Submitted:
28 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
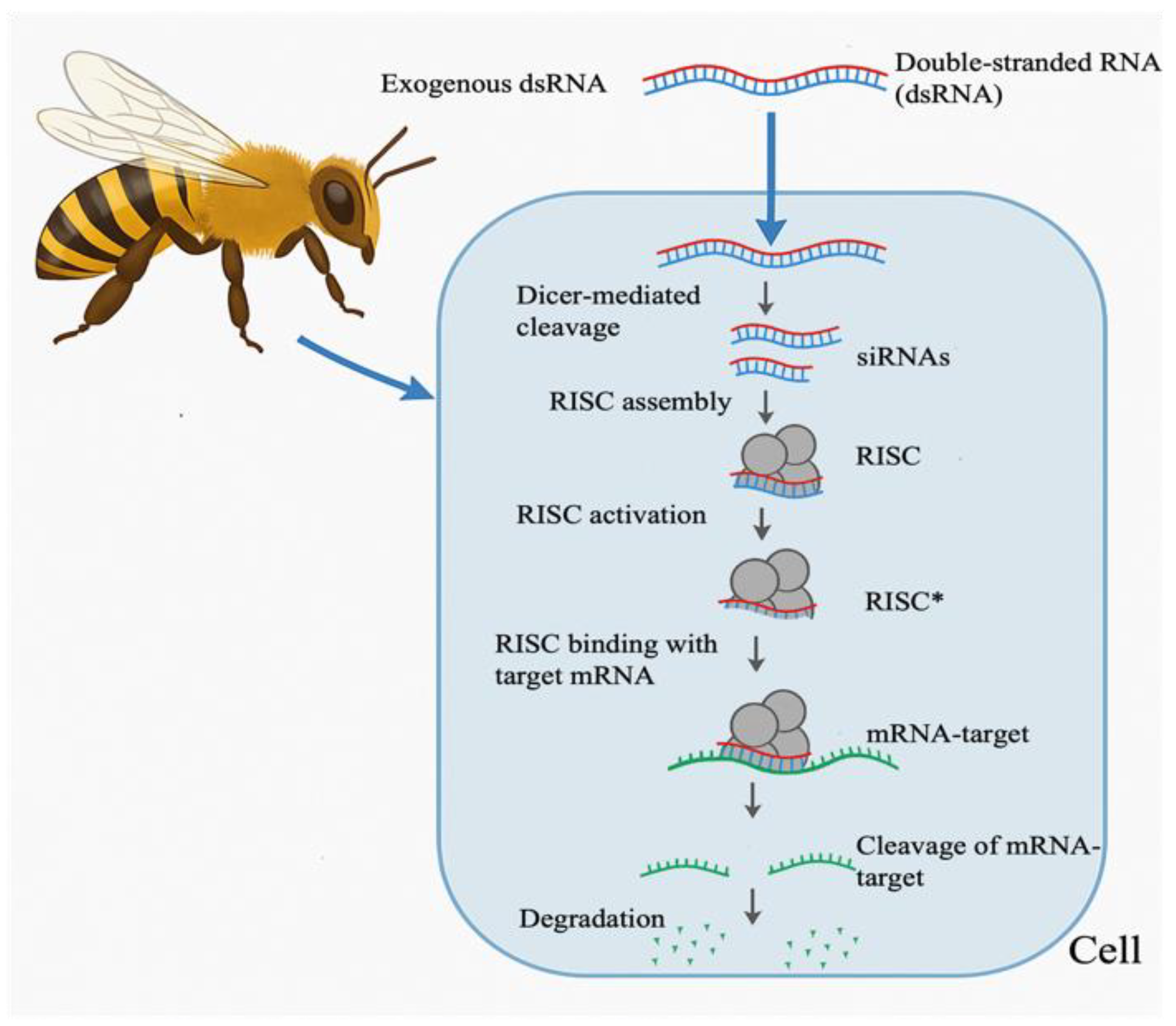
2. Mechanism of RNAi in Honeybee
3. Utilizing RNAi for the Treatment of Viral Infections in Honeybees
4. RNAi for Nosema Disease Treatment
5. RNAi for Varroa Mite Control
6. RNAi for Small Hive Beetle Control
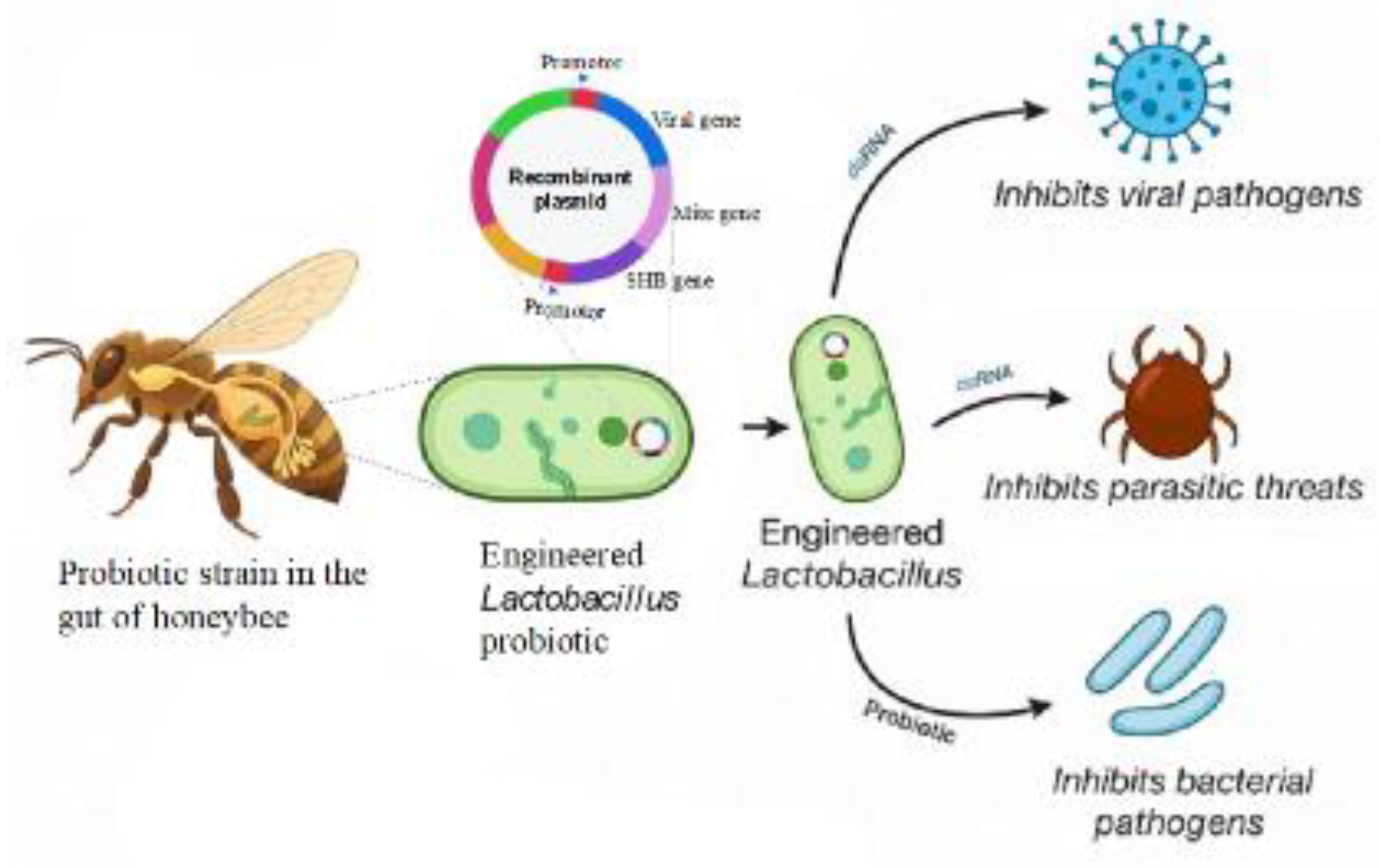
7. Engineered Endosymbionts Producing RNAi Offer a Promising Method for Controlling Pathogens and Parasites in Honeybees
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| RNAi | RNA interference |
| dsRNA | double-stranded RNA |
| DWV | Deformed Wing Virus |
| SHB | Small hive beetles |
| RISC | RNA-Induced Silencing Complex |
| mRNA | Messenger RNA |
| IAPV | Israeli Acute Paralysis Virus |
| BQCV | Black Queen Cell Virus |
| siRNAs | Small interfering RNAs |
| N. ceranae | Nosema ceranae |
| qPCR | Quantitative Polymerase Chain Reaction |
References
- Hung, K.-L.J.; Kingston, J.M.; Albrecht, M.; Holway, D.A.; Kohn, J.R. The Worldwide Importance of Honey Bees as Pollinators in Natural Habitats. Proc. R. Soc. B Biol. Sci. 2018, 285, 20172140. [Google Scholar] [CrossRef]
- DeGrandi-Hoffman, G.; Chen, Y.; Simonds, R. The Effects of Pesticides on Queen Rearing and Virus Titers in Honey Bees (Apis mellifera L.). Insects 2013, 4, 71–89. [Google Scholar] [CrossRef]
- Kulhanek, K.; Steinhauer, N.; Rennich, K.; Caron, D.M.; Sagili, R.R.; Pettis, J.S.; Ellis, J.D.; Wilson, M.E.; Wilkes, J.T.; Tarpy, D.R.; et al. A National Survey of Managed Honey Bee 2015–2016 Annual Colony Losses in the USA. J. Apic. Res. 2017, 56, 328–340. [Google Scholar] [CrossRef]
- Goulson, D.; Nicholls, E.; Botias, C.; Rotheray, E.L. Bee Declines Driven by Combined Stress from Parasites, Pesticides, and Lack of Flowers. Science 2015, 347, 1255957. [Google Scholar] [CrossRef] [PubMed]
- Rosenkranz, P.; Aumeier, P.; Ziegelmann, B. Biology and Control of Varroa destructor. J. Invertebr. Pathol. 2010, 103, S96–S119. [Google Scholar] [CrossRef]
- Brutscher, L.M.; Flenniken, M.L. RNAi and Antiviral Defense in the Honeybee. J. Immunol. 2015, 2015, 941897. [Google Scholar] [CrossRef]
- Brutscher, L.M.; Daughenbaugh, K.F.; Flenniken, M.L. Antiviral Defense Mechanisms in Honey Bees. Curr. Opin. Insect Sci. 2015, 10, 71–82. [Google Scholar] [CrossRef]
- Reybroeck, W. Residues of Antibiotics and Chemotherapeutics in Honey. J. Apic. Res. 2017, 57, 97–112. [Google Scholar] [CrossRef]
- Elzen, P.J.; Westervelt, D.; Lucas, R. Formic Acid Treatment for Control of Varroa destructor (Mesostigmata: Varroidae) and Safety to Apis mellifera (Hymenoptera: Apidae) under Southern United States Conditions. J. Econ. Entomol. 2004, 97, 1509–1512. [Google Scholar] [CrossRef]
- Whyard, S.; Singh, A.D.; Wong, S. Ingested Double-Stranded RNAs Can Act as Species-Specific Insecticides. Insect Biochem. Mol. Biol. 2009, 39, 824–832. [Google Scholar] [CrossRef]
- Baum, J.A.; Bogaert, T.; Clinton, W.; Heck, G.R.; Feldmann, P.; Ilagan, O.; Johnson, S.; Plaetinck, G.; Munyikwa, T.; Pleau, M.; Vaughn, T.; Roberts, J. Control of Coleopteran Insect Pests through RNA Interference. Nat. Biotechnol. 2007, 25, 1322–1326. [Google Scholar] [CrossRef] [PubMed]
- Huvenne, H.; Smagghe, G. Mechanisms of DsRNA Uptake in Insects and Potential of RNAi for Pest Control: A Review. J. Insect. Physiol. 2010, 56, 227–235. [Google Scholar] [CrossRef]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and Specific Genetic Interference by Double-Stranded RNA in Caenorhabditis Elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef]
- Hammond, S.M.; Bernstein, E.; Beach, D.; Hannon, G.J. An RNA-Directed Nuclease Mediates Post-Transcriptional Gene Silencing in Drosophila Cells. Nature 2000, 404, 293–296. [Google Scholar] [CrossRef]
- Azzami, K.; Ritter, W.; Tautz, J.; Beier, H. Infection of Honey Bees with Acute Bee Paralysis Virus Does Not Trigger Humoral or Cellular Immune Responses. Arch. Virol. 2012, 157, 689–702. [Google Scholar] [CrossRef]
- Maori, E.; Paldi, N.; Shafir, S.; Kalev, H.; Tsur, E.; Glick, E.; Sela, I. IAPV, a Bee-Affecting Virus Associated with Colony Collapse Disorder Can Be Silenced by DsRNA Ingestion. Insect Mol. Biol. 2009, 18, 55–60. [Google Scholar] [CrossRef]
- Smeele, Z.E.; Baty, J.W.; Lester, P.J. Effects of Deformed Wing Virus-Targeting DsRNA on Viral Loads in Bees Parasitised and Non-Parasitised by Varroa destructor. Viruses 2023, 15, 2259. [Google Scholar] [CrossRef] [PubMed]
- Grozinger, C.M.; Flenniken, M.L. Bee Viruses: Ecology, Pathogenicity, and Impacts. Annu. Rev. Entomol. 2019, 64, 205–226. [Google Scholar] [CrossRef]
- Ullah, A.; Tlak Gajger, I.; Majoros, A.; Dar, S.A.; Khan, S.; Kalimullah; Haleem Shah, A.; Nasir Khabir, M.; Hussain, R.; Khan, H.U.; Hameed, M.; Anjum, S.I. Viral Impacts on Honey Bee Populations: A Review. Saudi J. Biol. Sci. 2021, 28, 523–530. [Google Scholar] [CrossRef] [PubMed]
- Woodford, L.; Evans, D.J. Deformed Wing Virus: Using Reverse Genetics to Tackle Unanswered Questions about the Most Important Viral Pathogen of Honey Bees. FEMS Microbiol. Rev. 2021, 45, 1–20. [Google Scholar] [CrossRef]
- Sumpter, D.J.T.; Martin, S.J. The Dynamics of Virus Epidemics in Varroa-Infested Honey Bee Colonies. J. Anim. Ecol. 2004, 73, 51–63. [Google Scholar] [CrossRef]
- vanEngelsdorp, D.; Evans, J.D.; Saegerman, C.; Mullin, C.; Haubruge, E.; Nguyen, B.K.; Frazier, M.; Frazier, J.; Cox-Foster, D.; Chen, Y.; Underwood, R.; Tarpy, D.R.; Pettis, J.S. Colony Collapse Disorder: A Descriptive Study. PLoS ONE 2009, 4, e6481. [Google Scholar] [CrossRef]
- Dolezal, A.G.; Hendrix, S.D.; Scavo, N.A.; Carrillo-Tripp, J.; Harris, M.A.; Wheelock, M.J.; O’Neal, M.E.; Toth, A.L. Honey Bee Viruses in Wild Bees: Viral Prevalence, Loads, and Experimental Inoculation. PLoS ONE 2016, 11, e0166190. [Google Scholar] [CrossRef]
- McMenamin, A.J.; Genersch, E. Honey Bee Colony Losses and Associated Viruses. Curr. Opin. Insect Sci. 2015, 8, 121–129. [Google Scholar] [CrossRef]
- Tantillo, G.; Bottaro, M.; Di Pinto, A.; Martella, V.; Di Pinto, P.; Terio, V. Virus Infections of Honeybees Apis mellifera. Ital. J. Food Saf. 2015, 4, 5364. [Google Scholar] [CrossRef]
- Truong, A.-T.; Yoo, M.-S.; Yun, B.-R.; Kang, J.E.; Noh, J.; Hwang, T.J.; Seo, S.K.; Yoon, S.-S.; Cho, Y.S. Prevalence and Pathogen Detection of Varroa and Tropilaelaps Mites in Apis mellifera (Hymenoptera, Apidae) Apiaries in South Korea. J. Apic. Res. 2023, 62, 804–812. [Google Scholar] [CrossRef]
- Doublet, V.; Oddie, M.A.Y.; Mondet, F.; Forsgren, E.; Dahle, B.; Furuseth-Hansen, E.; Williams, G.R.; De Smet, L.; Natsopoulou, M.E.; Murray, T.E.; Semberg, E.; Yanez, O.; de Graaf, D.C.; Conte, Y.L.; Neumann, P.; Rimstad, E.; Paxton, R.J.; de Miranda, J.R. Shift in Virus Composition in Honeybees (Apis mellifera) Following Worldwide Invasion by the Parasitic Mite and Virus Vector Varroa destructor. R. Soc. Open. Sci. 2024, 11, 231529. [Google Scholar] [CrossRef] [PubMed]
- Gallai, N.; Salles, J.M.; Settele, J.; Vaissière, B. Economic Valuation of the Vulnerability of World Agriculture Confronted with Pollinator Decline. Ecol. Econ. 2009, 68, 810–821. [Google Scholar] [CrossRef]
- Chen, Y.P.; Siede, R. Honey Bee Viruses. Adv. Virus Res. 2007, 70, 33–80. [Google Scholar] [PubMed]
- Paudel, Y.; Mackereth, R.; Hanley, R.; Qin, W. Honey Bees (Apis mellifera L.) and Pollination Issues: Current Status, Impacts and Potential Drivers of Decline. J. Agric. Sci. 2015, 7, 2015. [Google Scholar] [CrossRef]
- Morfin, N.; Goodwin, P.H.; Guzman-Novoa, E. Varroa destructor and Its Impacts on Honey Bee Biology. Front. Bee Sci. 2023, 1, 1272937. [Google Scholar] [CrossRef]
- Martin, S.J. The Role of Varroa and Viral Pathogens in the Collapse of Honeybee Colonies: A Modelling Approach. J. Appl. Ecol. 2001, 38, 1082–1093. [Google Scholar] [CrossRef]
- Warner, S.; Pokhrel, L.R.; Akula, S.M.; Ubah, C.S.; Richards, S.L.; Jensen, H.; Kearney, G.D. A Scoping Review on the Effects of Varroa Mite (Varroa destructor) on Global Honey Bee Decline. Sci. Total Environ. 2024, 906, 167492. [Google Scholar] [CrossRef] [PubMed]
- Hunter, W.; Ellis, J.; vanEngelsdorp, D.; Hayes, J.; Westervelt, D.; Glick, E.; Williams, M.; Sela, I.; Maori, E.; Pettis, J.; Cox-Foster, D.; Paldi, N. Large-Scale Field Application of RNAi Technology Reducing Israeli Acute Paralysis Virus Disease in Honey Bees (Apis mellifera, Hymenoptera: Apidae). PLoS Pathog. 2010, 6, e1001160. [Google Scholar] [CrossRef]
- Desai, S.D.; Eu, Y. -J.; Whyard, S.; Currie, R.W. Reduction in Deformed Wing Virus Infection in Larval and Adult Honey Bees (Apis mellifera L.) by Double-stranded RNA Ingestion. Insect Mol. Biol. 2012, 21, 446–455. [Google Scholar] [CrossRef]
- Yoo, M.S.; Truong, A.T.; Jeong, H.; Hahn, D.H.; Lee, J.S.; Yoon, S.S.; Youn, S.Y.; Cho, Y.S. Large-Scale Application of Double-Stranded RNA Shows Potential for Reduction of Sacbrood Virus Disease in Apis cerana Apiaries. Viruses 2023, 15, 897. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, Z.; Wang, Z.-L.; Zhang, L.-Z.; Zeng, Z.-J. A Comparison of RNA Interference via Injection and Feeding in Honey Bees. Insects 2022, 13, 928. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Xu, X.; Zhao, H.; Yang, S.; Wang, X.; Zhao, D.; Diao, Q.; Hou, C. Diverse Factors Affecting Efficiency of RNAi in Honey Bee Viruses. Front. Genet. 2018, 9, 384. [Google Scholar] [CrossRef]
- Ferrufino, C.; Scannapieco, A.; Russo, R.M.; Gonzalez, F.N.; Salvador, R.; Dus Santos, M.J. Reduction in Acute Bee Paralysis Virus Infection and Mortality in Honey Bees (Apis Mellifera) by RNA Interference Technology. Insects 2025, 16, 453. [Google Scholar] [CrossRef] [PubMed]
- Brutscher, L.M.; Daughenbaugh, K.F.; Flenniken, M.L. Virus and DsRNA-Triggered Transcriptional Responses Reveal Key Components of Honey Bee Antiviral Defense. Sci. Rep. 2017, 7, 6448. [Google Scholar] [CrossRef]
- Zhu, K.Y.; Palli, S.R. Mechanisms, Applications, and Challenges of Insect RNA Interference. Annu. Rev. Entomol. 2020, 65, 293–311. [Google Scholar] [CrossRef]
- Botías, C.; Martín-Hernández, R.; Barrios, L.; Meana, A.; Higes, M. Nosema Spp. Infection and Its Negative Effects on Honey Bees (Apis mellifera iberiensis) at the Colony Level. Vet. Res. 2013, 44, 25. [Google Scholar] [CrossRef]
- Ghramh, H.A.; Khan, K.A. Current Insight into Nosema Disease of Honeybees and Their Future Prospective. Pak. J. Zool. 2025, 57, 951–960. [Google Scholar] [CrossRef]
- Paris, L.; Peghaire, E.; Moné, A.; Diogon, M.; Debroas, D.; Delbac, F.; El Alaoui, H. Honeybee Gut Microbiota Dysbiosis in Pesticide/Parasite Co-Exposures Is Mainly Induced by Nosema ceranae. J. Invertebr. Pathol. 2020, 172, 107348. [Google Scholar] [CrossRef]
- Goblirsch, M. Nosema Ceranae Disease of the Honey Bee (Apis mellifera). Apidologie 2018, 49, 131–150. [Google Scholar] [CrossRef]
- Marín-García, P.J.; Peyre, Y.; Ahuir-Baraja, A.E.; Garijo, M.M.; Llobat, L. The Role of Nosema ceranae (Microsporidia: Nosematidae) in Honey Bee Colony Losses and Current Insights on Treatment. Vet. Sci. 2022, 9, 130. [Google Scholar] [CrossRef]
- Panek, J.; Paris, L.; Roriz, D.; Moné, A.; Dubuffet, A.; Delbac, F.; Diogon, M.; Alaoui, H.E. Impact of the Microsporidian Nosema ceranae on the Gut Epithelium Renewal of the Honeybee, Apis mellifera. J. Invertebr. Pathol. 2018, 159, 121–128. [Google Scholar] [CrossRef]
- Huang, W.F.; Solter, L.F.; Yau, P.M.; Imai, B.S. Nosema Ceranae Escapes Fumagillin Control in Honey Bees. PLoS Pathog. 2013, 9, e1003185. [Google Scholar] [CrossRef]
- Williams, G.R.; Shafer, A.B.A.; Rogers, R.E.L.; Shutler, D.; Stewart, D.T. First Detection of Nosema ceranae, a Microsporidian Parasite of European Honey Bees (Apis mellifera) in Canada and Central USA. J. Invertebr. Pathol. 2008, 97, 189–192. [Google Scholar] [CrossRef] [PubMed]
- Higes, M.; Nozal, M.J.; Alvaro, A.; Barrios, L.; Meana, A.; Martín-Hernández, R.; Bernal, J.L.; Bernal, J. The Stability and Effectiveness of Fumagillin in Controlling Nosema ceranae (Microsporidia) Infection in Honey Bees (Apis mellifera) under Laboratory and Field Conditions. Apidologie 2011, 42, 364–377. [Google Scholar] [CrossRef]
- Porrini, M.P.; Sarlo, E.G.; Medici, S.K.; Garrido, P.M.; Porrini, D.P.; Damiani, N.; Eguaras, M.J. Nosema ceranae Development in Apis mellifera: Influence of Diet and Infective Inoculum. J. Apic. Res. 2011, 50, 35–41. [Google Scholar] [CrossRef]
- Kim, I.H.; Kim, D.J.; Gwak, W.S.; Woo, S.D. Increased Survival of the Honey Bee Apis mellifera Infected with the Microsporidian Nosema ceranae by Effective Gene Silencing. Arch. Insect Biochem. Physiol. 2020, 105, e21734. [Google Scholar] [CrossRef] [PubMed]
- Qi, Y.; Wang, C.; Lang, H.; Wang, Y.; Wang, X.; Zheng, H.; Lu, Y. Liposome-Based RNAi Delivery in Honeybee for Inhibiting Parasite Nosema ceranae. Synth. Syst. Biotechnol. 2024, 9, 853–860. [Google Scholar] [CrossRef] [PubMed]
- He, N.; Zhang, Y.; Duan, X.L.; Li, J.H.; Huang, W.F.; Evans, J.D.; DeGrandi-Hoffman, G.; Chen, Y.P.; Huang, S.K. RNA Interference-Mediated Knockdown of Genes Encoding Spore Wall Proteins Confers Protection against Nosema ceranae Infection in the European Honey Bee, Apis mellifera. Microorganisms 2021, 9, 505. [Google Scholar] [CrossRef]
- Huang, Q.; Lariviere, P.J.; Powell, J.E.; Moran, N.A. Engineered Gut Symbiont Inhibits Microsporidian Parasite and Improves Honey Bee Survival. Proc. Natl. Acad. Sci. U.S. A. 2023, 120, e2220922120. [Google Scholar] [CrossRef]
- Lang, H.; Wang, H.; Wang, H.; Zhong, Z.; Xie, X.; Zhang, W.; Guo, J.; Meng, L.; Hu, X.; Zhang, X.; et al. Engineered Symbiotic Bacteria Interfering Nosema Redox System Inhibit Microsporidia Parasitism in Honeybees. Nat. Commun. 2023, 14, 2778. [Google Scholar] [CrossRef]
- Huang, Q.; Chen, Y.; Neumann, P.; Li, W.; Evans, J.D. Effective Silencing of Dicer Decreases Spore Load of the Honey Bee Parasite Nosema ceranae. Fungal Genet. Biol. 2016, 6, 1000144. [Google Scholar] [CrossRef]
- Noël, A.; Le Conte, Y.; Mondet, F. Varroa destructor: How Does It Harm Apis mellifera Honey Bees and What Can Be Done about It? Emerg. Top. Life Sci. 2020, 4, 45–57. [Google Scholar] [CrossRef]
- Ayan, A.; Tutun, H.; Aldemir, O.S. Control Methods against Varroa Mites. Int. J. Adv. Study Res. Work 2019, 2, 19–23. [Google Scholar] [CrossRef]
- Jack, C.J.; Ellis, J.D. Integrated Pest Management Control of Varroa destructor (Acari: Varroidae), the Most Damaging Pest of (Apis mellifera L. (Hymenoptera: Apidae)) Colonies. J. Insect Sci. 2021, 21, 6. [Google Scholar] [CrossRef]
- Calderón, R.A.; Ramírez, M.; Ramírez, F.; Villalobos, E. Effectiveness of Formic Acid and Thymol in the Control of Varroa destructor in Africanized Honey Bee Colonies. Agron. Costarric. 2014, 38, 175–188. [Google Scholar]
- Narciso, L.; Topini, M.; Ferraiuolo, S.; Ianiro, G.; Marianelli, C. Effects of Natural Treatments on the Varroa Mite Infestation Levels and Overall Health of Honey Bee (Apis mellifera) Colonies. PLoS ONE 2024, 19, e0302846. [Google Scholar] [CrossRef]
- Lipiński, Z.; Szubstarski, J. Resistance of Varroa destructor to Most Commonly Used Synthetic Acaricides. Pol. J. Vet. Sci. 2007, 10, 289–294. [Google Scholar]
- Nekoei, S.; Rezvan, M.; Khamesipour, F.; Mayack, C.; Molento, M.B.; Revainera, P.D. A Systematic Review of Honey Bee (Apis mellifera, Linnaeus, 1758) Infections and Available Treatment Options. Vet. Med. Sci. 2023, 9, 1848–1860. [Google Scholar] [CrossRef] [PubMed]
- McGruddy, R.A.; Smeele, Z.E.; Manley, B.; Masucci, J.D.; Haywood, J.; Lester, P.J. RNA Interference as a Next-Generation Control Method for Suppressing Varroa destructor Reproduction in Honey Bee (Apis mellifera) Hives. Pest. Manag. Sci. 2024, 80, 4770–4778. [Google Scholar] [CrossRef]
- Bortolin, F.; Rigato, E.; Perandin, S.; Granato, A.; Zulian, L.; Millino, C.; Pacchioni, B.; Mutinelli, F.; Fusco, G. First Evidence of the Effectiveness of a Field Application of RNAi Technology in Reducing Infestation of the Mite Varroa destructor in the Western Honey Bee (Apis mellifera). Parasit. Vectors 2025, 18, 28. [Google Scholar] [CrossRef]
- Niu, J.; Shen, G.; Christiaens, O.; Smagghe, G.; He, L.; Wang, J. Beyond Insects: Current Status and Achievements of RNA Interference in Mite Pests and Future Perspectives. Pest. Manag. Sci. 2018, 74, 2680–2687. [Google Scholar] [CrossRef] [PubMed]
- Cuthbertson, A.G.S.; Wakefield, M.E.; Powell, M.E.; Marris, G.A.; Anderson, H.; Budge, G.E.; Brown, M.A. The Small Hive Beetle Aethina tumida: A Review of Its Biology and Control Measures. Curr. Zool. 2013, 59, 644–653. [Google Scholar] [CrossRef]
- Hood, W.M. The Small Hive Beetle, Aethina tumida: A Review. Bee World 2004, 85, 51–59. [Google Scholar] [CrossRef]
- Neumann, P.; Pirk, C.W.; Hepburn, H.R.; Solbrig, A.J.; Ratnieks, F.L.; Elzen, P.J.; Baxter, J.R. Social Encapsulation of Beetle Parasites by Cape Honeybee Colonies (Apis mellifera capensis Esch.). Sci. Nat. 2001, 88, 214–216. [Google Scholar] [CrossRef]
- Tutun, H.; Sekercİ, Y.; Sevin, S. Future Effects of Small Hive Beetle, Aethina tumida, on Honey Bee Colony in Turkey Based on Temperature Factor Using a Mathematical Model. Eur. Zool. J. 2022, 89, 1259–1270. [Google Scholar] [CrossRef]
- Gela, A.A.B.T.N. Investigating the Effect and Control of Small Hive Beetle, Aethina tumida (Murray) on Honeybee Colonies in Ethiopia. Int. J. Res. Stud. Biosci. 2018, 6, 1–6. [Google Scholar]
- Ellis, J.D.; Delaplane, K.S. Small Hive Beetle (Aethina tumida) Oviposition Behaviour in Sealed Brood Cells with Notes on the Removal of the Cell Contents by European Honey Bees (Apis mellifera). J. Apic. Res. 2008, 47, 210–215. [Google Scholar] [CrossRef]
- Powell, M.E.; Bradish, H.M.; Gatehouse, J.A.; Fitches, E.C. Systemic RNAi in the Small Hive Beetle Aethina tumida Murray (Coleoptera: Nitidulidae), a Serious Pest of the European Honey Bee Apis mellifera. Pest. Manag. Sci. 2017, 73, 53–63. [Google Scholar] [CrossRef]
- Kim, K.; Kim, S.H.; Yoon, K.A.; Cho, Y.S.; Yoo, M.-S.; Lee, S.H. Characterization of the Small Hive Beetle Transcriptome Focused on the Insecticide Target Site and RNA Interference Genes. J. Asia. Pac. Entomol. 2018, 21, 1256–1261. [Google Scholar] [CrossRef]
- Lariviere, P.J.; Leonard, S.P.; Horak, R.D.; Powell, J.E.; Barrick, J.E. Honey Bee Functional Genomics Using Symbiont-Mediated RNAi. Nat. Protoc. 2023, 18, 902–928. [Google Scholar] [CrossRef]
- Leonard, S.P.; Powell, J.E.; Perutka, J.; Geng, P.; Heckmann, L.C.; Horak, R.D.; Davies, B.W.; Ellington, A.D.; Barrick, J.E.; Moran, N.A. Engineered Symbionts Activate Honey Bee Immunity and Limit Pathogens. Science 2020, 367, 573–576. [Google Scholar] [CrossRef] [PubMed]
- Chege, M.; Kinyua, J.; Paredes, J.C. Lactobacillus kunkeei Impacts the Health of Honey Bees, Apis mellifera scutellata, and Protects the Bees against the Opportunistic Pathogen Serratia marcescens. Int. J. Trop. Insect Sci. 2023, 43, 1947–1955. [Google Scholar] [CrossRef]
- Mojgani, N.; Bagheri, M.; Ashique, S.; Islam, A.; Moharrami, M.; Modirrousta, H.; Hussain, A. Honeybee Defense Mechanisms: Role of Honeybee Gut Microbiota and Antimicrobial Peptides in Maintaining Colony Health and Preventing Diseases. Microb. Pathog. 2025, 198, 107161. [Google Scholar] [CrossRef] [PubMed]
- Peirotén, Á.; Landete, J.M. Natural and Engineered Promoters for Gene Expression in Lactobacillus Species. Appl. Microbiol. Biotechnol. 2020, 104, 3797–3805. [Google Scholar] [CrossRef] [PubMed]
- Heiss, S.; Hörmann, A.; Tauer, C.; Sonnleitner, M.; Egger, E.; Grabherr, R.; Heinl, S. Evaluation of Novel Inducible Promoter/Repressor Systems for Recombinant Protein Expression in Lactobacillus plantarum. Microb. Cell. Fact. 2016, 15, 50. [Google Scholar] [CrossRef] [PubMed]
- Kazi, T.A.; Acharya, A.; Mukhopadhyay, B.C.; Mandal, S.; Arukha, A.P.; Nayak, S.; Biswas, S.R. Plasmid-Based Gene Expression Systems for Lactic Acid Bacteria: A Review. Microorganisms 2022, 10, 1132. [Google Scholar] [CrossRef]
- Daisley, B.A.; Pitek, A.P.; Chmiel, J.A.; Gibbons, S.; Chernyshova, A.M.; Al, K.F.; Faragalla, K.M.; Burton, J.P.; Thompson, G.J.; Reid, G. Lactobacillus spp. Attenuate Antibiotic-Induced Immune and Microbiota Dysregulation in Honey Bees. Commun. Biol. 2020, 3, 534. [Google Scholar] [CrossRef] [PubMed]
- Daisley, B.A.; Pitek, A.P.; Torres, C.; Lowery, R.; Adair, B.A.; Al, K.F.; Niño, B.; Burton, J.P.; Allen-Vercoe, E.; Thompson, G.J.; Niño, E. Delivery Mechanism Can Enhance Probiotic Activity against Honey Bee Pathogens. ISME J. 2023, 17, 1382–1395. [Google Scholar] [CrossRef] [PubMed]
- Iorizzo, M.; Letizia, F.; Ganassi, S.; Testa, B.; Petrarca, S.; Albanese, G.; Di Criscio, D.; De Cristofaro, A. Functional Properties and Antimicrobial Activity from Lactic Acid Bacteria as Resources to Improve the Health and Welfare of Honey Bees. Insects 2022, 13, 308. [Google Scholar] [CrossRef]
| Target | RNAi Application | Delivery Method | Effect/Outcome | References |
|---|---|---|---|---|
| Viral Diseases | Silencing viral genes (e.g., DWV, IAPV, SBV, KBV) using virus-specific dsRNA or siRNA | Injection, feeding (e.g., Remebee-I), engineered symbionts | Reduced viral load, increased survival, suppression of replication, improved colony health | [16,34,37,39,41,76] |
| Nosema spp. | Silencing of essential parasite genes (e.g., SWP, mitosome-related genes), redox system | Direct feeding with dsRNA, engineered gut symbionts (Snodgrassella alvi) | Reduced spore load, increased survival, improved immunity, colony-level protection via symbiont transfer | [52,56] |
| Varroa destructor | Targeting reproductive and metabolic genes in mites (e.g., vitellogenin, aquaporins) | Feeding bees with dsRNA (larvae/adults), engineered symbionts | Reduced mite reproduction, decreased infestation, environmentally friendly, 60%+ reduction in studies | [65,67,76,77] |
| Small Hive Beetle | Silencing essential genes (e.g., V-ATPase, laccase2) for development and survival | Injection or feeding (larvae/adults) | High larval mortality (up to 100% via injection), no adverse effects on bees, specificity confirmed | [74,75] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




