Submitted:
28 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Projects
2.2. Research Papers
2.3. Data Processing and Statistical Analysis
3. Results and Discussions
3.1. Evolution of Soil-Related Projects
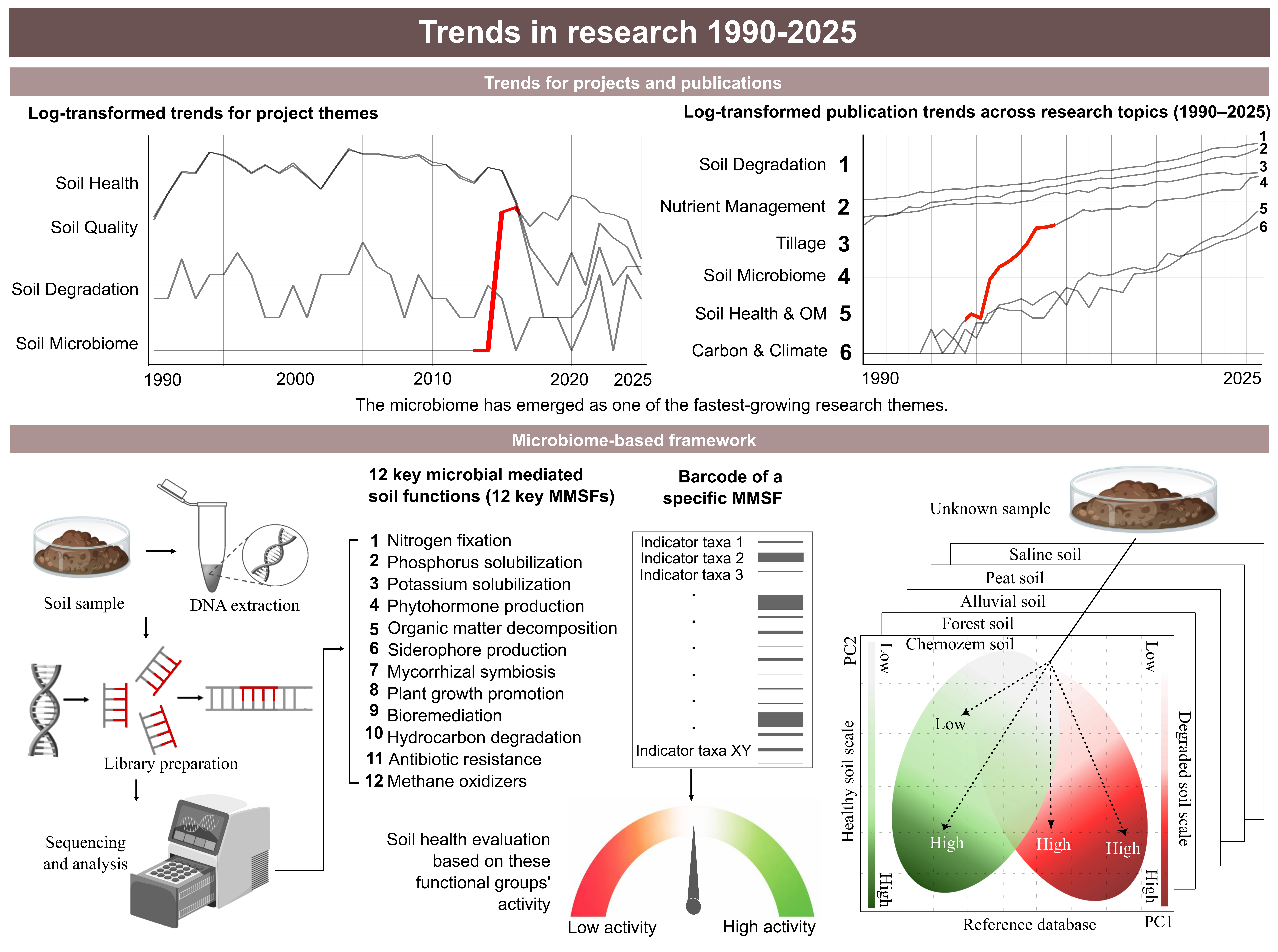
3.2. Three-Decade Trends in Soil Research
3.3. Dominant Research Themes Revealed by Keyword Analysis
3.4. Geographical Distribution and Growth Trends in Soil Research
3.5. Methodological Approaches in Soil Microbiome Research
3.6. Microbial Indicators Based on Key Microbially Mediated Soil Functions (MMSFs)
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Telo da Gama, J. The Role of Soils in Sustainability, Climate Change, and Ecosystem Services: Challenges and Opportunities. Ecologies 2023, 4, 552–567. [Google Scholar] [CrossRef]
- Anthony, M.A.; Bender, S.F.; van der Heijden, M.G.A. Enumerating Soil Biodiversity. Proc. Natl. Acad. Sci. 2023, 120, e2304663120. [Google Scholar] [CrossRef] [PubMed]
- FAO. FAO Highlights the Vital Role of Soil: 95 Percent of Our Food Is Directly Produced on Soils. Available online: https://www.fao.org/countryprofiles/news-archive/detail-news/en/c/1729967/ (accessed on 23 February 2026).
- FAO. FAO Warns 90 per Cent of Earth’s Topsoil at Risk by 2050 | UN News. Available online: https://news.un.org/en/story/2022/07/1123462 (accessed on 22 February 2026).
- Robinson, J.M.; Liddicoat, C.; Muñoz-Rojas, M.; Breed, M.F. Restoring Soil Biodiversity. Curr. Biol. 2024, 34, R393–R398. [Google Scholar] [CrossRef]
- Penuelas, J.; Coello, F.; Sardans, J. A Better Use of Fertilizers Is Needed for Global Food Security and Environmental Sustainability. Agric. Food Secur. 2023, 12, 5. [Google Scholar] [CrossRef]
- Jin, H.; Huang, S.; Shi, D.; Li, J.; Li, J.; Li, Y.; Zhu, H. Effects of Different Tillage Practices on Soil Stability and Erodibility for Red Soil Sloping Farmland in Southern China. Agronomy 2023, 13, 1310. [Google Scholar] [CrossRef]
- Sünnemann, M.; Beugnon, R.; Breitkreuz, C.; Buscot, F.; Cesarz, S.; Jones, A.; Lehmann, A.; Lochner, A.; Orgiazzi, A.; Reitz, T.; et al. Climate Change and Cropland Management Compromise Soil Integrity and Multifunctionality. Commun. Earth Env. 2023, 4, 394. [Google Scholar] [CrossRef]
- Kabato, W.; Hailegnaw, N.; Mutum, L.; Molnar, Z. Managing Soil Health for Climate Resilience and Crop Productivity in a Changing Environment. Sci. Total Environ. 2025, 1000, 180460. [Google Scholar] [CrossRef] [PubMed]
- UN, U. World Population Projected to Reach 9.8 Billion in 2050, and 11.2 Billion in 2100. Available online: https://www.un.org/en/desa/world-population-projected-reach-98-billion-2050-and-112-billion-2100 (accessed on 23 February 2026).
- UN World Must Sustainably Produce 70 per Cent More Food by Mid-Century – UN Report | UN News. Available online: https://news.un.org/en/story/2013/12/456912 (accessed on 23 February 2026).
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 Statement: An Updated Guideline for Reporting Systematic Reviews. Syst. Rev. 2021, 10, 89. [Google Scholar] [CrossRef]
- World Bank Projects. Available online: https://projects.worldbank.org/en/projects-operations/projects-list (accessed on 16 February 2026).
- CORDIS Projects & Results | CORDIS | European Commission. Available online: https://cordis.europa.eu/projects (accessed on 16 February 2026).
- Elsevier Scopus. Available online: https://www.scopus.com/pages/home#basic (accessed on 16 February 2026).
- Clarivate Smart Search - Web of Science Core Collection. Available online: https://www.webofscience.com/wos/woscc/smart-search (accessed on 16 February 2026).
- National Library of Medicine PubMed. Available online: https://pubmed.ncbi.nlm.nih.gov/ (accessed on 16 February 2026).
- FAO Voluntary Guidelines for Sustainable Soil Management; FAO: Rome, Italy, 2017.
- FAO Status of the World’s Soil Resources: Main Report, 1st ed.; FAO: Rome, Italy, 2015; ISBN 978-92-5-109004-6.
- Weil, R.; Brady, N. The Nature and Properties of Soils; Pearson Education, Inc.: Columbus, 2017; Vol. 15, ISBN 978-0-13-325448-8. [Google Scholar]
- R Core Team The R Project for Statistical Computing 2024.
- Wickham, H.; François, R.; Henry, L.; Müller, K.; Vaughan, D.; Software, P. PBC Dplyr: A Grammar of Data Manipulation 2026.
- Wickham, H.; Vaughan, D.; Girlich, M.; Ushey, K.; Software, P. PBC Tidyr: Tidy Messy Data 2025.
- UNCED Summary Report 3–14 June 1992. Available online: https://enb.iisd.org/events/unced/summary-report-3-14-june-1992 (accessed on 16 February 2026).
- UNCCD United Nations Convention to Combat Dsertification in Those Countries Experiencing Serious Drought and/or Deserrtification Particularly in Africa; The United Nations Convention to Combat Desertification: Bonn, Germany, 1994.
- IPCC Climate Change 2022 Mitigation of Climate Change; Intergovernmental Panel on Climate Change: Geneva, Switzerland, 2022.
- UNCCD Sustainable Land Management Contribution to Successful Land-Based Climate Change Adaptation and Mitigation. Available online: https://www.unccd.int/resources/reports/sustainable-land-management-contribution-successful-land-based-climate-change (accessed on 16 February 2026).
- UNFCCC United Nations Framework Convention on Climate Change (UNFCCC) | Environmental Sciences | Research Starters | EBSCO Research. Available online: https://www.ebsco.com (accessed on 21 February 2026).
- UN THE 17 GOALS | Sustainable Development. Available online: https://sdgs.un.org/goals (accessed on 21 February 2026).
- Iese, V.; Wairiu, M.; Hickey, G.M.; Ugalde, D.; Hinge Salili, D.; Walenenea, J.; Tabe, T.; Keremama, M.; Teva, C.; Navunicagi, O.; et al. Impacts of COVID-19 on Agriculture and Food Systems in Pacific Island Countries (PICs): Evidence from Communities in Fiji and Solomon Islands. Agric. Syst. 2021, 190, 103099. [Google Scholar] [CrossRef]
- Koloma, Y.; Kemeze, F.H. COVID-19 and Perceived Effects on Agricultural Financing in Africa: Evidence and Policy Implications. Afr. Dev. Rev. 2022, 34, S63–S79. [Google Scholar] [CrossRef]
- Das, S.; Dutta, S.; Roy Choudhury, M.; Garai, S.; Mukherjee, S.; Sengupta, S.; Jana, S.; Dey, S.; Dhar, A.; Dutta, S.; et al. Regenerating Rural Soil and Ecosystems: A 15-Year Systematic Review of Emerging Methods and Technologies. Sci. Total Environ. 2025, 990, 179926. [Google Scholar] [CrossRef]
- Bhunia, G.S.; Sengupta, D. Cultivating Resilience: Bibliometric Analysis of Soil Health for a Sustainable Future. Discov. Soil. 2025, 2, 95. [Google Scholar] [CrossRef]
- Omer, E.; Szlatenyi, D.; Csenki, S.; Tünde, G.; Chhetri, G.; Veres, Z.; Láng, V. Soil Health for Sustainable Agriculture: A Bibliometric Review of EU Current Scientific Findings and Research Trends. Soil. Adv. 2026, 5, 100097. [Google Scholar] [CrossRef]
- FAO. UNEP Spatial Distribution of Soil Pollution in Asia and the Pacific. In Global assessment of soil pollution: Report; FAO and UNEP: Rome, Italy, 2021; p. 846. ISBN 978-92-5-134469-9. [Google Scholar]
- Alexandra, J.; Carter, G.; Delaney, M.; Hadobas, S.; Howden, M.; Maharjan, K.; Mahani, M.E.; Mudaliar, L.; Prinsley, R.; Rimon, A.; et al. Inquiry into Australia’s Response to the Priorities of Pacific Island Countries and the Pacific Region 2024.
- SADC; IUCN. UNCCD Drought Risk Management and Mitigation Strategy (DRIMMS) 2022-2032; Botswana, 2022. [Google Scholar]
- GEF REPORT OF THE GLOBAL ENVIRONMENT FACILITY TO THE FIFTEENTH SESSION OF THE CONFERENCE OF THE PARTIES TO THE UNITED NATIONS CONVENTION TO COMBAT DESERTIFICATION 2022.
- UNCCD Great Green Wall Initiative. Available online: https://www.unccd.int/our-work/ggwi (accessed on 21 February 2026).
- Yimer, A.H.; Tarnawa, A. Advancing Nutrient Management Strategies for Sustainable Crop Productivity in a Changing Climate: A Systematic Review. ScientificWorldJournal 2025, 2025, 7101060. [Google Scholar] [CrossRef] [PubMed]
- Ge, G.; Chen, X.; Ma, H.; Zhang, X.; Shi, J.; Wang, X.; Zhao, X.; Wang, M.; Xian, F.; Lu, Z.; et al. Bibliometric Analysis of Research Trends in Agricultural Soil Organic Carbon Components from 2000 to 2023. Front. Plant Sci. 2024, 15. [Google Scholar] [CrossRef]
- Reznikova, D.A.; Barannikova, M.V.; Shnakhova, L.M.; Mitkin, N.A.; Vatlin, A.A. Next-Generation Sequencing Approaches for Soil Microbiome Research. Front. Soil. Sci. 2026, 5. [Google Scholar] [CrossRef]
- Romero, F.; Labouyrie, M.; Orgiazzi, A.; Ballabio, C.; Panagos, P.; Jones, A.; Tedersoo, L.; Bahram, M.; Eisenhauer, N.; Sünnemann, M.; et al. The Soil Microbiome as an Indicator of Ecosystem Multifunctionality in European Soils. Nat. Commun. 2025, 17, 705. [Google Scholar] [CrossRef] [PubMed]
- Mc, G.; Kn, S.; Yadahalli, V.R. Emerging Techniques in Soil Microbiome Analysis for Sustainable Agriculture. Int. J. Adv. Biochem. Res. 2025, 9, 350–355. [Google Scholar] [CrossRef]
- Jesus, E. da C.; Coelho, M.R.R.; Chaer, G.M.; Mendes, I. de C.; de Oliveira, M.I.L.; Malaquias, J.V.; da Silva, O.D.D.; Oliveira, I. von der W.A.; Eugenio, N.R.; Brown, G.G. A Bibliometric Analysis on Soil Microbial Diversity and Processes: Global Trends and Methodologies. Soil. Org. 2025, 97, 65–77. [Google Scholar] [CrossRef]
- Zhao, S.; Zhang, J. Microplastics in Soils during the COVID-19 Pandemic: Sources, Migration and Transformations, and Remediation Technologies. Sci. Total Env. 2023, 883, 163700. [Google Scholar] [CrossRef]
- Lal, R. Soil Health and Carbon Management. Food Energy Secur. 2016, 5, 212–222. [Google Scholar] [CrossRef]
- Delgado-Baquerizo, M.; Eldridge, D.J.; Liu, Y.-R.; Liu, Z.-W.; Coleine, C.; Trivedi, P. Soil Biodiversity and Function under Global Change. PLoS Biol. 2025, 23, e3003093. [Google Scholar] [CrossRef]
- Rinot, O.; Levy, G.J.; Steinberger, Y.; Svoray, T.; Eshel, G. Soil Health Assessment: A Critical Review of Current Methodologies and a Proposed New Approach. Sci. Total Environ. 2019, 648, 1484–1491. [Google Scholar] [CrossRef] [PubMed]
- Cherubin, M.R.; Pinheiro Junior, C.R.; Souza, L.F.N.; Canisares, L.P.; Ferreira, T.O.; Cerri, C.E.P.; Minasny, B.; Smith, P. Global Blind Spots in Soil Health Research Overlap with Environmental Vulnerability Hotspots. Commun. Earth Env. 2025, 6, 651. [Google Scholar] [CrossRef]
- Sellami, M.H.; Mori, M.; Terribile, F. Mapping the Conceptual and Intellectual Structure of Soil Health Research (1996–2021): A Terms Co-Occurrence and Co-Cited Reference Network Analysis. Front. Soil. Sci. 2025, 5. [Google Scholar] [CrossRef]
- Delgado-Baquerizo, M.; Hu, H.-W.; Maestre, F.T.; Guerra, C.A.; Eisenhauer, N.; Eldridge, D.J.; Zhu, Y.-G.; Chen, Q.-L.; Trivedi, P.; Du, S.; et al. The Global Distribution and Environmental Drivers of the Soil Antibiotic Resistome. Microbiome 2022, 10, 219. [Google Scholar] [CrossRef]
- Li, Y.; Duan, Y.; Zhang, J.; Petropoulos, E.; Zhao, J.; Wu, F.; Wang, L.; Chen, Y.; Wang, X. Structure and Function of the Topsoil Microbiome in Chinese Terrestrial Ecosystems. Front Microbiol. 2025, 16, 1595810. [Google Scholar] [CrossRef]
- Böttcher, H.; Zell-Ziegler, C.; Herold, A.; Siemons, A. EU LULUCF Regulation Explained Summary of Core Provisions and Expected Effects; Berlin, Germany, 2019. [Google Scholar]
- EU Regulation (EU) 2024/3012 of the European Parliament and of the Council of 27 November 2024 Establishing a Union Certification Framework for Permanent Carbon Removals, Carbon Farming and Carbon Storage in Products. 2024.
- Huang, B.; Gao, J.; Zhang, W.; Gan, C.; Bao, X.; Xu, M.; Wu, L. Impacts of Conservation Tillage on Soil Organic Carbon and Crop Yield in Black Soil Region of Northeast China: Integrated Regulation by Climate, Management and Soil Properties. J. Agric. Food Res. 2025, 24, 102388. [Google Scholar] [CrossRef]
- Leite, M.F.A.; van den Broek, S.W.E.B.; Kuramae, E.E. Current Challenges and Pitfalls in Soil Metagenomics. Microorganisms 2022, 10, 1900. [Google Scholar] [CrossRef]
- Ferrocino, I.; Ferrara, M.; Garello, M.; Turchetti, B.; Andreolli, M.; Arena, M.P.; Alduina, R.; Aloi, F.; Bevivino, A.; Bruno, E.; et al. Standardizing Microbiome Research: Interlaboratory Validation of SOPs for Sample Preparation and DNA Extraction from Food and Environmental Ecosystems. Env. Microbiome 2025, 21, 14. [Google Scholar] [CrossRef] [PubMed]
- Sergaki, C.; Lagunas, B.; Lidbury, I.; Gifford, M.L.; Schäfer, P. Challenges and Approaches in Microbiome Research: From Fundamental to Applied. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Hill, G.T.; Mitkowski, N.A.; Aldrich-Wolfe, L.; Emele, L.R.; Jurkonie, D.D.; Ficke, A.; Maldonado-Ramirez, S.; Lynch, S.T.; Nelson, E.B. Methods for Assessing the Composition and Diversity of Soil Microbial Communities. Appl. Soil. Ecol. 2000, 15, 25–36. [Google Scholar] [CrossRef]
- Martiny, A.C. High Proportions of Bacteria Are Culturable across Major Biomes. ISME J. 2019, 13, 2125–2128. [Google Scholar] [CrossRef] [PubMed]
- FAO Standard Operating Procedure for Soil Microbial Biomass (Carbon): Chloroform Fumigation-Extraction Method; FAO: Rome, Italy, 2024.
- Joergensen, R.G. Microbial Biomass Estimation by Fumigation Extraction and ATP – A Review in Memoriam of Phil Brookes. Biol. Fertil. Soils 2025, 61, 1287–1299. [Google Scholar] [CrossRef]
- Quideau, S.A.; McIntosh, A.C.S.; Norris, C.E.; Lloret, E.; Swallow, M.J.B.; Hannam, K. Extraction and Analysis of Microbial Phospholipid Fatty Acids in Soils. J. Vis. Exp. 2016, 54360. [Google Scholar] [CrossRef] [PubMed]
- van Galen, L.G.; Smith, G.R.; Margenot, A.J.; Waldrop, M.P.; Crowther, T.W.; Peay, K.G.; Jackson, R.B.; Yu, K.; Abrahão, A.; Ahmed, T.A.; et al. A Global Database of Soil Microbial Phospholipid Fatty Acids and Enzyme Activities. Sci. Data 2025, 12, 1568. [Google Scholar] [CrossRef]
- Matchado, M.S.; Rühlemann, M.; Reitmeier, S.; Kacprowski, T.; Frost, F.; Haller, D.; Baumbach, J.; List, M. On the Limits of 16S rRNA Gene-Based Metagenome Prediction and Functional Profiling. Microb. Genom. 2024, 10, 001203. [Google Scholar] [CrossRef]
- Microbiome Insights 16S rRNA Gene Sequencing vs. Shotgun Metagenomic Sequencing. Available online: https://blog.microbiomeinsights.com/16s-rrna-sequencing-vs-shotgun-metagenomic-sequencing (accessed on 31 March 2026).
- Lugli, G.A.; Ventura, M. A Breath of Fresh Air in Microbiome Science: Shallow Shotgun Metagenomics for a Reliable Disentangling of Microbial Ecosystems. Microbiome Res. Rep. 2022, 1, 8. [Google Scholar] [CrossRef]
- Terrón-Camero, L.C.; Gordillo-González, F.; Salas-Espejo, E.; Andrés-León, E. Comparison of Metagenomics and Metatranscriptomics Tools: A Guide to Making the Right Choice. Genes 2022, 13, 2280. [Google Scholar] [CrossRef]
- Josephs-Spaulding, J.; Rettig, H.C.; Zimmermann, J.; Chkonia, M.; Mischnik, A.; Franzenburg, S.; Graspeuntner, S.; Rupp, J.; Kaleta, C. Metatranscriptomics-Based Metabolic Modeling of Patient-Specific Urinary Microbiome during Infection. Npj Biofilms Microbiomes 2025, 11, 183. [Google Scholar] [CrossRef]
- Ascandari, A.; Aminu, S.; Safdi, N.E.H.; El Allali, A.; Daoud, R. A Bibliometric Analysis of the Global Impact of Metaproteomics Research. Front. Microbiol. 2023, 14. [Google Scholar] [CrossRef] [PubMed]
- Tartaglia, M.; Bastida, F.; Sciarrillo, R.; Guarino, C. Soil Metaproteomics for the Study of the Relationships Between Microorganisms and Plants: A Review of Extraction Protocols and Ecological Insights. Int. J. Mol. Sci. 2020, 21, 8455. [Google Scholar] [CrossRef]
- Johnson, C.H.; Ivanisevic, J.; Siuzdak, G. Metabolomics: Beyond Biomarkers and towards Mechanisms. Nat. Rev. Mol. Cell. Biol. 2016, 17, 451–459. [Google Scholar] [CrossRef]
- Song, Y.; Yao, S.; Li, X.; Wang, T.; Jiang, X.; Bolan, N.; Warren, C.R.; Northen, T.R.; Chang, S.X. Soil Metabolomics: Deciphering Underground Metabolic Webs in Terrestrial Ecosystems. Eco-Environ. Health 2024, 3, 227–237. [Google Scholar] [CrossRef] [PubMed]
- Divyasorubini, S.; Kandage, S.M.; Liyanage, S.; Rajapakse, C.; Silva, G.N. Microbial Biosensors for Real-Time Monitoring of the Bioremediation Processes. In Bioremediation of Environmental Pollutants: Emerging Trends and Strategies; Suyal, D.C., Soni, R., Eds.; Springer International Publishing: Cham, 2022; pp. 111–144. ISBN 978-3-030-86169-8. [Google Scholar]
- Huang, C.-W.; Lin, C.; Nguyen, M.K.; Hussain, A.; Bui, X.-T.; Ngo, H.H. A Review of Biosensor for Environmental Monitoring: Principle, Application, and Corresponding Achievement of Sustainable Development Goals. Bioengineered 2023, 14, 58–80. [Google Scholar] [CrossRef]
- Araz, M.K.; Tentori, A.M.; Herr, A.E. Microfluidic Multiplexing in Bioanalyses. SLAS Technol. 2013, 18, 350–366. [Google Scholar] [CrossRef]
- Hariharasudha, H.K.R.S.; Chandrasekaran, M.; Baskaran, C.M.; Govindarajan, D.K.; Kandaswamy, K. Lab-on-a-Chip Devices for Bacterial Systems: Tracking Biofilm Growth and Antimicrobial Interactions. Total Environ. Microbiol. 2025, 1, 100042. [Google Scholar] [CrossRef]
- Fierer, N.; Wood, S.A.; Bueno de Mesquita, C.P. How Microbes Can, and Cannot, Be Used to Assess Soil Health. Soil. Biol. Biochem. 2021, 153, 108111. [Google Scholar] [CrossRef]
- Nicholls, C.I.; Altieri, M.A.; Dezanet, A.; Lana, M.; Feistauer, D.; Ouriques, M. A Rapid, Farmer-Friendly Agroecological Method to Estimate Soil Quality and Crop Health in Vineyard Systems. 2004. [Google Scholar]
- Wu, S.; Jie, H.; Jie, Y. Role of Rhizosphere Soil Microbes in Adapting Ramie (Boehmeria Nivea L.) Plants to Poor Soil Conditions through N-Fixing and P-Solubilization. Agronomy 2021, 11, 2096. [Google Scholar] [CrossRef]
- Lomtatidze, Z.; Gioshvili, I. Distribution Characteristics of Free-Living Nitrogen-Fixing Bacteria in Solonetz Soil Types. Asian J. Agric. Hortic. Res. 2025, 12, 149–155. [Google Scholar] [CrossRef]
- Silva, D.F.; Cardoso, E.J.B.N.; Huang, L.; Erikson, C.; Silva, A.M.M.; Araujo, V.L.V.P.; Silva, D.E.O.; Melo, V.M.M.; Araujo, A.S.F.; Pereira, A.P.A.; et al. Functional Genes Related to N and P Cycling in Degraded and Restored Areas from Brazilian Drylands. Appl. Soil. Ecol. 2024, 196, 105316. [Google Scholar] [CrossRef]
- Bouffaud, M.-L.; Renoud, S.; Moënne-Loccoz, Y.; Muller, D. Is Plant Evolutionary History Impacting Recruitment of Diazotrophs and nifH Expression in the Rhizosphere? Sci. Rep. 2016, 6, 21690. [Google Scholar] [CrossRef]
- Jia, R.; Wang, K.; Li, L.; Qu, Z.; Shen, W.; Qu, D. Abundance and Community Succession of Nitrogen-Fixing Bacteria in Ferrihydrite Enriched Cultures of Paddy Soils Is Closely Related to Fe(III)-Reduction. Sci. Total Environ. 2020, 720, 137633. [Google Scholar] [CrossRef]
- Rodrigues, E.P.; Rodrigues, L. da S.; de Oliveira, A.L.M.; Divan Baldani, V.L.; Teixeira, K.R. dos S.; Urquiaga, S.; Reis, V.M. Azospirillum Amazonense Inoculation: Effects on Growth, Yield and N2 Fixation of Rice (Oryza Sativa L.). Plant Soil. 2009, 316, 323–323. [Google Scholar] [CrossRef]
- Okon, Y.; Heytler, P.G.; Hardy, R.W.F. N2 Fixation by Azospirillum Brasilense and Its Incorporation into Host Setaria Italica. Appl. Environ. Microbiol. 1983, 46, 694–697. [Google Scholar] [CrossRef] [PubMed]
- Reinhold, B.; Hurek, T.; Fendrik, I.; Pot, B.; Gillis, M.; Kersters, K.; Thielemans, S.; De Ley, J. Azospirillum Halopraeferens Sp. Nov., a Nitrogen-Fixing Organism Associated with Roots of Kallar Grass (Leptochloa Fusca (L.) Kunth). Int. J. Syst. Evol. Microbiol. 1987, 37, 43–51. [Google Scholar] [CrossRef]
- Karpati, E.; Kiss, P.; Ponyi, T.; Fendrik, I.; de Zamaroczy, M.; Orosz, L. Interaction of Azospirillum Lipoferum with Wheat Germ Agglutinin Stimulates Nitrogen Fixation. J. Bacteriol. 1999, 181, 3949–3955. [Google Scholar] [CrossRef]
- Kargar, M.; Kargar, M.; Noerooznejad, M.; Ayazpour, K. The Role of Azotobacter Chrococcum and Azotobacter Paspali on Wheat Growth in Fars Province. Adv. Environ. Biol. 2014, 8, 837–841. [Google Scholar]
- Hamilton, T.L.; Ludwig, M.; Dixon, R.; Boyd, E.S.; Dos Santos, P.C.; Setubal, J.C.; Bryant, D.A.; Dean, D.R.; Peters, J.W. Transcriptional Profiling of Nitrogen Fixation in Azotobacter Vinelandii▿. J. Bacteriol. 2011, 193, 4477–4486. [Google Scholar] [CrossRef] [PubMed]
- Salas, A.; Tortosa, G.; Hidalgo-García, A.; Delgado, A.; Bedmar, E.J.; Richardson, D.J.; Gates, A.J.; Delgado, M.J. The Hemoglobin Bjgb From Bradyrhizobium Diazoefficiens Controls NO Homeostasis in Soybean Nodules to Protect Symbiotic Nitrogen Fixation. Front Microbiol. 2020, 10, 2915. [Google Scholar] [CrossRef]
- Bitire, T.D.; Abberton, M.; Oyatomi, O.; Babalola, O.O. Effect of Bradyrhizobium Japonicum Strains and Inorganic Nitrogen Fertilizer on the Growth and Yield of Bambara Groundnut (Vigna Subterranea (L.) Verdc) Accessions. Front. Sustain. Food Syst. 2022, 6. [Google Scholar] [CrossRef]
- Li, Z.; Henawy, A.R.; Halema, A.A.; Fan, Q.; Duanmu, D.; Huang, R. A Wild Rice Rhizobacterium Burkholderia Cepacia BRDJ Enhances Nitrogen Use Efficiency in Rice. Int. J. Mol. Sci. 2022, 23, 10769. [Google Scholar] [CrossRef] [PubMed]
- Potrikus, C.J.; Breznak, J.A. Nitrogen-Fixing Enterobacter Agglomerans Isolated from Guts of Wood-Eating Termites. Appl. Environ. Microbiol. 1977, 33, 392–399. [Google Scholar] [CrossRef]
- Ji, C.; Liu, Z.; Hao, L.; Song, X.; Wang, C.; Liu, Y.; Li, H.; Li, C.; Gao, Q.; Liu, X. Effects of Enterobacter Cloacae HG-1 on the Nitrogen-Fixing Community Structure of Wheat Rhizosphere Soil and on Salt Tolerance. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef]
- Alloisio, N.; Kucho, K.; Pujic, P.; Normand, P. The Frankia Alni Symbiotic Transcriptome. In Biological Nitrogen Fixation; John Wiley & Sons, Ltd, 2015; pp. 757–768. ISBN 978-1-119-05309-5. [Google Scholar]
- Kucho, K.; Taniyama, K. Characterization of Frankia Casuarinae Mutants Defective in Vesicle Envelope Development. Microbes Environ. 2025, 40. [Google Scholar] [CrossRef]
- Tian, G.; Pauls, K.; Dong, Z.; Reid, L.; Tian, L. Colonization of the Nitrogen-Fixing Bacterium Gluconacetobacter Diazotrophicus in a Large Number of Canadian Corn Plants. Can. J. Plant Sci.-CAN. J. PLANT SCI. 2009, 89, 1009–1016. [Google Scholar] [CrossRef]
- Pedrolo, A.M.; Matteoli, F.P.; Soares, C.R.F.S.; Arisi, A.C.M. Comparative Genomics Reveal the High Conservation and Scarce Distribution of Nitrogen Fixation Nif Genes in the Plant-Associated Genus Herbaspirillum. Microb. Ecol. 2023, 86, 563–574. [Google Scholar] [CrossRef]
- Klassen, G.; de Oliveira Pedrosa, F.; de Souza, E.M.; Yates, M.G.; Rigo, L.U. Nitrogenase Activity of Herbaspirillum Seropedicae Grown under Low Iron Levels Requires the Products of nifXorf1 Genes. FEMS Microbiol. Lett. 2003, 224, 255–259. [Google Scholar] [CrossRef] [PubMed]
- Temme, K.; Zhao, D.; Voigt, C.A. Refactoring the Nitrogen Fixation Gene Cluster from Klebsiella Oxytoca. Proc. Natl. Acad. Sci. U S A 2012, 109, 7085–7090. [Google Scholar] [CrossRef] [PubMed]
- Benjelloun, I.; Thami Alami, I.; El Khadir, M.; Douira, A.; Udupa, S.M. Co-Inoculation of Mesorhizobium Ciceri with Either Bacillus Sp. or Enterobacter Aerogenes on Chickpea Improves Growth and Productivity in Phosphate-Deficient Soils in Dry Areas of a Mediterranean Region. Plants 2021, 10, 571. [Google Scholar] [CrossRef]
- Shimoda, Y.; Hirakawa, H.; Sato, S.; Saeki, K.; Hayashi, M. Whole-Genome Sequence of the Nitrogen-Fixing Symbiotic Rhizobium Mesorhizobium Loti Strain TONO. Genome Announc. 2016, 4, 10.1128/genomea.01016-16. [Google Scholar] [CrossRef]
- Zhivotchenko, A.G.; Nikonova, E.S.; Jørgensen, M.H. Effect of Fermentation Conditions on N2 Fixation by Methylococcus Capsulatus. Bioprocess Eng. 1995, 14, 9–15. [Google Scholar] [CrossRef]
- Canto-Encalada, G.; Tec-Campos, D.; Tibocha-Bonilla, J.D.; Zengler, K.; Zepeda, A.; Zuñiga, C. Flux Balance Analysis of the Ammonia-Oxidizing Bacterium Nitrosomonas Europaea ATCC19718 Unravels Specific Metabolic Activities While Degrading Toxic Compounds. PLoS Comput. Biol. 2022, 18, e1009828. [Google Scholar] [CrossRef]
- Stępień, A.; Wojtkowiak, K.; Kolankowska, E. Effect of Commercial Microbial Preparations Containing Paenibacillus Azotofixans, Bacillus Megaterium and Bacillus Subtilis on the Yield and Photosynthesis of Winter Wheat and the Nitrogen and Phosphorus Content in the Soil. Appl. Sci. 2022, 12, 12541. [Google Scholar] [CrossRef]
- Daud, N.S.; Mohd Din, A.R.J.; Rosli, M.A.; Azam, Z.M.; Othman, N.Z.; Sarmidi, M.R. Paenibacillus Polymyxa Bioactive Compounds for Agricultural and Biotechnological Applications. Biocatal. Agric. Biotechnol. 2019, 18, 101092. [Google Scholar] [CrossRef]
- Soren, T.; Sarkar, A.; Mondal, T.; Ghosh, S.K.; Mondal, S.; Pal, P.; Maiti, T.K. The Rhizospheric Strain Paraburkholderia Kururiensis WP2T3 Efficiently Degrades Para-Nitrophenol and Exhibits Plant Growth-Promoting Characters. The Microbe 2025, 9, 100550. [Google Scholar] [CrossRef]
- Esmaeel, Q.; Miotto, L.; Rondeau, M.; Leclère, V.; Clément, C.; Jacquard, C.; Sanchez, L.; Barka, E.A. Paraburkholderia Phytofirmans PsJN-Plants Interaction: From Perception to the Induced Mechanisms. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef]
- Casas-Román, A.; Lorite, M.-J.; Werner, M.; Muñoz, S.; Gallegos, M.-T.; Sanjuán, J. The Gap Gene of Rhizobium Etli Is Required for Both Free Life and Symbiosis with Common Beans. Microbiol. Res. 2024, 284, 127737. [Google Scholar] [CrossRef]
- Janczarek, M.; Kozieł, M.; Adamczyk, P.; Buczek, K.; Kalita, M.; Gromada, A.; Mordzińska-Rak, A.; Polakowski, C.; Bieganowski, A. Symbiotic Efficiency of Rhizobium Leguminosarum Sv. Trifolii Strains Originating from the Subpolar and Temperate Climate Regions. Sci. Rep. 2024, 14, 6264. [Google Scholar] [CrossRef]
- Demtröder, L.; Pfänder, Y.; Masepohl, B. Rhodobacter Capsulatus AnfA Is Essential for Production of Fe-Nitrogenase Proteins but Dispensable for Cofactor Biosynthesis and Electron Supply. MicrobiologyOpen 2020, 9, e1033. [Google Scholar] [CrossRef]
- Arashida, H.; Kugenuma, T.; Watanabe, M.; Maeda, I. Nitrogen Fixation in Rhodopseudomonas Palustris Co-Cultured with Bacillus Subtilis in the Presence of Air. J. Biosci. Bioeng. 2019, 127, 589–593. [Google Scholar] [CrossRef] [PubMed]
- Temprano-Vera, F.; Rodríguez-Navarro, D.N.; Acosta-Jurado, S.; Perret, X.; Fossou, R.K.; Navarro-Gómez, P.; Zhen, T.; Yu, D.; An, Q.; Buendía-Clavería, A.M.; et al. Sinorhizobium Fredii Strains HH103 and NGR234 Form Nitrogen Fixing Nodules With Diverse Wild Soybeans (Glycine Soja) From Central China but Are Ineffective on Northern China Accessions. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef]
- Marcela, D.; Fourment, J.; Lucas, A.; Bergès, H.; Kahn, D. Nitrogen Regulation in Sinorhizobium Meliloti Probed with Whole Genome Arrays. FEMS Microbiol. Lett. 2004, 241, 33–40. [Google Scholar] [CrossRef]
- Peix, A.; Rivas, R.; Santa-Regina, I.; Mateos, P.F.; Martínez-Molina, E.; Rodríguez-Barrueco, C.; Velázquez, E. Pseudomonas Lutea Sp. Nov., a Novel Phosphate-Solubilizing Bacterium Isolated from the Rhizosphere of Grasses. Int. J. Syst. Evol. Microbiol. 2004, 54, 847–850. [Google Scholar] [CrossRef]
- Widawati, S. Diversity and Phosphate Solubilization by Bacteria Isolated from Laki Island Coastal Ecosystem. Biodiversitas J. Biol. Divers. 2011, 12. [Google Scholar] [CrossRef]
- Zhao, X.; Zhang, Y.; Cui, Z.; Peng, L.; Cao, C. Dynamics of phoD- and Gcd-Harboring Microbial Communities Across an Age Sequence of Biological Soil Crusts Under Sand-Fixation Plantation. Front. Microbiol. 2022, 13. [Google Scholar] [CrossRef] [PubMed]
- Wan, W.; Qin, Y.; Wu, H.; Zuo, W.; He, H.; Tan, J.; Wang, Y.; He, D. Isolation and Characterization of Phosphorus Solubilizing Bacteria With Multiple Phosphorus Sources Utilizing Capability and Their Potential for Lead Immobilization in Soil. Front. Microbiol. 2020, 11. [Google Scholar] [CrossRef]
- Azziz, G.; Bajsa, N.; Haghjou, T.; Taulé, C.; Valverde, Á.; Igual, J.M.; Arias, A. Abundance, Diversity and Prospecting of Culturable Phosphate Solubilizing Bacteria on Soils under Crop–Pasture Rotations in a No-Tillage Regime in Uruguay. Appl. Soil. Ecol. 2012, 61, 320–326. [Google Scholar] [CrossRef]
- Zhang, L.; Ge, A.-H.; Tóth, T.; Yang, F.; Wang, Z.; An, F. Enrichment of Keystone Fungal Taxa after Flue Gas Desulphurization Gypsum Application Drives Reclamation of the Saline-Sodic Soil. Land Degrad. Dev. 2023, 34, 2276–2287. [Google Scholar] [CrossRef]
- Zampieri, E.; Pesenti, M.; Nocito, F.F.; Sacchi, G.A.; Valè, G. Rice Responses to Water Limiting Conditions: Improving Stress Management by Exploiting Genetics and Physiological Processes. Agriculture 2023, 13, 464. [Google Scholar] [CrossRef]
- Dobbratz, M.; Jungers, J.M.; Gutknecht, J.L.M. Seasonal Plant Nitrogen Use and Soil N Pools in Intermediate Wheatgrass (Thinopyrum Intermedium). Agriculture 2023, 13, 468. [Google Scholar] [CrossRef]
- Rodriguez, H.; Gonzalez, T.; Goire, I.; Bashan, Y. Gluconic Acid Production and Phosphate Solubilization by the Plant Growth-Promoting Bacterium Azospirillum Spp. Naturwissenschaften 2004, 91, 552–555. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Kumar Behl, R.; Narula, N. Establishment of Phosphate-Solubilizing Strains of Azotobacter Chroococcum in the Rhizosphere and Their Effect on Wheat Cultivars under Green House Conditions. Microbiol. Res. 2001, 156, 87–93. [Google Scholar] [CrossRef]
- Ciopińska, J.; Bezak-Mazur, E. PHOSPHORUS SOLUBILIZING BACTERIA – REVIEW ARTICLE. Struct. Environ. 2018, 10, 278–287. [Google Scholar] [CrossRef]
- Velazquez, L.E.; Perez-Rios, R.; Munguia, N.E.; Zepeda, D.S.; Neudert, C.; Esquer, J. Environmental Approach for Energy Audits on Government Buildings: A Mexican Case for the Air Conditioning System. J. Environ. Prot. 2018, 9, 278–293. [Google Scholar] [CrossRef]
- Mohamed, E.A.H.; Farag, A.G.; Youssef, S.A. Phosphate Solubilization by Bacillus Subtilis and Serratia Marcescens Isolated from Tomato Plant Rhizosphere. J. Environ. Prot. 2018, 9, 266–277. [Google Scholar] [CrossRef]
- Grodzicki, P.; Bacia, J.; Piątkowska, K.; Piechowicz, B. Do Glycogen Content and Thermal Preference in Worker Bees Vary Depending on Geographical Origin? A Comparison of Carniolan Honeybees, Central European Honeybees and Caucasian Honeybees. Agriculture 2023, 13, 492. [Google Scholar] [CrossRef]
- Piepel, M.-F.; Olfs, H.-W. Development of a Physicochemical Test Kit for On-Farm Measurement of Nutrients in Liquid Organic Manures. Agriculture 2023, 13, 477. [Google Scholar] [CrossRef]
- Saeid, A.; Prochownik, E.; Dobrowolska-Iwanek, J. Phosphorus Solubilization by Bacillus Species. Molecules 2018, 23, 2897. [Google Scholar] [CrossRef]
- Davison, N.; Brown, A.; Ross, A. Potential Greenhouse Gas Mitigation from Utilising Pig Manure and Grass for Hydrothermal Carbonisation and Anaerobic Digestion in the UK, EU, and China. Agriculture 2023, 13, 479. [Google Scholar] [CrossRef]
- Sawicka, M.; Latocha, P.; Łata, B. Peel to Flesh Bioactive Compounds Ratio Affect Apple Antioxidant Potential and Cultivar Functional Properties. Agriculture 2023, 13, 478. [Google Scholar] [CrossRef]
- Yan, Z.; Wang, M.; Li, X.; Jiang, H. Impact of African Swine Fever Epidemic on the Cost Intensity of Pork Production in China. Agriculture 2023, 13, 497. [Google Scholar] [CrossRef]
- Méndez-León, J.R.; Camarena-Gómez, D.M.J.; Salgado-Beltrán, L. Competitive Positioning of Mexican Pork in Japan. Agriculture 2023, 13, 505. [Google Scholar] [CrossRef]
- Wu, B.; Wu, Y.; Dong, R.; Sato, K.; Ikuno, S.; Nishimura, S.; Jin, Q. Behavioral Analysis of Mowing Workers Based on Hilbert–Huang Transform: An Auxiliary Movement Analysis of Manual Mowing on the Slopes of Terraced Rice Fields. Agriculture 2023, 13, 489. [Google Scholar] [CrossRef]
- Chi, L.; Zhu, M.; Shen, C.; Zhang, J.; Xing, L.; Zhou, X. Does the Winner Take All in E-Commerce of Agricultural Products under the Background of Platform Monopoly? Agriculture 2023, 13, 476. [Google Scholar] [CrossRef]
- Hashemi, F.; Zalaghi, R.; Enayatizamir, N. Using Zeolite, Molasses, and PGP Microorganisms to Improve Apatite Solubility and Increase Phosphorus Uptake by Sorghum Bicolor L. (Speedfed Cultivar). Sci. Rep. 2025, 15, 19352. [Google Scholar] [CrossRef]
- Borham, A.; Belal, E.; Salem, M. El-gremy, shokry Phosphate Solubilization by Enterobacter Cloacae and Its Impact on Growth and Yield of Wheat Plants. J. Sustain. Agric. Sci. 2017, 43, 89–103. [Google Scholar] [CrossRef]
- Geneva, M.; Hristozkova, M.; Kirova, E.; Sichanova, M.; Stancheva, I. Response to Drought Stress of In Vitro and In Vivo Propagated Physalis Peruviana L. Plants Inoculated with Arbuscular Mycorrhizal Fungi. Agriculture 2023, 13, 472. [Google Scholar] [CrossRef]
- Bieniek, A.; Bieniek, A.; Bielska, N. Variation in Fruit and Seed Morphology of Selected Biotypes and Cultivars of Elaeagnus Multiflora Thunb. in North-Eastern Europe. Agriculture 2023, 13, 495. [Google Scholar] [CrossRef]
- Cheng, T.; Cao, X.; Wang, Y.; Zhao, L. Development of CRISPR-Cas9-Based Genome Editing Tools for Non-Model Microorganism Erwinia Persicina. Synth. Syst. Biotechnol. 2025, 10, 555–563. [Google Scholar] [CrossRef]
- da Silva, L.I.; Pereira, M.C.; de Carvalho, A.M.X.; Buttrós, V.H.; Pasqual, M.; Dória, J. Phosphorus-Solubilizing Microorganisms: A Key to Sustainable Agriculture. Agriculture 2023, 13, 462. [Google Scholar] [CrossRef]
- Correa-Garcia, S.; Constant, P.; Yergeau, E. The Forecasting Power of the Microbiome. Trends Microbiol. 2023, 31, 444–452. [Google Scholar] [CrossRef]
- Liu, R.; Liang, B.; Zhao, H.; Zhao, Y. Impacts of Various Amendments on the Microbial Communities and Soil Organic Carbon of Coastal Saline–Alkali Soil in the Yellow River Delta. Front. Microbiol. 2023, 14. [Google Scholar] [CrossRef]
- Khalid, M.; Sarfraz, M.S.; Iqbal, U.; Aftab, M.U.; Niedbała, G.; Rauf, H.T. Correction: Khalid et al. Real-Time Plant Health Detection Using Deep Convolutional Neural Networks. Agriculture;Agriculture 2023, 13 15, 510 38. [Google Scholar] [CrossRef]
- Gabilondo, R.; Sánchez, J.; Muñoz, P.; Montero-Muñoz, I.; Mauri, P.V.; Marín, J.; Mostaza-Colado, D. Evaluation of Biostimulatory Activity of Commercial Formulations on Three Varieties of Chickpea. Agriculture 2023, 13, 474. [Google Scholar] [CrossRef]
- Hao, M.; Lu, C.; Zhou, X.; Xu, J. How Agricultural Farmers Respond to Risks during the COVID-19 Pandemic: An Exploration through the Dual Social Capitals Approach. Agriculture 2023, 13, 485. [Google Scholar] [CrossRef]
- Sharma, S.; Gupta, S.; Kumari, D.; Kothari, S.L.; Jain, R.; Kachhwaha, S. Exploring Plant Tissue Culture and Steviol Glycosides Production in Stevia Rebaudiana (Bert.) Bertoni: A Review. Agriculture 2023, 13, 475. [Google Scholar] [CrossRef]
- Pei, J.; Liu, P.; Fang, H.; Gao, X.; Pan, B.; Li, H.; Guo, H.; Zhang, F. Estimating Yield and Economic Losses Induced by Ozone Exposure in South China Based on Full-Coverage Surface Ozone Reanalysis Data and High-Resolution Rice Maps. Agriculture 2023, 13, 506. [Google Scholar] [CrossRef]
- Lal, M.; Munda, S.; Begum, T.; Pandey, S.K. Identification and Registration of the Novel High-Rhizome-Yielding Variety Bharamputra-1 of Kaempferia Galanga L. Agriculture 2023, 13, 482. [Google Scholar] [CrossRef]
- Mubeen, S.; Ni, W.; He, C.; Yang, Z. Agricultural Strategies to Reduce Cadmium Accumulation in Crops for Food Safety. Agriculture 2023, 13, 471. [Google Scholar] [CrossRef]
- Hu, J.; Li, G.; Mo, H.; Lv, Y.; Qian, T.; Chen, M.; Lu, S. Crop Node Detection and Internode Length Estimation Using an Improved YOLOv5 Model. Agriculture 2023, 13, 473. [Google Scholar] [CrossRef]
- Wang, Y.; Guo, Y.; Wang, X.; Song, C.; Song, Y.; Liu, Z.; Wang, S.; Gao, S.; Ma, G. Mineral Protection Controls Soil Organic Carbon Stability in Permafrost Wetlands. Sci. Total Env. 2023, 869, 161864. [Google Scholar] [CrossRef] [PubMed]
- Tang, F.; Li, Q.; Yue, J.; Ge, F.; Li, F.; Liu, Y.; Zhang, D.; Tian, J. Penicillium Oxalicum Augments Soil Lead Immobilization by Affecting Indigenous Microbial Community Structure and Inorganic Phosphate Solubilization Potential during Microbial-Induced Phosphate Precipitation. Environ. Pollut. 2023, 319, 120953. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, T.M.; Nadeem, M.; Iftikhar, J.; Salim-ur-Rehman; Ibrahim, S.M.; Majeed, F.; Sultan, M. Effect of Traditional Spices on the Quality and Antioxidant Potential of Paneer Prepared from Buffalo Milk. Agriculture 2023, 13, 491. [Google Scholar] [CrossRef]
- Tomatis, F.; Egerer, M.; Correa-Guimaraes, A.; Navas-Gracia, L.M. Urban Gardening in a Changing Climate: A Review of Effects, Responses and Adaptation Capacities for Cities. Agriculture 2023, 13, 502. [Google Scholar] [CrossRef]
- Shruti; Shukla, A.; Rahman, S.S.; Suneja, P.; Yadav, R.; Hussain, Z.; Singh, R.; Yadav, S.K.; Rana, J.C.; Yadav, S.; et al. Developing an NIRS Prediction Model for Oil, Protein, Amino Acids and Fatty Acids in Amaranth and Buckwheat. Agriculture 2023, 13, 469. [Google Scholar] [CrossRef]
- Park, K. -H.; Lee, C. -Y.; Son, H. -J. Mechanism of Insoluble Phosphate Solubilization by Pseudomonas Fluorescens RAF15 Isolated from Ginseng Rhizosphere and Its Plant Growth-promoting Activities. Lett. Appl. Microbiol. 2009, 49, 222–228. [Google Scholar] [CrossRef]
- Osei, E.; Jafri, S.H.; Gassman, P.W.; Saleh, A.; Gallego, O. Climate Change Impacts on Surface Runoff and Nutrient and Sediment Losses in Buchanan County, Iowa. Agriculture 2023, 13, 470. [Google Scholar] [CrossRef]
- Costa-Gutierrez, S.B.; Adler, C.; Espinosa-Urgel, M.; de Cristóbal, R.E. Pseudomonas Putida and Its Close Relatives: Mixing and Mastering the Perfect Tune for Plants. Appl. Microbiol. Biotechnol. 2022, 106, 3351–3367. [Google Scholar] [CrossRef]
- López-Ortega, M.D.P.; Criollo-Campos, P.J.; Gómez-Vargas, R.M.; Camelo-Runsinque, M.; Estrada-Bonilla, G.; Garrido-Rubiano, M.F.; Bonilla-Buitrago, R. Characterization of Diazotrophic Phosphate Solubilizing Bacteria as Growth Promoters of Maize Plants. Rev. Colomb. De. Biotecnol. 2013, 15, 115–123. [Google Scholar] [CrossRef]
- Abril, A.; Zurdo-Piñeiro, J.L.; Peix, A.; Rivas, R.; Velázquez, E. Solubilization of Phosphate by a Strain of Rhizobium Leguminosarum Bv. Trifolii Isolated from Phaseolus Vulgaris in El Chaco Arido Soil (Argentina). In First International Meeting on Microbial Phosphate Solubilization; Velázquez, E., Rodríguez-Barrueco, C., Eds.; Developments in Plant and Soil Sciences; Springer Netherlands: Dordrecht, 2007; Vol. 102, pp. 135–138. ISBN 978-1-4020-4019-1. [Google Scholar]
- Marra, L.M.; de Oliveira, S.M.; Soares, C.R.F.S.; Moreira, F.M. de S. Solubilisation of Inorganic Phosphates by Inoculant Strains from Tropical Legumes. Sci. agric. 2011, 68, 603–609. [Google Scholar] [CrossRef]
- Akhtar, M.N.; Ansari, E.; Alhady, S.S.N.; Abu Bakar, E. Leveraging on Advanced Remote Sensing- and Artificial Intelligence-Based Technologies to Manage Palm Oil Plantation for Current Global Scenario: A Review. Agriculture 2023, 13, 504. [Google Scholar] [CrossRef]
- Ropelewska, E.; Rutkowski, K.P. The Classification of Peaches at Different Ripening Stages Using Machine Learning Models Based on Texture Parameters of Flesh Images. Agriculture 2023, 13, 498. [Google Scholar] [CrossRef]
- Piechowicz, B.; Kwiatek, A.; Sadło, S.; Zaręba, L.; Koziorowska, A.; Kloc, D.; Balawejder, M. Use of Gas Chromatography and SPME Extraction for the Differentiation between Healthy and Paenibacillus Larvae Infected Colonies of Bee Brood—Preliminary Research. Agriculture 2023, 13, 487. [Google Scholar] [CrossRef]
- Gidey, E.; Dikinya, O.; Sebego, R.; Segosebe, E.; Zenebe, A.; Mussa, S.; Mhangara, P.; Birhane, E. Land Use and Land Cover Change Determinants in Raya Valley, Tigray, Northern Ethiopian Highlands. Agriculture 2023, 13, 507. [Google Scholar] [CrossRef]
- Kan, H.; Qu, S.; Dong, K.; Wang, S.; Xu, C.; Wang, Y.; Hua, S. Integrated Metabolome and Transcriptome Analysis Unveils the Underlying Molecular Response of Panax Ginseng Plants to the Phytophthora Cactorum Infection. Agriculture 2023, 13, 509. [Google Scholar] [CrossRef]
- Chen, N.; Zhao, X.; Dou, S.; Deng, A.; Zheng, C.; Cao, T.; Song, Z.; Zhang, W. The Tradeoff between Maintaining Maize (Zea Mays L.) Productivity and Improving Soil Quality under Conservation Tillage Practice in Semi-Arid Region of Northeast China. Agriculture 2023, 13, 508. [Google Scholar] [CrossRef]
- V, K.; R, J.; M, S. Effect of Potassium Solubilizing Bacteria and Foliar Application of Potassium on Soil Nutrient Status and Soil Biological Properties in Paddy (Oryza Sativa) in Coastal Acid Soils of Karnataka. Int. J. Res. Agron. 2025, 8, 403–408. [Google Scholar] [CrossRef]
- Sani, S.; Bairwa, R.; Abdulkadir, A.I.; Lawal, A.U.; Gurjar, O.P.; Regar, C.L.; Meena, V.S.; Alhassan, S. A Comparative Assessment of Potassium Mineralization from Waste Mica by Application of KSB and FYM in Alluvial and Black Soils 2025.
- Zheng, J.; Shi, G.; Dini-Andreote, F.; Yang, Y.; Jiang, Y. Root-Derived Low Molecular Weight Organic Acids Modulate Keystone Microbial Taxa Impacting Plant Phosphorus Acquisition. J. Adv. Res. 2026, 81, 95–109. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; Lv, L.; Wang, W.; Liu, Y.; Yin, C.; Xu, Q.; Yan, H.; Fu, J.; Liu, X. Differences in Distribution of Potassium-Solubilizing Bacteria in Forest and Plantation Soils in Myanmar. Int. J. Environ. Res. Public Health 2019, 16, 700. [Google Scholar] [CrossRef] [PubMed]
- Pratama, D.; Anas, I.; Suwarno. Ability of Potassium-Solubilissing Microbes to Solubilise Feldspar and Their Effects on Sorghum Growth. Malaysian Journal of Soil Science 2016, 20, 163–175. [Google Scholar]
- Zhang, C.; Kong, F. Isolation and Identification of Potassium-Solubilizing Bacteria from Tobacco Rhizospheric Soil and Their Effect on Tobacco Plants. Appl. Soil. Ecol. 2014, 82, 18–25. [Google Scholar] [CrossRef]
- Lian, B.; Wang, B.; Pan, M.; Liu, C.; Teng, H.H. Microbial Release of Potassium from K-Bearing Minerals by Thermophilic Fungus Aspergillus Fumigatus. Geochim. Et. Cosmochim. Acta 2008, 72, 87–98. [Google Scholar] [CrossRef]
- Lopes-Assad, M.L.; Avansini, S.H.; Rosa, M.M.; de Carvalho, J.R.P.; Ceccato-Antonini, S.R. The Solubilization of Potassium-Bearing Rock Powder by Aspergillus Niger in Small-Scale Batch Fermentations. Can. J. Microbiol. 2010, 56, 598–605. [Google Scholar] [CrossRef]
- Ashrafi-Saiedlou, S.; Rasouli-Sadaghiani, M.; Samadi, A.; Barin, M.; Sepehr, E. Aspergillus Niger as an Eco-Friendly Agent for Potassium Release from K- Bearing Minerals: Isolation, Screening and Culture Medium Optimization Using Plackett-Burman Design and Response Surface Methodology. Heliyon 2024, 10, e29117. [Google Scholar] [CrossRef]
- Gao, G.; Li, X.; Ma, J.; Cui, Y.; Ying, M.; Huang, L.; Li, M. Bacillus Amyloliquefaciens SQ-2 and Biochar: A Promising Combination for Enhancing Rice Growth in Pb/Al-Contaminated Acidic Soils. Microorganisms 2025, 13, 1556. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Jiang, W.; Chen, G.; Wang, X.; Li, T. Gypsum and Organic Materials Improved Soil Quality and Crop Production in Saline-Alkali on the Loess Plateau of China. Front. Environ. Sci. 2024, 12. [Google Scholar] [CrossRef]
- Etesami, H.; Emami, S.; Alikhani, H.A. Potassium Solubilizing Bacteria (KSB):: Mechanisms, Promotion of Plant Growth, and Future Prospects A Review. J. Soil. Sci. Plant Nutr. 2017, 17, 897–911. [Google Scholar] [CrossRef]
- Romero-Munar, A.; Aroca, R. A Non-K+-Solubilizing PGPB (Bacillus Megaterium) Increased K+ Deprivation Tolerance in Oryza Sativa Seedlings by up-Regulating Root K+ Transporters. Plant Physiol. Biochem. 2023, 196, 774–782. [Google Scholar] [CrossRef]
- Nguyen, N.K.; Vo, D.T.V.; Le, T.X.; Morton, L.W.; Tran, H.-T.; Robatjazi, J.; Lasar, H.G.W.; Tecimen, H.B. Isolation, and Selection of Indigenous Potassium Solubilizing Bacteria from Vietnam Mekong Delta Rhizospheric Soils and Their Effects on Diverse Cropping Systems. Biocatal. Agric. Biotechnol. 2024, 58, 103200. [Google Scholar] [CrossRef]
- Kumar, P.; Singh, A.B.; Singh, R. Seasonal Variation and Size Distribution in the Airborne Indoor Microbial Concentration of Residential Houses in Delhi and Its Impact on Health. Aerobiologia 2021, 37, 719–732. [Google Scholar] [CrossRef]
- Boubekri, K.; Soumare, A.; Mardad, I.; Lyamlouli, K.; Hafidi, M.; Ouhdouch, Y.; Kouisni, L. The Screening of Potassium- and Phosphate-Solubilizing Actinobacteria and the Assessment of Their Ability to Promote Wheat Growth Parameters. Microorganisms 2021, 9, 470. [Google Scholar] [CrossRef]
- Ai Min, Z.; Gang Yong, Z.; Tong Guo, G.; Wen, W.; Jia, L.; Shuang Feng, Z.; Bao Cheng, Z. Solubilization of Insoluble Potassium and Phosphate by Paenibacillus Kribensis CX-7: A Soil Microorganism with Biological Control Potential. Afr. J. Microbiol. Res. 2013, 7, 41–47. [Google Scholar] [CrossRef]
- Zhuang, J.; Liu, C.; Wang, X.; Xu, T.; Yang, H. Penicillium Simplicissimum NL-Z1 Induced an Imposed Effect to Promote the Leguminous Plant Growth. Front Microbiol. 2021, 12, 738734. [Google Scholar] [CrossRef] [PubMed]
- Ashrafi-Saeidlou, S.; Samadi, A.; Rasouli-Sadaghiani, M.; Barin, M.; Sepehr, E. Optimization of Potassium Dissolution by Pseudomonas Fluorescens Using Placket-Burman Design and Response Surface Methodology. 2019. [Google Scholar] [CrossRef]
- Yaghoubi Khanghahi, M.; Pirdashti, H.; Rahimian, H.; Nematzadeh, G.; Ghajar Sepanlou, M. The Role of Potassium Solubilizing Bacteria (KSB) Inoculations on Grain Yield, Dry Matter Remobilization and Translocation in Rice (Oryza Sativa L.). J. Plant Nutr. 2019, 42, 1165–1179. [Google Scholar] [CrossRef]
- Chouyia, F.E.; Ventorino, V.; Pepe, O. Diversity, Mechanisms and Beneficial Features of Phosphate-Solubilizing Streptomyces in Sustainable Agriculture: A Review. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- El-Egami, H.M.; Hegab, R.H.; Montaser, H.; El-Hawary, M.M.; Hasanuzzaman, M. Impact of Potassium-Solubilizing Microorganisms with Potassium Sources on the Growth, Physiology, and Productivity of Wheat Crop under Salt-Affected Soil Conditions. Agronomy 2024, 14, 423. [Google Scholar] [CrossRef]
- M.d.h.m, K.; A.d.s.n.p, A.; H.k.i.s, V.; R.m.g, R. Potassium Solubilizers, Aspergillus Sp. and Trichoderma Sp. Promote Growth in Tomato. Proc. Int. For. Environ. Symp. 2023, 27. [Google Scholar] [CrossRef]
- Sl, E.; Km, L.; Padamyar; Ho, K.; Ss, Y. Study on IAA Producing Rhizobacterial Isolates and Their Effect in Talc-Based Carrier on Some Plants. J. Soil. Sci. Plant Health 2000, 2, 1–7. [Google Scholar]
- Yuliatin, E.; Ardyati, T.; Suharjono, S. Effect of Soil Physicochemical Properties on PGPR Density at A Coffee Plantation in Malang, Indonesia. IOP Conf. Ser. Earth Environ. Sci. 2019, 391, 012071. [Google Scholar] [CrossRef]
- Gruet, C.; Abrouk, D.; Börner, A.; Muller, D.; Moënne-Loccoz, Y. D Genome Acquisition and Breeding Have Had a Significant Impact on Interaction of Wheat with ACC Deaminase Producers in Soil or ACC Deaminase Potential Activity in the Rhizosphere. Soil. Biol. Biochem. 2024, 193, 109392. [Google Scholar] [CrossRef]
- Uzma, M.; Nisar, A.; Iqbal, A.; Hasnain, S.; Mahmoud, M.H.; Rahim, M.A.; Gull, T.; Castro-Muñoz, R.; Zongo, E. Exploring the Efficacy of Drought Tolerant, IAA-Producing Plant Growth-Promoting Rhizobacteria for Sustainable Agriculture. Plant Signal Behav. 2025, 20, 2452331. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Chi, Y.; Song, S. Important Soil Microbiota’s Effects on Plants and Soils: A Comprehensive 30-Year Systematic Literature Review. Front. Microbiol. 2024, 15. [Google Scholar] [CrossRef]
- Egamberdieva, D.; Wirth, S.J.; Alqarawi, A.A.; Abd_Allah, E.F.; Hashem, A. Phytohormones and Beneficial Microbes: Essential Components for Plants to Balance Stress and Fitness. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef]
- Neelam, A.; Tabassum, S. Optical Sensing Technologies to Elucidate the Interplay between Plant and Microbes. Micromachines 2023, 14, 195. [Google Scholar] [CrossRef]
- Zaheer, M.S.; Ali, H.H.; Iqbal, M.A.; Erinle, K.O.; Javed, T.; Iqbal, J.; Hashmi, M.I.U.; Mumtaz, M.Z.; Salama, E.A.A.; Kalaji, H.M.; et al. Cytokinin Production by Azospirillum Brasilense Contributes to Increase in Growth, Yield, Antioxidant, and Physiological Systems of Wheat (Triticum Aestivum L.). Front. Microbiol. 2022, 13. [Google Scholar] [CrossRef]
- Nghia, N.K.; Robatjazi, J.; Vy, V.D.T.; Tecimen, H.B.; Lasar, H.G.W.; Lesueur, D.; Bai, S.H.; Tran, H.-T.; Thien, N.H.; Luan, D.T. Effect of Organic Farming Practices on Soil Health Improvement of Coconut Farms. Environ. Technol. Innov. 2025, 38, 104067. [Google Scholar] [CrossRef]
- Allingham, S.M.; Drake, S.J.; Ramsey, A.; Field, C.D.; Nwaishi, F.C.; Elliott, D.R. Changes in Nitrogen Functional Genes and Microbial Populations in Soil Profiles of a Peatland under Different Burning Regimes. Appl. Soil. Ecol. 2024, 200, 105426. [Google Scholar] [CrossRef]
- Cretoiu, M.S.; Korthals, G.W.; Visser, J.H.M.; van Elsas, J.D. Chitin Amendment Increases Soil Suppressiveness toward Plant Pathogens and Modulates the Actinobacterial and Oxalobacteraceal Communities in an Experimental Agricultural Field. Appl. Env. Microbiol. 2013, 79, 5291–5301. [Google Scholar] [CrossRef]
- Jiang, L.; Xu, B.; Wang, Q. Functional Characteristics and Cellulose Degradation Genes of the Microbial Community in Soils with Different Initial pH Values. Agriculture 2025, 15, 1068. [Google Scholar] [CrossRef]
- Li, G.; Wei, M.; Wei, G.; Chen, Z.; Shao, Z. Efficient Heterotrophic Nitrification by a Novel Bacterium Sneathiella Aquimaris 216LB-ZA1-12T Isolated from Aquaculture Seawater. Ecotoxicol. Environ. Saf. 2023, 266, 115588. [Google Scholar] [CrossRef]
- Kalwasinska, A.; Kesy, J.; Donderski, W.; Lalke-Porczyk, E. Biodegradation of Carbendazim by Planktonic and Benthic Bacteria of Eutrophic Lake Chełmżyńskie. Pol. J. Environ. Stud. 2008, 17, 515–523. [Google Scholar]
- Singh, R.; Upadhyay, S.K. A Study on the Plant Litter Decomposition Using Mycoflora for Sustainable Environment. Plan. Sci. 2019, 2, 11–14. [Google Scholar] [CrossRef]
- Fang, W.; Latgé, J.-P. Microbe Profile: Aspergillus Fumigatus: A Saprotrophic and Opportunistic Fungal Pathogen. Microbiology 2018, 164, 1009–1011. [Google Scholar] [CrossRef]
- Heidarzadeh, M.H.; Amani, H.; Javadian, B. Improving Municipal Solid Waste Compost Process by Cycle Time Reduction through Inoculation of Aspergillus Niger. J. Env. Health Sci. Eng. 2019, 17, 295–303. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Tang, M.; Chen, J.; Yang, Y. Identification and Degradation Characteristics of Bacillus Cereus Strain WD-2 Isolated from Prochloraz-Manganese-Contaminated Soils. PLoS ONE 2019, 14, e0220975. [Google Scholar] [CrossRef]
- Su, J.; Zhou, K.; Chen, W.; Xu, S.; Feng, Z.; Chang, Y.; Ding, X.; Zheng, Y.; Tao, X.; Zhang, A.; et al. Enhanced Organic Degradation and Microbial Community Cooperation by Inoculating Bacillus Licheniformis in Low Temperature Composting. J. Environ. Sci. 2024, 143, 189–200. [Google Scholar] [CrossRef]
- Alshelmani, M.I.; Loh, T.C.; Foo, H.L.; Lau, W.H.; Sazili, A.Q. Characterization of Cellulolytic Bacterial Cultures Grown in Different Substrates. [CrossRef]
- Sobolczyk-Bednarek, J.; Choińska-Pulit, A.; Łaba, W. Composting Poultry Feathers with Keratinolytic Bacillus Subtilis: Effects on Degradation Efficiency and Compost Maturity. Materials 2025, 18, 4667. [Google Scholar] [CrossRef] [PubMed]
- Zafar, H.; Peleato, N.; Roberts, D. Bioaugmentation with Bacillus Subtilis and Cellulomonas Fimi to Enhance the Biodegradation of Complex Carbohydrates in MFC-Fed Fruit Waste. Biomass Bioenergy 2023, 174, 106843. [Google Scholar] [CrossRef]
- Bowen, R.M.; Harper, S.H.T. Decomposition of Wheat Straw and Related Compounds by Fungi Isolated from Straw in Arable Soils. Soil. Biol. Biochem. 1990, 22, 393–399. [Google Scholar] [CrossRef]
- Li, X.; Han, C.; Li, W.; Chen, G.; Wang, L. Insights into the Cellulose Degradation Mechanism of the Thermophilic Fungus Chaetomium Thermophilum Based on Integrated Functional Omics. Biotechnol. Biofuels 2020, 13, 143. [Google Scholar] [CrossRef] [PubMed]
- Ha-Tran, D.M.; Nguyen, T.T.M.; Huang, C.-C. Clostridium Thermocellum as a Promising Source of Genetic Material for Designer Cellulosomes: An Overview. Catalysts 2021, 11, 996. [Google Scholar] [CrossRef]
- Olajuyigbe, F.M.; Nlekerem, C.M.; Ogunyewo, O.A. Production and Characterization of Highly Thermostable β-Glucosidase during the Biodegradation of Methyl Cellulose by Fusarium Oxysporum. Biochem. Res. Int. 2016, 2016, 3978124. [Google Scholar] [CrossRef]
- Ladevèze, S.; Haon, M.; Villares, A.; Cathala, B.; Grisel, S.; Herpoël-Gimbert, I.; Henrissat, B.; Berrin, J.-G. The Yeast Geotrichum Candidum Encodes Functional Lytic Polysaccharide Monooxygenases. Biotechnol. Biofuels 2017, 10, 215. [Google Scholar] [CrossRef]
- Zhu, N.; Zhu, Y.; Li, B.; Jin, H.; Dong, Y. Increased Enzyme Activities and Fungal Degraders by Gloeophyllum Trabeum Inoculation Improve Lignocellulose Degradation Efficiency during Manure-Straw Composting. Bioresour. Technol. 2021, 337, 125427. [Google Scholar] [CrossRef] [PubMed]
- Bhairamkar, S.; Kadam, P.; Anjulal, H.; Joshi, A.; Chaudhari, R.; Bagul, D.; Javdekar, V.; Zinjarde, S. Comprehensive Updates on the Biological Features and Metabolic Potential of the Versatile Extremophilic Actinomycete Nocardiopsis Dassonvillei. Res. Microbiol. 2024, 175, 104171. [Google Scholar] [CrossRef]
- Waeonukul, R.; Kyu, K.L.; Sakka, K.; Ratanakhanokchai, K. Isolation and Characterization of a Multienzyme Complex (Cellulosome) of the Paenibacillus Curdlanolyticus B-6 Grown on Avicel under Aerobic Conditions. J. Biosci. Bioeng. 2009, 107, 610–614. [Google Scholar] [CrossRef]
- Wolski, E.A. The Versatility of Penicillium Species to Degrade Organic Pollutants and Its Use for Wastewater Treatment. SIF 2023, 8, 1–10. [Google Scholar] [CrossRef]
- Martinez, D.; Larrondo, L.F.; Putnam, N.; Gelpke, M.D.S.; Huang, K.; Chapman, J.; Helfenbein, K.G.; Ramaiya, P.; Detter, J.C.; Larimer, F.; et al. Genome Sequence of the Lignocellulose Degrading Fungus Phanerochaete Chrysosporium Strain RP78. Nat. Biotechnol. 2004, 22, 695–700. [Google Scholar] [CrossRef]
- Li, F.; Ma, F.; Zhao, H.; Zhang, S.; Wang, L.; Zhang, X.; Yu, H. A Lytic Polysaccharide Monooxygenase from a White-Rot Fungus Drives the Degradation of Lignin by a Versatile Peroxidase. Appl. Environ. Microbiol. 2019, 85, e02803-18. [Google Scholar] [CrossRef]
- Kameshwar, A.K.S.; Qin, W. Molecular Networks of Postia Placenta Involved in Degradation of Lignocellulosic Biomass Revealed from Metadata Analysis of Open Access Gene Expression Data. Int. J. Biol. Sci. 2018, 14, 237–252. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Fransen, A.; de Winde, H. Lignin-Degrading Enzymes and the Potential of Pseudomonas Putida as a Cell Factory for Lignin Degradation and Valorization. Microorganisms 2025, 13, 935. [Google Scholar] [CrossRef]
- Sabar, M.A.; Ali, M.I.; Fatima, N.; Malik, A.Y.; Jamal, A.; Farman, M.; Huang, Z.; Urynowicz, M. Degradation of Low Rank Coal by Rhizopus Oryzae Isolated from a Pakistani Coal Mine and Its Enhanced Releases of Organic Substances. Fuel 2019, 253, 257–265. [Google Scholar] [CrossRef]
- Ahmad, M.; Roberts, J.N.; Hardiman, E.M.; Singh, R.; Eltis, L.D.; Bugg, T.D.H. Identification of DypB from Rhodococcus Jostii RHA1 as a Lignin Peroxidase. Biochemistry 2011, 50, 5096–5107. [Google Scholar] [CrossRef]
- Chater, K.F.; Biró, S.; Lee, K.J.; Palmer, T.; Schrempf, H. The Complex Extracellular Biology of Streptomyces. FEMS Microbiol. Rev. 2010, 34, 171–198. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Pérez, G.; Goma, G.; Rols, J.L. Enhanced Degradation of Lignosulfonated Compounds by Streptomyces Viridosporus. Water Sci. Technol. 1998, 38, 289–297. [Google Scholar] [CrossRef]
- Zainudin, M.H.M.; Gan, H.M.; Tokura, M. Genome Sequence of Thermophilic Cellulolytic Actinobacterium Thermobifida Fusca Strain UPMC 901 Producing Thermostable Cellulases. Gene Rep. 2024, 35, 101900. [Google Scholar] [CrossRef]
- Tišma, M.; Žnidaršič-Plazl, P.; Šelo, G.; Tolj, I.; Šperanda, M.; Bucić-Kojić, A.; Planinić, M. Trametes Versicolor in Lignocellulose-Based Bioeconomy: State of the Art, Challenges and Opportunities. Bioresour. Technol. 2021, 330, 124997. [Google Scholar] [CrossRef] [PubMed]
- Li, J.-X.; Zhang, F.; Jiang, D.-D.; Li, J.; Wang, F.-L.; Zhang, Z.; Wang, W.; Zhao, X.-Q. Diversity of Cellulase-Producing Filamentous Fungi From Tibet and Transcriptomic Analysis of a Superior Cellulase Producer Trichoderma Harzianum LZ117. Front. Microbiol. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Reza, M.M. 1987- ASSESSING THE EFFECT OF SIDEROPHORE PRODUCING BACTERIA FOR THE IRON NUTRITION IN LENTIL AND PEA. 2017. [Google Scholar]
- Yi, X.; Wang, M.; Zhou, Z. The Potential Impact of Naturally Produced Antibiotics, Environmental Factors, and Anthropogenic Pressure on the Occurrence of Erm Genes in Urban Soils. Environ. Pollut. 2019, 245, 282–289. [Google Scholar] [CrossRef] [PubMed]
- Le, T.-H.; Sivachidambaram, V.; Yi, X.; Li, X.; Zhou, Z. Quantification of Polyketide Synthase Genes in Tropical Urban Soils Using Real-Time PCR. J. Microbiol. Methods 2014, 106, 135–142. [Google Scholar] [CrossRef]
- Majewska, M.; Słomka, A.; Hanaka, A. Siderophore-Producing Bacteria from Spitsbergen Soils—Novel Agents Assisted in Bioremediation of the Metal-Polluted Soils. Env. Sci. Pollut. Res. 2024, 31, 32371–32381. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.; Mishra, V.; Rau, N.; Sharma, R.S. Increased Iron-Stress Resilience of Maize through Inoculation of Siderophore-Producing Arthrobacter Globiformis from Mine. J. Basic Microbiol. 2016, 56, 719–735. [Google Scholar] [CrossRef]
- Xie, B.; Wei, X.; Wan, C.; Zhao, W.; Song, R.; Xin, S.; Song, K. Exploring the Biological Pathways of Siderophores and Their Multidisciplinary Applications: A Comprehensive Review. Molecules 2024, 29, 2318. [Google Scholar] [CrossRef]
- Timofeeva, A.M.; Galyamova, M.R.; Sedykh, S.E. Bacterial Siderophores: Classification, Biosynthesis, Perspectives of Use in Agriculture. Plants 2022, 11, 3065. [Google Scholar] [CrossRef]
- Suneja, S.; Lakshminarayana, K.; Narula, N. Optimization of Cultural Conditions for Hydroxamate Type of Siderophore Production by Azotobacter Chroococcum. Microbiol. Res. 1994, 149, 385–390. [Google Scholar] [CrossRef]
- Nouioui, I.; Cortés-albayay, C.; Carro, L.; Castro, J.F.; Gtari, M.; Ghodhbane-Gtari, F.; Klenk, H.-P.; Tisa, L.S.; Sangal, V.; Goodfellow, M. Genomic Insights Into Plant-Growth-Promoting Potentialities of the Genus Frankia. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef]
- Ferreira, C.M.H.; Vilas-Boas, Â.; Sousa, C.A.; Soares, H.M.V.M.; Soares, E.V. Comparison of Five Bacterial Strains Producing Siderophores with Ability to Chelate Iron under Alkaline Conditions. AMB. Express 2019, 9, 78. [Google Scholar] [CrossRef]
- Barona-Gómez, F.; Lautru, S.; Francou, F.-X.; Leblond, P.; Pernodet, J.-L.; Challis, G.L. Multiple Biosynthetic and Uptake Systems Mediate Siderophore-Dependent Iron Acquisition in Streptomyces Coelicolor A3(2) and Streptomyces Ambofaciens ATCC 23877. Microbiology 2006, 152, 3355–3366. [Google Scholar] [CrossRef]
- Bolaji, A.S.; Liasu, M.O.; Ayandele, A.A.; Amao, J.A. Assessing the Synergistic Relationship of Mycorrhiza and Bacteria for the Degradation of Spent Engine Oil in Maize Grown Soil. Bull. Natl. Res. Cent. 2020, 44, 43. [Google Scholar] [CrossRef]
- Park, S.H.; Kang, B.R.; Kim, J.; Lee, Y.; Nam, H.S.; Lee, T.K. Enhanced Soil Fertility and Carbon Dynamics in Organic Farming Systems: The Role of Arbuscular Mycorrhizal Fungal Abundance. J. Fungi 2024, 10, 598. [Google Scholar] [CrossRef]
- Zou, G.; Liu, Y.; Kong, F.; Liao, L.; Deng, G.; Jiang, X.; Cai, J.; Liu, W. Depression of the Soil Arbuscular Mycorrhizal Fungal Community by the Canopy Gaps in a Japanese Cedar (Cryptomeria Japonica) Plantation on Lushan Mountain, Subtropical China. PeerJ 2021, 9, e10905. [Google Scholar] [CrossRef]
- Pawar, P.B.; Melo, J.S.; Kotkar, H.M.; Kulkarni, M.V. Role of Indigenous Mycorrhizal Species in Enhancing Physiological and Biochemical Status, Nutrient Acquisition and Yield Pattern of Groundnut (Arachis Hypogaea L.). J. Crop Sci. Biotechnol. 2018, 21, 23–33. [Google Scholar] [CrossRef]
- Bücking, H.; Liepold, E.; Ambilwade, P. The Role of the Mycorrhizal Symbiosis in Nutrient Uptake of Plants and the Regulatory Mechanisms Underlying These Transport Processes. In Plant Science; IntechOpen, 2012; ISBN 978-953-51-0905-1. [Google Scholar]
- Chen, X.; Song, F.; Liu, F.; Tian, C.; Liu, S.; Xu, H.; Zhu, X. Effect of Different Arbuscular Mycorrhizal Fungi on Growth and Physiology of Maize at Ambient and Low Temperature Regimes. ScientificWorldJournal 2014, 2014, 956141. [Google Scholar] [CrossRef]
- Sauter, M.; Hager, A. The Mycorrhizal Fungus Amanita Muscaria Induces Chitinase Activity in Roots and in Suspension-Cultured Cells of Its Host Picea Abies. Planta 1989, 179, 61–66. [Google Scholar] [CrossRef]
- Sun, Q.; Fu, Z.; Finlay, R.; Lian, B. Transcriptome Analysis Provides Novel Insights into the Capacity of the Ectomycorrhizal Fungus Amanita Pantherina To Weather K-Containing Feldspar and Apatite. Appl. Environ. Microbiol. 2019, 85, e00719-19. [Google Scholar] [CrossRef]
- Aučina, A.; Rudawska, M.; Leski, T.; Skridaila, A.; Riepšas, E.; Iwanski, M. Growth and Mycorrhizal Community Structure of Pinus Sylvestris Seedlings Following the Addition of Forest Litter. Appl. Environ. Microbiol. 2007, 73, 4867–4873. [Google Scholar] [CrossRef] [PubMed]
- Nasslahsen, B.; Prin, Y.; Ferhout, H.; Smouni, A.; Duponnois, R. Mycorrhizae Helper Bacteria for Managing the Mycorrhizal Soil Infectivity. Front. Soil. Sci. 2022, 2. [Google Scholar] [CrossRef]
- Aguilera, P.; Becerra, N.; Alvear, M.; Ortiz, N.; Turrini, A.; Azcón-Aguilar, C.; López-Gómez, M.; Romero, J.K.; Massri, M.; Seguel, A.; et al. Arbuscular Mycorrhizal Fungi from Acidic Soils Favors Production of Tomatoes and Lycopene Concentration. J. Sci. Food Agric. 2022, 102, 2352–2358. [Google Scholar] [CrossRef]
- Brambilla, S.; Stritzler, M.; Soto, G.; Ayub, N. A Synthesis of Functional Contributions of Rhizobacteria to Growth Promotion in Diverse Crops. Rhizosphere 2022, 24, 100611. [Google Scholar] [CrossRef]
- Castillo-Esparza, J.F.; Bandala, V.M.; Ramos, A.; Desgarennes, D.; Carrión, G.; César, E.; Montoya, L.; Ortiz-Castro, R. Pisolithus Tinctorius Extract Affects the Root System Architecture through Compound Production with Auxin-like Activity in Arabidopsis Thaliana. Rhizosphere 2021, 19, 100397. [Google Scholar] [CrossRef]
- Leguminous Trees for Sustainable Tropical Agroforestry. In Advances in Legumes for Sustainable Intensification; Academic Press, 2022; pp. 483–504.
- Frey, S.D. Mycorrhizal Fungi as Mediators of Soil Organic Matter Dynamics. Annu. Rev. Ecol. Evol. Syst. 2019, 50, 237–259. [Google Scholar] [CrossRef]
- Gan, X.; Janus, J.; Willbold, S.; Dombinov, V.; Kuhn, A.J.; Amelung, W.; Reinecke, D.; Calahan, D.; Nedbal, L.; Klose, H.; et al. Wheat Growth and Phosphorus Uptake from Polyculture Algal Biofilms Are Synergistically Modulated by Arbuscular Mycorrhizal Fungi and Serendipita Vermifera. Plant Soil. 2025, 514, 1923–1940. [Google Scholar] [CrossRef]
- Chemistry of Plant Microbe Synergy in the Rhizosphere. In Unravelling Plant-Microbe Synergy; Academic Press, 2023; pp. 283–294.
- Hambleton, S.; Sigler, L. Meliniomyces, a New Anamorph Genus for Root-Associated Fungi with Phylogenetic Affinities to Rhizoscyphus Ericae (Â%¡ Hymenoscyphus Ericae), Leotiomycetes. Stud. Mycol. 2005, 53, 1–27. [Google Scholar] [CrossRef]
- Hilszczanska, D.; Sierota, Z. Persistence of Ectomycorrhizas by Thelephora Terrestris on Outplanted Scots Pine Seedlings. Acta Mycol. 2013, 41, 313–318. [Google Scholar] [CrossRef]
- Jangir, P.; Shekhawat, P.K.; Bishnoi, A.; Ram, H.; Soni, P. Role of Serendipita Indica in Enhancing Drought Tolerance in Crops. Physiol. Mol. Plant Pathol. 2021, 116, 101691. [Google Scholar] [CrossRef]
- Jha, S.S. Evaluating Mycorrhizal Association of Funneliformis Mosseae and Glomus Macrocarpum with Solanum Melongena L. (Brinjal) Using Proteomics and System Biology Approaches 2025, 2025.03.17.643673.
- Klinsukon, C.; Ekprasert, J.; Boonlue, S. Using Arbuscular Mycorrhizal Fungi (Gigaspora Margarita) as a Growth Promoter and Biocontrol of Leaf Blight Disease in Eucalyptus Seedlings Caused by Cylindrocladium Quinqueseptatum. Rhizosphere 2021, 20, 100450. [Google Scholar] [CrossRef]
- Kusakabe, R.; Yamato, M. Isolation and Identification of an Arbuscular Mycorrhizal Fungus Specifically Associated with Mycoheterotrophic Seedlings of Gentiana Zollingeri (Gentianaceae). Mycoscience 64 55–62. [CrossRef]
- Ladeyn, I.; Plassard, C.; Staunton, S. Mycorrhizal Association of Maritime Pine, Pinus Pinaster, with Rhizopogon Roseolus Has Contrasting Effects on the Uptake from Soil and Root-to-Shoot Transfer of 137Cs, 85Sr and 95mTc. J. Environ. Radioact. 2008, 99, 853–863. [Google Scholar] [CrossRef]
- Malicka, M.; Magurno, F.; Posta, K.; Chmura, D.; Piotrowska-Seget, Z. Differences in the Effects of Single and Mixed Species of AMF on the Growth and Oxidative Stress Defense in Lolium Perenne Exposed to Hydrocarbons. Ecotoxicol. Environ. Saf. 2021, 217, 112252. [Google Scholar] [CrossRef]
- Mello, A.; Lumini, E.; Napoli, C.; Bianciotto, V.; Bonfante, P. Arbuscular Mycorrhizal Fungal Diversity in the Tuber Melanosporum Brûlé. Fungal Biol. 2015, 119, 518–527. [Google Scholar] [CrossRef]
- Midgley, D.J.; Rosewarne, C.P.; Greenfield, P.; Li, D.; Vockler, C.J.; Hitchcock, C.J.; Sawyer, N.A.; Brett, R.; Edwards, J.; Pitt, J.I.; et al. Genomic Insights into the Carbohydrate Catabolism of Cairneyella Variabilis Gen. Nov. Sp. Nov., the First Reports from a Genome of an Ericoid Mycorrhizal Fungus from the Southern Hemisphere. Mycorrhiza 2016, 26, 345–352. [Google Scholar] [CrossRef] [PubMed]
- Mikheev, V.S.; Struchkova, I.V.; Churkina, L.M.; Brilkina, A.A.; Berezina, E.V. Several Characteristics of Oidiodendron Maius G.L. Barron Important for Heather Plants’ Controlled Mycorrhization. J. Fungi 2023, 9, 728. [Google Scholar] [CrossRef]
- Oliveira, T.C.; Cabral, J.S.R.; Santana, L.R.; Tavares, G.G.; Santos, L.D.S.; Paim, T.P.; Müller, C.; Silva, F.G.; Costa, A.C.; Souchie, E.L.; et al. The Arbuscular Mycorrhizal Fungus Rhizophagus Clarus Improves Physiological Tolerance to Drought Stress in Soybean Plants. Sci. Rep. 2022, 12, 9044. [Google Scholar] [CrossRef] [PubMed]
- Orfanoudakis, M.; Wheeler, C.T.; Hooker, J.E. Both the Arbuscular Mycorrhizal Fungus Gigaspora Rosea and Frankia Increase Root System Branching and Reduce Root Hair Frequency in Alnus Glutinosa. Mycorrhiza 2010, 20, 117–126. [Google Scholar] [CrossRef] [PubMed]
- Pagano, M.C.; Cabello, M.N.; Scotti, M.R. Arbuscular Mycorrhizal Colonization and Growth of Eremanthus Incanus Less. in a Highland Field. Plant Soil. Environ. 2010, 56, 412–418. [Google Scholar] [CrossRef]
- Parise, A.G.; De Oliveira, V.H.; Tibbett, M.; Pickles, B.J. The Pitfalls of Ectomycorrhizal Microcosms: Lessons Learnt for Future Success. Plant Signal. Behav. 2025, 20, 2527378. [Google Scholar] [CrossRef]
- Perotto, S.; Daghino, S.; Martino, E. Ericoid Mycorrhizal Fungi and Their Genomes: Another Side to the Mycorrhizal Symbiosis? New Phytol. 2018, 220, 1141–1147. [Google Scholar] [CrossRef]
- Peterson, R.L.; Currah, R.S. Synthesis of Mycorrhizae between Protocorms of Goodyera Repens (Orchidaceae) and Ceratobasidium Cereale. Can. J. Bot. 1990, 68, 1117–1125. [Google Scholar] [CrossRef]
- Pujasatria, G.C.; Miura, C.; Yamaguchi, K.; Shigenobu, S.; Kaminaka, H. Colonization by Orchid Mycorrhizal Fungi Primes Induced Systemic Resistance against Necrotrophic Pathogen. Front Plant Sci. 2024, 15, 1447050. [Google Scholar] [CrossRef]
- Rineau, F.; Roth, D.; Shah, F.; Smits, M.; Johansson, T.; Canbäck, B.; Olsen, P.B.; Persson, P.; Grell, M.N.; Lindquist, E.; et al. The Ectomycorrhizal Fungus Paxillus Involutus Converts Organic Matter in Plant Litter Using a Trimmed Brown-Rot Mechanism Involving Fenton Chemistry. Env. Microbiol. 2012, 14, 1477–1487. [Google Scholar] [CrossRef]
- Sun, X.; Chen, W.; Ivanov, S.; MacLean, A.M.; Wight, H.; Ramaraj, T.; Mudge, J.; Harrison, M.J.; Fei, Z. Genome and Evolution of the Arbuscular Mycorrhizal Fungus Diversispora Epigaea (Formerly Glomus Versiforme) and Its Bacterial Endosymbionts. New Phytol. 2019, 221, 1556–1573. [Google Scholar] [CrossRef]
- Timonen, S.; Peterson, R.L. Cytoskeleton in Mycorrhizal Symbiosis. Plant Soil. 2002, 244, 199–210. [Google Scholar] [CrossRef]
- Wang, P.; Kuzyakov, Y.; Wang, Y.; Liu, Y.; Liu, J.; Qi, Z.; He, Y.; Jiang, Q. Quantifying Microbial Necromass Contributions to Soil Carbon Sequestration under Diverse Cropland Management Practices: A Meta-Analysis. J. Environ. Manag. 2025, 388, 126008. [Google Scholar] [CrossRef]
- Wu, Q.-S.; Zou, Y.-N.; Huang, Y.-M. The Arbuscular Mycorrhizal Fungus Diversispora Spurca Ameliorates Effects of Waterlogging on Growth, Root System Architecture and Antioxidant Enzyme Activities of Citrus Seedlings. Fungal Ecol. 2013, 6, 37–43. [Google Scholar] [CrossRef]
- Jeon, J.; Lee, S.; Kim, H.; Ahn, T.-S.; Song, H.-G. Plant Growth Promotion in Soil by Some Inoculated Microorganisms. J. Microbiol. 2003, 41, 271–276. [Google Scholar]
- Bouffaud, M.-L.; Renoud, S.; Dubost, A.; Moënne-Loccoz, Y.; Muller, D. 1-Aminocyclopropane-1-Carboxylate Deaminase Producers Associated to Maize and Other Poaceae Species. Microbiome 2018, 6, 114. [Google Scholar] [CrossRef]
- Alexandre, F.S.; Della Flora, L.V.; Henrique, I.G.; da Silva, D.C.; Mercedes, A.P.; Cardoso Silva, A.; Silva de Oliveira, A.; Bondespacho da Silva, M.P.; Formelh Ronning, B.P.; Dreher, D.R.; et al. Arbuscular Mycorrhizal Fungi (Rhizophagus Clarus) and Rhizobacteria (Bacillus Subtilis) Can Improve the Clonal Propagation and Development of Teak for Commercial Plantings. Front. Plant Sci. 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Pseudomonas Fluorescens: A Plant-Growth-Promoting Rhizobacterium (PGPR) With Potential Role in Biocontrol of Pests of Crops. In Crop Improvement Through Microbial Biotechnology; Elsevier, 2018; pp. 221–243.
- Du, Y.; Wang, T.; Jiang, J.; Wang, Y.; Lv, C.; Sun, K.; Sun, J.; Yan, B.; Kang, C.; Guo, L.; et al. Biological Control and Plant Growth Promotion Properties of Streptomyces Albidoflavus St-220 Isolated from Salvia Miltiorrhiza Rhizosphere. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Fanai, A.; Bohia, B.; Lalremruati, F.; Lalhriatpuii, N.; Lalrokimi; Lalmuanpuii, R.; Singh, P.K. Zothanpuia Plant Growth Promoting Bacteria (PGPB)-Induced Plant Adaptations to Stresses: An Updated Review. PeerJ 2024, 12, e17882. [Google Scholar] [CrossRef]
- Kakabouki, I.; Mavroeidis, A.; Tataridas, A.; Kousta, A.; Efthimiadou, A.; Karydogianni, S.; Katsenios, N.; Roussis, I.; Papastylianou, P. Effect of Rhizophagus Irregularis on Growth and Quality of Cannabis Sativa Seedlings. Plants 2021, 10, 1333. [Google Scholar] [CrossRef]
- Ma, N.; Yokoyama, K.; Marumoto, T. Promotion of Host Plant Growth and Infection of Roots with Arbuscular Mycorrhizal Fungus Gigaspora Margarita by the Application of Peat. Soil. Sci. Plant Nutr. 2006, 52, 162–167. [Google Scholar] [CrossRef]
- Magesh, S.; Hurley, A.I.; Nepper, J.F.; Chevrette, M.G.; Schrope, J.H.; Li, C.; Beebe, D.J.; Handelsman, J. Surface Colonization by Flavobacterium Johnsoniae Promotes Its Survival in a Model Microbial Community. mBio 2024, 15, e03428-23. [Google Scholar] [CrossRef] [PubMed]
- Noel, T.C.; Sheng, C.; Yost, C.K.; Pharis, R.P.; Hynes, M.F. Rhizobium Leguminosarum as a Plant Growth-Promoting Rhizobacterium: Direct Growth Promotion of Canola and Lettuce. Can. J. Microbiol. 1996, 42, 279–283. [Google Scholar] [CrossRef]
- Oleńska, E.; Małek, W.; Wójcik, M.; Swiecicka, I.; Thijs, S.; Vangronsveld, J. Beneficial Features of Plant Growth-Promoting Rhizobacteria for Improving Plant Growth and Health in Challenging Conditions: A Methodical Review. Sci. Total Environ. 2020, 743, 140682. [Google Scholar] [CrossRef] [PubMed]
- Pallucchini, M.; Franchini, M.; El-Ballat, E.M.; Narraidoo, N.; Pointer-Gleadhill, B.; Palframan, M.J.; Hayes, C.J.; Dent, D.; Cocking, E.C.; Perazzolli, M.; et al. Gluconacetobacter Diazotrophicus AZ0019 Requires Functional nifD Gene for Optimal Plant Growth Promotion in Tomato Plants. Front Plant Sci. 2024, 15, 1469676. [Google Scholar] [CrossRef]
- Ruiz-Güereca, D.A.; Sánchez-Saavedra, M. del P. Growth and Phosphorus Removal by Synechococcus Elongatus Co-Immobilized in Alginate Beads with Azospirillum Brasilense. J. Appl. Phycol. 2016, 28, 1501–1507. [Google Scholar] [CrossRef]
- Stassinos, P.M.; Rossi, M.; Borromeo, I.; Capo, C.; Beninati, S.; Forni, C. Amelioration of Salt Stress Tolerance in Rapeseed (Brassica Napus) Cultivars by Seed Inoculation with Arthrobacter Globiformis. Plant Biosystems-An International Journal Dealing with all Aspects of Plant Biology 2022, 156, 370–383. [Google Scholar] [CrossRef]
- Viscardi, S.; Ventorino, V.; Duran, P.; Maggio, A.; De Pascale, S.; Mora, M.L.; Pepe, O. Assessment of Plant Growth Promoting Activities and Abiotic Stress Tolerance of Azotobacter Chroococcum Strains for a Potential Use in Sustainable Agriculture. J. Soil. Sci. Plant Nutr. 2016, 16, 848–863. [Google Scholar] [CrossRef]
- Yadav, M.; Divyanshu, K.; Dubey, M.K.; Rai, A.; Kumar, S.; Tripathi, Y.N.; Shukla, V.; Upadhyay, R.S. Plant Growth Promotion and Differential Expression of Defense Genes in Chilli Pepper against Colletotrichum Truncatum Induced by Trichoderma Asperellum and T. Harzianum. BMC Microbiol. 2023, 23, 54. [Google Scholar] [CrossRef]
- Zhang, J.; Diao, F.; Hao, B.; Xu, L.; Jia, B.; Hou, Y.; Ding, S.; Guo, W. Multiomics Reveals Claroideoglomus Etunicatum Regulates Plant Hormone Signal Transduction, Photosynthesis and La Compartmentalization in Maize to Promote Growth under La Stress. Ecotoxicol. Environ. Saf. 2023, 262, 115128. [Google Scholar] [CrossRef] [PubMed]
- Hashem, A.; Tabassum, B.; Fathi Abd_Allah, E. Bacillus Subtilis: A Plant-Growth Promoting Rhizobacterium That Also Impacts Biotic Stress. Saudi J. Biol. Sci. 2019, 26, 1291–1297. [Google Scholar] [CrossRef]
- Nozari, M.; Samaei, M.; Dehghani, M.; Ebrahimi, A. Bioremediation of Alkane Hydrocarbons Using Bacterial Consortium From Soil. Health Scope In Press. 2018. [Google Scholar] [CrossRef]
- Helmy, Q.; Kardena, E. Enhancing Field-Scale Bioremediation of Weathered Petroleum Oil-Contaminated Soil with Biocompost as a Bulking Agent. Case Stud. Chem. Environ. Eng. 2024, 9, 100735. [Google Scholar] [CrossRef]
- Park, J.-W.; Crowley, D.E. Dynamic Changes in nahAc Gene Copy Numbers during Degradation of Naphthalene in PAH-Contaminated Soils. Appl. Microbiol. Biotechnol. 2006, 72, 1322–1329. [Google Scholar] [CrossRef]
- Sun, Y.; Polishchuk, E.A.; Radoja, U.; Cullen, W.R. Identification and Quantification of arsC Genes in Environmental Samples by Using Real-Time PCR. J. Microbiol. Methods 2004, 58, 335–349. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Nostrand, J.D.V.; Wu, L.; He, Z.; Qin, Y.; Zhao, F.-J.; Zhou, J. Bacterial Community and Arsenic Functional Genes Diversity in Arsenic Contaminated Soils from Different Geographic Locations. PLoS ONE 2017, 12, e0176696. [Google Scholar] [CrossRef]
- Kim, J.-W.; Hong, Y.-K.; Kim, H.-S.; Oh, E.-J.; Park, Y.-H.; Kim, S.-C. Metagenomic Analysis for Evaluating Change in Bacterial Diversity in TPH-Contaminated Soil after Soil Remediation. Toxics 2021, 9, 319. [Google Scholar] [CrossRef] [PubMed]
- Ayilara, M.S.; Babalola, O.O. Bioremediation of Environmental Wastes: The Role of Microorganisms. Front. Agron. 2023, 5. [Google Scholar] [CrossRef]
- Dell’ Anno, F.; Rastelli, E.; Sansone, C.; Brunet, C.; Ianora, A.; Dell’ Anno, A. Bacteria, Fungi and Microalgae for the Bioremediation of Marine Sediments Contaminated by Petroleum Hydrocarbons in the Omics Era. Microorganisms 2021, 9, 1695. [Google Scholar] [CrossRef]
- Jangra, B.; Dwibedi, V.; Kaur, G.; Bhadrecha, P. Decoding Genomic Potential of Streptomyces Spp. for Sustainable Environment and Industries. South Afr. J. Bot. 2025, 184, 480–494. [Google Scholar] [CrossRef]
- Zeng, Y.; Feng, R.; Huang, C.; Liu, J.; Yang, F. Antibiotic Resistance Genes in Agricultural Soils: A Comprehensive Review of the Hidden Crisis and Exploring Control Strategies. Toxics 2025, 13, 239. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Cai, Z.; Zhou, Q.; Zhang, Z.; Chen, C. Bioelectrochemical Stimulation of Petroleum Hydrocarbon Degradation in Saline Soil Using U-Tube Microbial Fuel Cells. Biotechnol. Bioeng. 2012, 109, 426–433. [Google Scholar] [CrossRef]
- Fukuhara, Y.; Horii, S.; Matsuno, T.; Matsumiya, Y.; Mukai, M.; Kubo, M. Distribution of Hydrocarbon-Degrading Bacteria in the Soil Environment and Their Contribution to Bioremediation. Appl. Biochem Biotechnol. 2013, 170, 329–339. [Google Scholar] [CrossRef] [PubMed]
- Kloos, K.; Munch, J.C.; Schloter, M. A New Method for the Detection of Alkane-Monooxygenase Homologous Genes (alkB) in Soils Based on PCR-Hybridization. J. Microbiol. Methods 2006, 66, 486–496. [Google Scholar] [CrossRef]
- Liu, Q.; Tang, J.; Bai, Z.; Hecker, M.; Giesy, J.P. Distribution of Petroleum Degrading Genes and Factor Analysis of Petroleum Contaminated Soil from the Dagang Oilfield, China. Sci. Rep. 2015, 5, 11068. [Google Scholar] [CrossRef] [PubMed]
- Tuomi, P.M.; Salminen, J.M.; Jørgensen, K.S. The Abundance of nahAc Genes Correlates with the 14C-Naphthalene Mineralization Potential in Petroleum Hydrocarbon-Contaminated Oxic Soil Layers. FEMS Microbiol. Ecol. 2004, 51, 99–107. [Google Scholar] [CrossRef]
- Ezemba, C.C.; Ekwealor, C.C.; Ekwealor, I.A.; Ozokpo, C.A.; Chukwujekwu, C.E.; Anakwenze, V.N.; Anaukwu, C.G.; Archibong, E.J. Utilization of Hydrocarbons by Bacteria Isolated from Oil-Contaminated and Uncontaminated Soil in South-East Nigeria 2015.
- Abo-State, M.A.M.; Osman, M.E.; Khattab, O.H.; El-Kelani, T.A.; Abdel-Rahman, Z.M. Degradative Pathways of Polycyclic Aromatic Hydrocarbons (PAHs) by Phanerochaete Chrysosporium under Optimum Conditions. J. Radiat. Res. Appl. Sci. 2021, 14, 507–520. [Google Scholar] [CrossRef]
- Al-Otibi, F.; Al-Zahrani, R.M.; Marraiki, N. Biodegradation of Selected Hydrocarbons by Fusarium Species Isolated from Contaminated Soil Samples in Riyadh, Saudi Arabia. J. Fungi 2023, 9, 216. [Google Scholar] [CrossRef]
- Asemoloye, M.D.; Tosi, S.; Daccò, C.; Wang, X.; Xu, S.; Marchisio, M.A.; Gao, W.; Jonathan, S.G.; Pecoraro, L. Hydrocarbon Degradation and Enzyme Activities of Aspergillus Oryzae and Mucor Irregularis Isolated from Nigerian Crude Oil-Polluted Sites. Microorganisms 2020, 8, 1912. [Google Scholar] [CrossRef]
- Barathi, S.; Vasudevan, N. Utilization of Petroleum Hydrocarbons by Pseudomonas Fluorescens Isolated from a Petroleum-Contaminated Soil. Environ. Int. 2001, 26, 413–416. [Google Scholar] [CrossRef] [PubMed]
- Brown, W.A.; Cooper, D.G. Hydrocarbon Degradation by Acinetobacter Calcoaceticus RAG-1 Using the Self-Cycling Fermentation Technique. Biotechnol. Bioeng. 1992, 40, 797–805. [Google Scholar] [CrossRef] [PubMed]
- Chikwem, I.; Christopher N, U.; Ekwenye, U.; Stanley, O.; Irene, E.; Duru, I. Biodegradation Potential of Aspergillus Niger on Crude Oil Contaminated Soil. 2017. [Google Scholar]
- Dyksterhouse, S.E.; Gray, J.P.; Herwig, R.P.; Lara, J.C.; Staley, J.T. Cycloclasticus Pugetii Gen. Nov., Sp. Nov., an Aromatic Hydrocarbon-Degrading Bacterium from Marine Sediments. Int. J. Syst. Bacteriol. 1995, 45, 116–123. [Google Scholar] [CrossRef]
- Fredslund, L.; Sniegowski, K.; Wick, L.Y.; Jacobsen, C.S.; De Mot, R.; Springael, D. Surface Motility of Polycyclic Aromatic Hydrocarbon (PAH)-Degrading Mycobacteria. Res. Microbiol. 2008, 159, 255–262. [Google Scholar] [CrossRef]
- Hu, K.; Peris, A.; Torán, J.; Eljarrat, E.; Sarrà, M.; Blánquez, P.; Caminal, G. Exploring the Degradation Capability of Trametes Versicolor on Selected Hydrophobic Pesticides through Setting Sights Simultaneously on Culture Broth and Biological Matrix. Chemosphere 2020, 250, 126293. [Google Scholar] [CrossRef]
- Ibrahim, S.Y.; Abdelhamid, E.A.; El-Hagrassi, A.M.; Kamal, N.M. Statistical Optimization of Crude Oil Bioremediation Using Streptomyces Aurantiogriseus Isolated from Egypt’s Western Desert. Biodegradation 2025, 36, 68. [Google Scholar] [CrossRef]
- Kadri, T.; Magdouli, S.; Rouissi, T.; Brar, S.K. Ex-Situ Biodegradation of Petroleum Hydrocarbons Using Alcanivorax Borkumensis Enzymes. Biochem. Eng. J. 2018, 132, 279–287. [Google Scholar] [CrossRef]
- Khan, S.R.; Nirmal, J.I.K.; Kumar, R.N.; Patel, J.G. Biodegradation of Kerosene: Study of Growth Optimization and Metabolic Fate of P. Janthinellum SDX7. Braz. J. Microbiol. 2015, 46, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.-J.; Kweon, O.; Jones, R.C.; Edmondson, R.D.; Cerniglia, C.E. Genomic Analysis of Polycyclic Aromatic Hydrocarbon Degradation in Mycobacterium Vanbaalenii PYR-1. Biodegradation 2008, 19, 859–881. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Yang, Y.; Zhang, D.; Li, Y.; Zhang, H.; Luo, J.; Jones, K.C. Evaluating the Simulated Toxicities of Metal Mixtures and Hydrocarbons Using the Alkane Degrading Bioreporter Acinetobacter Baylyi ADPWH_recA. J. Hazard. Mater. 2021, 419, 126471. [Google Scholar] [CrossRef]
- Li, L.; Liu, Y.; Zhang, Y.; Wang, X.; Liu, H.; Liu, Q.; Chen, S.; Zhou, G. Dynamic Responses in Pseudomonas Stutzeri M3 Bioaugmentation of Crude-Oil-Contaminated Soil: Hydrocarbons, Microbial Community Structures, and Functional Genes. J. Environ. Chem. Eng. 2024, 12, 111863. [Google Scholar] [CrossRef]
- Lopez, E.S.; Elufisan, T.O.; Bustos, P.; Charles, C.P.M.; Mendoza-Herrera, A.; Guo, X. Complete Genome Report of a Hydrocarbon-Degrading Sphingobium Yanoikuyae S72. Appl. Sci. 2022, 12, 6201. [Google Scholar] [CrossRef]
- Mayans, B.; Antón-Herrero, R.; García-Delgado, C.; Delgado-Moreno, L.; Guirado, M.; Pérez-Esteban, J.; Escolástico, C.; Eymar, E. Bioremediation of Petroleum Hydrocarbons Polluted Soil by Spent Mushroom Substrates: Microbiological Structure and Functionality. J. Hazard. Mater. 2024, 473, 134650. [Google Scholar] [CrossRef]
- Mounier, J.; Camus, A.; Mitteau, I.; Vaysse, P.-J.; Goulas, P.; Grimaud, R.; Sivadon, P. The Marine Bacterium Marinobacter hydrocarbonoclasticusSP17 Degrades a Wide Range of Lipids and Hydrocarbons through the Formation of Oleolytic Biofilms with Distinct Gene Expression Profiles. FEMS Microbiol. Ecol. 2014, 90, 816–831. [Google Scholar] [CrossRef] [PubMed]
- Oh, K.-B.; Mar, W.; Chang, I.-M. Biodegradation of Hydrocarbons by an Organic Solvent-Tolerant Fungus, Cladosporium Resinae NK-1. J. Microbiol. Biotechnol. 2001, 11, 56–60. [Google Scholar]
- Okoro, C.; Amund, O. Biodegradation of Produced Water Hydrocarbons by Aspergillus Fumigatus. J. Am. Sc. 2009, 6. [Google Scholar]
- Lamprea Pineda, P.A.; Demeestere, K.; Alvarado-Alvarado, A.A.; Devlieghere, F.; Boon, N.; Van Langenhove, H.; Walgraeve, C. Degradation of Gaseous Hydrocarbons in Aerated Stirred Bioreactors Inoculated with Rhodococcus Erythropolis: Effect of the Carbon Source and SIFT-MS Method Development. J. Environ. Sci. 2025, 147, 268–281. [Google Scholar] [CrossRef]
- Purnomo, A.S.; Rizqi, H.D.; Harmelia, L.; Anggraeni, S.D.; Melati, R.E.; Damayanti, Z.H.; Shafwah, O.M.; Kusuma, F.C. Biodegradation of Crude Oil by Ralstonia Pickettii under High Salinity Medium. Malays. J. Fundam. Appl. Sci. 2019, 15, 377–380. [Google Scholar] [CrossRef]
- Radwan, S.S.; Barabás, G.; Sorkhoh, N.A.; Damjanovich, S.; Szabó, I.; Szöllősi, J.; Matkó, J.; Penyige, A.; Hirano, T.; Szabó, I.M. Hydrocarbon Uptake by Streptomyces. FEMS Microbiol. Lett. 1998, 169, 87–94. [Google Scholar] [CrossRef]
- Ramdass, A.C.; Rampersad, S.N. Draft Genome of Hydrocarbon-Degrader Burkholderia Cepacia BCTT, Isolated from Soil Chronically Polluted with Crude Oil in Trinidad. Microbiol. Resour. Announc. 2024, 13, e00902-24. [Google Scholar] [CrossRef]
- Romanowska, I.; Kwapisz, E.; Mitka, M.; Bielecki, S. Isolation and Preliminary Characterization of a Respiratory Nitrate Reductase from Hydrocarbon-Degrading Bacterium Gordonia Alkanivorans S7. J. Ind. Microbiol. Biotechnol. 2010, 37, 625–629. [Google Scholar] [CrossRef]
- Sonia, S.; Abdelkader, B.; Zoulikha, B.; Vogel, T.; Navarro, E.; Sonia, I. THE DIVERSITY OF CULTIVABLE HYDROCARBON-DEGRADING BACTERIA ISOLATED FROM CRUDE OIL CONTAMINATED SOIL AND SLUDGE FROM ARZEW REFINERY IN ALGERIA. J. Microbiol. Biotechnol. Food Sci. 2019, 9. [Google Scholar] [CrossRef]
- Trögl, J.; Esuola, C.O.; Kříženecká, S.; Kuráň, P.; Seidlová, L.; Veronesi-Dáňová, P.; Popelka, J.; Babalola, O.O.; Hrabák, P.; Czinnerová, M.; et al. Biodegradation of High Concentrations of Aliphatic Hydrocarbons in Soil from a Petroleum Refinery: Implications for Applicability of New Actinobacterial Strains. Appl. Sci. 2018, 8, 1855. [Google Scholar] [CrossRef]
- Xu, X.; Liu, W.; Tian, S.; Wang, W.; Qi, Q.; Jiang, P.; Gao, X.; Li, F.; Li, H.; Yu, H. Petroleum Hydrocarbon-Degrading Bacteria for the Remediation of Oil Pollution Under Aerobic Conditions: A Perspective Analysis. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Zhang, J.; Liu, Y.; Feng, W. Biodegradation of Hydrocarbons by Purpureocillium Lilacinum and Penicillium Chrysogenum from Heavy Oil Sludge and Their Potential for Bioremediation of Contaminated Soils. Int. Biodeterior. Biodegrad. 2023, 178, 105566. [Google Scholar] [CrossRef]
- Zylstra, G.J.; Kim, E. Aromatic Hydrocarbon Degradation by Sphingomonas Yanoikuyae B1. J. Ind. Microbiol. Biotechnol. 1997, 19, 408–414. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Dawson, R.A.; Bradley, J.A.; Hernández, M. Prevalence and Dynamics of Antimicrobial Resistance in Pioneer and Developing Arctic Soils. BMC Microbiol. 2025, 25, 50. [Google Scholar] [CrossRef]
- Meyers, M.A.; Durso, L.M.; Gilley, J.E.; Waldrip, H.M.; Castleberry, L.; Millmier-Schmidt, A. Antibiotic Resistance Gene Profile Changes in Cropland Soil after Manure Application and Rainfall. J. Env. Qual. 2020, 49, 754–761. [Google Scholar] [CrossRef] [PubMed]
- Symochko, L.; Demyanyuk, O.; Symochko, V.; Grulova, D.; Fejer, J.; Mariychuk, R. The Spreading of Antibiotic-Resistant Bacteria in Terrestrial Ecosystems and the Formation of Soil Resistome. Land 2023, 12, 769. [Google Scholar] [CrossRef]
- Crowley, P.D.; Vaillant, J.J.; Shirley, J.D.; Wengenack, N.L.; Jo Kasten, M. Rapidly Growing Knowledge of Mycobacterium Smegmatis: A Case Series and Review of Antimicrobial Susceptibility Patterns. J. Clin. Tuberc. Other Mycobact. Dis. 2024, 37, 100489. [Google Scholar] [CrossRef]
- Holmes, A.H.; Govan, J.; Goldstein, R. Agricultural Use of Burkholderia (Pseudomonas) Cepacia: A Threat to Human Health? Emerg. Infect. Dis. J.-CDC 1998, 4. [Google Scholar] [CrossRef]
- Jensen, G.B.; Hansen, B.M.; Eilenberg, J.; Mahillon, J. The Hidden Lifestyles of Bacillus Cereus and Relatives. Environ. Microbiol. 2003, 5, 631–640. [Google Scholar] [CrossRef]
- Kisková, J.; Juhás, A.; Galušková, S.; Maliničová, L.; Kolesárová, M.; Piknová, M.; Pristaš, P. Antibiotic Resistance and Genetic Variability of Acinetobacter Spp. from Wastewater Treatment Plant in Kokšov-Bakša (Košice, Slovakia). Microorganisms 2023, 11, 840. [Google Scholar] [CrossRef]
- Kyselková, M.; Chroňáková, A.; Němec, J.; Kotrbová, L.; Elhottová, D. ISOLATION AND ANTIBIOTIC SUSCEPTIBILITY TESTING OF RAPIDLY-GROWING MYCOBACTERIA FROM GRASSLAND SOILS. J. Microbiol. Biotechnol. Food Sci. 2013, 3, 76–80. [Google Scholar]
- Lewandowska, W.; Mahillon, J.; Drewnowska, J.M.; Swiecicka, I. Insight into the Phylogeny and Antibiotic Resistance of Pseudomonas Spp. Originating from Soil of the Białowieża National Park in Northeastern Poland. Front. Microbiol. 2025, 16. [Google Scholar] [CrossRef] [PubMed]
- Mahapatra, S.; Yadav, R.; Ramakrishna, W. Bacillus Subtilis Impact on Plant Growth, Soil Health and Environment: Dr. Jekyll and Mr. Hyde. J. Appl. Microbiol. 2022, 132, 3543–3562. [Google Scholar] [CrossRef]
- Nirwati, H.; Sinanjung, K.; Fahrunissa, F.; Wijaya, F.; Napitupulu, S.; Hati, V.P.; Hakim, M.S.; Meliala, A.; Aman, A.T.; Nuryastuti, T. Biofilm Formation and Antibiotic Resistance of Klebsiella Pneumoniae Isolated from Clinical Samples in a Tertiary Care Hospital, Klaten, Indonesia. BMC Proc. 2019, 13, 20. [Google Scholar] [CrossRef]
- Peleg, A.Y.; Seifert, H.; Paterson, D.L. Acinetobacter Baumannii: Emergence of a Successful Pathogen. Clin. Microbiol. Rev. 2008, 21, 538–582. [Google Scholar] [CrossRef] [PubMed]
- Silverio, M.P.; Schultz, J.; Parise, M.T.D.; Parise, D.; Viana, M.V.C.; Nogueira, W.; Ramos, R.T.J.; Góes-Neto, A.; Azevedo, V.A.D.C.; Brenig, B.; et al. Genomic and Phenotypic Insight into Antimicrobial Resistance of Pseudomonas Fluorescens from King George Island, Antarctica. Front. Microbiol. 2025, 16. [Google Scholar] [CrossRef]
- Twomey, E.; Fields, D.; Begley, M. Investigating Soil Bacteria for Novel Antimicrobials Which Inhibit the Opportunistic Pathogen, Staphylococcus Epidermidis. Access Microbiol. 2019, 1, 778. [Google Scholar] [CrossRef]
- Zhang, X.; Ma, Z.; Hao, P.; Ji, S.; Gao, Y. Characteristics and Health Impacts of Bioaerosols in Animal Barns: A Comprehensive Study. Ecotoxicol. Env. Saf. 2024, 278, 116381. [Google Scholar] [CrossRef]
- Zhao, Y.; Li, L.; Huang, Y.; Xu, X.; Liu, Z.; Li, S.; Zhu, L.; Hu, B.; Zhang, T. Global Soil Antibiotic Resistance Genes Are Associated with Increasing Risk and Connectivity to Human Resistome. Nat. Commun. 2025, 16, 7141. [Google Scholar] [CrossRef]
- Svenning, M.M.; Wartiainen, I.; Hestnes, A.G.; Binnerup, S.J. Isolation of Methane Oxidising Bacteria from Soil by Use of a Soil Substrate Membrane System. FEMS Microbiol. Ecol. 2003, 44, 347–354. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.; Frossard, V.; Perga, M.-E.; Tofield-Pasche, N.; Hofmann, H.; Dubois, N.; Belkina, N.; Zobkova, M.; Robert, S.; Lyautey, E. Lateral Variations and Vertical Structure of the Microbial Methane Cycle in the Sediment of Lake Onego (Russia). Inland Waters 2019, 9, 205–226. [Google Scholar] [CrossRef]
- Deng, Y.; Che, R.; Wang, F.; Conrad, R.; Dumont, M.; Yun, J.; Wu, Y.; Hu, A.; Fang, J.; Xu, Z.; et al. Upland Soil Cluster Gamma Dominates Methanotrophic Communities in Upland Grassland Soils. Sci. Total Environ. 2019, 670, 826–836. [Google Scholar] [CrossRef]
- de Groot, T.R.; Engelmann, J.C.; Ramond, P.; Dorigo, J.; van Bleijswijk, J.; Niemann, H. Adaptation of Methane-Oxidizing Bacteria to Environmental Changes: Implications for Coastal Methane Dynamics. Biogeosciences 2025, 22, 5173–5191. [Google Scholar] [CrossRef]
- van Teeseling, M.C.F.; Pol, A.; Harhangi, H.R.; van der Zwart, S.; Jetten, M.S.M.; Op den Camp, H.J.M.; van Niftrik, L. Expanding the Verrucomicrobial Methanotrophic World: Description of Three Novel Species of Methylacidimicrobium Gen. Nov. Applied Environ. Microbiol.> 2014, 80, 6782–6791. [Google Scholar] [CrossRef]
- Picone, N.; Mohammadi, S.S.; Waajen, A.C.; van Alen, T.A.; Jetten, M.S.M.; Pol, A.; Op den Camp, H.J.M. More Than a Methanotroph: A Broader Substrate Spectrum for Methylacidiphilum Fumariolicum SolV. Front. Microbiol. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Hou, S.; Makarova, K.S.; Saw, J.H.; Senin, P.; Ly, B.V.; Zhou, Z.; Ren, Y.; Wang, J.; Galperin, M.Y.; Omelchenko, M.V.; et al. Complete Genome Sequence of the Extremely Acidophilic Methanotroph Isolate V4, Methylacidiphilum Infernorum, a Representative of the Bacterial Phylum Verrucomicrobia. Biol. Direct 2008, 3, 26. [Google Scholar] [CrossRef]
- Ratnadevi, C.M.; Erikstad, H.-A.; Kruse, T.; Birkeland, N.-K. Methylacidiphilum Kamchatkense Gen. Nov., Sp. Nov., an Extremely Acidophilic and Moderately Thermophilic Methanotroph Belonging to the Phylum Verrucomicrobiota. Int. J. Syst. Evol. Microbiol. 2023, 73, 006060. [Google Scholar] [CrossRef] [PubMed]
- Smith, G.J.; Angle, J.C.; Solden, L.M.; Borton, M.A.; Morin, T.H.; Daly, R.A.; Johnston, M.D.; Stefanik, K.C.; Wolfe, R.; Gil, B.; et al. Members of the Genus Methylobacter Are Inferred To Account for the Majority of Aerobic Methane Oxidation in Oxic Soils from a Freshwater Wetland. mBio 2018, 9. [Google Scholar] [CrossRef]
- Feisthauer, S.; Vogt, C.; Modrzynski, J.; Szlenkier, M.; Krüger, M.; Siegert, M.; Richnow, H.-H. Different Types of Methane Monooxygenases Produce Similar Carbon and Hydrogen Isotope Fractionation Patterns during Methane Oxidation. Geochim. Et. Cosmochim. Acta 2011, 75, 1173–1184. [Google Scholar] [CrossRef]
- Delherbe, N.A.; Pearce, D.; But, S.Y.; Murrell, J.C.; Khmelenina, V.N.; Kalyuzhnaya, M.G. Genomic Insights into Moderately Thermophilic Methanotrophs of the Genus Methylocaldum. Microorganisms 2024, 12, 469. [Google Scholar] [CrossRef]
- Schmider, T.; Hestnes, A.G.; Brzykcy, J.; Schmidt, H.; Schintlmeister, A.; Roller, B.R.K.; Teran, E.J.; Söllinger, A.; Schmidt, O.; Polz, M.F.; et al. Physiological Basis for Atmospheric Methane Oxidation and Methanotrophic Growth on Air. Nat. Commun. 2024, 15, 4151. [Google Scholar] [CrossRef]
- Peterse, I.F.; Pol, A.; Cremers, G.; Berben, T.; van Alen, T.A.; Op den Camp, H.J.M.; Veraart, A.J.; Lücker, S. Hydrogen Oxidation Benefits Alphaproteobacterial Methanotrophs Under Severe Methane Limitation. Env. Microbiol. 2025, 27, e70163. [Google Scholar] [CrossRef] [PubMed]
- Im, J.; Lee, S.-W.; Yoon, S.; DiSpirito, A.A.; Semrau, J.D. Characterization of a Novel Facultative Methylocystis Species Capable of Growth on Methane, Acetate and Ethanol. Environ. Microbiol. Rep. 2011, 3, 174–181. [Google Scholar] [CrossRef]
- Bordel, S.; Rodríguez, Y.; Hakobyan, A.; Rodríguez, E.; Lebrero, R.; Muñoz, R. Genome Scale Metabolic Modeling Reveals the Metabolic Potential of Three Type II Methanotrophs of the Genus Methylocystis. Metab. Eng. 2019, 54, 191–199. [Google Scholar] [CrossRef] [PubMed]
- del Cerro, C.; García, J.M.; Rojas, A.; Tortajada, M.; Ramón, D.; Galán, B.; Prieto, M.A.; García, J.L. Genome Sequence of the Methanotrophic Poly-β-Hydroxybutyrate Producer Methylocystis Parvus OBBP. J. Bacteriol. 2012, 194, 5709–5710. [Google Scholar] [CrossRef]
- Cáceres, M.; Gentina, J.C.; Aroca, G. Oxidation of Methane by Methylomicrobium Album and Methylocystis Sp. in the Presence of H2S and NH3. Biotechnol. Lett. 2014, 36, 69–74. [Google Scholar] [CrossRef]
- Akberdin, I.R.; Thompson, M.; Hamilton, R.; Desai, N.; Alexander, D.; Henard, C.A.; Guarnieri, M.T.; Kalyuzhnaya, M.G. Methane Utilization in Methylomicrobium Alcaliphilum 20ZR: A Systems Approach. Sci. Rep. 2018, 8, 2512. [Google Scholar] [CrossRef]
- Stone, K.; Hilliard, M.; Badr, K.; Bradford, A.; He, Q.P.; Wang, J. Comparative Study of Oxygen-Limited and Methane-Limited Growth Phenotypes of Methylomicrobium Buryatense 5GB1. Biochem. Eng. J. 2020, 161, 107707. [Google Scholar] [CrossRef]
- Sly, L.I.; Bryant, L.J.; Cox, J.M.; Anderson, J.M. Development of a Biofilter for the Removal of Methane from Coal Mine Ventilation Atmospheres. Appl. Microbiol. Biotechnol. 1993, 39, 400–404. [Google Scholar] [CrossRef]
- Sang, Y.; Hao, Q.; Zhang, Y.; Wang, O.; Zheng, S.; Liu, F. Simultaneous Removal of Methane and High Nitrite from the Wastewater by Methylomonas Sp. with Soluble Methane Monooxygenase. Bioresour. Technol. 2025, 418, 131972. [Google Scholar] [CrossRef]
- Park, Y.R.; Yoo, H.S.; Song, M.Y.; Lee, D.-H.; Lee, S.J. Biocatalytic Oxidations of Substrates through Soluble Methane Monooxygenase from Methylosinus Sporium 5. Catalysts 2018, 8, 582. [Google Scholar] [CrossRef]
- Poshvina, D.V.; Balkin, A.S.; Teslya, A.V.; Dilbaryan, D.S.; Stepanov, A.A.; Kravchenko, S.V.; Vasilchenko, A.S. Structural and Functional Differences in the Bacterial Community of Chernozem Soil Under Conventional and Organic Farming Conditions. Agriculture 2024, 14, 2127. [Google Scholar] [CrossRef]
- Khrapai, E.S.; Kolesnikov, S.I.; Kuzina, A.A.; Kazeev, K.Sh.; Minnikova, T.V.; Demin, K.A.; Kocharovskaya, Yu.N.; Delegan, Ya.A.; Bogun, A.G.; Minkina, T.M.; et al. Metagenomic Profiling of the Soil Microbiological Community in the Impact Area of the Urup Mining and Processing Plant Tailings (Karachay-Cherkess Republic). Eurasian Soil. Sc. 2025, 58, 119. [Google Scholar] [CrossRef]
- Wang, C.; Lin, W.; Jia, S.; Chen, S.; Xiong, D.; Xu, C.; Yang, Z.; Liu, X.; Yang, Y. Effects of Litter and Root Inputs on Soil Microbial Community Structure in Subtropical Natural and Plantation Forests. Plant Soil. 2025, 513, 823–838. [Google Scholar] [CrossRef]
- Yao, R.; Gao, Q.; Liu, Y.; Li, H.; Yang, J.; Bai, Y.; Zhu, H.; Wang, X.; Xie, W.; Zhang, X. Deep Vertical Rotary Tillage Mitigates Salinization Hazards and Shifts Microbial Community Structure in Salt-Affected Anthropogenic-Alluvial Soil. Soil. Tillage Res. 2023, 227, 105627. [Google Scholar] [CrossRef]
- Liang, X.; Zhong, D.; Zhang, C.; Pan, Y.; Zhang, C.; Guo, H.; Zhu, X.; Li, X.; He, Y.; Huang, S.; et al. Effects of Different Cultivation Substrates on the Growth of Podocarpus Macrophyllus and the Rhizosphere Soil Microbial Community Structure. Agronomy 2025, 15, 1055. [Google Scholar] [CrossRef]
- Bautista, D.; García, D.; Dávila, L.; Caro-Quintero, A.; Cotes, A.M.; González, A.; Zuluaga, A.P. Studying the Microbiome of Suppressive Soils against Vascular Wilt, Caused by Fusarium Oxysporum in Cape Gooseberry (Physalis Peruviana). [CrossRef]
- Sun, Y.; Ji, L.; Chang, J.; Li, Y.; Wang, H.; Lu, D.; Tian, C. Compositional and Functional Succession of Soil Bacterial Communities during Long-Term Rice Cultivation on Saline-Alkali Soils: Insights Derived from a New Perspective on Core Bacterial Taxa. Pedosphere 2025, 35, 641–654. [Google Scholar] [CrossRef]
- Li, Y.; Gu, M.; Xu, W.; Zhu, J.; Chu, M.; Tang, Q.; Yi, Y.; Zhang, L.; Li, P.; Zhang, Y.; et al. Whole-Genome Analysis of Halomonas Sp. H5 Revealed Multiple Functional Genes Relevant to Tomato Growth Promotion, Plant Salt Tolerance, and Rhizosphere Soil Microecology Regulation. Microorganisms 2025, 13, 1781. [Google Scholar] [CrossRef]
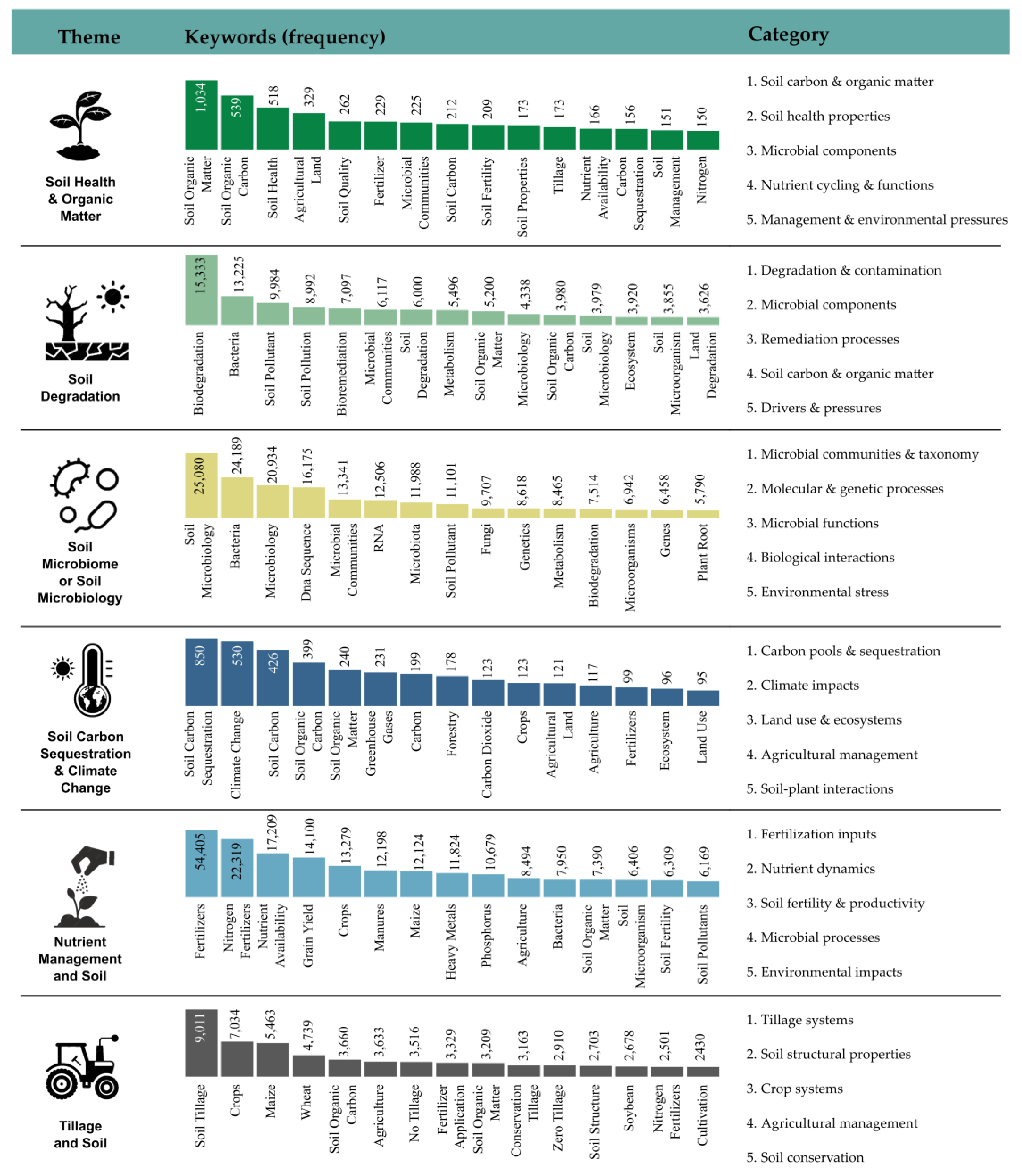
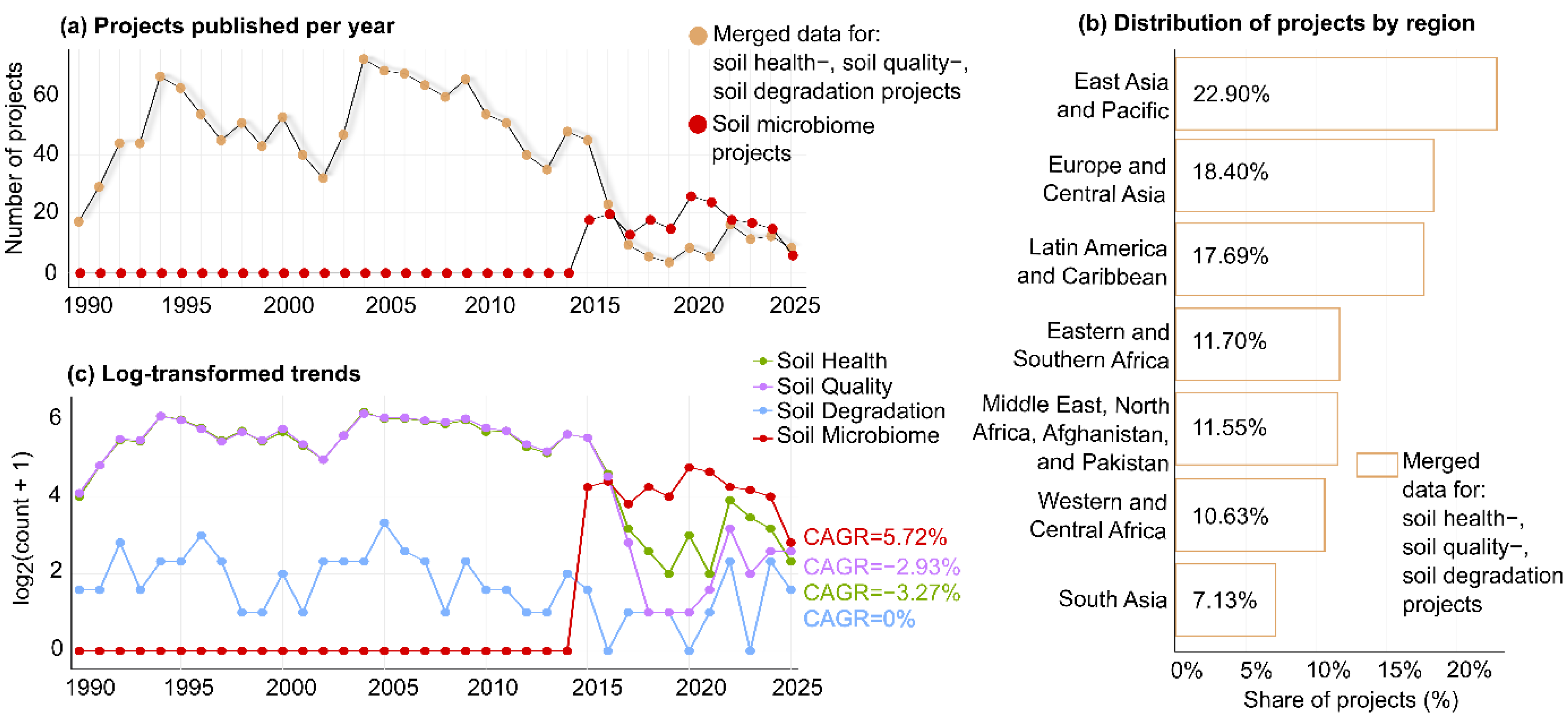
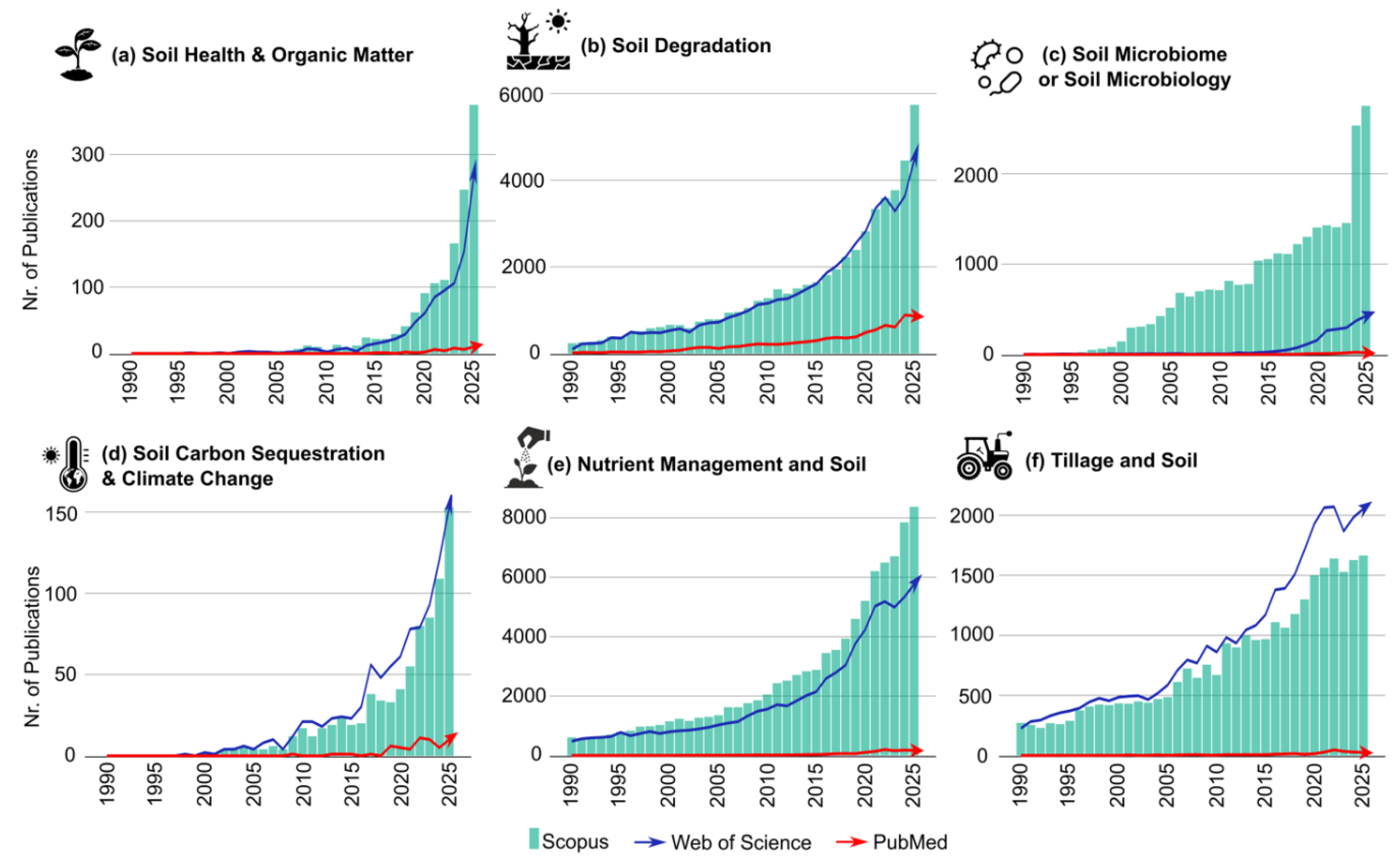

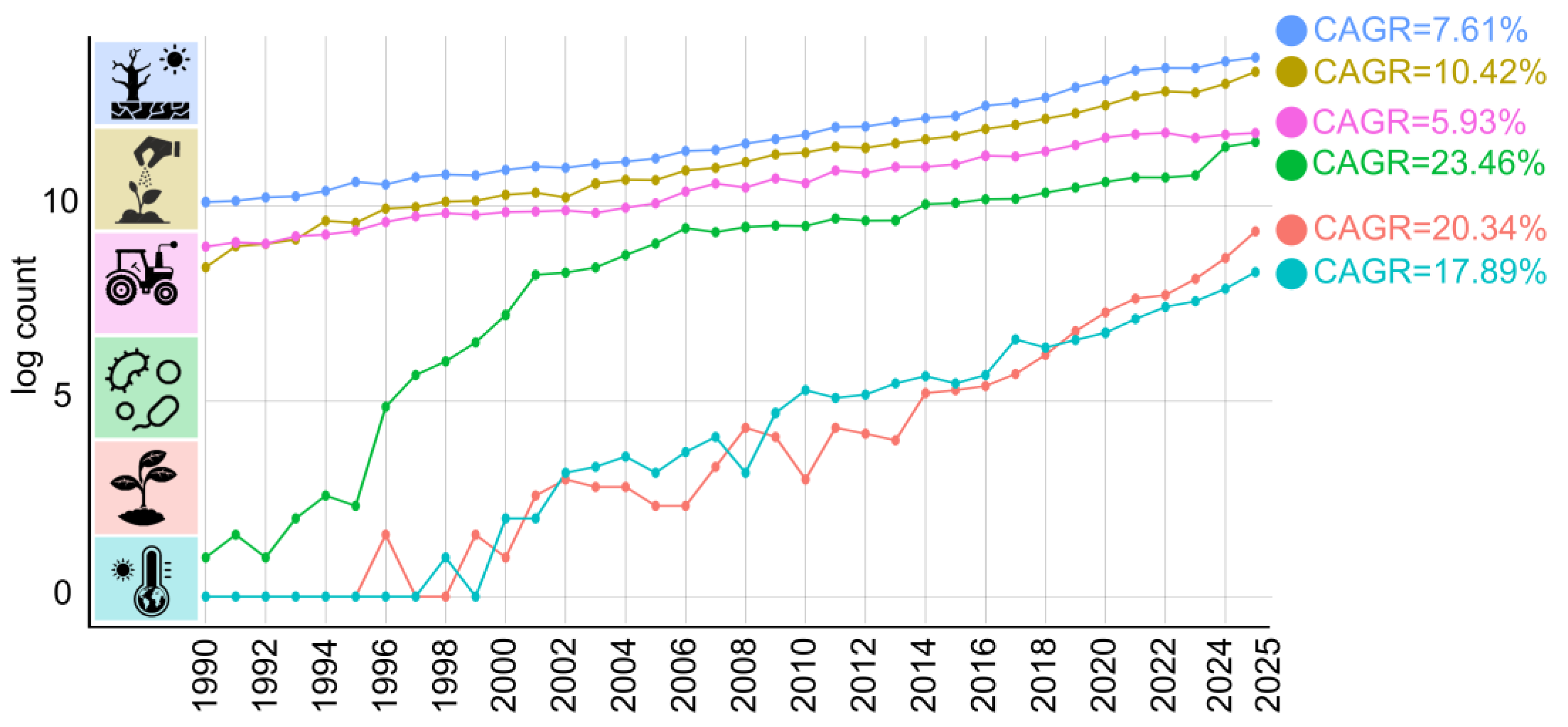
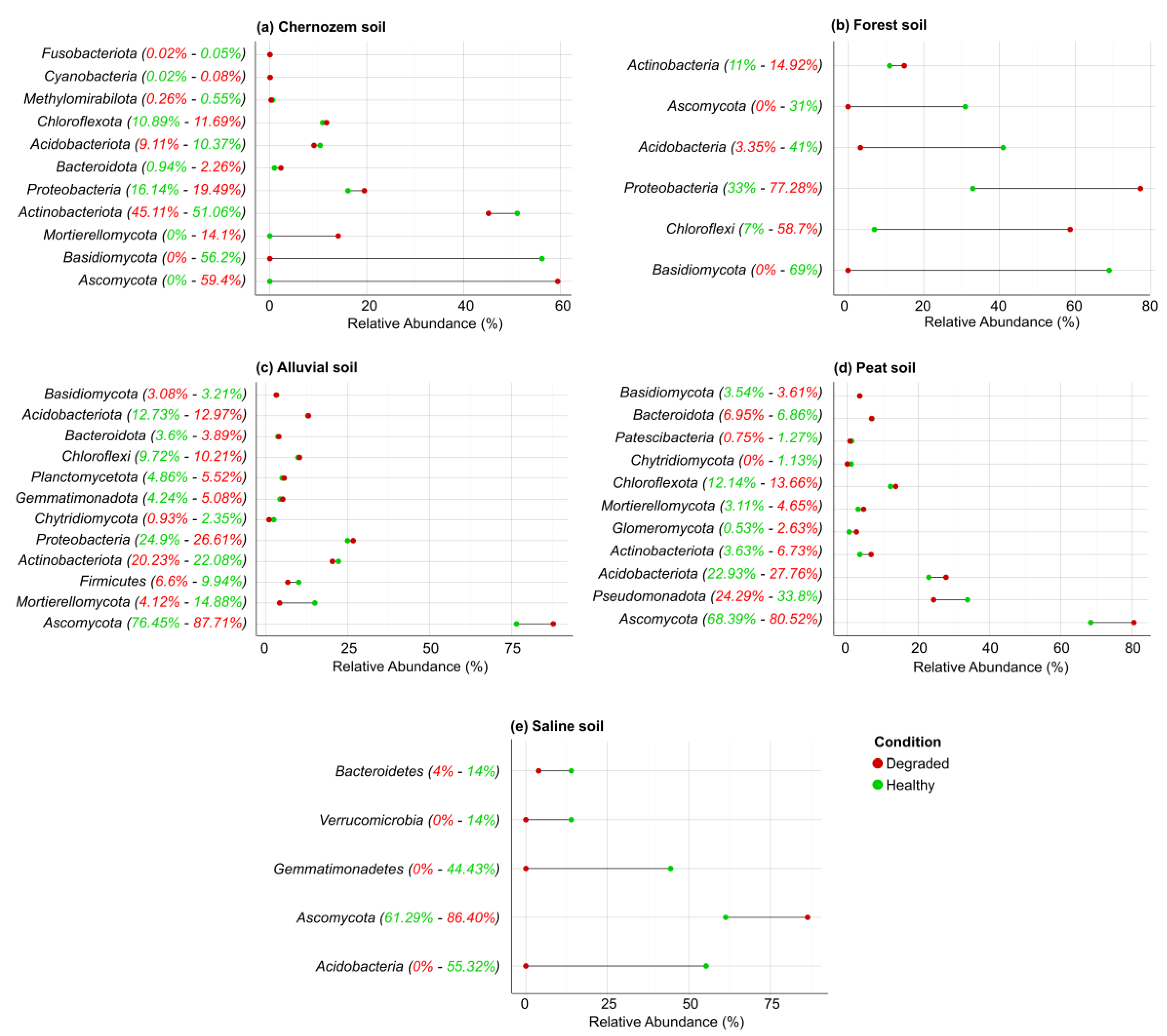
| Method category | Method | Research question | Key strengths | Key limitations | References |
|---|---|---|---|---|---|
| Classical & biomass-based methods | Culture-based methods (CFU counts) | How many viable, culturable microorganisms are present? | Simple; enables strain isolation | <1 of microbes are culturable | [60,61] |
| Microbial biomass C/N (fumigation–extraction) | How much living microbial biomass is present in soil? | Integrative estimate of living microbial biomass | No taxonomic or functional resolution | [62,63] | |
| Phospholipid fatty acid (PLFA) analysis | How large is the living microbial community and what are its major groups? | Quantitative; functional group resolution | Low taxonomic specificity | [64,65] | |
| Molecular & sequencing methods | Metabarcoding / targeted amplicon sequencing (16S rRNA, ITS) | Who is there? | High throughput; widely comparable; cost-effective | Provides relative, not absolute abundance; limited functional insight | [66,67] |
| Shotgun metagenomics | What can they do (genetic potential)? | Unbiased, stable DNA; broad functional coverage | High cost; complex data analysis, does not indicate activity | [68,69] | |
| Metatranscriptomics | What genes are being expressed? | Reflects real-time microbial activity | RNA instability; high technical complexity | [69,70] | |
| Metaproteomics | What proteins are produced? | Direct evidence of functional expression | Protein extraction, interference from humic substances, identification of low-abundance proteins | [71,72] | |
| Metabolomics | What are the biochemical outputs? | Captures final functional outcomes | Difficult compound identification | [73,74] | |
| Emerging & field-deployable methods | Biosensors (aptamer-, enzyme-, or microbe-based) | What specific process or compound is active now? | Real-time, in situ monitoring | Still under development | [75,76] |
| Portable qPCR / lab-on-chip systems | How much of a specific gene or pathogen is present? | Rapid field diagnostics | Limited multiplexing | [77,78] | |
| Soil health test kits (integrated indicators) | Is the soil generally functioning well? | Farmer-friendly | Low microbial specificity | [79,80] |
| Function* | Indicator | Degraded soil | Healthy soil |
|---|---|---|---|
| Nitrogen fixation | CFU [81,82] | 1.1×102 | 3.88 × 106 |
| nifH gene [83,84] | 2.88 × 10⁵ copy/ g of dry soil | 1.0–4.6 × 107 copies/ g of soil | |
| Relative abundance [85] | No data available | 21.5% | |
| Representative species (Supplementary Table S2) | [86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116] | ||
| Phosphorus solubilization | CFU [117,118] | 10-2 | 4 × 108 |
| phoD gene [119] | 1.33 × 105 copies/ g of dry soil | 5.80 × 107 copy/ g of dry soil | |
| gcd gene [119,120] | 1.46 × 104 copy/ g of dry soil | 2.19 × 108 copies/ g of soil | |
| Relative abundance [121] | 0.18% | 13.13% | |
| Representative species (Supplementary Table S2) | [109,122,123,124,125,126,127,128,129,130,131,132,133,134,135,135,136,137,138,139,140,141,142,143,144,145,146,147,148,149,150,151,152,153,154,155,156,157,158,159,160,161,162,163,164,165,166,167,168,169,170,171] | ||
| Potassium solubilization | CFU [172,173] | 4 × 10² | 75 × 104 |
| pqqC gene [174] | 3.25 × 104 copies/ g of dry soil | 1.44 × 109 copies/ g of dry soil | |
| Relative abundance [175] | 5.46% | 12.25% | |
| Representative species (Supplementary Table S2) | [155,176,177,178,179,180,181,182,183,184,185,186,187,187,188,189,190,191,192,193,194] | ||
| Phytohormone production | CFU [195,196] | 1 × 105 | 1.5×105 |
| acdS gene [197] | 1.0 × 101.91 copies/ g of soil | 1.0 × 10¹² copies/ g of soil | |
| Relative abundance [198,199] | 7% | 80% | |
| Representative species (Supplementary Table S2) | [94,200,201,202] | ||
| Organic matter decomposition | CFU [203] | 6.6 × 103 | 1.5 × 104 |
| chiA gene [204,205] | 1.70 × 105 copies/ g of dry soil | 3.5 × 108 copies/ g of dry soil | |
| cbhI gene [206,207] | 0.17 × 104 copies/ g of soil | 7.2 × 104 copies/ g of soil | |
| GH48 gene [206,207] | 1.04 × 103 copies/ g of soil | 5.96 × 105 copies/ g of soil | |
| Relative abundance | No data available | No data available | |
| Representative species (Supplementary Table S2) | [208,209,210,211,212,213,214,214,215,216,217,217,217,218,219,220,221,222,223,224,225,226,227,228,229,230,231,232,233,234,235,236] | ||
| Siderophore production | CFU [237] | 2.6 × 10² | 2 × 10⁶ |
| PKS type I [238] | 5.77 × 102 copies/ g of dry weight of soil | 9.39 × 106 copies/ g of dry weight of soil | |
| Relative abundance [240] | 47% | 85% | |
| Representative species (Supplementary Table S2) | [241,242,243,244,245,246,247] | ||
| Mycorrhizal symbiosis (Nutrient uptake enhancement) | CFU [248] | 3.0 × 106 | 1.18 × 108 |
| AMF-specific 18S rRNA gene [249] | 1.2 × 10⁶ copies/ g of soil | 2.1 × 10⁸ copies/ g of soil | |
| Relative abundance [250] | 0.06% | 98.16% | |
| Representative species (Supplementary Table S2) | [119,251,251,252,253,254,254,255,255,256,257,257,258,259,260,261,262,263,264,265,266,267,268,269,270,271,272,273,274,275,276,277,278,279,280,281,282,283,284,285,286,287] | ||
| Plant growth promotion | CFU [288] | 105 copies/ g of soil | 5.9×107 copies/ g of soil |
| phoD gene [119] | 1.33 × 105 copies/ g of dry soil | 5.80 × 107 copies/ g of dry soil | |
| gcd gene [119,120] | 1.46 × 104 copies/ g of dry soil | 2.19 × 108 copies/ g of dry soil | |
| acdS gene [197,289] | 8.1 × 10¹ copies/ g of soil | 1 × 10¹² copies/ g of soil | |
| Relative abundance | No data available | No data available | |
| Representative species (Supplementary Table S2) | [94,108,109,139,229,290,291,292,293,294,295,296,297,298,299,300,301,302,303,304,305] | ||
| Bioremediation | CFU [306,307] | 103 | 7.32 × 106 |
| nahAc gene [308] | 5 x 103 copies/ g of soil | 107 copies/ g of soil | |
| arsC gene [309] | 0.88 x 104 copies/ng total DNA | 1.56 x 105 copies/ng total DNA | |
| arsM gene [310] | 0.4 × 107 copies/ g of dry soil | 2.3 × 107 copies/ g of dry soil | |
| Relative abundance [311] | 8.63% | 0.01% | |
| Representative species (Supplementary Table S2) | [208,312,313,314,315] | ||
| Hydrocarbon degradation | CFU [316] | 373 ± 56 × 103 | 8 ± 2 × 103 |
| alkB gene [317,318] | 5.0 × 108 cells/g-soil | 0.9 × 104 copies per nanogram of soil | |
| Nah gene [319,320] | 1.1 × 108 copies/ g of dry mass soil | Not detectable | |
| Relative abundance [321] | 10% | 1% | |
| Representative species (Supplementary Table S2) | [322,323,324,325,326,327,328,329,330,331,332,333,334,335,336,337,338,339,340,341,342,343,344,345,346,347,348,349,350,351] | ||
| Antibiotic resistance | CFU [352] | > 2.0 × 10⁴ | 6 |
| sull gene [353] | 4.37 × 107 | 2.12 × 103 | |
| erm(B) [353] | 9.57 × 108 | Not detectable | |
| intI1 gene [353] | 4.12 × 107 | 3.17 × 103 | |
| Relative abundance [354] | 74% | 3% | |
| Representative species (Supplementary Table S2) | [185,315,355,356,357,358,359,360,361,362,363,364,365,366,367] | ||
| Methane oxidizers | CFU [368] | 1 × 10² | 1 × 109 |
| pmoA gene [369,370] | 103 | 1.1 × 108 | |
| Relative abundance [286,371] | 0% | 57% | |
| Representative species (Supplementary Table S2) | [372,373,374,375,376,377,377,378,379,379,380,381,382,383,384,385,386,387,388,388,389] | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




