Submitted:
28 April 2026
Posted:
29 April 2026
You are already at the latest version
Abstract
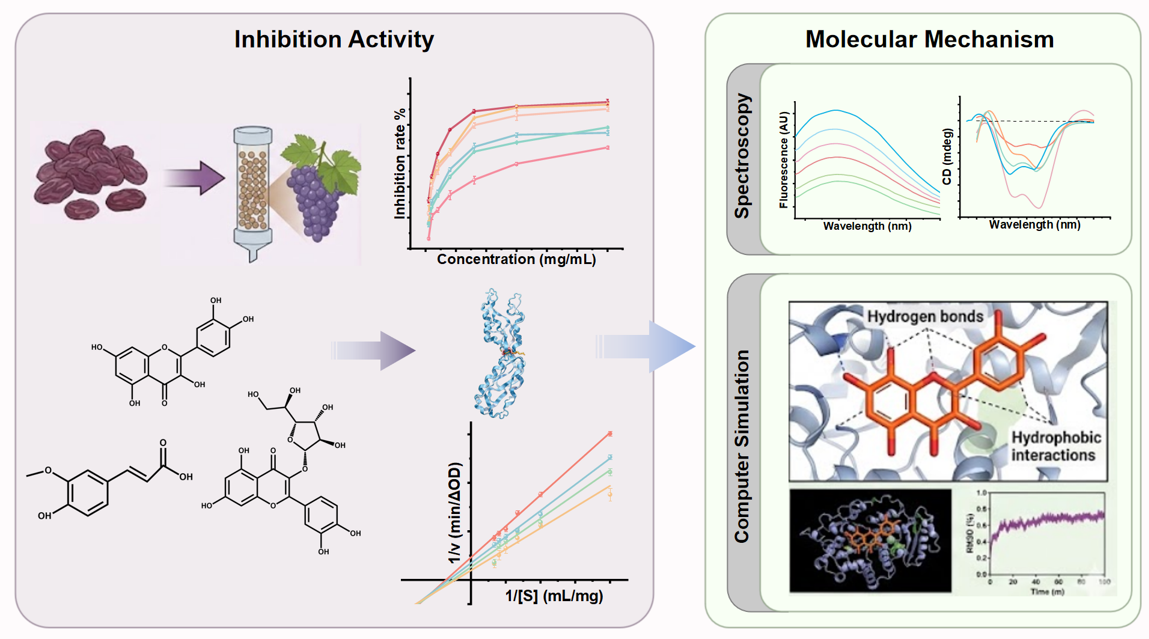
Keywords:
1. Introduction
2. Materials and Methods
2.1. Chemicals and Materials
2.2. Adsorption Kinetic Curves and Fitting Models
2.3. Adsorption Isotherms and Fitting Models
2.4. Dynamic Adsorption and Desorption Curves
2.5. UHPLC-MS/MS Identification of Raisin Polyphenols
2.6. Assessment of Antioxidant Capacity of Raisin Polyphenols
2.7. Inhibitory Capacity of Raisin Polyphenols toward PPA
2.8. Kinetic Analysis of α-Amylase Inhibition
2.9. Fluorescence Quenching Assay of α-Amylase
2.10. CD Analysis
2.11. Molecular Docking
2.12. MD Simulations
3. Results and Discussion
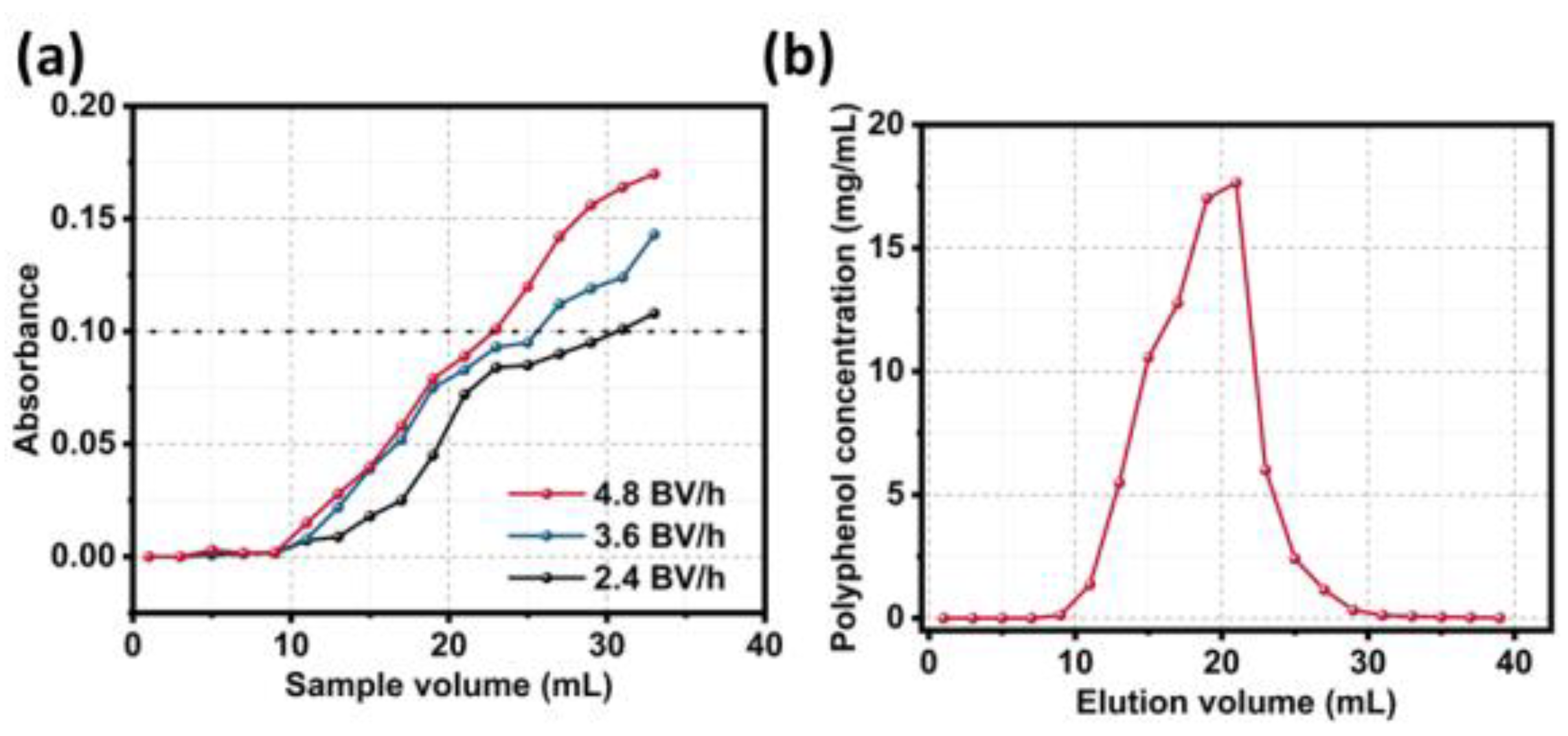
3.1. Screening of Macroporous Resins and Process Optimization
3.2. Adsorption Kinetics and Fitting Models
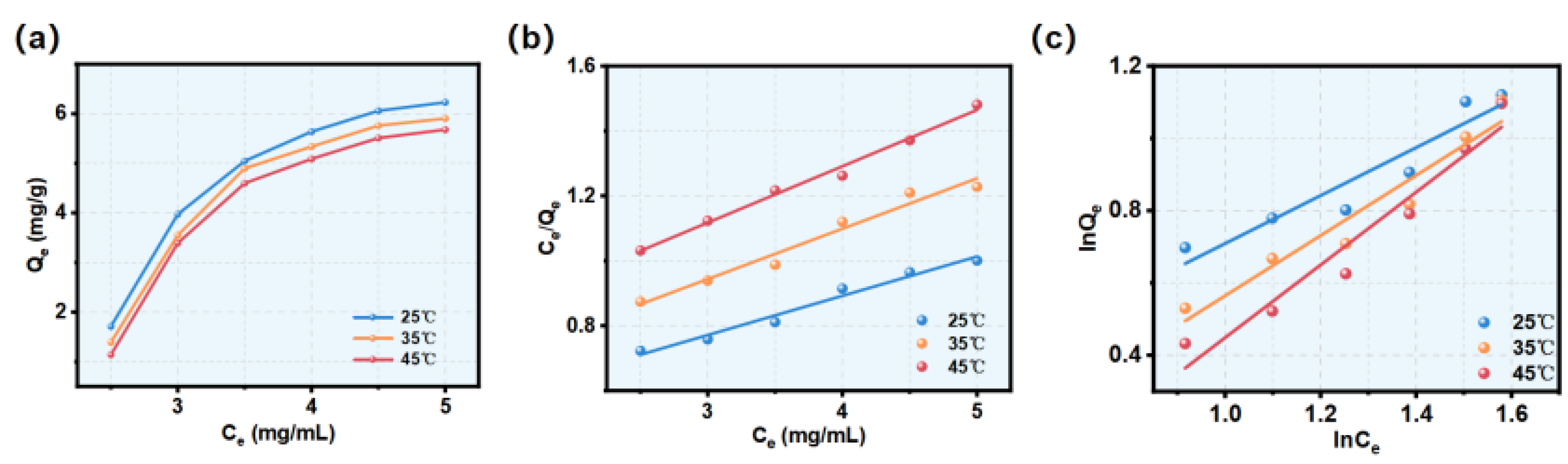
3.3. Adsorption Isotherms
3.4. Dynamic Adsorption and Desorption Curves
3.5. Identification of Polyphenols in Dried Apricots by UHPLC-QTOF-MS/MS
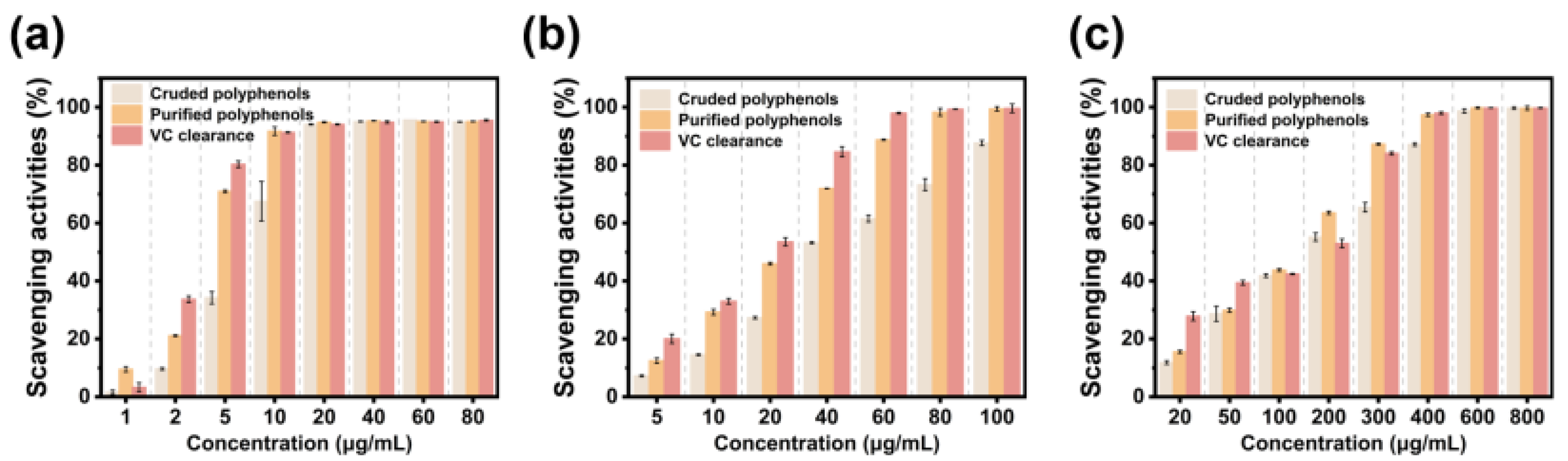
3.6. Evaluation of the Antioxidant Capacity of Raisin Polyphenols
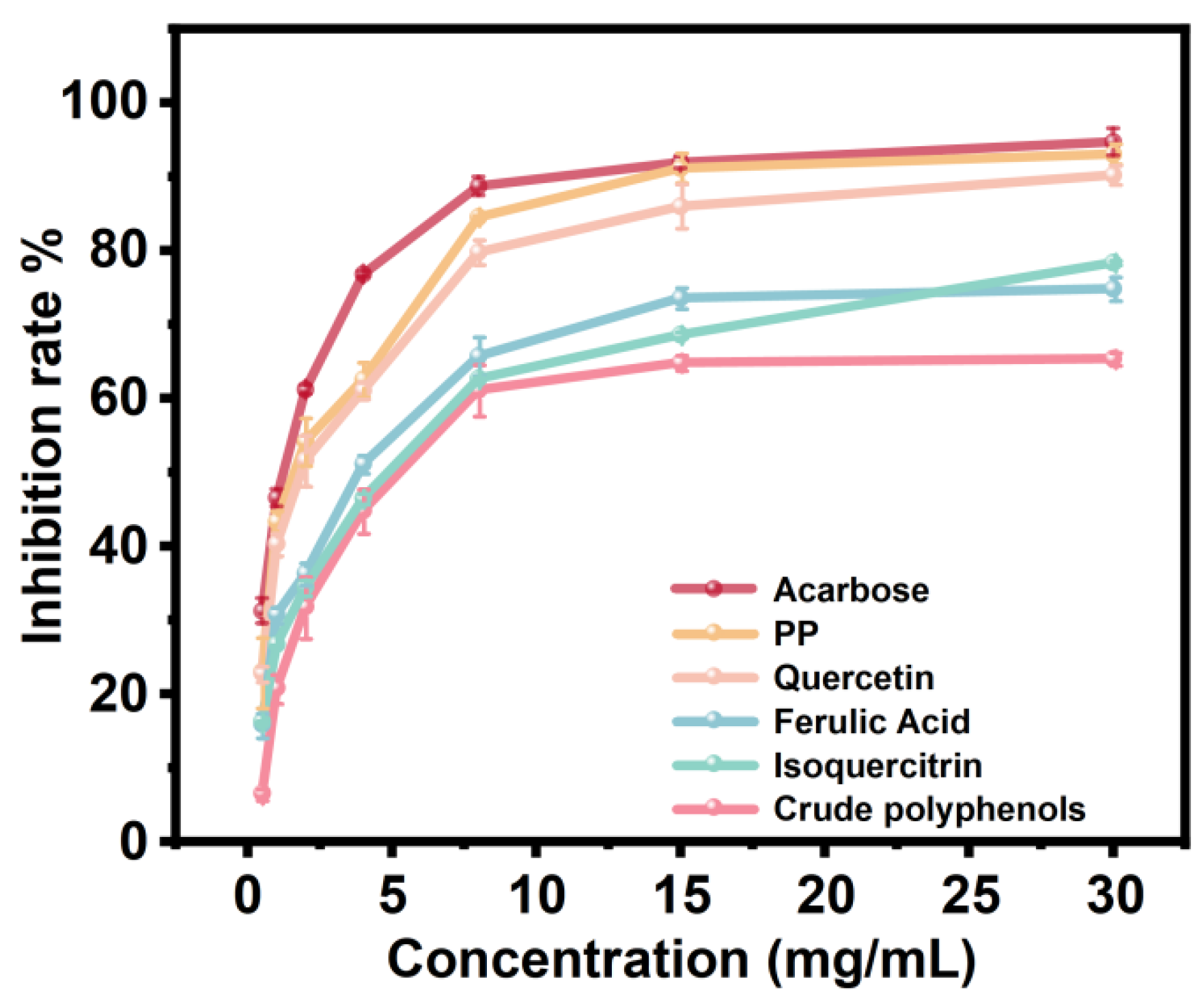
3.7. Inhibitory Activity of Raisin Polyphenols against α-Amylase
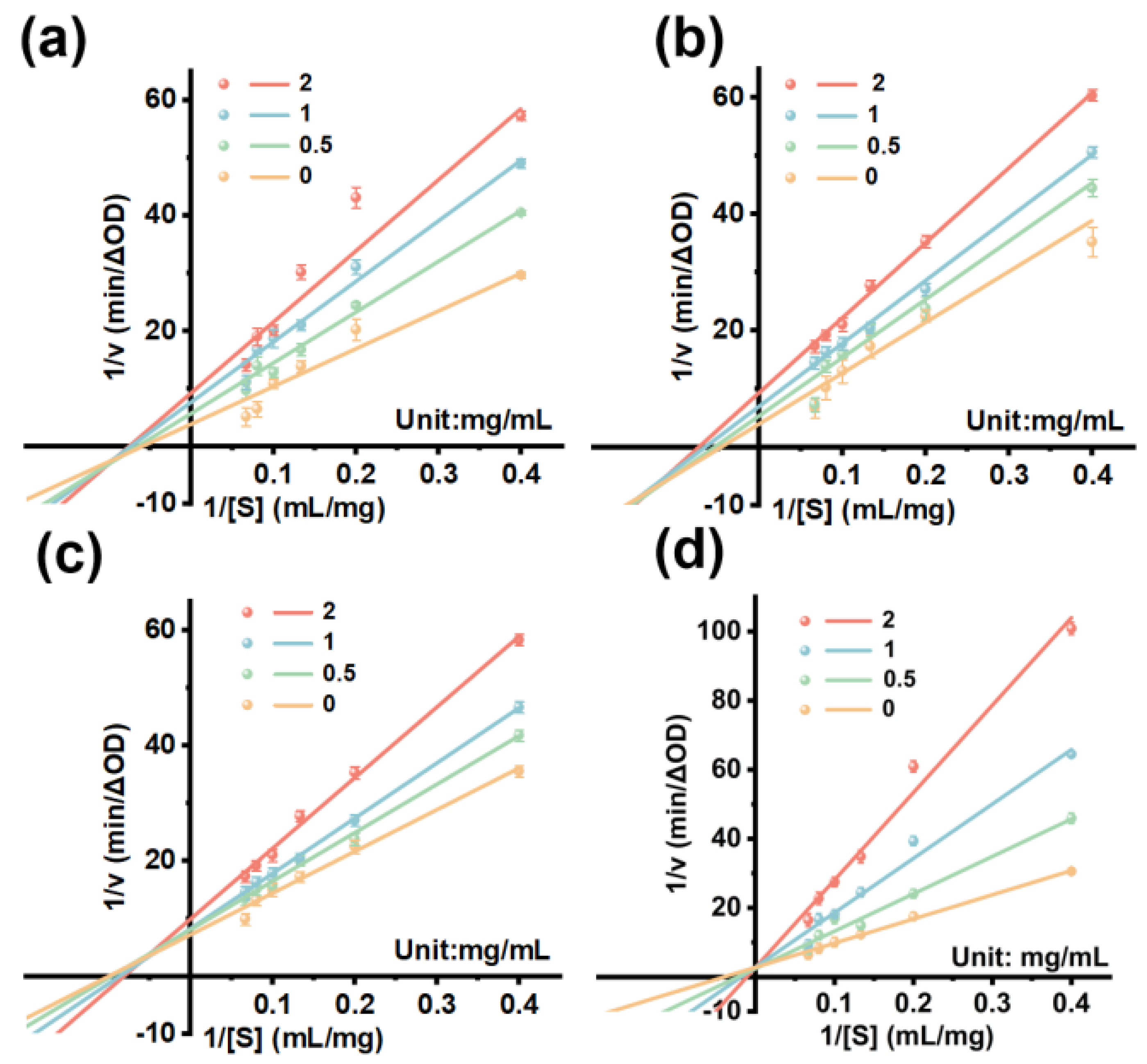
3.8. Kinetic Analysis of Enzyme Inhibition
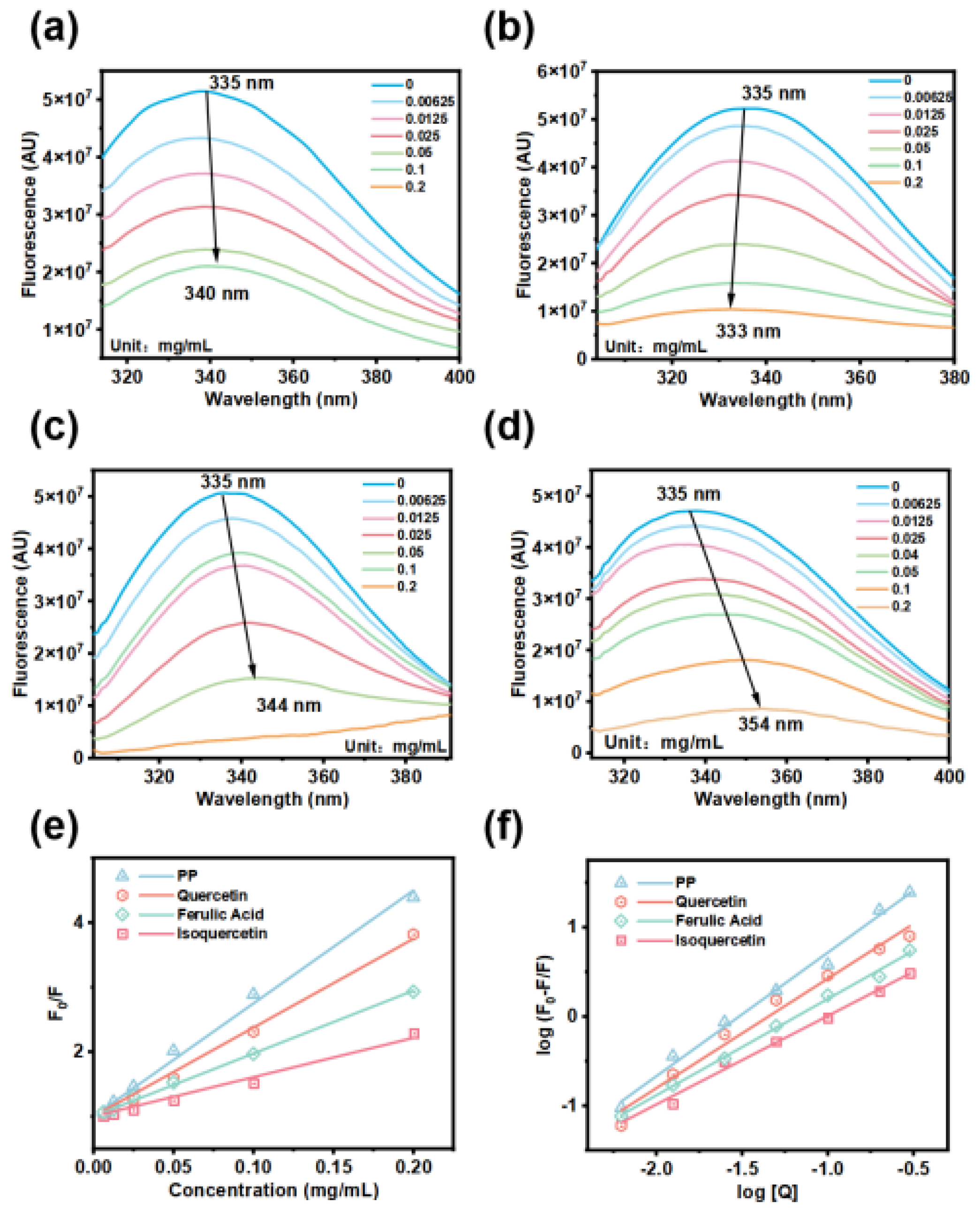
3.9. Mechanisms of Fluorescence Quenching
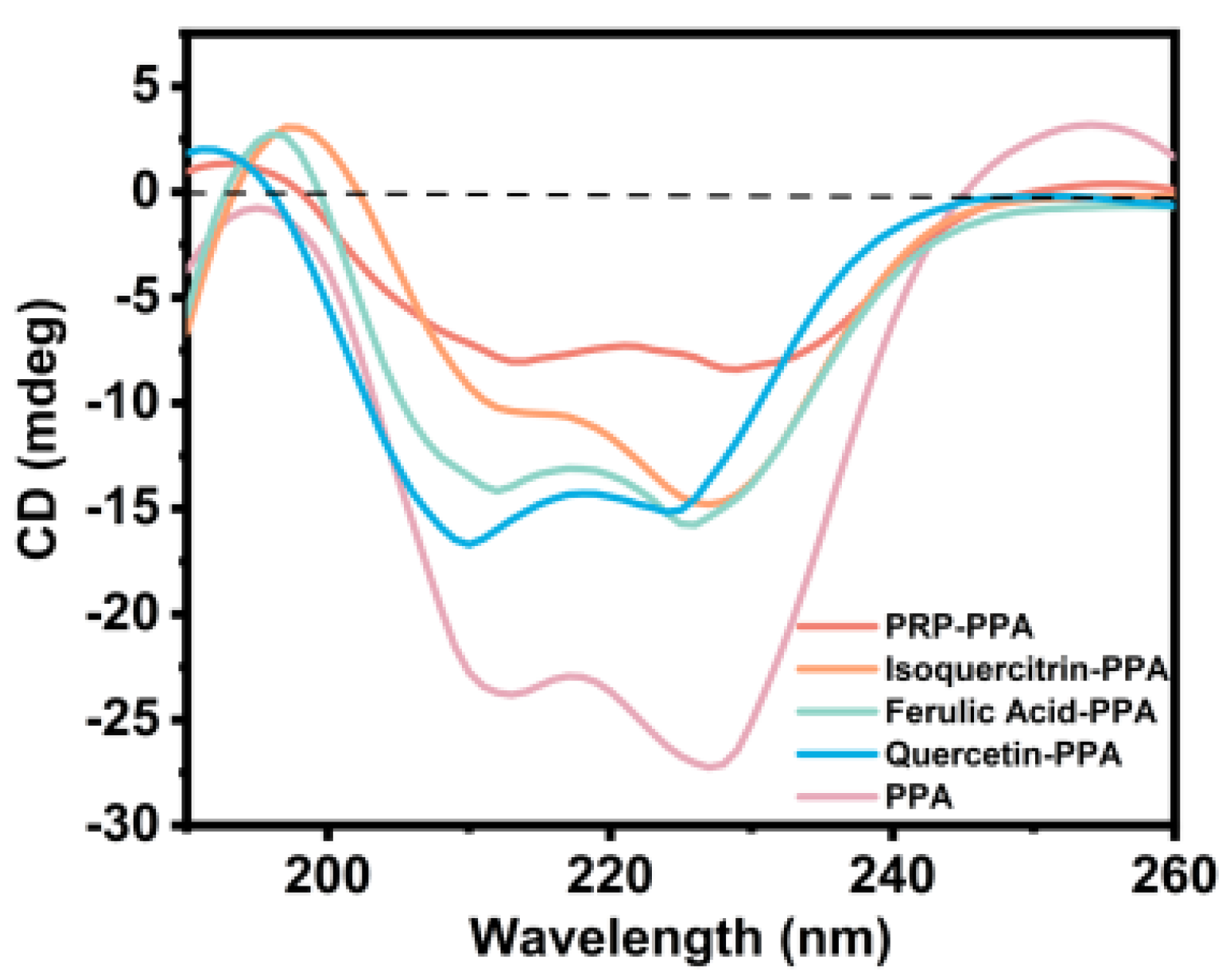
3.10. CD Analysis
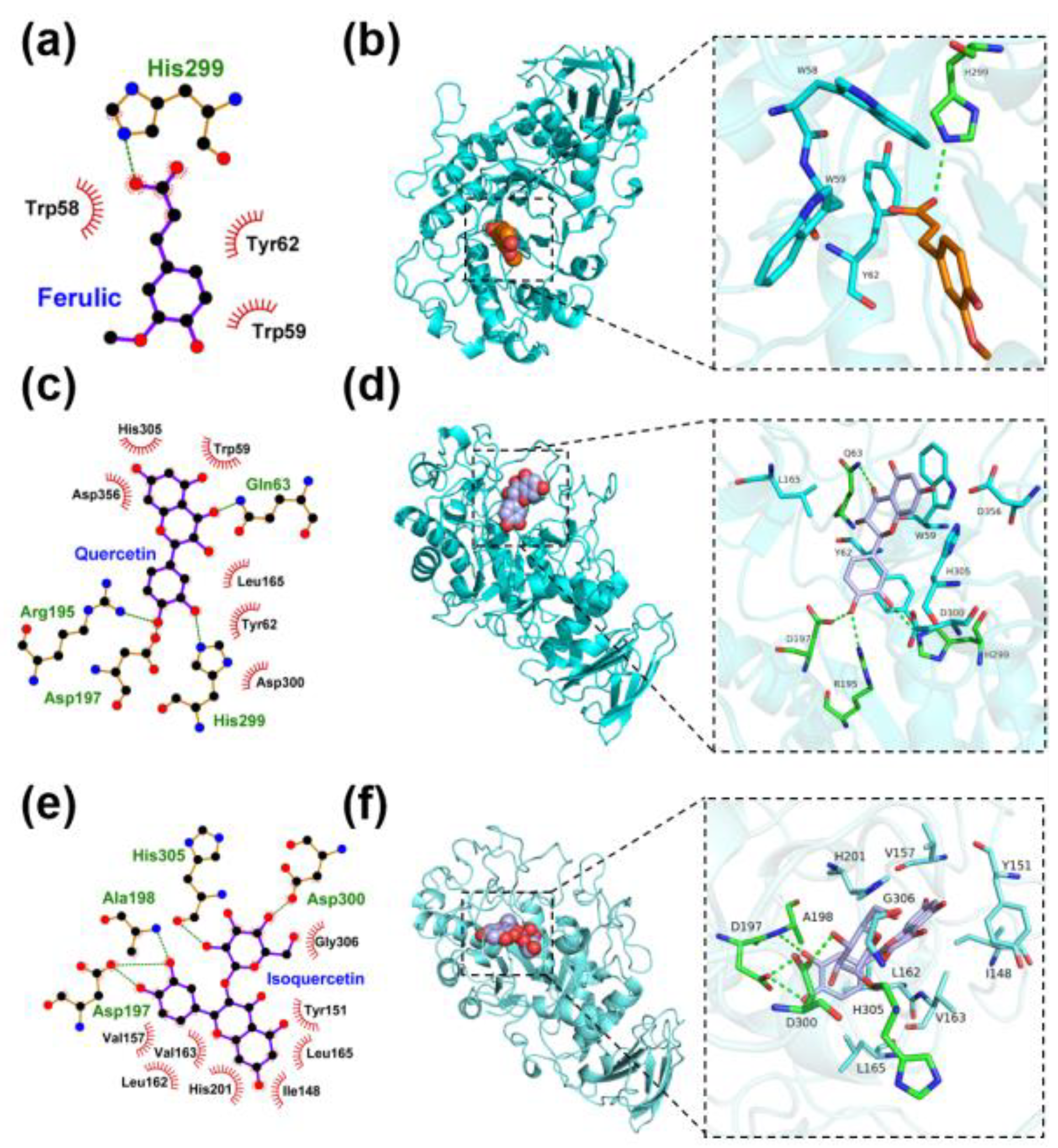
3.11. Molecular Docking Simulation
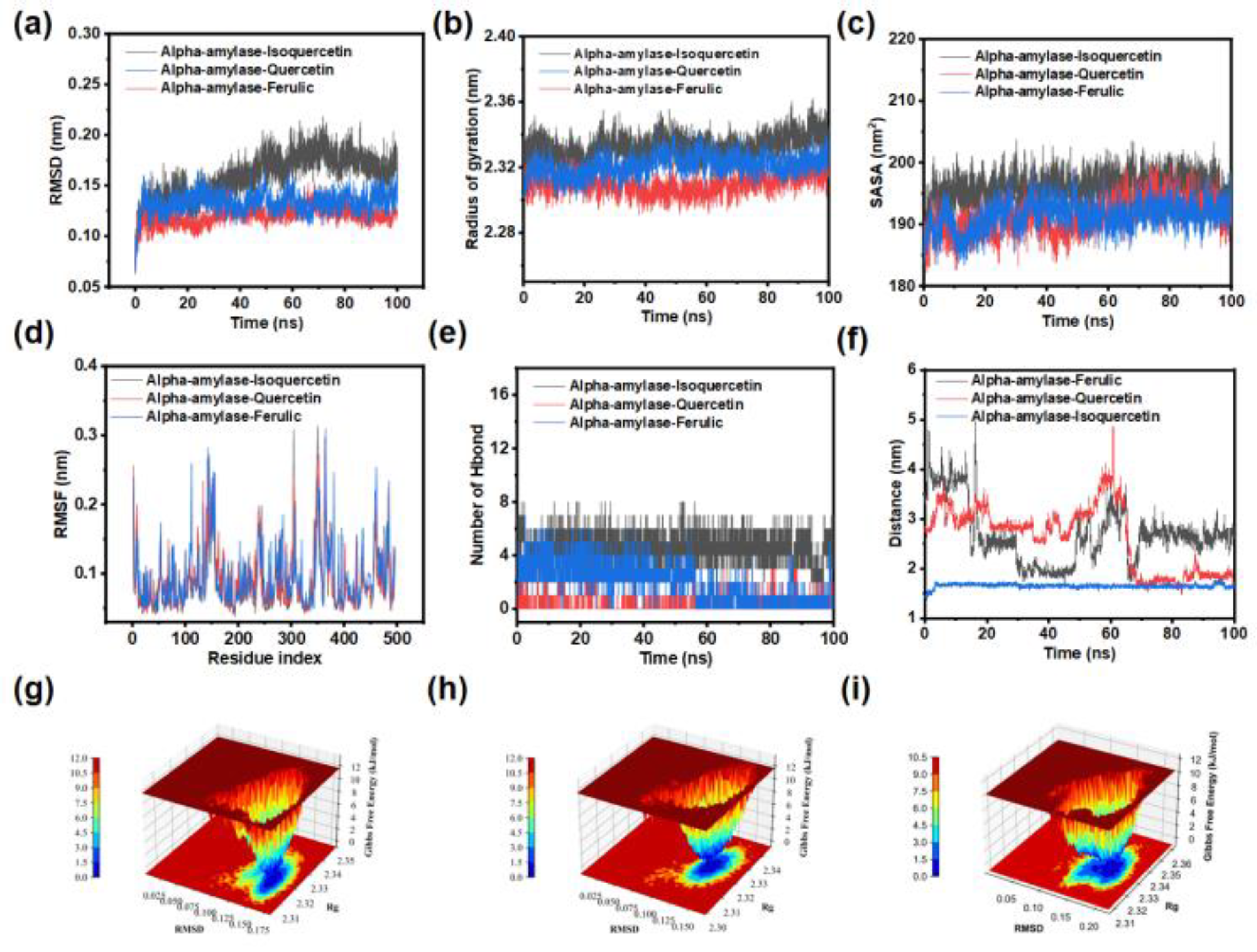
3.12. MD Simulations
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ogle, G. D.; Wang, F.; Haynes, A.; Gregory, G. A.; King, T. W.; Deng, K.; Dabelea, D.; James, S.; Jenkins, A. J.; Li, X. Global type 1 diabetes prevalence, incidence, and mortality estimates 2025: Results from the International diabetes Federation Atlas, and the T1D Index Version 3.0. Diabetes Res. Clin. Pract. 2025(225), 112277. [CrossRef]
- Prama, T. T.; Rahman, M. J.; Zaman, M.; Sarker, F.; Mamun, K. A. DiaBD: A diabetes dataset for enhanced risk analysis and research in Bangladesh. Data Brief. 2025, 61, 111746. [Google Scholar] [CrossRef]
- Im, E.-K. Utility of casual postprandial glucose levels in type 2 diabetes management. Diabetes Care 2004, 27, 334–339. [Google Scholar] [CrossRef]
- Kirkman, M. S.; Shankar, R. R.; Shankar, S.; Shen, C.; Brizendine, E.; Baron, A.; McGill, J. Treating postprandial hyperglycemia does not appear to delay progression of early type 2 diabetes: the Early Diabetes Intervention Program. Diabetes Care 2006, 29((9)), 2095–2101. [Google Scholar] [CrossRef]
- Dong, Y.; Sui, L.; Yang, F.; Ren, X.; Xing, Y.; Xiu, Z. Reducing the intestinal side effects of acarbose by baicalein through the regulation of gut microbiota: An in vitro study. Food Chem. 2022, 394, 133561. [Google Scholar] [CrossRef]
- Mabate, B.; Daub, C. D.; Malgas, S.; Edkins, A. L.; Pletschke, B. I. A combination approach in inhibiting type 2 diabetes-related enzymes using Ecklonia radiata fucoidan and acarbose. Pharmaceutics 2021, 13((11)), 1979. [Google Scholar] [CrossRef]
- Sun, L.; Wang, Y.; Miao, M. Inhibition of α-amylase by polyphenolic compounds: Substrate digestion, binding interactions and nutritional intervention. Trends Food Sci. Technol. 2020, 104, 190–207. [Google Scholar] [CrossRef]
- Wu, H.; Liu, X.; Xie, S.; Zhou, J.; Corradini, M. G.; Pan, Y.; Cui, X. Inhibitory effects of phenolic compounds from blueberry leaf on α-amylase and α-glucosidase: kinetics, mode of action, and molecular interactions. J. Sci. Food Agric. 2025, 105((8)), 4524–4536. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Lian, W.; Li, K.; Han, C.; Liu, W.; Japar, A.; Lei, J. Quality Evolution of Turpan Dry Red Wine During Aging Under Controlled Micro-Oxygenation. J. Food Sci. 2025, 90((8)), e70468. [Google Scholar] [CrossRef]
- Zhang, B.; Deng, Y.; Song, Y.; Yang, F.; He, Y.; Guo, T. Inhibition effects of xanthohumol on α-amylase and α-glucosidase: Kinetics, multi-spectral and molecular docking. Int. J. Biol. Macromol. 2025, 311, 143676. [Google Scholar] [CrossRef] [PubMed]
- Gupta, J.; Das, K.; Rajamani, P. Size-responsive differential modulation in α-amylase by MPA-CdSe QDs: multispectroscopy and molecular docking study. J. Nanoparticle Res. 2021, 23((8)), 190. [Google Scholar] [CrossRef]
- Liu, Q.-Z.; Zhang, H.; Dai, H.-Q.; Zhao, P.; Mao, Y.-F.; Chen, K.-X.; Chen, Z.-X. Inhibition of starch digestion: The role of hydrophobic domain of both α-amylase and substrates. Food Chem. 2021, 341, 128211. [Google Scholar] [CrossRef]
- Shen, H.; Wang, J.; Ao, J.; Hou, Y.; Xi, M.; Cai, Y.; Li, M.; Luo, A. Structure-activity relationships and the underlying mechanism of α-amylase inhibition by hyperoside and quercetin: Multi-spectroscopy and molecular docking analyses. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2023, 285, 121797. [Google Scholar] [CrossRef]
- Oyewusi, H. A.; Wu, Y.-S.; Safi, S. Z.; Wahab, R. A.; Hatta, M. H. M.; Batumalaie, K. Molecular dynamics simulations reveal the inhibitory mechanism of Withanolide A against α-glucosidase and α-amylase. J. Biomol. Struct. Dyn. 2023, 41((13)), 6203–6218. [Google Scholar] [CrossRef]
- Jiang, C.; Wang, L.; Shao, J.; Jing, H.; Ye, X.; Jiang, C.; Wang, H.; Ma, C. Screening and identifying of α-amylase inhibitors from medicine food homology plants: Insights from computational analysis and experimental studies. J. Food Biochem. 2020, 44((12)), e13536. [Google Scholar] [CrossRef]
- Zhu, X.; Yan, K.; Ye, X.; Xiang, X.; Ru, Y.; Niu, Y.; Liu, Y.; Xi, J.; Liu, K. Polyphenols from Flos Trollii inhibit α-amylase and α-glucosidase: Kinetic analysis and mechanistic insights. LWT 2025, 225, 117940. [Google Scholar] [CrossRef]
- Liao, S.-M.; Shen, N.-K.; Liang, G.; Lu, B.; Lu, Z.-L.; Peng, L.-X.; Zhou, F.; Du, L.-Q.; Wei, Y.-T.; Zhou, G.-P. Inhibition of α-amylase activity by Zn2+: Insights from spectroscopy and molecular dynamics simulations. Med. Chem. 2019, 15((5)), 510–520. [Google Scholar] [CrossRef] [PubMed]
- Avwioroko, O. J.; Anigboro, A. A.; Atanu, F. O.; Otuechere, C. A.; Alfred, M. O.; Abugo, J. N.; Omorogie, M. O. Investigation of the binding interaction of α-amylase with Chrysophyllum albidum seed extract and its silver nanoparticles: a multi-spectroscopic approach. Chem. Data Collect. 2020, 29, 100517. [Google Scholar] [CrossRef]
- Lu, Y.; Liu, Y.; Chang, X.; Bai, L.; Bai, Y.; Qi, H. Efficient purification of polyphenols from Ascophyllum nodosum using Macroporous Resin: Adsorption Mechanism, Kinetics, and enhanced bioactivity. Sep. Purif. Technol. 2025, 366, 132704. [Google Scholar] [CrossRef]
- Yang, Y.; Zhu, Q.; Yan, X.; Zhou, Z.; Zhao, J.; Xia, J.; Gao, X.; Ye, M.; Deng, L.; Zeng, Z. Enrichment of polyphenols from Cinnamomum camphora seed kernel by macroporous adsorption resins and evaluation of its antioxidant and enzyme inhibitory activities. Ind. Crop. Prod. 2024, 222, 119486. [Google Scholar] [CrossRef]
- Guo, L.; Tan, D.-C.; Bao, R.-J.; Sun, Q.; Xiao, K.-M.; Xu, Y.; Wang, J.-M.; Hua, Y. Purification and antioxidant activities of polyphenols from Boletus edulis Bull.: Fr. J. Food Meas. Charact. 2020, 14((2)), 649–657. [Google Scholar] [CrossRef]
- Abramovič, H.; Grobin, B.; Poklar Ulrih, N.; Cigić, B. Relevance and standardization of in vitro antioxidant assays: ABTS, DPPH, and Folin–Ciocalteu. J. Chem. 2018, 2018((1)), 4608405. [Google Scholar] [CrossRef]
- Sahin, S.; Isik, E.; Aybastier, O.; Demir, C. Orthogonal signal correction-based prediction of total antioxidant activity using partial least squares regression from chromatograms. J. Chemom. 2012, 26((7)), 390–399. [Google Scholar] [CrossRef]
- Xie, J.; Liu, S.; Dong, R.; Xie, J.; Chen, Y.; Peng, G.; Liao, W.; Xue, P.; Feng, L.; Yu, Q. Bound polyphenols from insoluble dietary fiber of defatted rice bran by solid-state fermentation with trichoderma viride: profile, activity, and release mechanism. J. Agric. Food Chem. 2021, 69((17)), 5026–5039. [Google Scholar] [CrossRef]
- Liu, S.; Yu, J.; Guo, S.; Fang, H.; Chang, X. Inhibition of pancreatic α-amylase by Lonicera caerulea berry polyphenols in vitro and their potential as hyperglycemic agents. LWT 2020, 126, 109288. [Google Scholar] [CrossRef]
- Huang, Y.; Condict, L.; Richardson, S. J.; Brennan, C. S.; Kasapis, S. Exploring the inhibitory mechanism of p-coumaric acid on α-amylase via multi-spectroscopic analysis, enzymatic inhibition assay and molecular docking. Food Hydrocoll. 2023, 139, 108524. [Google Scholar] [CrossRef]
- Liu, D.; Cao, X.; Kong, Y.; Mu, T.; Liu, J. Inhibitory mechanism of sinensetin on α-glucosidase and non-enzymatic glycation: Insights from spectroscopy and molecular docking analyses. Int. J. Biol. Macromol. 2021, 166, 259–267. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Kumar Singh, V.; Kayastha, A. M. Studies on α-amylase inhibition by acarbose and quercetin using fluorescence, circular dichroism, docking, and dynamics simulations. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2024, 314, 124160. [Google Scholar] [CrossRef]
- Eberhardt, J.; Santos-Martins, D.; Tillack, A. F.; Forli, S. AutoDock Vina 1.2.0: New Docking Methods, Expanded Force Field, and Python Bindings. J. Chem. Inf. Model. 2021, 61((8)), 3891–3898. [Google Scholar] [CrossRef]
- Van Der Spoel, D.; Lindahl, E.; Hess, B.; Groenhof, G.; Mark, A. E.; Berendsen, H. J. C. GROMACS: Fast, flexible, and free. J. Comput. Chem. 2005, 26((16)), 1701–1718. [Google Scholar] [CrossRef]
- Cava, C.; Bertoli, G.; Castiglioni, I. Potential Drugs Against COVID-19 Revealed By Gene Expression profile, Molecular Docking and Molecular Dynamic Simulation. Future Virol. 2021, 16((8)), 527–542. [Google Scholar] [CrossRef]
- Shen, D.; Labreche, F.; Wu, C.; Fan, G.; Li, T.; Dou, J.; Zhu, J. Ultrasound-assisted adsorption/desorption of jujube peel flavonoids using macroporous resins. Food Chem. 2022, 368, 130800. [Google Scholar] [CrossRef]
- Gao, F.; Muhammad, T.; Bakri, M.; Yang, W.-W.; Pataer, P.; Yang, X.-X. In-situ measurement of the adsorption thermodynamics of rutin on macroporous adsorption resins by fiber-optic sensing. Instrum. Sci. Technol. 2019, 47((2)), 170–184. [Google Scholar] [CrossRef]
- Huang, Q.; Wen, T.; Fang, T.; Lao, H.; Zhou, X.; Wei, T.; Luo, Y.; Xie, C.; Huang, Z.; Li, K. A comparative evaluation of the composition and antioxidant activity of free and bound polyphenols in sugarcane tips. Food Chem. 2025, 463, 141510. [Google Scholar] [CrossRef]
- Chi, C.; Shi, M.; Zhao, Y.; Chen, B.; He, Y.; Wang, M. Dietary compounds slow starch enzymatic digestion: A review. Front. Nutr. 2022, 9–2022. [Google Scholar] [CrossRef]
- Sun, L.; Warren, F. J.; Gidley, M. J. Natural products for glycaemic control: Polyphenols as inhibitors of alpha-amylase. Trends Food Sci. Technol. 2019, 91, 262–273. [Google Scholar] [CrossRef]
- Zheng, Y.; Yang, W.; Sun, W.; Chen, S.; Liu, D.; Kong, X.; Tian, J.; Ye, X. Inhibition of porcine pancreatic α-amylase activity by chlorogenic acid. J. Funct. Foods 2020, 64, 103587. [Google Scholar] [CrossRef]
- Man, Z.; Feng, Y.; Xiao, J.; Yang, H.; Wu, X. Structural changes and molecular mechanism study on the inhibitory activity of epigallocatechin against α-glucosidase and α-amylase. Front. Nutr. 2022. [Google Scholar] [CrossRef]
- Huang, M.; Xiao, Q.; Li, Y.; Ahmad, M.; Tang, J.; Liao, Q.; Tan, C. Inhibition of α-amylase activity by quercetin via multi-spectroscopic and molecular docking approaches. Food Biosci. 2024, 61, 104951. [Google Scholar] [CrossRef]
- Sun, N.; Xie, J.; Zheng, B.; Xie, J.; Chen, Y.; Hu, X.; Yu, Q. The inhibition mechanism of bound polyphenols extracted from mung bean coat dietary fiber on porcine pancreatic α-amylase: kinetic, spectroscopic, differential scanning calorimetric and molecular docking. Food Chem. 2024, 436, 137749. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; An, J.; Yao, S.; Zhang, C.; Zhou, Y.; Li, L.; Li, H. Effect of Sugarcane Polyphenol Extract on α-Amylase Inhibition and Mechanism Exploration. Foods 2025, Vol. 14. [Google Scholar] [CrossRef] [PubMed]
- Lai, M.; Chen, H.; Liu, X.; Li, F.; Liu, F.; Li, Y.; Yan, J.; Lin, L. Exploration of phenolic profile from mangosteen (Garcinia mangostana L.) pericarp and their contribution to inhibitory effects on α-amylase and α-glucosidase. LWT 2024, 203, 116350. [Google Scholar] [CrossRef]
- Liu, S.; Meng, F.; Guo, S.; Yuan, M.; Wang, H.; Chang, X. Inhibition of α-amylase digestion by a Lonicera caerulea berry polyphenol starch complex revealed via multi-spectroscopic and molecular dynamics analyses. Int. J. Biol. Macromol. 2024, 260, 129573. [Google Scholar] [CrossRef] [PubMed]
- Le, D. T.; Kumar, G.; Williamson, G.; Devkota, L.; Dhital, S. Molecular interactions between polyphenols and porcine α-amylase: An inhibition study on starch granules probed by kinetic, spectroscopic, calorimetric and in silico techniques. Food Hydrocoll. 2024, 151, 109821. [Google Scholar] [CrossRef]
- Yu, M.; Zhu, S.; Huang, D.; Tao, X.; Li, Y. Inhibition of starch digestion by phenolic acids with a cinnamic acid backbone: Structural requirements for the inhibition of α-amylase and α-glucosidase. Food Chem. 2024, 435, 137499. [Google Scholar] [CrossRef]
- Liu, T.; Teng, X.; Tian, Y.; Yao, Y.; Wang, Y.; Wang, W. Chemical composition analysis and antioxidant activity evaluation of leaf, calyx,and fruit of Physalis peruviana L. Food Ferment. Ind. 2024, 50((20)), 243–251. [Google Scholar]
- Wu, X.; Xu, L.-Y.; Li, E.-M.; Dong, G. Application of molecular dynamics simulation in biomedicine. Chem. Biol. Drug. Des. 2022, 99((5)), 789–800. [Google Scholar] [CrossRef] [PubMed]
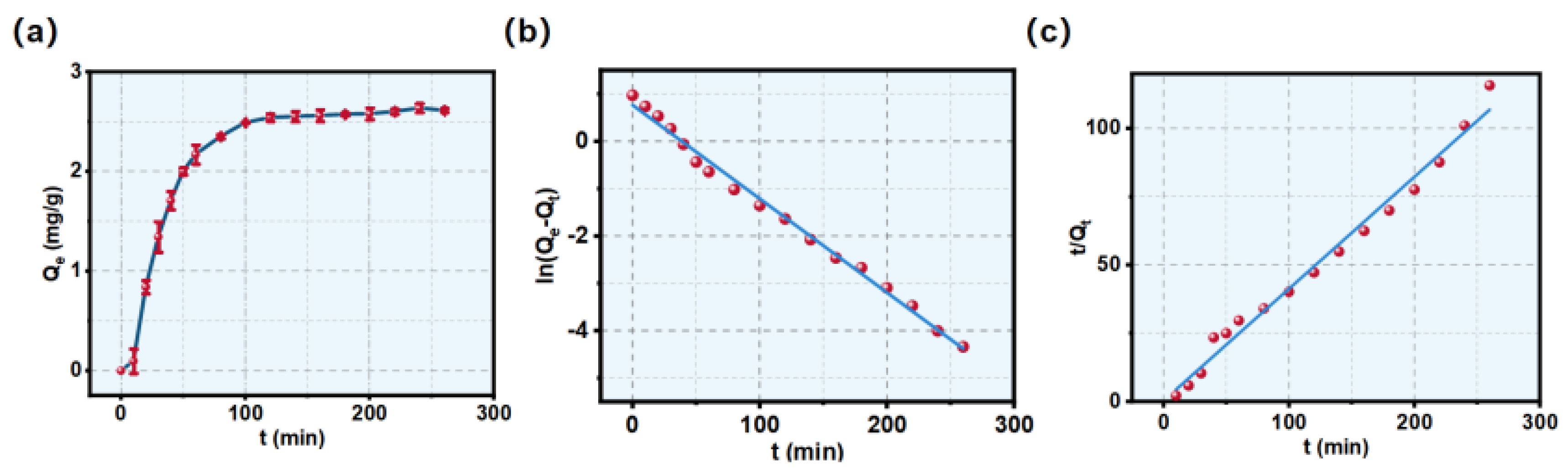
| Models | Equations | Parameters | ||
| Pseudo-first-order | ln(Qe-Qt)=-0.02 t+0.95 | k1=0.02 min-1 | R2=0.993 | Qe=2.57 mg/g |
| Pseudo-second-order | t/Qt=0.41 t+0.23 | k2=0.039 | R2=0.984 | Qe=4.29 mg/g |
| Models | T(℃) | Equations | Parameters | ||
| KL (mL/mg) | Qm | R2 | |||
| Langmuir | 25 | Ce/Qe=0.12Ce+0.40 | 0.29 | 8.33 | 0.99002 |
| 35 | Ce/Qe=0.16Ce+0.48 | 0.32 | 6.45 | 0.96842 | |
| 45 | Ce/Qe=0.17Ce+0.59 | 0.29 | 5.76 | 0.97659 | |
| KF | 1/n | R2 | |||
| Freundlich | 25 | lnQe=0.63lnCe+0.046 | 1.047 | 0.63 | 0.87688 |
| 35 | lnQe=0.83lnCe-0.267 | 0.766 | 0.83 | 0.92198 | |
| 45 | lnQe=0.99lnCe-0.557 | 0.573 | 0.99 | 0.93380 | |
| polyphenols | Kq (×1012 L·mg-1·S-1) | Ksv (×103 L·mg-1) | Ka(mL·mg-1) | n | IC50 (mL·mg-1) |
| Purified polyphenols | 5.89 | 17.48 | 21.99 | 1.39 | 3.15±0.16 |
| Quercetin | 4.62 | 13.74 | 18.04 | 1.23 | 4.44±0.21 |
| Ferulic Acid | 3.27 | 9.73 | 17.50 | 1.08 | 6.02±0.21 |
| Isoquercetin | 2.08 | 6.07 | 16.95 | 0.99 | 11.20±0.56 |
| Crude polyphenols | - | - | - | - | 24.55±0.23 |
| Content | polyphenol species | ||||
| PPA | Quercetin | Isoquercetin | Ferulic Acid | PRP | |
| α-Helix | 30.7 | 32.5 | 18.0 | 35.7 | 25.8 |
| β-Sheet | 20.5 | 14.8 | 31.2 | 15.9 | 28.4 |
| β-Turn | 17.3 | 25.3 | 19.0 | 16.8 | 18.5 |
| Random Coil | 31.5 | 27.4 | 31.8 | 31.6 | 27.3 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




