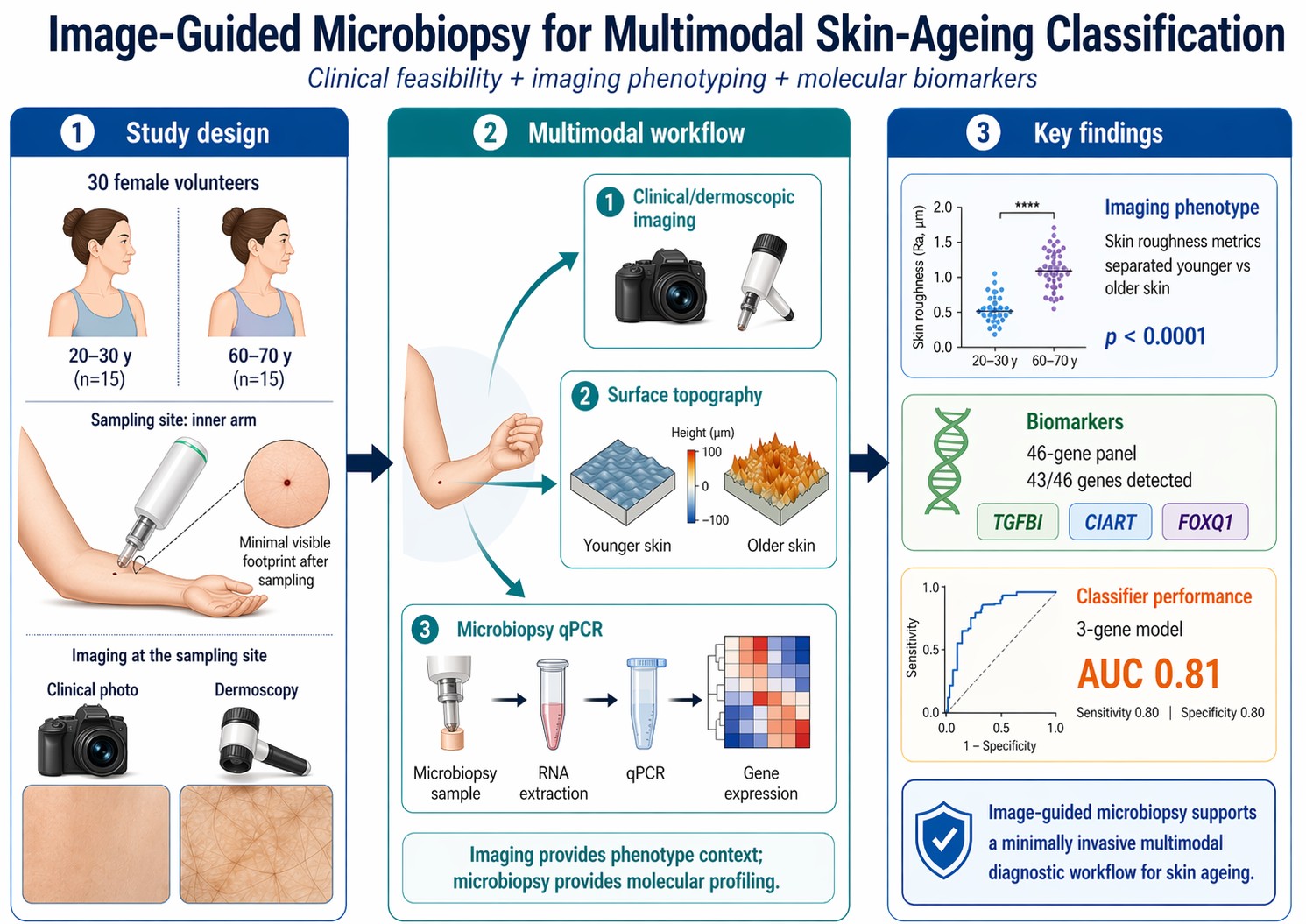
Background/Objectives: Microbiopsy enables minimally invasive molecular sampling of skin, but diagnostic use will likely depend on integrating molecular readouts with imaging-based phenotype context. This manuscript frames the supplied Beiersdorf–UniSA/Prow volunteer study as a proof-of-concept diagnostic workflow for skin ageing and removes the ancillary shipping experiment to focus on clinical feasibility and classifier performance. Methods: Thirty volunteers were recruited into younger (20–30 years, n=15) and older (60–70 years, n=15) cohorts. Clinical photographs, dermoscopic images, skin-mold impressions, and inner-arm microbiopsy samples were collected. A 46-gene TaqMan qPCR panel was analyzed, with candidate biomarkers assessed for detectability, between-group separation, intra- versus interindividual variation, and receiver operating characteristic (ROC) performance. The manuscript was reconstructed from the supplied presentation and should therefore be regarded as a submission-ready draft derived from the available source material. Results: Pre- and post-procedure images showed only small punctate marks after sampling, supporting clinical feasibility of repeated, image-guided microbiopsy. Skin-surface roughness measures derived from mold imaging separated the younger and older cohorts on both displayed metrics (p<0.0001). Of 46 assayed genes, 43 were detected in at least one volunteer sample. TGFBI, CIART, and FOXQ1 were highlighted as the strongest age-associated markers. Single-gene ROC analysis ranked TGFBI (AUC 0.76), CIART (AUC 0.75), and FOXQ1 (AUC 0.73) highest. A three-gene model combining TGFBI, CIART, and FOXQ1 reached AUC 0.81 with sensitivity, specificity, positive predictive value, and negative predictive value all reported as 0.80. An exploratory four-gene model reached AUC 0.84, specificity 0.93, sensitivity 0.80, negative predictive value 0.82, and positive predictive value 0.92; the identity of the fourth marker should be confirmed from