Submitted:
27 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
Keywords:
I. Introduction
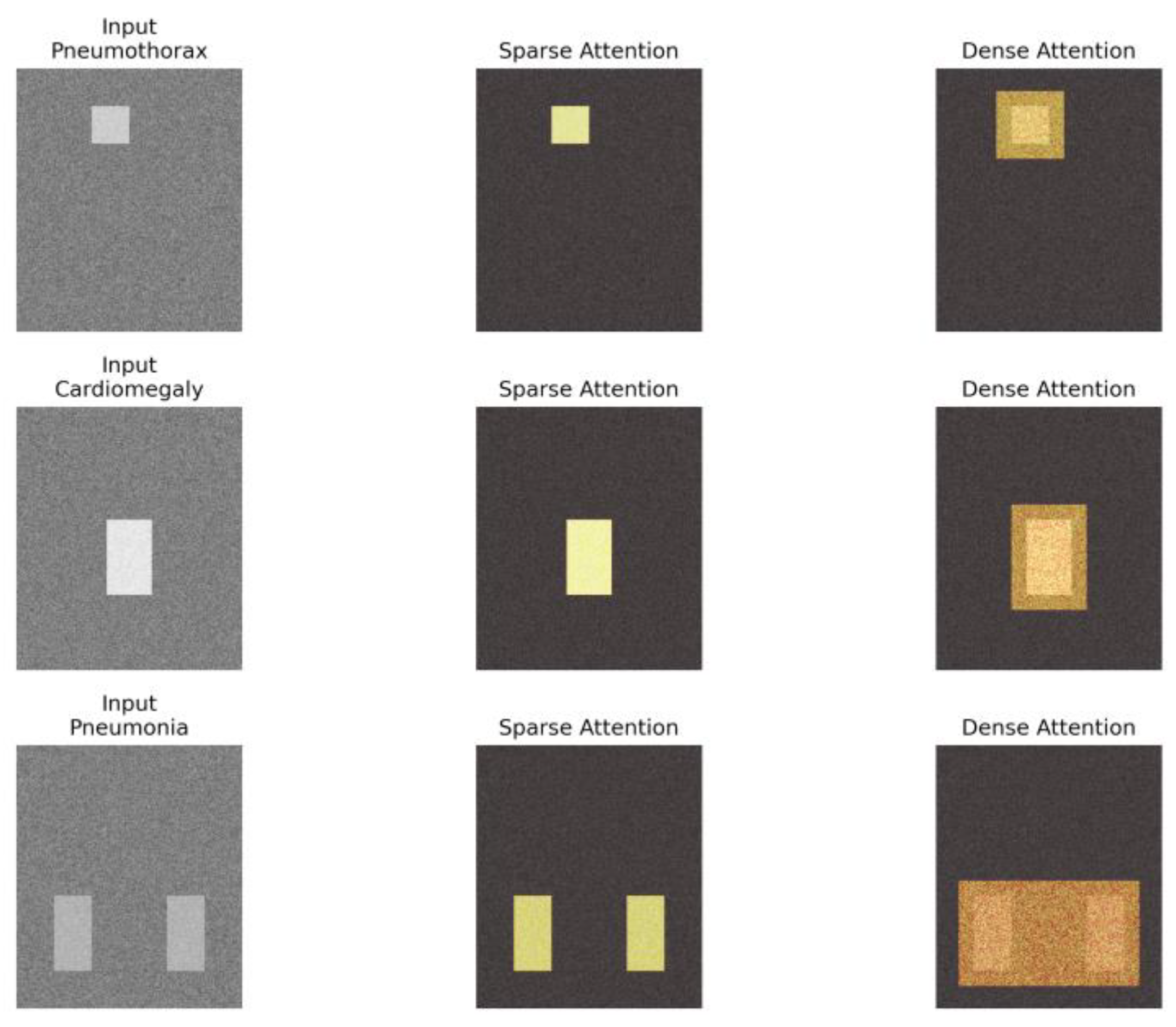
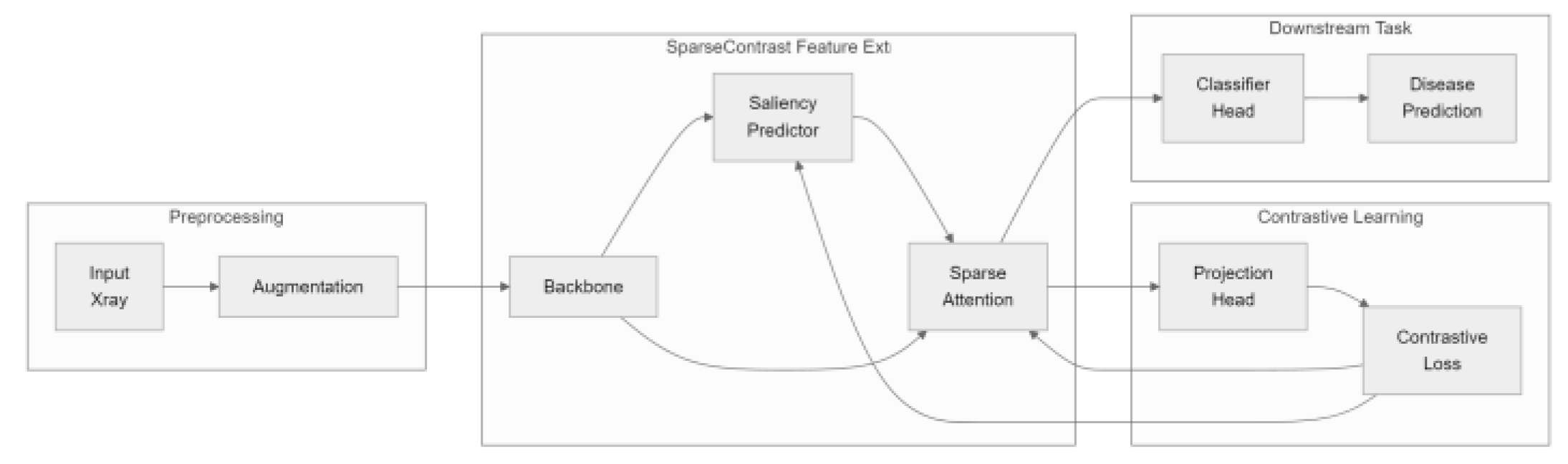
- We introduce a dynamic sparse attention approach for contrastive learning in medical imaging, which markedly lowers computational expenses without compromising diagnostic performance.
- We propose a lightweight saliency predictor that jointly optimizes sparsity and feature quality, which permits the model to adaptively focus on diagnostically relevant regions.
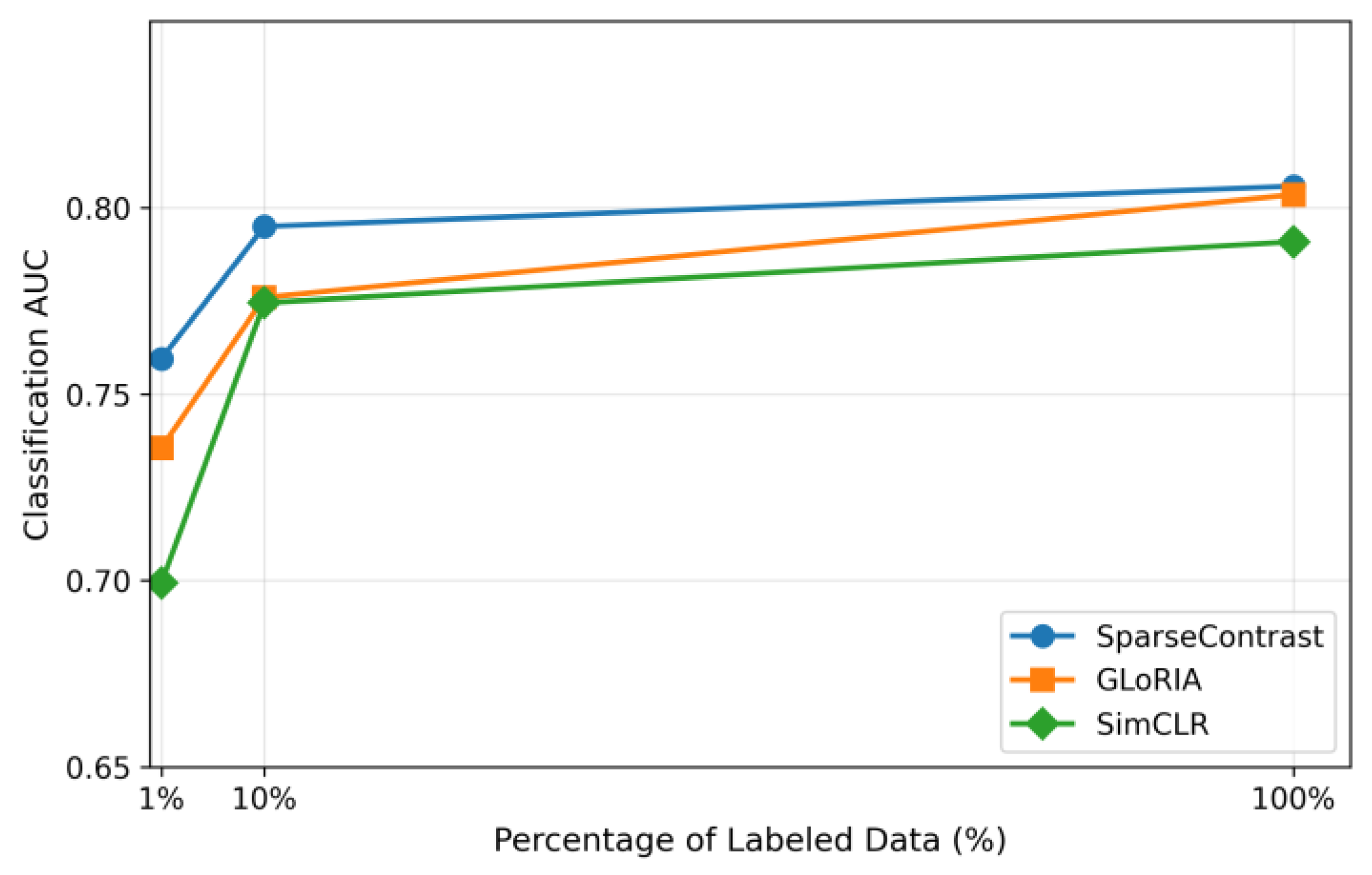
- The efficacy of SparseContrast is established on chest X-ray disease detection tasks, with results indicating superior performance compared to current approaches in both efficiency and accuracy, especially under limited-data conditions.
II. Related Work
A. Contrastive Learning in Medical Imaging
B. Sparse Attention Mechanisms
C. Hybrid Approaches
III. Preliminaries on Contrastive Learning and Attention in Medical Imaging
A. Attention Mechanisms in Vision
B. Medical Imaging Data Characteristics
IV. Sparse Token Attention for Efficient Contrastive Learning
A. Dynamic N:M Fine-Grained Sparse Attention Implementation
B. Integration of Sparse Attention into Contrastive Loss Computation
C. Downstream Adaptation with Reused Sparse Attention Maps
D. Joint Optimization of Sparsity and Feature Quality
V. Experiments
A. Experimental Setup
B. Main Results
C. Ablation Studies
D. Low-Data Regime Performance
VI. Discussion and Future Work
A. Limitations of SparseContrast
B. Potential Application Scenarios of SparseContrast
C. Ethical Considerations in SparseContrast
VII. Conclusion
References
- K. Shung, M. Smith, and B. Tsui, “Principles of medical imaging,” books.google.com, 2012.
- H. Kasban, M. El-Bendary, et al., “A comparative study of medical imaging techniques,” Unable to determine complete publication venue, 2015.
- Suzuki, K. Overview of deep learning in medical imaging. In Radiological physics and technology; 2017. [Google Scholar]
- J. A. B. Rubak, K. Naveed, S. Jain, L. Esterle, A. Iosifidis, and R. Pauwels, “Impact of labeling inaccuracy and image noise on tooth segmentation in panoramic radiographs using federated, centralized and local learning,” Dentomaxillofacial Radiology, p. twag001, 2026.
- B. Khalil, M. Baraka, S. Haghighat, S. Jain, N. Manila, R. Ramani, A. Tichy, E. Tolstaya, F. Schwendicke, and R. Pauwels, “Synthetic imaging in dentistry: A narrative review of deep learning techniques and applications,” Journal of Dentistry, p. 106274, 2025.
- Z. Chen, Z. Qu, Y. Quan, L. Liu, Y. Ding, and Y. Xie, “Dynamic n: M fine-grained structured sparse attention mechanism,” in Proceedings of the 28th ACM SIGKDD conference on knowledge discovery and data mining, 2023.
- Y. Vu, R. Wang, N. Balachandar, et al., “Medaug: Contrastive learning leveraging patient metadata improves representations for chest x-ray interpretation,” in Proceedings of machine learning research, 2021.
- A. Jaiswal, T. Li, C. Zander, Y. Han, et al., “Scalp-supervised contrastive learning for cardiopulmonary disease classification and localization in chest x-rays using patient metadata,” in IEEE international conference on data mining, 2021.
- S. Rizvi, R. Tang, X. Jiang, X. Ma, et al., “Local contrastive learning for medical image recognition,” in American medical informatics association annual symposium, 2024.
- Y. Han, C. Chen, A. Tewfik, Y. Ding, et al., “Pneumonia detection on chest x-ray using radiomic features and contrastive learning,” in 2021 IEEE 18th international symposium on biomedical imaging (ISBI), 2021.
- J. Li et al., “Multi-task contrastive learning for automatic CT and x-ray diagnosis of COVID-19,” Pattern recognition, 2021.
- J. Yuan, H. Gao, D. Dai, J. Luo, L. Zhao, et al., “Native sparse attention: Hardware-aligned and natively trainable sparse attention,” in Annual conference of the association for computational linguistics, 2025.
- Y. Zhang et al., “A medical image segmentation method based on adaptive graph sparse algorithm under contrastive learning framework,” Displays, 2025.
- Öksüz, C.; Urhan, O.; Güllü, M. An integrated convolutional neural network with attention guidance for improved performance of medical image classification. In Neural Computing and Applications; 2024. [Google Scholar]
- Ren, H.; Dai, H.; Dai, Z.; Yang, M.; et al. Combiner: Full attention transformer with sparse computation cost. Advances in neural information processing systems, 2021. [Google Scholar]
- Shen, G.; Zhao, J.; Chen, Q.; Leng, J.; Li, C.; et al. SALO: An efficient spatial accelerator enabling hybrid sparse attention mechanisms for long sequences. In Proceedings of the 59th annual design automation conference, 2022. [Google Scholar]
- Li, X.; et al. “Deep learning attention mechanism in medical image analysis: Basics and beyonds,” Unable to determine the complete publication venue; 2023. [Google Scholar]
- Parmar, C.; Barry, J.; Hosny, A.; et al. Data analysis strategies in medical imaging. In Clinical Cancer Research; 2018. [Google Scholar]
- Li, J.; et al. A systematic collection of medical image datasets for deep learning. ACM Computing Surveys, 2023. [Google Scholar]
- I. Dimitrovski, D. Kocev, S. Loskovska, and S. Džeroski, “Hierarchical annotation of medical images,” Pattern Recognition, 2011.
- J. Irvin, P. Rajpurkar, M. Ko, Y. Yu, S. Ciurea-Ilcus, et al., “Chexpert: A large chest radiograph dataset with uncertainty labels and expert comparison,” in Proceedings of the aaai conference on artificial intelligence, 2019.
- Johnson, A.; Pollard, T.; Greenbaum, N.; et al. MIMIC-CXR-JPG, a large publicly available database of labeled chest radiographs. arXiv 2019, arXiv:1901.07042. [Google Scholar]
- Yanar, E.; Kutan, F.; Ayturan, K.; Kutbay, U.; Algın, O.; et al. A comparative analysis of the mamba, transformer, and CNN architectures for multi-label chest x-ray anomaly detection in the NIH chestX-Ray14 dataset. Diagnostics 2025. [Google Scholar] [CrossRef] [PubMed]
- I. Baltruschat, H. Nickisch, M. Grass, T. Knopp, et al., “Comparison of deep learning approaches for multilabel chest x-ray classification,” Scientific reports, 2019.
- Dosovitskiy, A. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv 2020, arXiv:2010.11929. [Google Scholar]
- D. Kingma, “Adam: A method for stochastic optimization,” arXiv preprint arXiv:1412.6980, 2014, 1412. arXiv:1412.6980.
- Chen, T.; Kornblith, S.; Norouzi, M.; et al. A simple framework for contrastive learning of visual representations. International conference on machine learning, 2020. [Google Scholar]
- Chen, X.; Xie, S.; He, K. An empirical study of training self-supervised vision transformers. In Proceedings of the IEEE/CVF international conference on computer vision, 2021. [Google Scholar]
- Ryoo, M.; Piergiovanni, A.; Arnab, A.; et al. Tokenlearner: Adaptive space-time tokenization for videos. Advances in neural information processing systems, 2021. [Google Scholar]
- Rao, Y.; Zhao, W.; Liu, B.; Lu, J.; Zhou, J.; et al. Dynamicvit: Efficient vision transformers with dynamic token sparsification. Advances in neural information processing systems, 2021. [Google Scholar]
- Zhang, Y.; Jiang, H.; Miura, Y.; et al. Contrastive learning of medical visual representations from paired images and text. Proceedings of machine learning research, 2022. [Google Scholar]
- Huang, S.; Shen, L.; Lungren, M.; et al. Gloria: A multimodal global-local representation learning framework for label-efficient medical image recognition. In Proceedings of the IEEE/CVF international conference on computer vision, 2021. [Google Scholar]
- Pritam, N. A.; Jain, S.; et al. SkinGenBench: Generative model and preprocessing effects for synthetic dermoscopic augmentation in melanoma diagnosis. arXiv 2025, arXiv:2512.17585. [Google Scholar]
- S. Pedersen, S. Jain, M. Chavez, V. Ladehoff, B. N. de Freitas, and R. Pauwels, “Pano-GAN: A deep generative model for panoramic dental radiographs,” Journal of Imaging, vol. 11, no. 2, p. 41, 2025. [CrossRef] [PubMed]
- Jain, S.; de Freitas, B. N.; Basse-OConnor, A.; Iosifidis, A.; Pauwels, R. PanoDiff-SR: Synthesizing dental panoramic radiographs using diffusion and super-resolution. arXiv 2025, arXiv:2507.09227. [Google Scholar]

| Method | CheXpert | MIMIC-CXR | NIH-14 | GFLOPs | Memory (GB) |
|---|---|---|---|---|---|
| SimCLR | 0.79 | 0.784 | 0.772 | 72.5 | 3.2 |
| MoCo v3 | 0.80 | 0.788 | 0.776 | 70.8 | 3.1 |
| TokenLearner | 0.80 | 0.781 | 0.768 | 58.4 | 2.6 |
| DynamicViT | 0.80 | 0.792 | 0.779 | 53.7 | 2.4 |
| ConVIRT | 0.80 | 0.790 | 0.774 | 65.2 | 2.9 |
| GLoRIA | 0.80 | 0.793 | 0.781 | 68.3 | 3.0 |
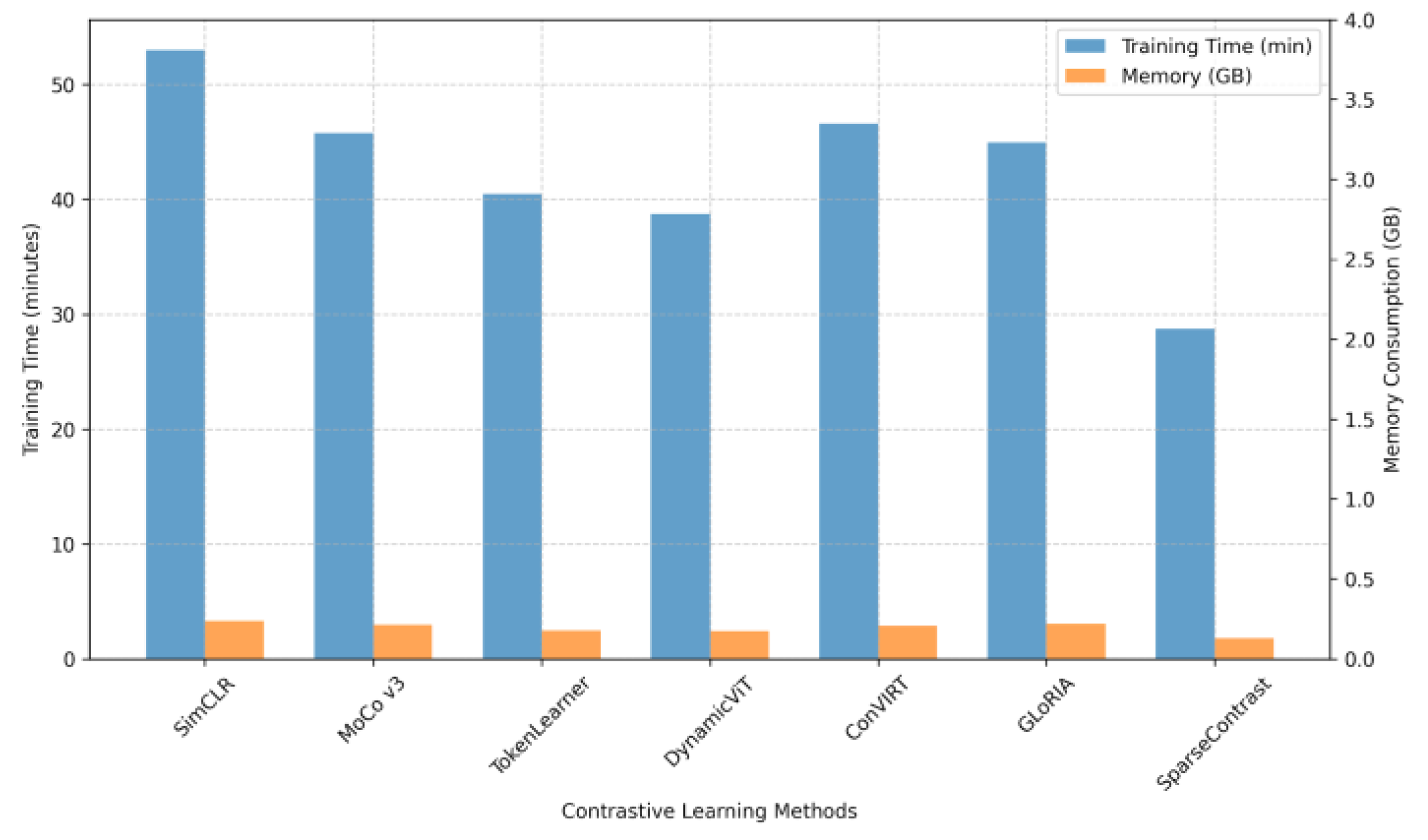
| SparseContrast | 0.81 | 0.797 | 0.785 | 41.2 | 1.9 |
| Variant | ρ=0.1 | ρ=0.3 | ρ=0.5 | Dense |
|---|---|---|---|---|
| Static | 0.782 | 0.792 | 0.799 | - |
| Dynamic | 0.789 | 0.812 | 0.808 | 0.798 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




