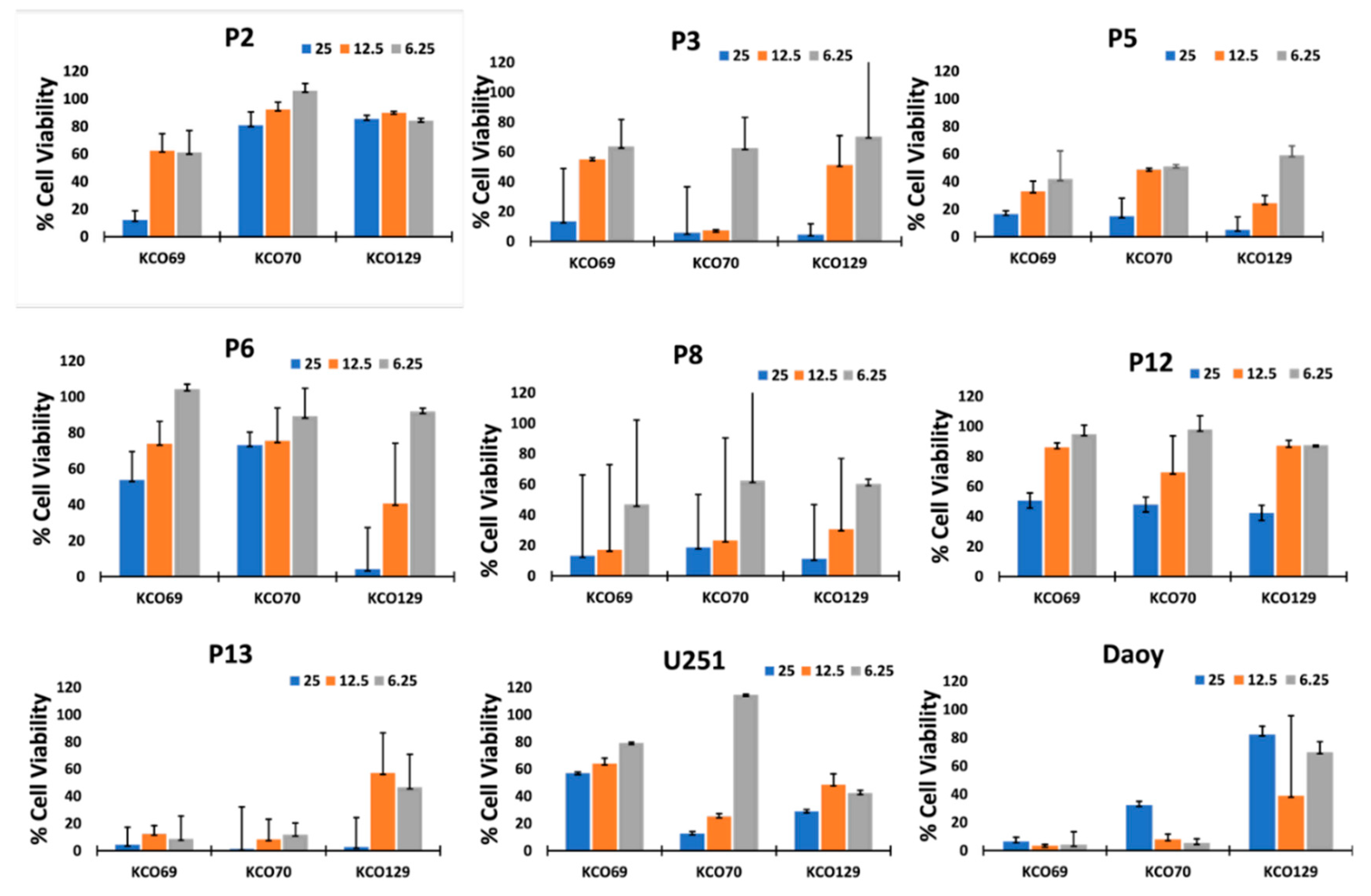
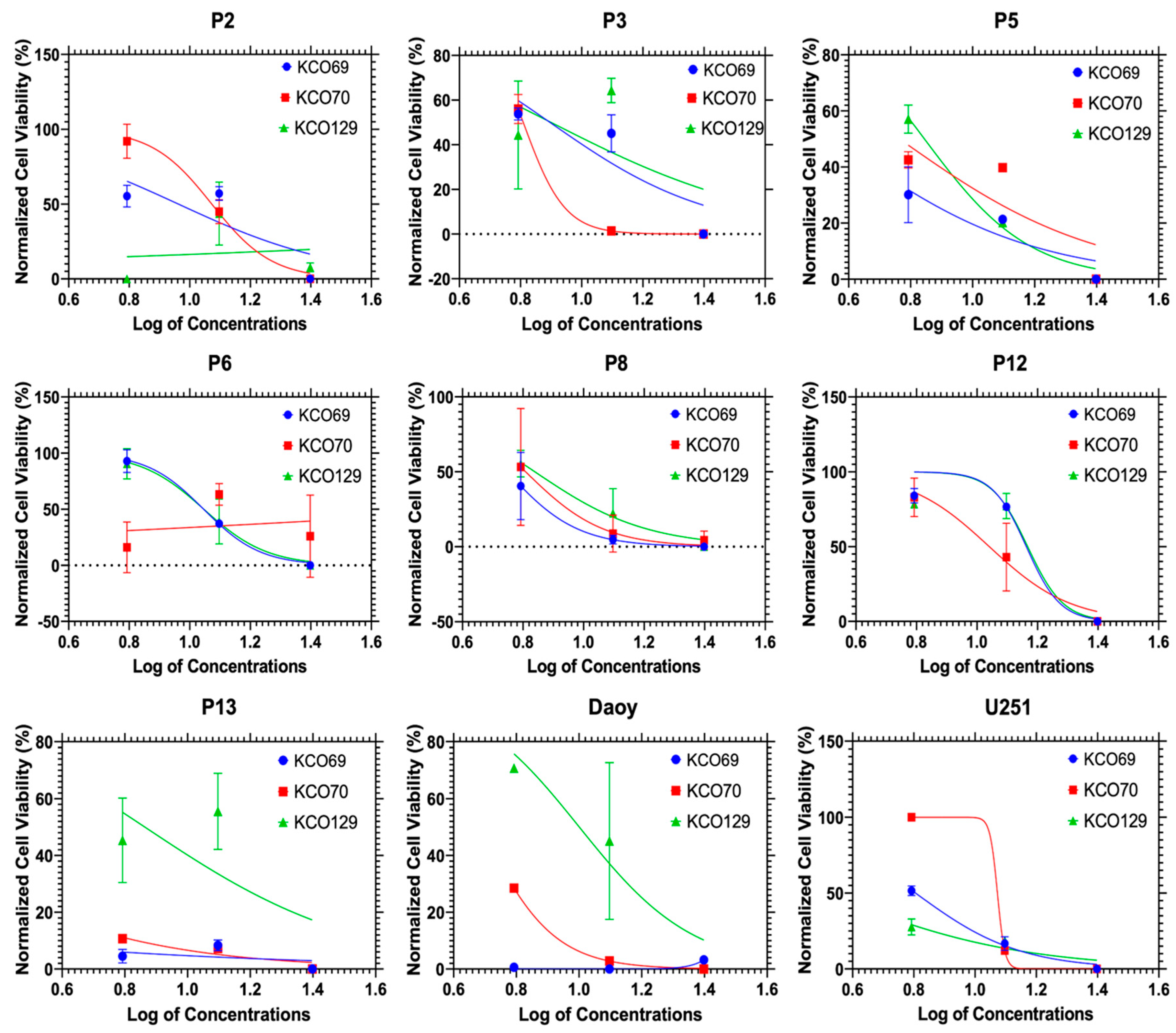
3. Materials and Methods
General Precautions
- All activities were performed in a physical containment 2 (PC2) laboratory, in a dedicated Class II B2 biological safety bio cabinet.
-
Strict aseptic techniques were employed under the following conditions:
- (a)
- Prior to use, the work surface of the biological safety cabinet was thoroughly wiped with 80% (v/v) ethanol. All materials and equipment were sprayed with 80% ethanol before being introduced into the cabinet.
- (b)
- Biological waste was disposed of in a sealed container containing a 10% bleach solution within the biological safety cabinet to minimize the risk of aerosol exposure.
- (c)
- Personnel wore a double layer of nitrile gloves during tissue processing, and care was taken to prevent skin punctures. The outer layer of gloves was discarded into a designated hazardous waste bag before the hands were withdrawn from the biological safety cabinet
The Source of the Tissue
Clinical History
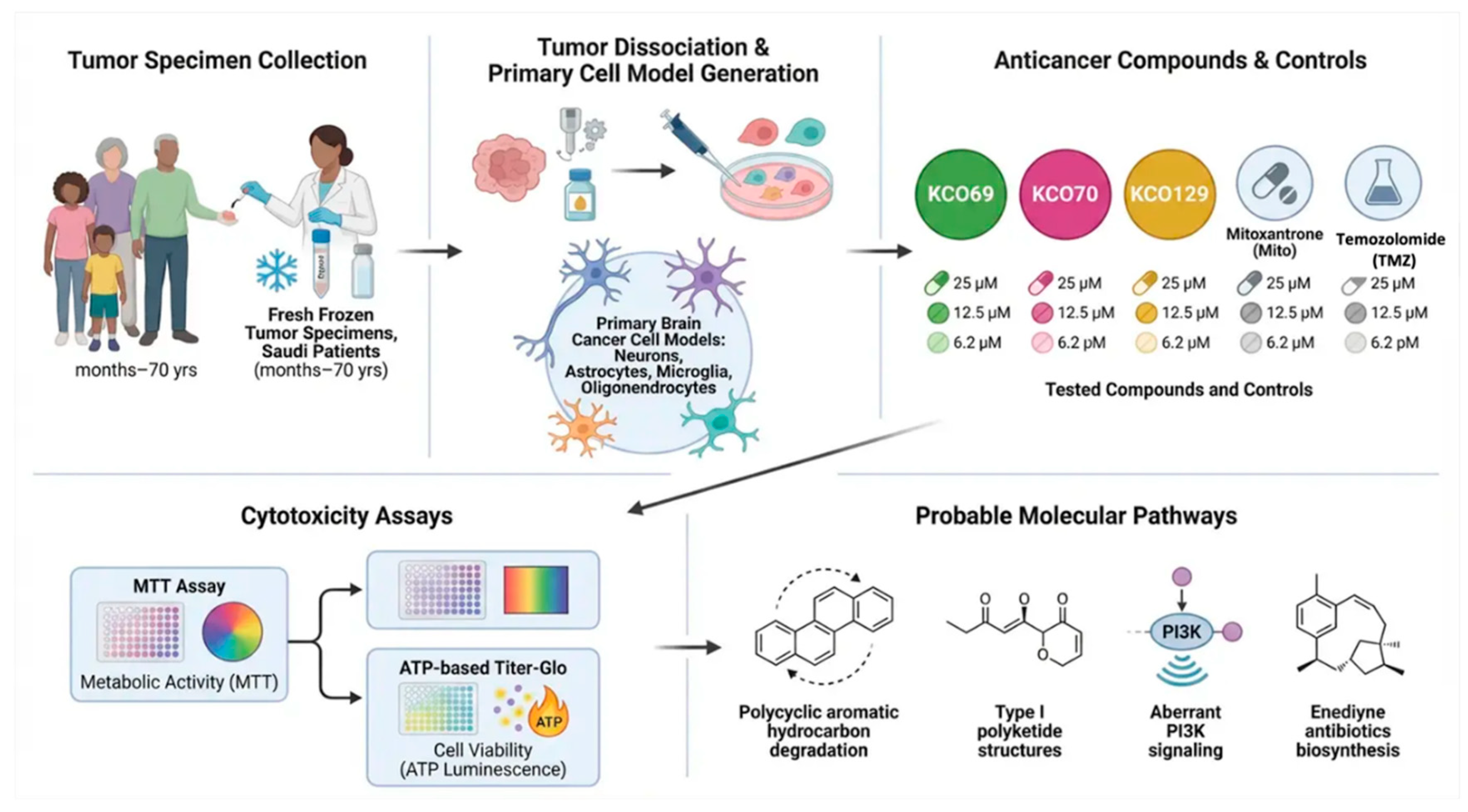
Experimental Design
Sample Collection (Brain Tissue)
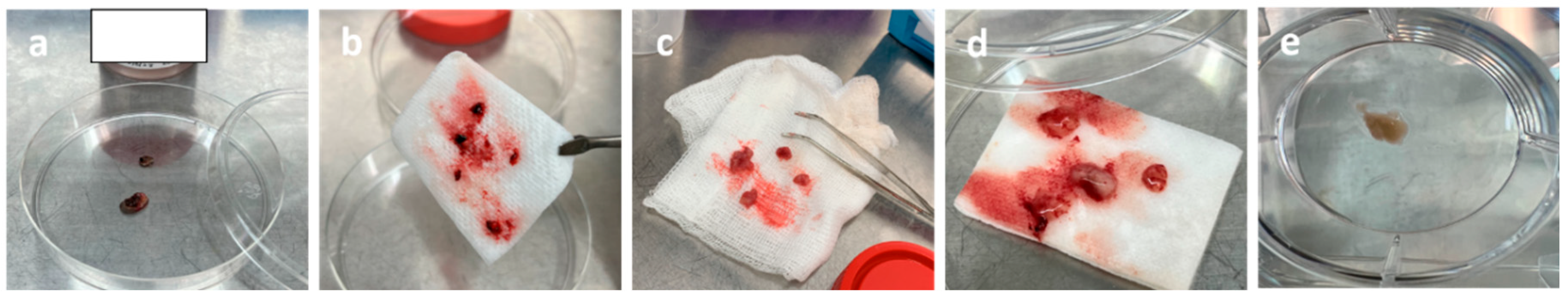
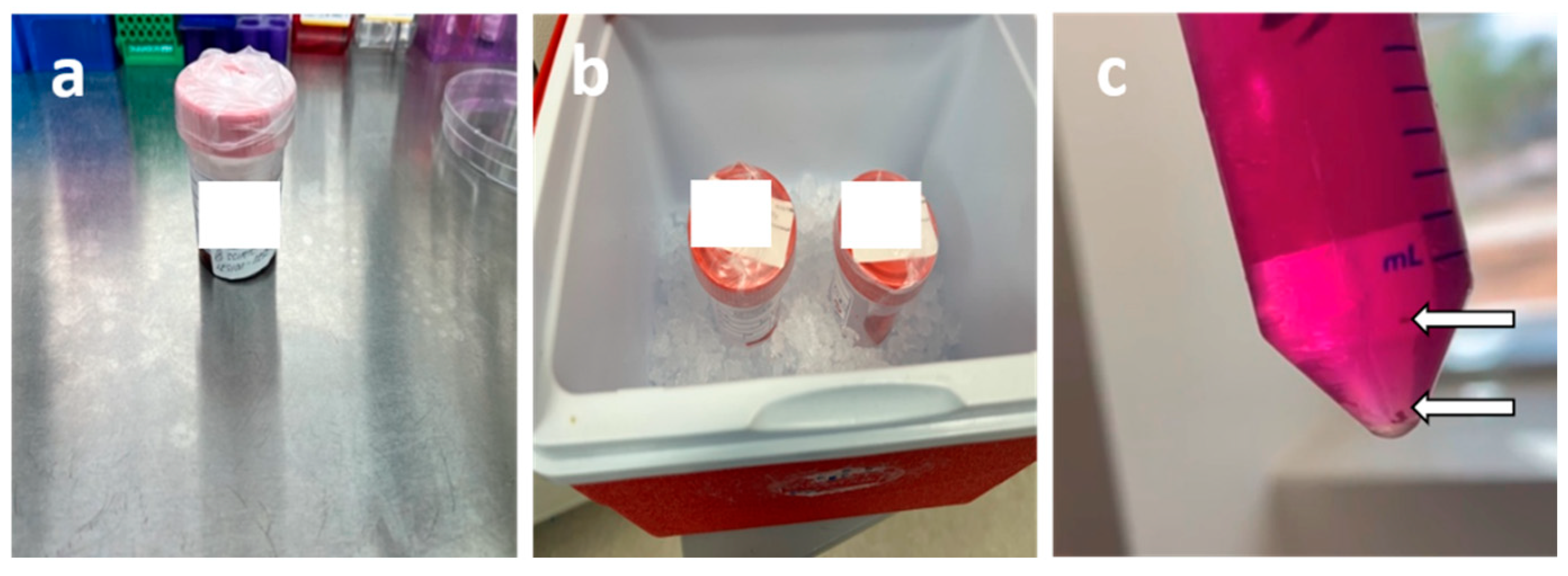
- Tissue fragments were washed in Hank's Balanced Salt Solution (HBSS) (Figure 12-a).
- After centrifugation, the supernatant was carefully discarded by manual aspiration, taking care not to dislodge the specimen.
- The sample was transferred to a sterile 10 cm tissue culture dish pre-cooled on ice.
- Working quickly, a scalpel blade was used to finely mince the tissue and cut it into small pieces to achieve coarse mechanical decomposition (Figure 12-b).
-
Once the tissue is finely minced, it is collected into a 50 mL Falcon tube:
- a.
- The minced tissue was transferred by tilting the dish at an angle and rinsing the dish with 10 mL of PBS with a 25 mL serological pipette.
- b.
- This process was repeated until all tissue had been collected into the conical tube.
- c.
- the sample was centrifuged at 300xg for 5 min at 4°C.
- Then the tissue sample was incubated for 1-2 h with enzyme dissociation medium (1 ml of HBSS + 1μl dispase1 + 0.1μl DNase 1 + 2.5μl collagenase) with manual flicking every 15 min to allow more cells to dissociate.
- Additionally, to enhance the digestion process, the tissue was processed with manual digestion tools (Figure 12-c).
- This is followed by centrifugation at 300xg for 3 min
- Discard the supernatant after centrifugation by carefully but swiftly decanting directly into the waste container without disturbing the pellet
- Using a 10 mL serological pipette, 3 mL of DMEM media was added, and the cell pellet was triturated until no visible large pieces were present. If large pieces remain, repeat enzymatic dissociation
- This is followed by centrifugation at 300×g for 3 min.
- The sediment is resuspended again in cell culture medium containing antibiotics Pen/Strep 5% and fetal bovine serum (FBS) 10%, and plated out into a 6 Well culture plate (Figure 12-d)
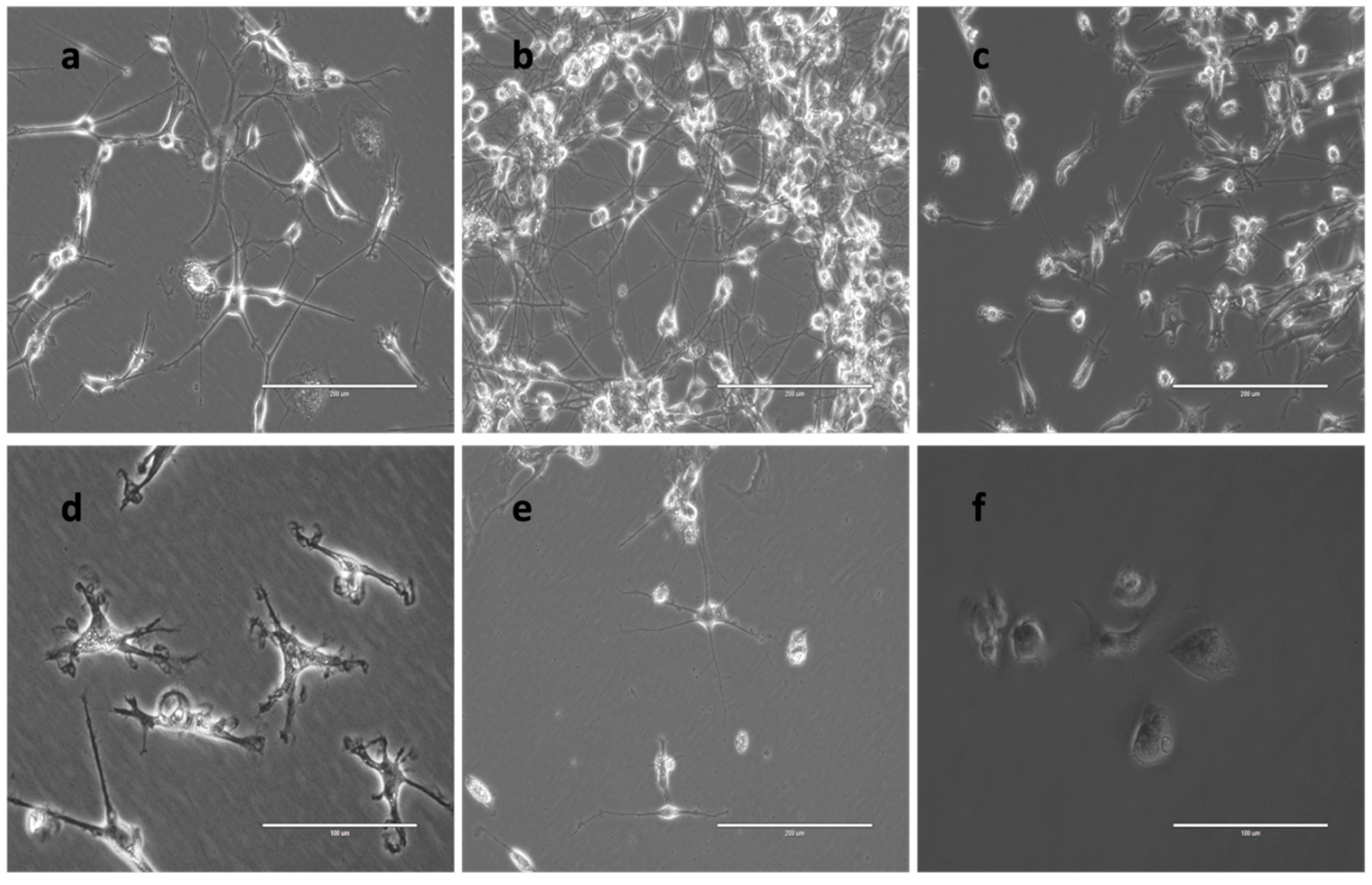
- The resulting cell suspension was incubated at 37 ◦C and 5% CO2 for approximately one month, resulting in preferential proliferation and survival of brain cells. The medium is normally changed twice a week. Explants were cultured in different media conditions to optimize viability (Figure 12-e)
- Cell culture media used for primary cells and their recipe (Table 4)
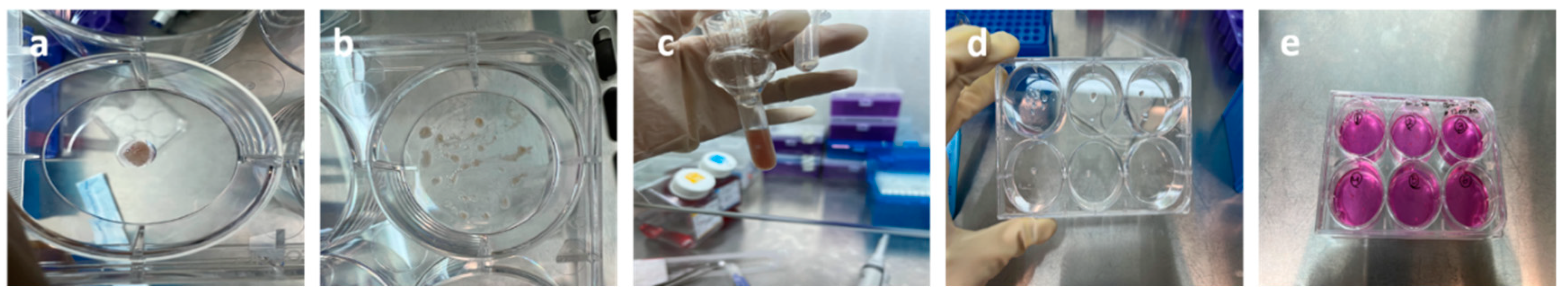

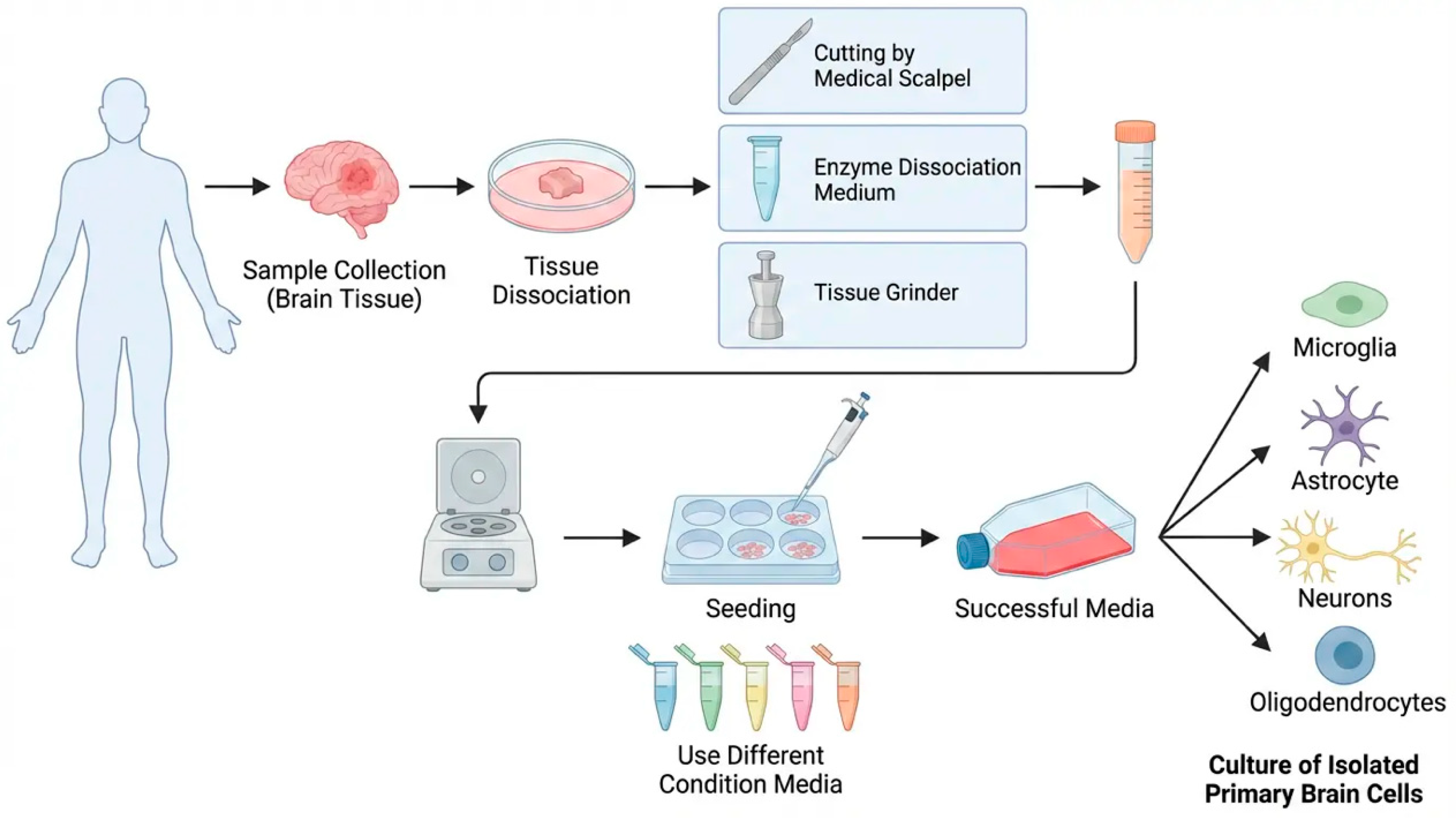
The Culture of Isolated Primary Brain cells
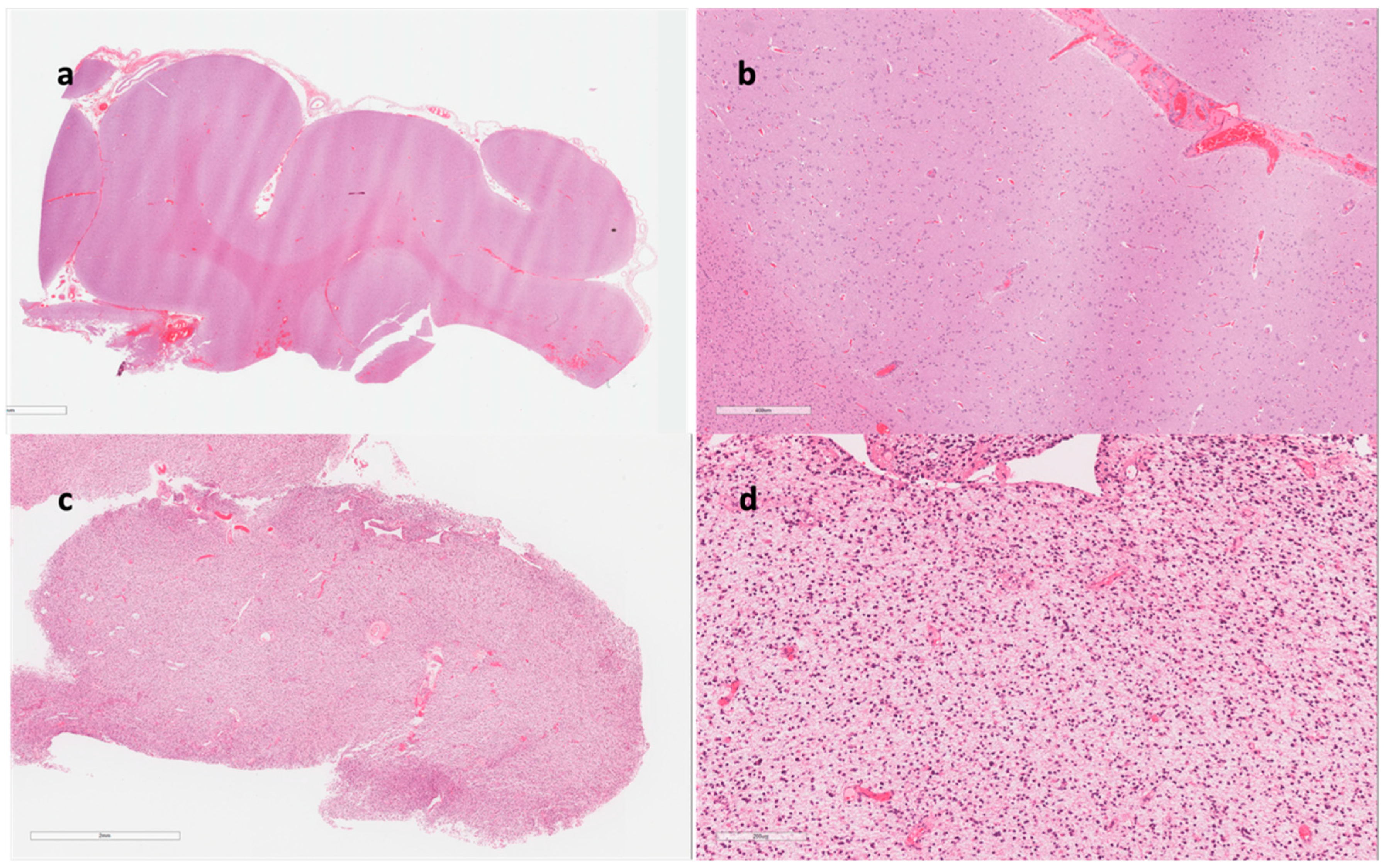
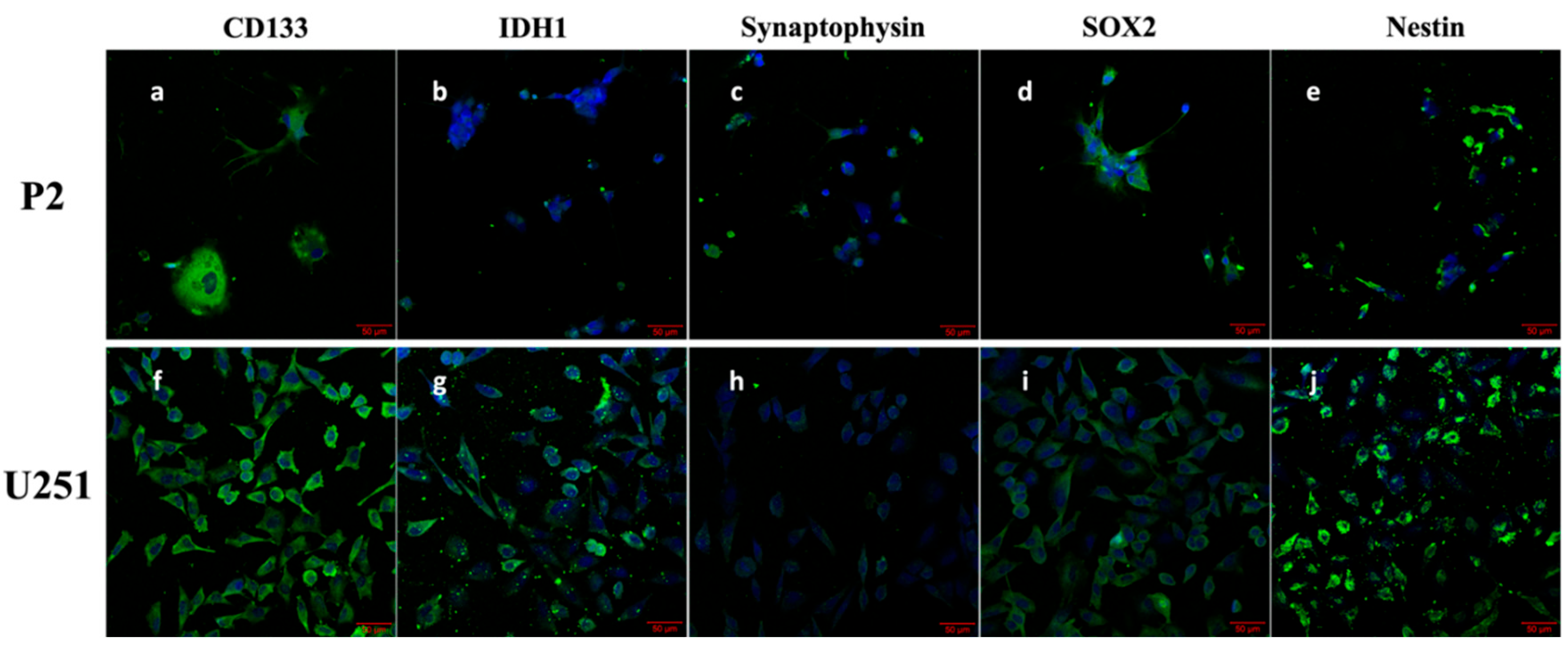
Cytology Analysis (Characterization)
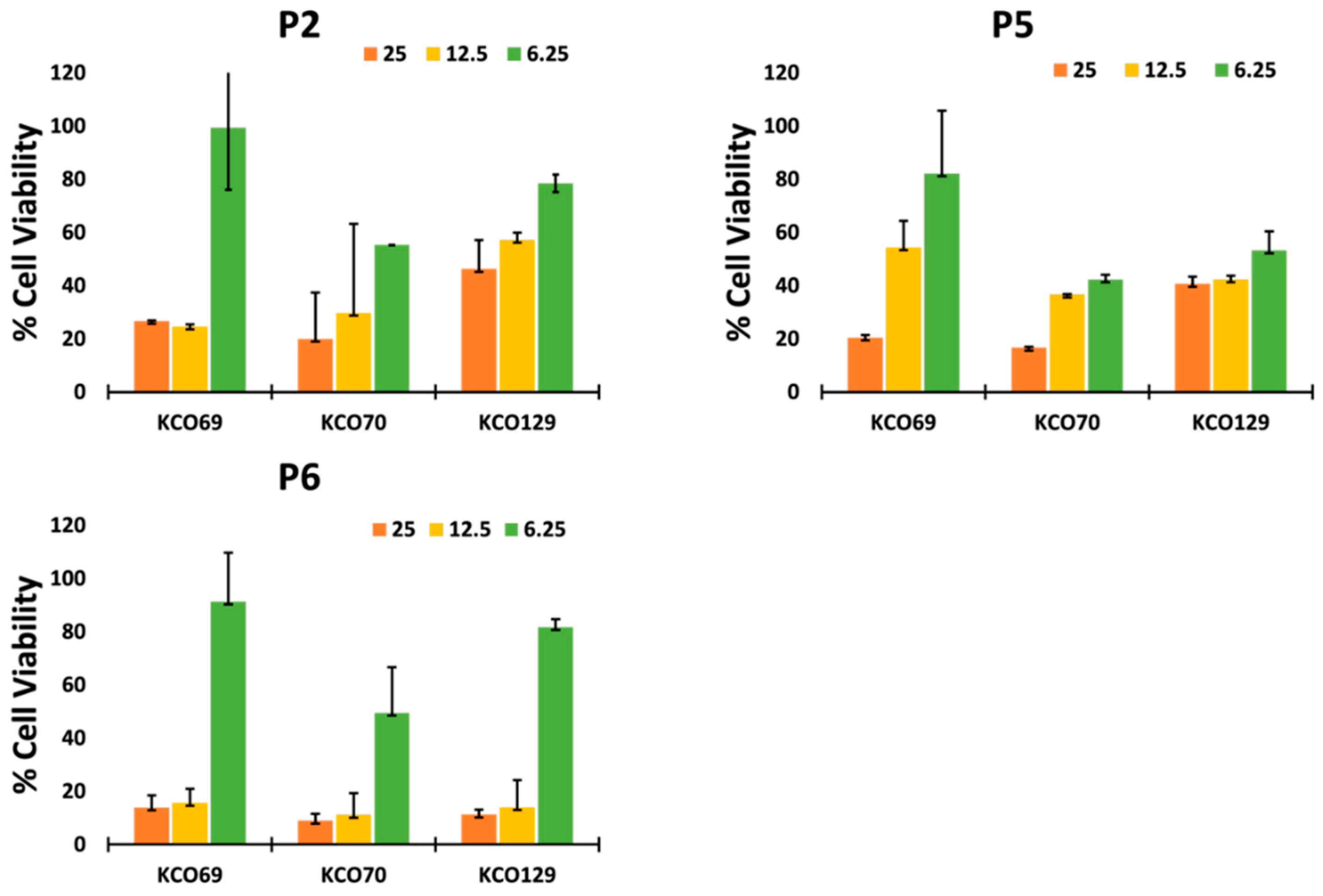
Cell Viability and Proliferation
Cell Culture and Treatment:
The CellTiter-Glo assay
Cell Culture and Treatment:
Immunocytochemistry (ICC)
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Roy, P. S.; Saikia, B. J. Cancer and cure: A critical analysis. Indian J. Cancer 2016, vol. 53(no. 3), 441–442. [Google Scholar] [CrossRef] [PubMed]
- Tandel, G. S. A Review on a Deep Learning Perspective in Brain Cancer Classification. Cancers 2019, vol. 11(no. 1). [Google Scholar] [CrossRef] [PubMed]
- Alessa, M.; Khan, A. S. Epidemiology of Colorectal Cancer in Saudi Arabia: A Review. Cureus 2024, vol. 16(no. 7). [Google Scholar] [CrossRef] [PubMed]
- Pichaivel, M. An Overview of Brain Tumor. In Brain Tumors; Jan 2022. [Google Scholar] [CrossRef]
- Hotchkiss, K. M.; Sampson, J. H. Temozolomide treatment outcomes and immunotherapy efficacy in brain tumor. J. Neurooncol. 2021, vol. 151(no. 1), 55–62. [Google Scholar] [CrossRef] [PubMed]
- Bellamy, C.; Hajal, C.; Ligon, K. L.; Touat, M. Revisiting an old therapy for new, promising combinations: Biology and perspectives of lomustine in glioma treatment. Neuro. Oncol. 2025, vol. 27(no. 12), 3043–3057. [Google Scholar] [CrossRef] [PubMed]
- Yamamuro, S. Lomustine and nimustine exert efficient antitumor effects against glioblastoma models with acquired temozolomide resistance. Cancer Sci. 2021, vol. 112(no. 11), 4736–4747. [Google Scholar] [CrossRef] [PubMed]
- Solimando, D. A.; Waddell, J. A. Procarbazine, Lomustine, and Vincristine (PCV) Regimen for Central Nervous System Tumors. Hosp. Pharm. 2017, vol. 52(no. 2), 98–104. [Google Scholar] [CrossRef] [PubMed]
- Mirza, Baig; et al. The use of Bevacizumab in the treatment of brain arteriovenous malformations: a systematic review. Neurosurg. Rev. 2025, vol. 48(no. 1). [Google Scholar] [CrossRef]
- Bombino; Magnani, M.; Conti, A. A Promising Breakthrough: The Potential of VORASIDENIB in the Treatment of Low-grade Glioma. Curr. Mol. Pharmacol. 2024, vol. 17. [Google Scholar] [CrossRef] [PubMed]
- Basu Roy, T.; Dokholyan, N. V. MT-125: A glimpse of hope for glioblastoma patients. Cell. Biomater. 2025, vol. 1(no. 8), 100160. [Google Scholar] [CrossRef]
- Kaur, G.; Dufour, J. M. Cell lines. Spermatogenesis 2012, vol. 2(no. 1), 1–5. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Margol, A.; Asgharzadeh, S.; Erdreich-Epstein, A. Pediatric brain tumor cell lines. J. Cell. Biochem. 2015, vol. 116(no. 2), 218–224. [Google Scholar] [CrossRef] [PubMed]
- Powley, R. Patient-derived explants (PDEs) as a powerful preclinical platform for anti-cancer drug and biomarker discovery. Br. J. Cancer 2020, vol. 122(no. 6), 735–744. [Google Scholar] [CrossRef] [PubMed]
- Jezierzański, M. Temozolomide (TMZ) in the Treatment of Glioblastoma Multiforme—A Literature Review and Clinical Outcomes. Curr. Oncol. 2024, vol. 31(no. 7), 3994. [Google Scholar] [CrossRef] [PubMed]
- Matada, S.; Pattanashettar, R.; Yernale, N. G. A comprehensive review on the biological interest of quinoline and its derivatives. Bioorg. Med. Chem. 2021, vol. 32. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, M. F. A.; Abuo-Rahma, G. E. D. A. Molecular targets and anticancer activity of quinoline–chalcone hybrids: literature review. RSC Adv. 2020, vol. 10(no. 52), 31139–31155. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, M. F. A.; Abuo-Rahma, G. E. D. A. Molecular targets and anticancer activity of quinoline–chalcone hybrids: literature review. RSC Adv. 2020, vol. 10(no. 52), 31139–31155. [Google Scholar] [CrossRef] [PubMed]
- Fabregat. The Reactome Pathway Knowledgebase. Nucleic Acids Res. 2018, vol. 46(no. D1), D649–D655. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Garcia, J. iFrag: A Protein–Protein Interface Prediction Server Based on Sequence Fragments. J. Mol. Biol. 2017, vol. 429(no. 3), 382–389. [Google Scholar] [CrossRef] [PubMed]
- Niu, J.; Li, M.; Wang, Y. Cell Proliferation and Cytotoxicity Assays, The Fundamentals for Drug Discovery; Scilight Press Pty. Ltd, 01 Sep 2024. [Google Scholar] [CrossRef]
- Adan; Kiraz, Y.; Baran, Y. The international international journal journal for timely for timely in-depth reviews in in-depth reviews in Pharmaceutical Pharmaceutical Biotechnology Impact Factor: 1.802 BENTHAM. Curr. Biotechnol. Pharm. 2016, vol. 17(no. 14), 1873–4316. [Google Scholar] [CrossRef]
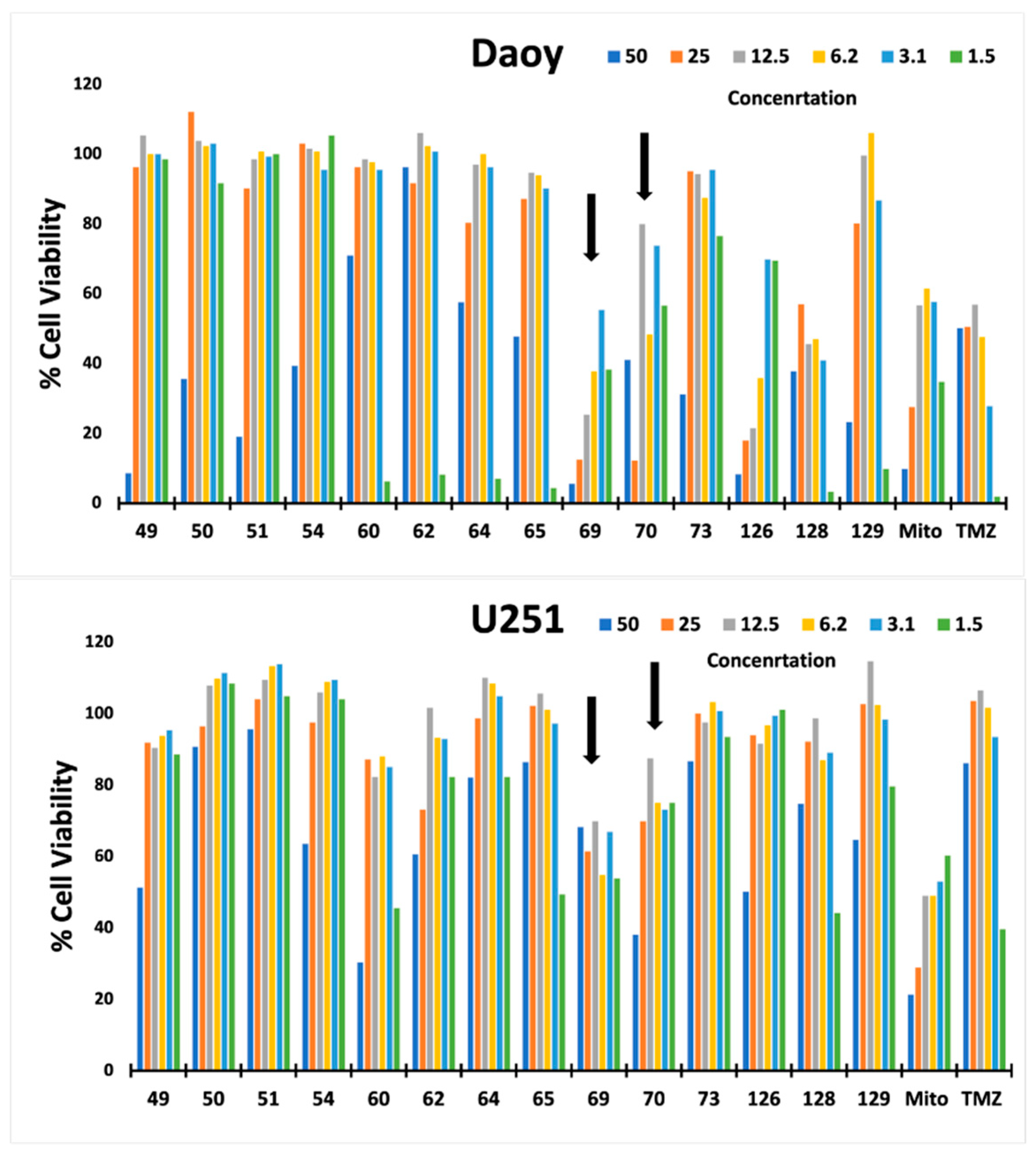
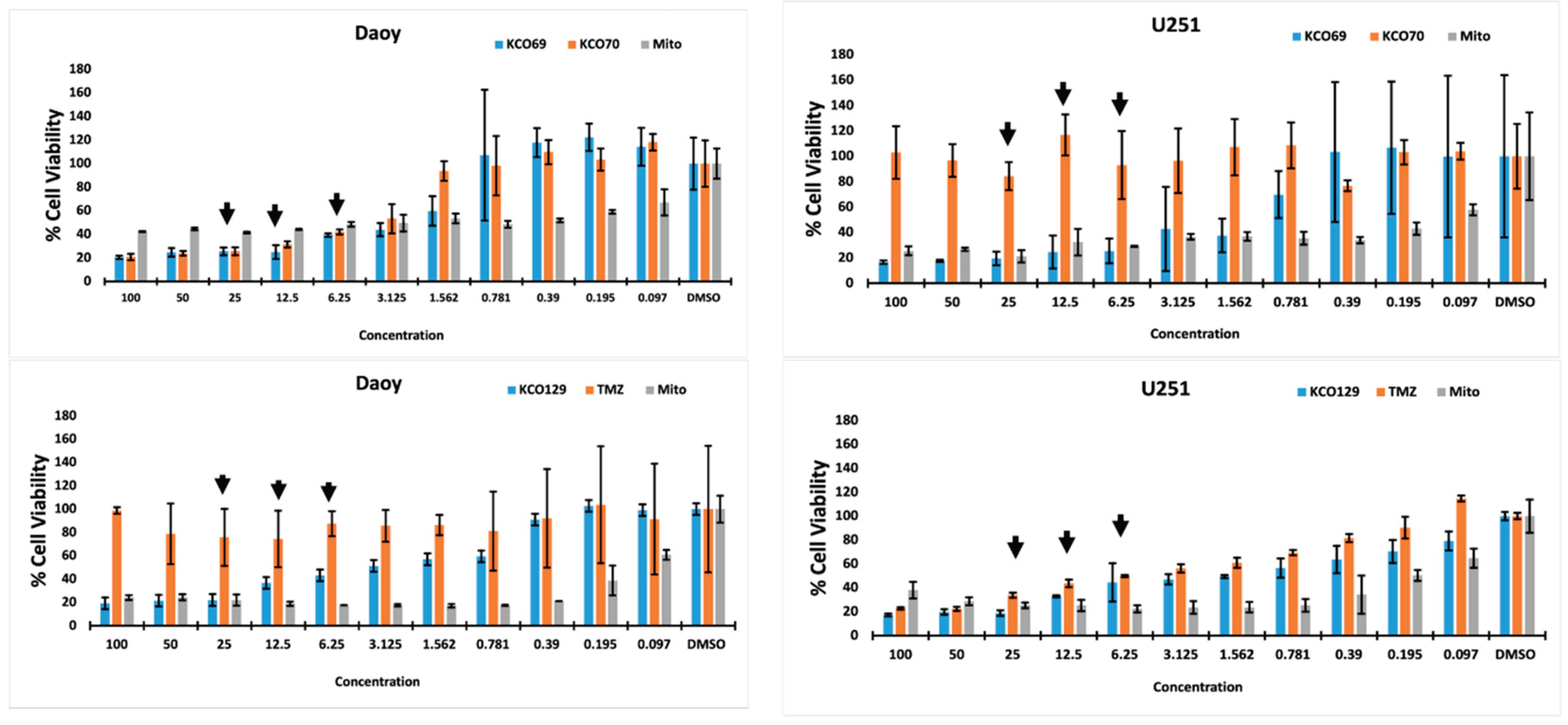
| KCO | U251 | Daoy | P2 | P3 | P5 | |||||
| IC50 | R2 | IC50 | R2 | IC50 | R2 | IC50 | R2 | IC50 | R2 | |
| 69 | 6 | 0.98 | 35 | 0.92 | 9 | 0.67 | 7 | 0.75 | 3 | 0.7 |
| 70 | 11 | 1.00 | 4 | 0.99 | 11 | 0.97 | 6 | 0.98 | 5 | 0.67 |
| 129 | 3 | 0.8 | 10 | 0.8 | 7 | 0.01 | 7 | 0.33 | 6 | 0.98 |
| KCO | P6 | P8 | P12 | P13 | ||||||
| IC50 | R2 | IC50 | R2 | IC50 | R2 | IC50 | R2 | |||
| 69 | 11 | 0.98 | 5 | 0.79 | 14 | 0.93 | 0.03 | 0.17 | ||
| 70 | 12 | 0.01 | 6 | 0.6 | 10 | 0.89 | 1.11 | 0.8 | ||
| 129 | 11 | 0.93 | 6 | 0.88 | 14 | 0.87 | 7 | 0.4 | ||
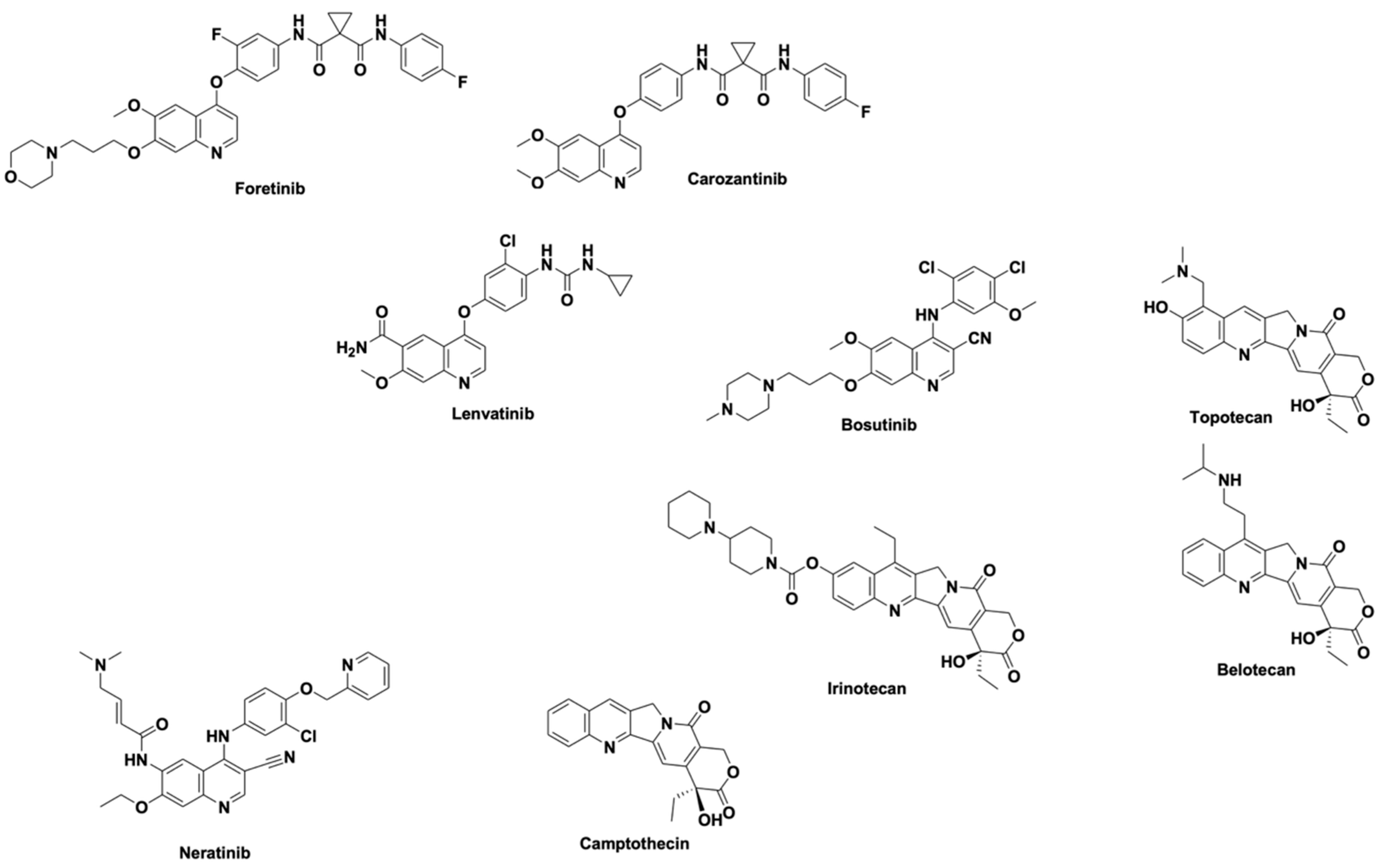
| Compound | Structure |
Molecular Pathways |
Database |
Confidence (p-value) |
| KCO69 | 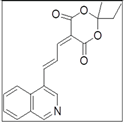 |
Polycyclic aromatic hydrocarbon degradation | KEGG Pathways | 0.0251 |
| Type I polyketide structures | 0.0362 | |||
| Constitutive Signaling by Aberrant PI3K in Cancer | Reactome | 0.0319 | ||
| KCO70 | 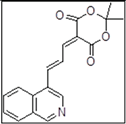 |
Polycyclic aromatic hydrocarbon degradation | KEGG Pathways | 0.0209 |
| Constitutive Signaling by Aberrant PI3K in Cancer | Reactome | 0.0289 | ||
| KCO129 | 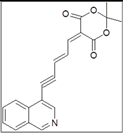 |
Biosynthesis of enediyne antibiotics | KEGG Pathways | 0.0225 |
| Constitutive Signaling by Aberrant PI3K in Cancer | Reactome | 0.0587 |
| Abbreviation | Diagnosis | Age | Gander | Location |
| P1 | Brain Tumor-Metastatic carcinoma | 68 | M | Right Frontal |
| P2 | Brain Tumor- High-Grade Astrocytoma, WHO grade4 | 33 | M | Left Frontal |
| P3 | Brain Tumor-Schwannoma, WHO grade 1 | 43 | M | Right Sciatic nerve tumor |
| P4 | Meningioma | 48 | F | Olfactory groove |
| P5 | Brain Tumor-Astrocytoma, WHO grade 2 | 38 | M | Right parietal lobe |
| P6 | Cerebral Tissue Control | 24 | M | Right frontal |
| P7 | Mammosomatotroph PitNET/adenoma | 39 | M | Pituitary region |
| P8 | Brain Tumor -Glioblastoma, WHO grade4 | 39 | M | Left temporal |
| P9 | Brain Tumor -Infant-type hemispheric glioma | 9M | F | Right Temporoparietal |
| P10 | Metastatic colonic adenocarcinoma | 61 | M | Left cerebellar tumor |
| P11 | Primary large B-cell lymphoma of the CNS | 63 | M | Left occipital |
| P12 | Medulloblastoma, WHO grade4 | 6y | F | Left cerebellar tumor |
| P13 | Anaplastic meningioma, WHO grade3 | 55 | M | Left frontal |
| P14 | Brain Tumor- Glioblastoma -grade 4 | 51 | F | Right frontal lobe |
| P15 | Brain Tumor-Meningioma, WHO grade1 | 55 | F | Left temporal |
| P16 | Brain Tumor-Glioblastoma WHO, grade 4 | 57 | M | Right hemispheric lesion |
| Basic Growth Media | |
| Component | Volume for 500 ml |
| A DMEM | 450 ml |
| FBS serum 10 % | 50 ml |
| Pen/Strep | 5 ml |
| Glutamine | 5 ml |
| Cell medium (1) | |
| DMEM/F-12 Complete | 22,5 ml |
| DMEM gultamax Complete | 22,5 ml |
| FBS serum 20 % | 5 ml |
| Gentamicin (50µg/ml) | 12,5µl |
| ITS 0.1% | 50µl |
| EGF stock | 50µl |
| HEPES 1 M | 550µl |
| EGF stock (1 µL in 100 µl H2O) 20ng/ml | |
| Cell medium (2) | |
| DMEM/F-12 Complete | 22,5 ml |
| DMEM gultamax Complete | 22,5 ml |
| B-27 | 1ml |
| Gentamicin (50µg/ml) | 12,5µl |
| ITS 0.1% | 50µl |
| EGF stock | 50µl |
| HEPES 1 M | 550µl |
| Cell medium (3) | |
| NeuroCult™ Neural Cell Culture Media |
Volume for 50 ml |
| BDNF (500ng/ml) | 25µl |
| FBS serum 10 % | 5ml |
| Pen/Strep | 500µl |
| Cell medium (4) | |
| Complete Advanced DMEM | Volume for 20 ml |
| Human Recombinant Brain-derived neurotrophic factor (BDNF) (500ng/ml) | 10µl |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




