Submitted:
28 April 2026
Posted:
29 April 2026
You are already at the latest version
Abstract
Keywords:
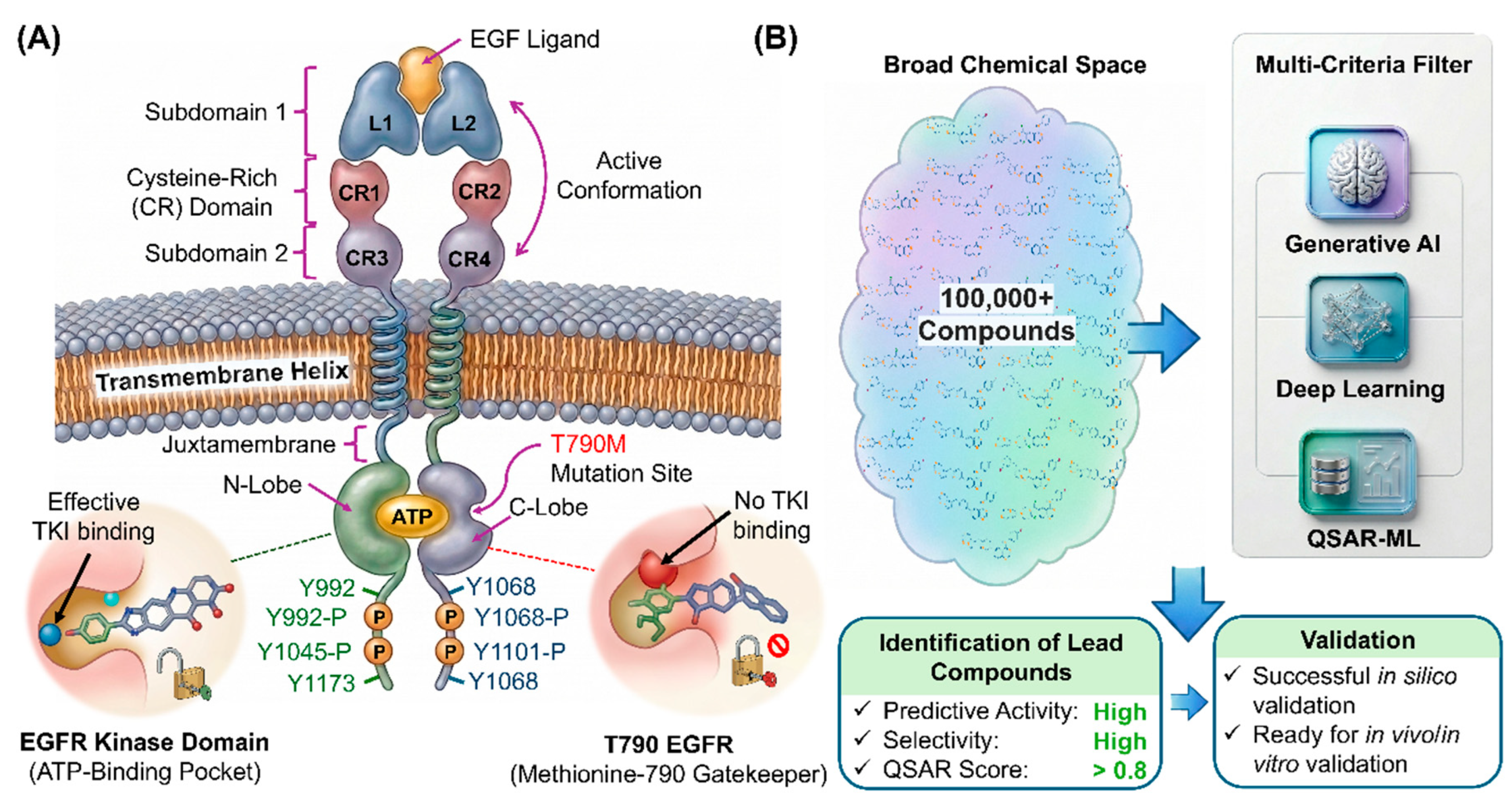
1. Introduction
2. Materials and Methods
2.1. Dataset Preparation and QSAR Model Construction
2.2. Molecular Generation and Hierarchical Chemical Space Refinement
2.3. Molecular Docking and Active Site Preparation
2.4. Molecular Dynamics (MD) Simulations
2.5. Binding Free Energy Calculation (MMPBSA)
2.6. Free Energy Landscape (FEL) Construction
2.8. Statistical Analysis and Data Validation
3. Results
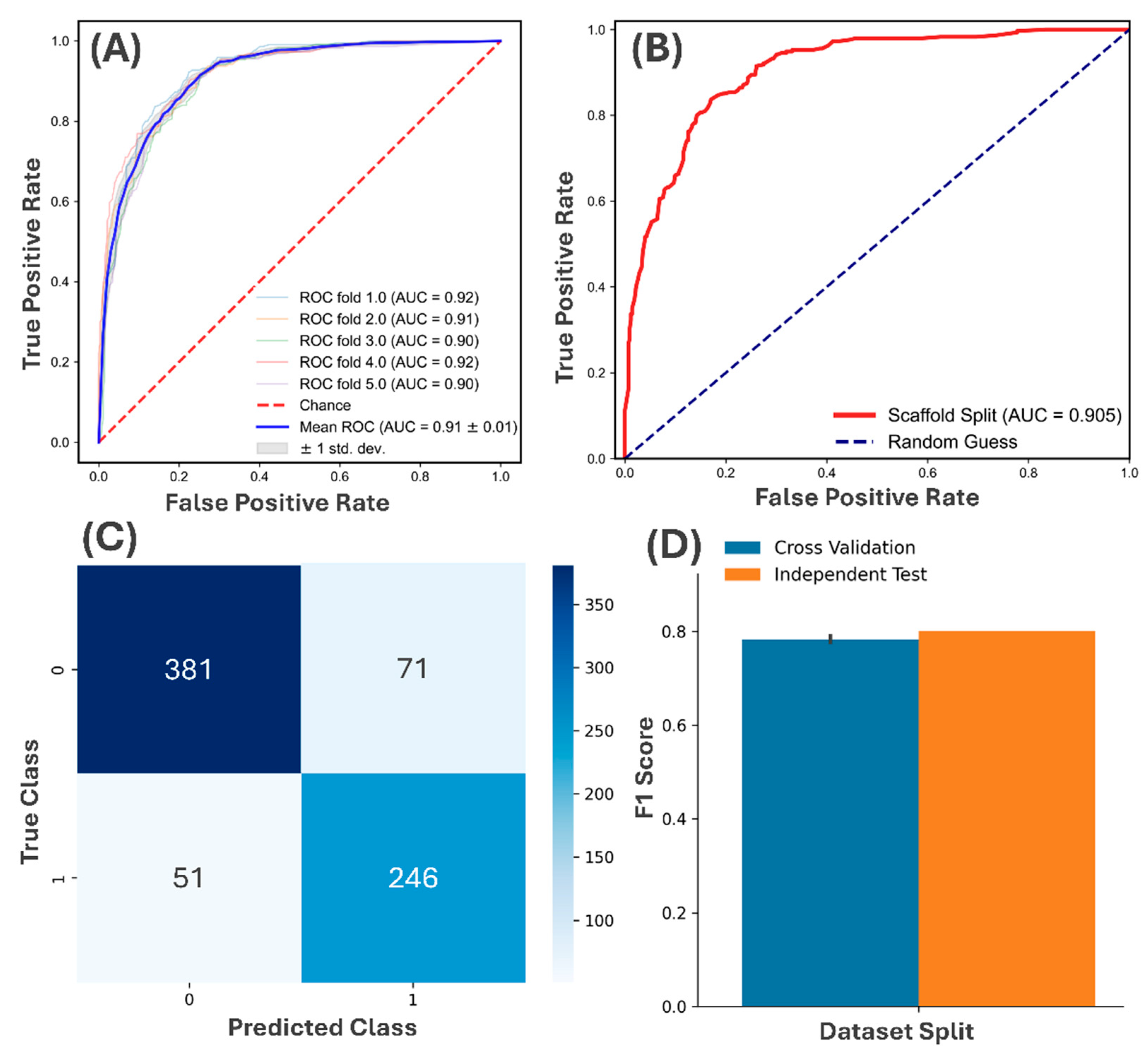
3.1. QSAR Model Validation and Generative Sampling Performance
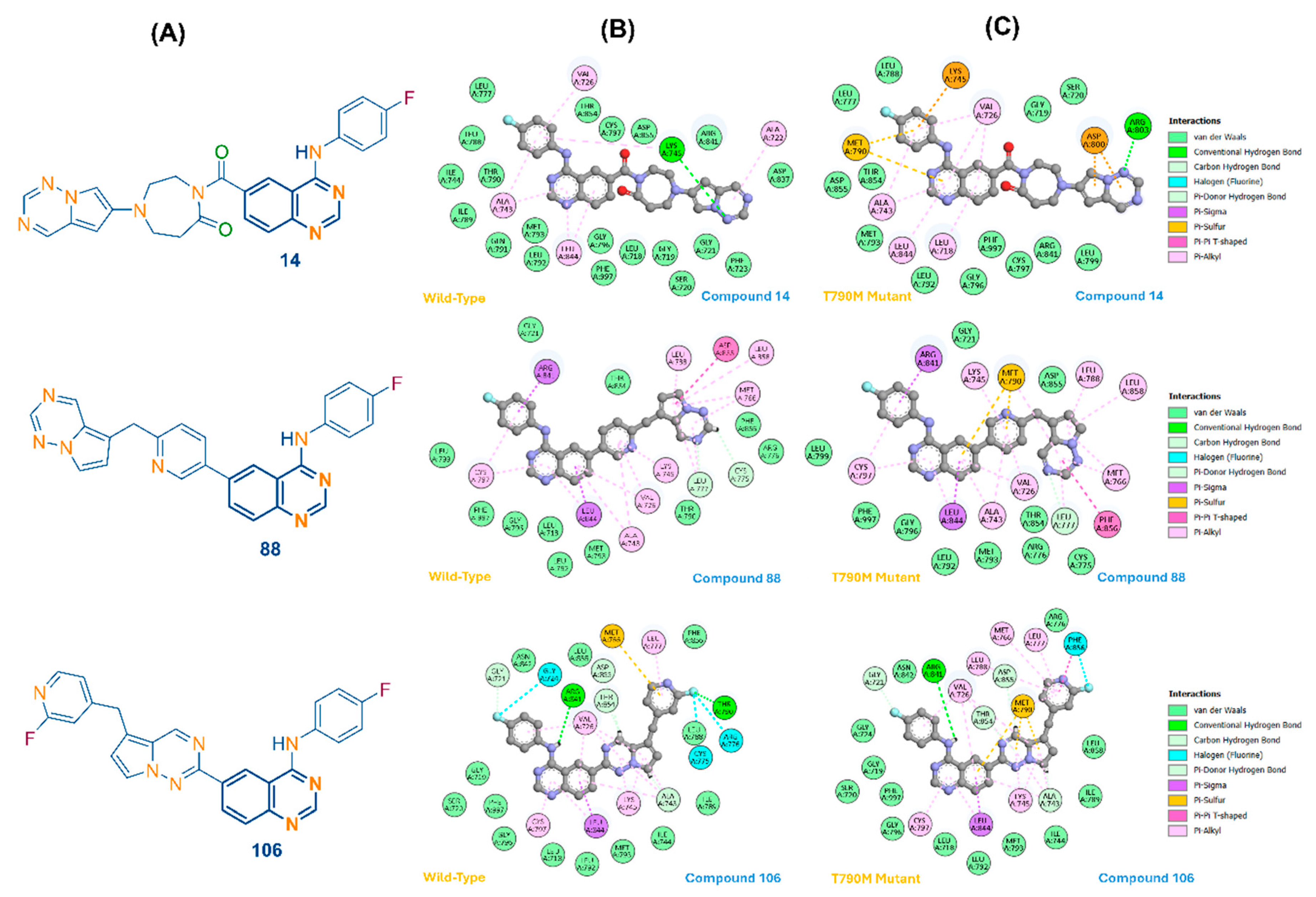
3.2. Molecular Docking and Active Site Comparison
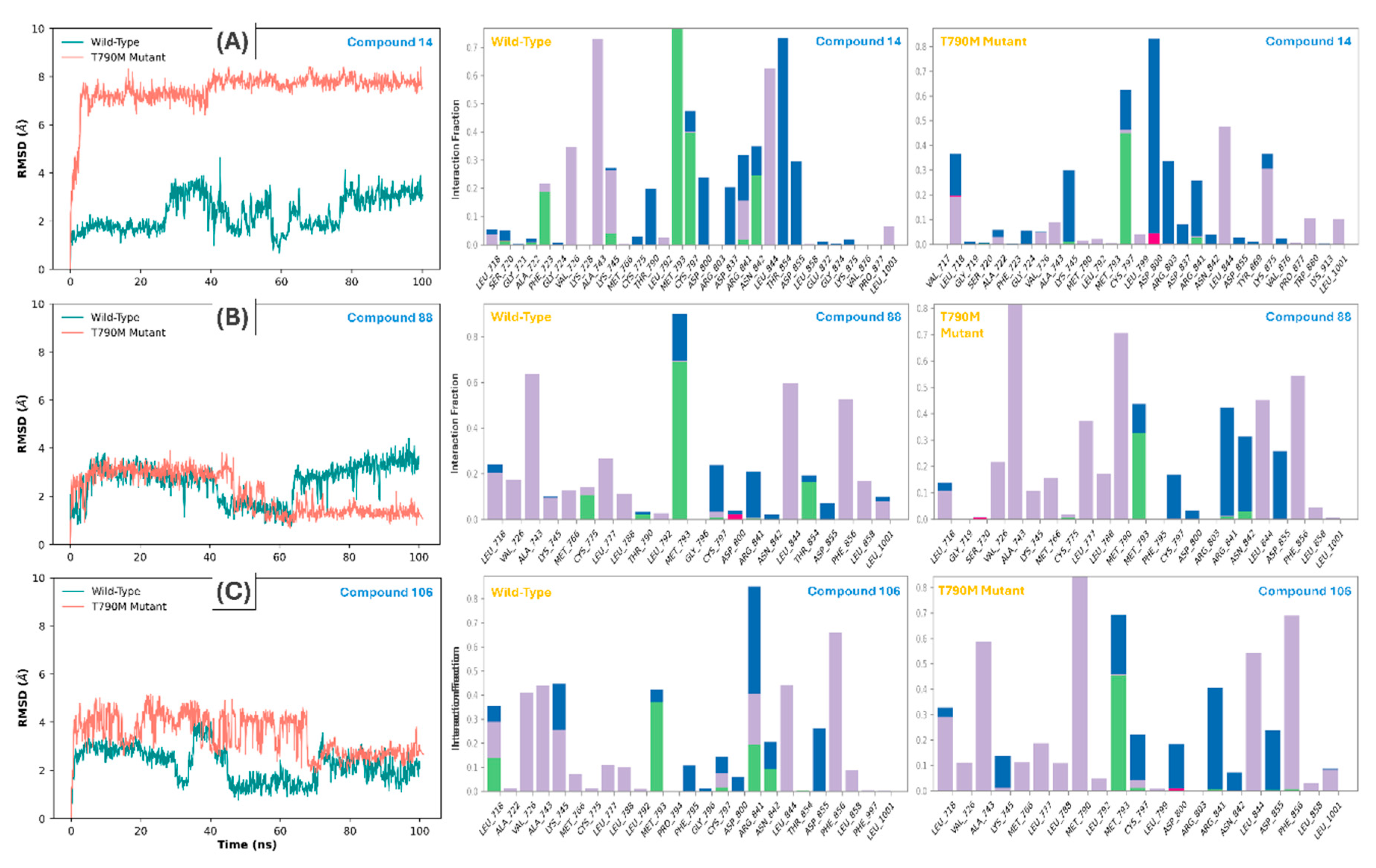
3.3. Molecular Dynamics and Interaction Persistence
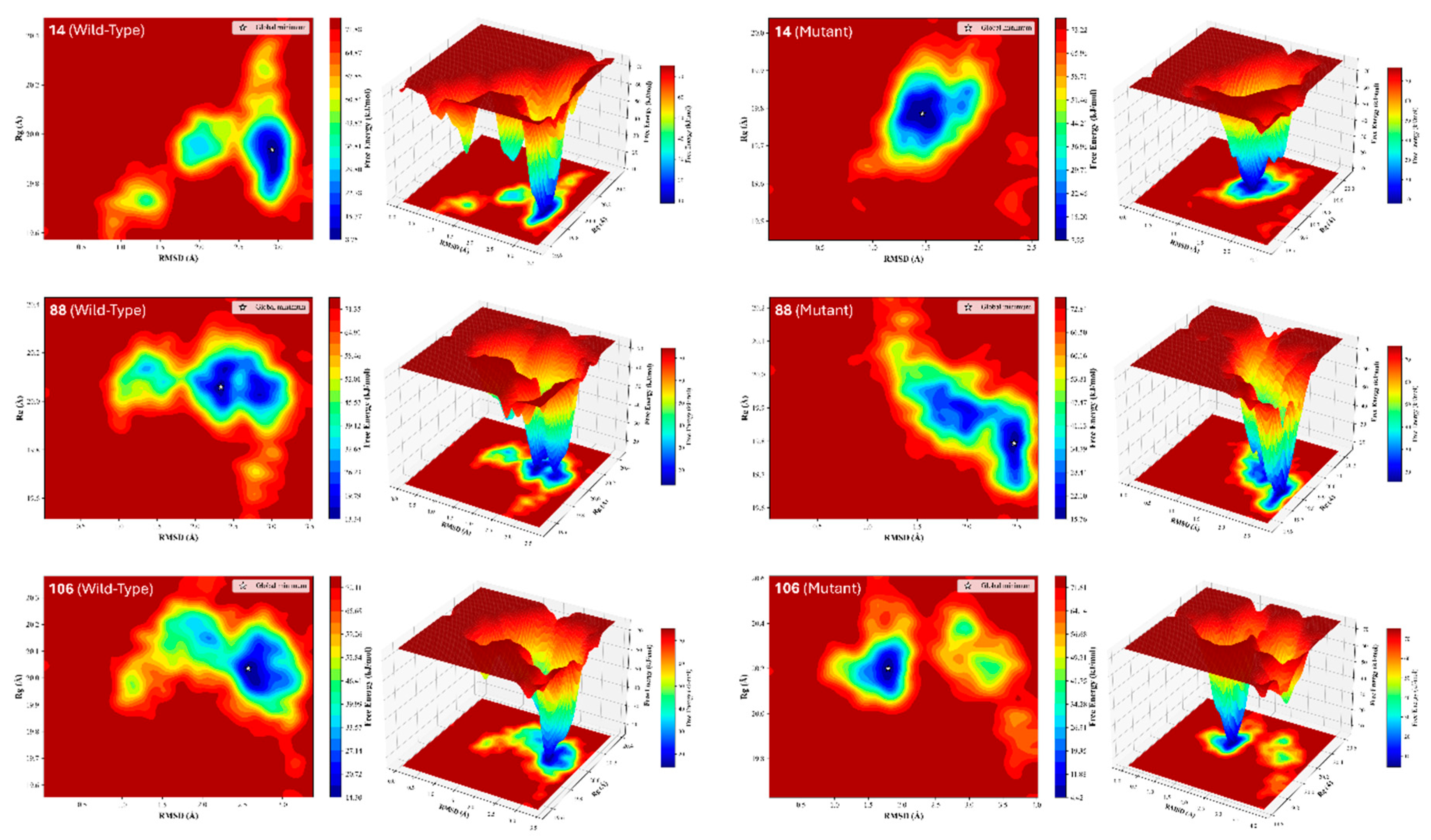
3.4. Thermodynamic Basins and Stereochemical Quality
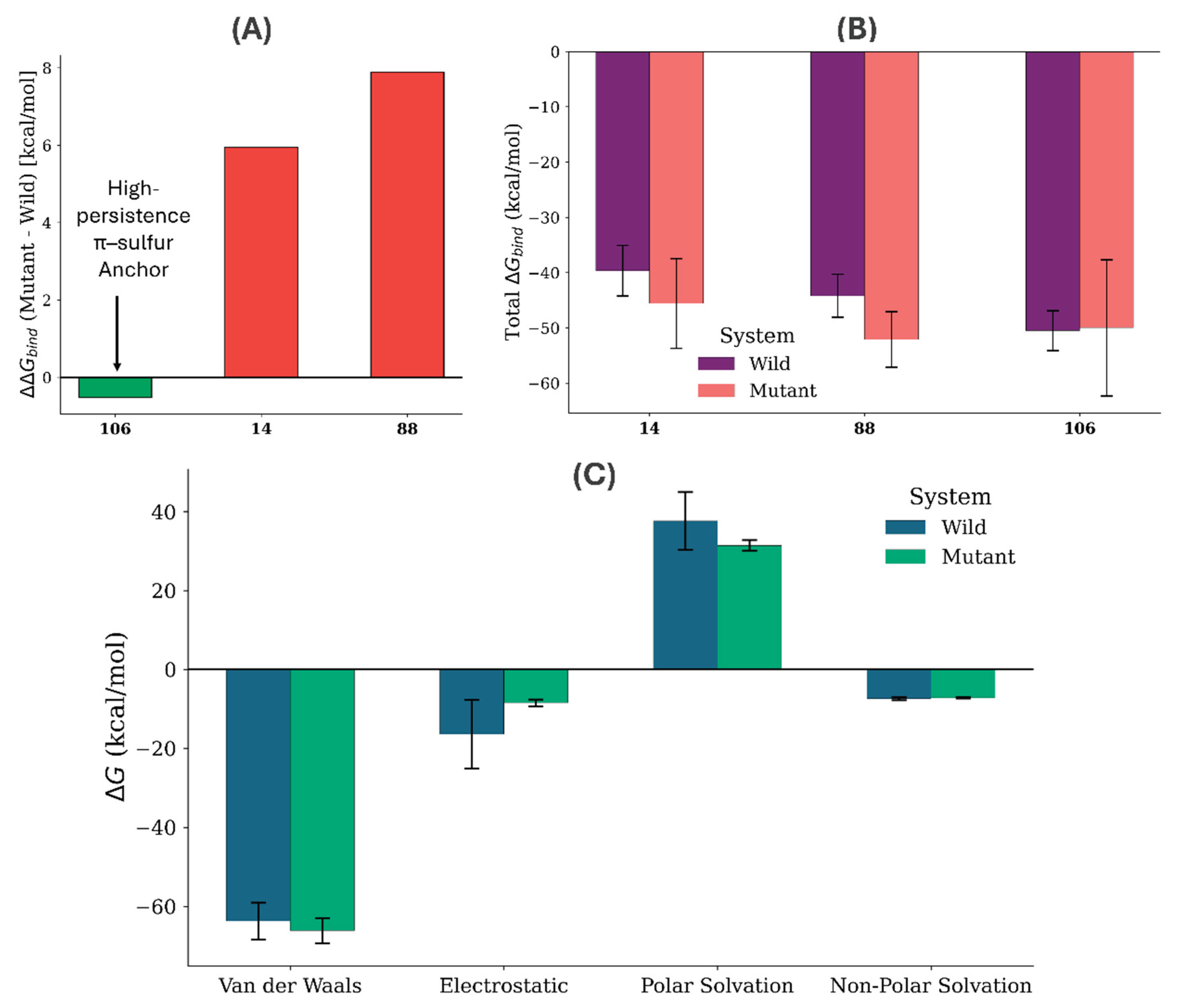
3.5. Ensemble-Based Binding Free Energy (MM-GBSA)
3.6. Statistical Validation and Trajectory Convergence
4. Discussion
4.1. Mechanistic Adaptation to the T790M Mutation
4.2. Comparison with Covalent Strategies and Resistance Bypass
4.3. Energetic Partitioning and the Strategic Roadmap for Lead Optimization
4.4. Strategic Directions for Experimental Translation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lynch, T.J.; Bell, D.W.; Sordella, R.; Gurubhagavatula, S.; Okimoto, R.A.; Brannigan, B.W.; Harris, P.L.; Haserlat, S.M.; Supko, J.G.; Haluska, F.G.; Louis, D.N.; Christiani, D.C.; Settleman, J.; Haber, D.A. Activating Mutations in the Epidermal Growth Factor Receptor Underlying Responsiveness of Non–Small-Cell Lung Cancer to Gefitinib. N. Engl. J. Med. 2004, 350, 2129–2139. [Google Scholar] [CrossRef]
- Paez, J.G.; Jänne, P.A.; Lee, J.C.; Tracy, S.; Greulich, H.; Gabriel, S.; Herman, P.; Kaye, F.J.; Lindeman, N.; Boggon, T.J.; Naoki, K.; Sasaki, H.; Fujii, Y.; Eck, M.J.; Sellers, W.R.; Johnson, B.E.; Meyerson, M. EGFR Mutations in Lung Cancer: Correlation with Clinical Response to Gefitinib Therapy. Science 2004, 304, 1497–1500. [Google Scholar] [CrossRef]
- Mok, T.S.; Wu, Y.-L.; Thongprasert, S.; Yang, C.-H.; Chu, D.-T.; Saijo, N.; Sunpaweravong, P.; Han, B.; Margono, B.; Ichinose, Y.; Nishiwaki, Y.; Ohe, Y.; Yang, J.-J.; Chewaskulyong, B.; Jiang, H.; Duffield, E.L.; Watkins, C.L.; Armour, A.A.; Fukuoka, M. Gefitinib or Carboplatin–Paclitaxel in Pulmonary Adenocarcinoma. N. Engl. J. Med. 2009, 361, 947–957. [Google Scholar] [CrossRef]
- Rosell, R.; Carcereny, E.; Gervais, R.; Vergnenegre, A.; Massuti, B.; Felip, E.; Palmero, R.; Garcia-Gomez, R.; Pallares, C.; Sanchez, J.M.; Porta, R.; Cobo, M.; Garrido, P.; Longo, F.; Moran, T.; Insa, A.; De Marinis, F.; Corre, R.; Bover, I.; Illiano, A.; Dansin, E.; de Castro, J.; Milella, M.; Reguart, N.; Altavilla, G.; Jimenez, U.; Provencio, M.; Moreno, M.A.; Terrasa, J.; Muñoz-Langa, J.; Valdivia, J.; Isla, D.; Domine, M.; Molinier, O.; Mazieres, J.; Baize, N.; Garcia-Campelo, R.; Robinet, G.; Rodriguez-Abreu, D.; Lopez-Vivanco, G.; Gebbia, V.; Ferrera-Delgado, L.; Bombaron, P.; Bernabe, R.; Bearz, A.; Artal, A.; Cortesi, E.; Rolfo, C.; Sanchez-Ronco, M.; Drozdowskyj, A.; Queralt, C.; de Aguirre, I.; Ramirez, J.L.; Sanchez, J.J.; Molina, M.A.; Taron, M.; Paz-Ares, L. Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): A multicentre, open-label, randomised phase 3 trial. Lancet Oncol. 2012, 13, 239–246. [Google Scholar] [CrossRef] [PubMed]
- Yun, C.-H.; Boggon, T.J.; Li, Y.; Woo, M.S.; Greulich, H.; Meyerson, M.; Eck, M.J. Structures of Lung Cancer-Derived EGFR Mutants and Inhibitor Complexes: Mechanism of Activation and Insights into Differential Inhibitor Sensitivity. Cancer Cell. 2007, 11, 217–227. [Google Scholar] [CrossRef] [PubMed]
- Rotow, J.; Bivona, T. Understanding and Targeting Resistance Mechanisms in NSCLC. Nat. Rev. Cancer 2017, 17, 637–658. [Google Scholar] [CrossRef]
- Camidge, D.; Pao, W.; Sequist, L. Acquired Resistance to TKIs in Solid Tumours: Learning from Lung Cancer. Nat. Rev. Clin. Oncol. 2014, 11, 473–481. [Google Scholar] [CrossRef]
- Pao, W.; Miller, V.A.; Politi, K.A.; Riely, G.J.; Somwar, R.; Zakowski, M.F.; Kris, M.G.; Varmus, H. Acquired Resistance of Lung Adenocarcinomas to Gefitinib or Erlotinib Is Associated with a Second Mutation in the EGFR Kinase Domain. PLoS Med. 2005, 2, e73. [Google Scholar] [CrossRef]
- Yu, H.A.; Arcila, M.E.; Rekhtman, N.; Sima, C.S.; Zakowski, M.F.; Pao, W.; Kris, M.G.; Miller, V.A.; Ladanyi, M.; Riely, G.J. Analysis of Tumor Specimens at the Time of Acquired Resistance to EGFR-TKI Therapy in 155 Patients with EGFR-Mutant Lung Cancers. Clin. Cancer Res. 2013, 19, 2240–2247. [Google Scholar] [CrossRef]
- Yun, C.-H.; Mengwasser, K.E.; Toms, A.V.; Woo, M.S.; Greulich, H.; Wong, K.-K.; Meyerson, M.; Eck, M.J. The T790M Mutation in EGFR Kinase Causes Drug Resistance by Increasing the Affinity for ATP. Proc. Natl. Acad. Sci. USA 2008, 105, 2070–2075. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, S.; Boggon, T.J.; Dayaram, T.; Jänne, P.A.; Kocher, O.; Meyerson, M.; Johnson, B.E.; Eck, M.J.; Tenen, D.G.; Halmos, B. EGFR Mutation and Resistance of Non–Small-Cell Lung Cancer to Gefitinib. N. Engl. J. Med. 2005, 352, 786–792. [Google Scholar] [CrossRef]
- Cross, D.A.E.; Ashton, S.E.; Ghiorghiu, S.; Eberlein, C.; Nebhan, C.A.; Spitzler, P.J.; Orme, J.P.; Finlay, M.R.V.; Ward, R.A.; Mellor, M.J.; Hughes, G.; Rahi, A.; Jacobs, V.N.; Red Brewer, M.; Ichihara, E.; Sun, J.; Jin, H.; Ballard, P.; Al-Kadhimi, K.; Rowlinson, R.; Klinowska, T.; Richmond, G.H.P.; Cantarini, M.; Kim, D.-W.; Ranson, M.R.; Pao, W. AZD9291, an Irreversible EGFR TKI, Overcomes T790M-Mediated Resistance to EGFR Inhibitors in Lung Cancer. Cancer Discov. 2014, 4, 1046–1061. [Google Scholar] [CrossRef]
- Jänne, P.A.; Yang, J.C.-H.; Kim, D.-W.; Planchard, D.; Ohe, Y.; Ramalingam, S.S.; Ahn, M.-J.; Kim, S.-W.; Su, W.-C.; Horn, L.; Haggstrom, D.; Felip, E.; Kim, J.-H.; Frewer, P.; Cantarini, M.; Brown, K.H.; Dickinson, P.A.; Ghiorghiu, S.; Ranson, M. AZD9291 in EGFR Inhibitor–Resistant Non–Small-Cell Lung Cancer. N. Engl. J. Med. 2015, 372, 1689–1699. [Google Scholar] [CrossRef]
- Thress, K.S.; Paweletz, C.P.; Felip, E.; Cho, B.C.; Stetson, D.; Dougherty, B.; Lai, Z.; Markovets, A.; Vivancos, A.; Kuang, Y.; Ercan, D.; Matthews, S.E.; Cantarini, M.; Barrett, J.C.; Jänne, P.A.; Oxnard, G.R. Acquired EGFR C797S Mutation Mediates Resistance to AZD9291 in Non–Small Cell Lung Cancer Harboring EGFR T790M. Nat. Med. 2015, 21, 560–562. [Google Scholar] [CrossRef]
- Wang, R.; Chen, Y.; Li, L.; Zhang, L.; Zhang, S. Osimertinib Acquired Resistance among Patients with EGFR-Mutated NSCLC: From Molecular Mechanisms to Clinical Therapeutic Strategies. Cancer Drug. Resist. 2025, 8, 61. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Piotrowska, Z.; Soo, R.; Cho, B.C.; Lim, S.M. Combatting Acquired Resistance to Osimertinib in EGFR-Mutant Lung Cancer. Ther. Adv. Med. Oncol. 2022, 14, 17588359221144099. [Google Scholar] [CrossRef]
- Romaniello, D.; Morselli, A.; Marrocco, I. Strategies to Overcome Resistance to Osimertinib in EGFR-Mutated Lung Cancer. Int. J. Mol. Sci. 2025, 26, 2957. [Google Scholar] [CrossRef]
- Shi, K.; Wang, G.; Pei, J.; Zhang, J.; Wang, J.; Ouyang, L.; Wang, Y.; Li, W. Emerging Strategies to Overcome Resistance to Third-Generation EGFR Inhibitors. J. Hematol. Oncol. 2022, 15, 94. [Google Scholar] [CrossRef]
- To, C.; Jang, J.; Chen, T.; Park, E.; Mushajiang, M.; De Clercq, D.J.H.; Xu, M.; Wang, S.; Cameron, M.D.; Heppner, D.E.; Shin, B.H.; Gero, T.W.; Yang, A.; Dahlberg, S.E.; Wong, K.K.; Eck, M.J.; Gray, N.S.; Jänne, P.A. Single and Dual Targeting of Mutant EGFR with an Allosteric Inhibitor. Cancer Discov. 2019, 9, 926–943. [Google Scholar] [CrossRef] [PubMed]
- Damghani, T.; Song, S.; Lin, K.S.; Li, J.; Heppner, D.E. Structural Studies of Fourth-Generation EGFR Inhibitors Reveal Insights into Selective T790M and C797S Targeting. ACS Med. Chem. Lett. 2026, in press. [Google Scholar] [CrossRef] [PubMed]
- Pranata, J. Sulfur–Aromatic Interactions: A Computational Study of the Dimethyl Sulfide–Benzene Complex. Bioorg. Chem. 1997, 25, 213–219. [Google Scholar] [CrossRef]
- Jin, Y.; Li, W.; Saragi, R.T.; Juanes, M.; Pérez, C.; Lesarri, A.; Feng, G. Sulfur–Arene Interactions: The S…π and S–H…π Interactions in the Dimers of Benzofuran…Sulfur Dioxide and Benzofuran…Hydrogen Sulfide. Phys. Chem. Chem. Phys. 2023, 25, 12174–12181. [Google Scholar] [CrossRef]
- Damghani, T.; Song, S.; Lin, K. S.; Li, J.; Heppner, D. E. Structural Studies of Fourth-Generation EGFR Inhibitors Reveal Insights into Selective T790M and C797S Targeting 10.1021/acsmedchemlett.5c00725. ACS medicinal chemistry letters 2026. Advance online publication. [Google Scholar] [CrossRef]
- Bissantz, C.; Kuhn, B.; Stahl, M. A Medicinal Chemist’s Guide to Molecular Interactions. J. Med. Chem. 2010, 53, 5061–5084. [Google Scholar] [CrossRef] [PubMed]
- Valley, C.C.; Cembran, A.; Perlmutter, J.D.; Lewis, A.K.; Labello, N.P.; Gao, J.; Sachs, J.N. The Methionine-Aromatic Motif Plays a Unique Role in Stabilizing Protein Structure. J. Biol. Chem. 2012, 287, 34979–34991. [Google Scholar] [CrossRef]
- Nagasaka, M.; Zhu, V.W.; Lim, S.M.; Greco, M.; Wu, F.; Ou, S.-H.I. Beyond Osimertinib: The Development of Third-Generation EGFR Tyrosine Kinase Inhibitors for Advanced EGFR+ NSCLC. J. Thorac. Oncol. 2021, 16, 740–763. [Google Scholar] [CrossRef]
- Dobson, C.M. Chemical Space and Biology. Nature 2004, 432, 824–828. [Google Scholar] [CrossRef] [PubMed]
- Restrepo, G. Expanding Diversity in Chemical Space. Nat. Chem. 2026, 18, 607–608. [Google Scholar] [CrossRef]
- Šícho, M.; Luukkonen, S.; van den Maagdenberg, H.W.; Schoenmaker, L.; Béquignon, O.J.M.; van Westen, G.J.P. DrugEx: Deep Learning Models and Tools for Exploration of Drug-Like Chemical Space. J. Chem. Inf. Model. 2023, 63, 3629–3636. [Google Scholar] [CrossRef] [PubMed]
- Bian, Y.; Xie, X.Q. Generative Chemistry: Drug Discovery with Deep Learning Generative Models. J. Mol. Model. 2021, 27, 71. [Google Scholar] [CrossRef]
- Cai, C.; Lin, M.; Li, W.; Chen, G.; Huang, D. A Unified Multi-Scale Deep Learning Framework for Molecular Property Prediction That Bridges Molecular Structures and Fingerprinting. Commun. Chem. 2026. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Mendez, D.; Gaulton, A.; Bento, A.P.; Chambers, J.; De Veij, M.; Félix, E.; Magariños, M.P.; Mosquera, J.F.; Mutowo, P.; Nowotka, M.; Gordillo-Marañón, M.; Hunter, F.; Junco, L.; Mugumbate, G.; Rodriguez-Lopez, M.; Atkinson, F.; Bosc, N.; Radoux, C.J.; Segura-Cabrera, A.; Hersey, A.; Leach, A.R. ChEMBL: Towards Direct Deposition of Bioassay Data. Nucleic Acids Res. 2019, 47, D930–D940. [Google Scholar] [CrossRef]
- Landrum, G. RDKit: Open-Source Cheminformatics Software. 2016. Available online: https://github.com/rdkit/rdkit.
- Rogers, D.; Hahn, M. Extended-Connectivity Fingerprints. J. Chem. Inf. Model. 2010, 50, 742–754. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Bemis, G.W.; Murcko, M.A. The Properties of Known Drugs. 1. Molecular Frameworks. J. Med. Chem. 1996, 39, 2887–2893. [Google Scholar] [CrossRef]
- Nilewar, S.S.; Chobe, S.; Dudhe, P.; Kumar, P.K.; Lodha, S.; Raut, A.D.; Fernández-Conde, D.; Farhan, M.; Muteeb, G.; Pawar, T.J. An Integrated QSAR-MD-DCCM Pipeline: A Predictive Computational Platform for the Rational Design and Dynamic Functional Validation of Dual-Target Directed Ligands. Pharmaceuticals 2026, 19, 249. [Google Scholar] [CrossRef]
- Özçelik, R.; Brinkmann, H.; Criscuolo, E.; Grisoni, F. Generative Deep Learning for de Novo Drug Design—A Chemical Space Odyssey. J. Chem. Inf. Model. 2025, 65, 7352–7372. [Google Scholar] [CrossRef] [PubMed]
- Sander, T.; Freyss, J.; von Korff, M.; Rufener, C. DataWarrior: An Open-Source Program for Chemistry-Aware Data Visualization and Analysis. J. Chem. Inf. Model. 2015, 55, 460–473. [Google Scholar] [CrossRef] [PubMed]
- Baell, J.B.; Holloway, G.A. New Substructure Filters for Removal of Pan Assay Interference Compounds (PAINS) from Screening Libraries and for Their Exclusion in Bioassays. J. Med. Chem. 2010, 53, 2719–2740. [Google Scholar] [CrossRef] [PubMed]
- Kawakita, Y.; Seto, M.; Ohashi, T.; Tamura, T.; Yusa, T.; Miki, H.; Iwata, H.; Kamiguchi, H.; Tanaka, T.; Sogabe, S.; Ohta, Y.; Ishikawa, T. Design and Synthesis of Novel Pyrimido [4,5-b]azepine Derivatives as HER2/EGFR Dual Inhibitors. Bioorg. Med. Chem. 2013, 21, 2250–2261. [Google Scholar] [CrossRef]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated Docking with Selective Receptor Flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed]
- Guex, N.; Peitsch, M.C. SWISS-MODEL and the Swiss-Pdb Viewer: An Environment for Comparative Protein Modeling. Electrophoresis 1997, 18, 2714–2723. [Google Scholar] [CrossRef]
- Koes, D.R.; Baumgartner, M.P.; Camacho, C.J. Lessons Learned in Empirical Scoring with Smina from the CSAR 2011 Benchmarking Exercise. J. Chem. Inf. Model. 2013, 53, 1893–1904. [Google Scholar] [CrossRef] [PubMed]
- Nilewar, S.S.; Chavan, A.D.; Pradhan, A.R.; Tripathy, A.A.; Bandaru, N.; Dudhe, P.B.; Kumar, P.K.; Lodha, S.; Muteeb, G.; Peredo-Valderrama, I.; et al. Dual-Site Acetylcholinesterase Inhibition and Multiscale Stability of Fused Quinoline Sulfonamides: A Chemoinformatic GA-MLR and Molecular Dynamics Study. Int. J. Mol. Sci. 2026, 27, 3286. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Wu, C.; Ghoreishi, D.; Chen, W.; Wang, L.; Damm, W.; Ross, G.A.; Dahlgren, M.K.; Russell, E.; von Bargen, C.D.; Abel, R.; Friesner, R.A.; Harder, E.D. OPLS4: Improving Force Field Accuracy on Challenging Regimes of Chemical Space. J. Chem. Theory Comput. 2021, 17, 4291–4300. [Google Scholar] [CrossRef]
- Nilewar, S.S.; Chobe, S.S.; Gurav, A.D.; Kureshi, S.B.; Palande, S.B.; Escobar-Cabrera, J.; Hernández-Rosas, F.; Pawar, T.J. Galloylation-Driven Anchoring of the Asp325-Asp336 Ridge: The Molecular Logic Behind the Superior Kinetic Stabilization of HMPV Fusion Protein by Green Tea Dimeric Catechins. Molecules 2026, 31, 821. [Google Scholar] [CrossRef]
| Stage | Process | Count | Rationale |
|---|---|---|---|
| I | Generative Sampling (DrugEx) | 100,000 | Enrichment for predicted EGFR activity. |
| II | Hierarchical ADMET/Liability Filtering | 1,601 | Optimization of solubility, polarity, and removal of toxicophores. |
| III | Structural Novelty Filtering | 136 | Exclusion of established scaffolds (Tanimoto <0.4). |
| Compound ID | WT Affinity (kcal/mol) |
T790M Affinity (kcal/mol) |
ΔAffinity (kcal/mol) |
Primary Mutant Anchor |
|---|---|---|---|---|
| 14 | −12.2 | −9.6 | 2.6 | Lys745 (H-bond) |
| 88 | −12.3 | −9.6 | 2.7 | Lys745 (H-bond) |
| 106 | −12.3 | −11.0 | 1.3 | Met790 (π-Sulfur) |
| W32 (Ref) | −12.1 | −10.8 | 1.3 | Met790 (Hydrophobic) |
| Compound | FEL Analysis | Ramachandran Analysis | ||||||
|---|---|---|---|---|---|---|---|---|
| RMSD (Å) | Rg (Å) |
ΔG (kJ/mol) | Region (%) | Disallowed Residues | ||||
| Favored | Allowed | Disallowed | ||||||
| 14 (Wild) | 2.922 | 19.937 | 8.751 | 91.91 | 6.8 | 1.29 | His835, Arg836, Lys879, Ala972 | |
| 14 (Mutant) | 1.467 | 19.785 | 7.953 | 91.43 | 7.62 | 0.95 | Glu734, Glu804, Val1011 | |
| 88 (Wild) | 2.335 | 20.059 | 13.343 | 90.88 | 8.14 | 0.98 | Asp855, Arg889, Thr993 | |
| 88 (Mutant) | 2.474 | 19.794 | 15.757 | 91.26 | 6.8 | 1.94 | Thr751, Lys806, Arg836, Asp855, Val897, Gln935 | |
| 106 (Wild) | 2.574 | 20.036 | 14.297 | 90.43 | 8.58 | 0.99 | Arg832, Pro848, Val980 | |
| 106 (Mutant) | 1.795 | 20.2 | 4.417 | 91 | 7.4 | 1.61 | Val769, Ser784, Val834, Leu883, His888 | |
| Compound | ΔGbind | ΔEvdw | ΔEelec | ΔGsolv |
|---|---|---|---|---|
| 14 (Wild) | -45.59 | -60.37 | -21.01 | 35.79 |
| 14 (Mutant) | -39.64 | -55.26 | -1.35 | 16.97 |
| 88 (Wild) | -52.08 | -66.63 | -11.38 | 25.92 |
| 88 (Mutant) | -44.19 | -65.31 | -9.37 | 30.49 |
| 106 (Wild) | -49.99 | -63.76 | -16.43 | 30.21 |
| 106 (Mutant) | -50.51 | -66.19 | -8.53 | 24.21 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




