Submitted:
24 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
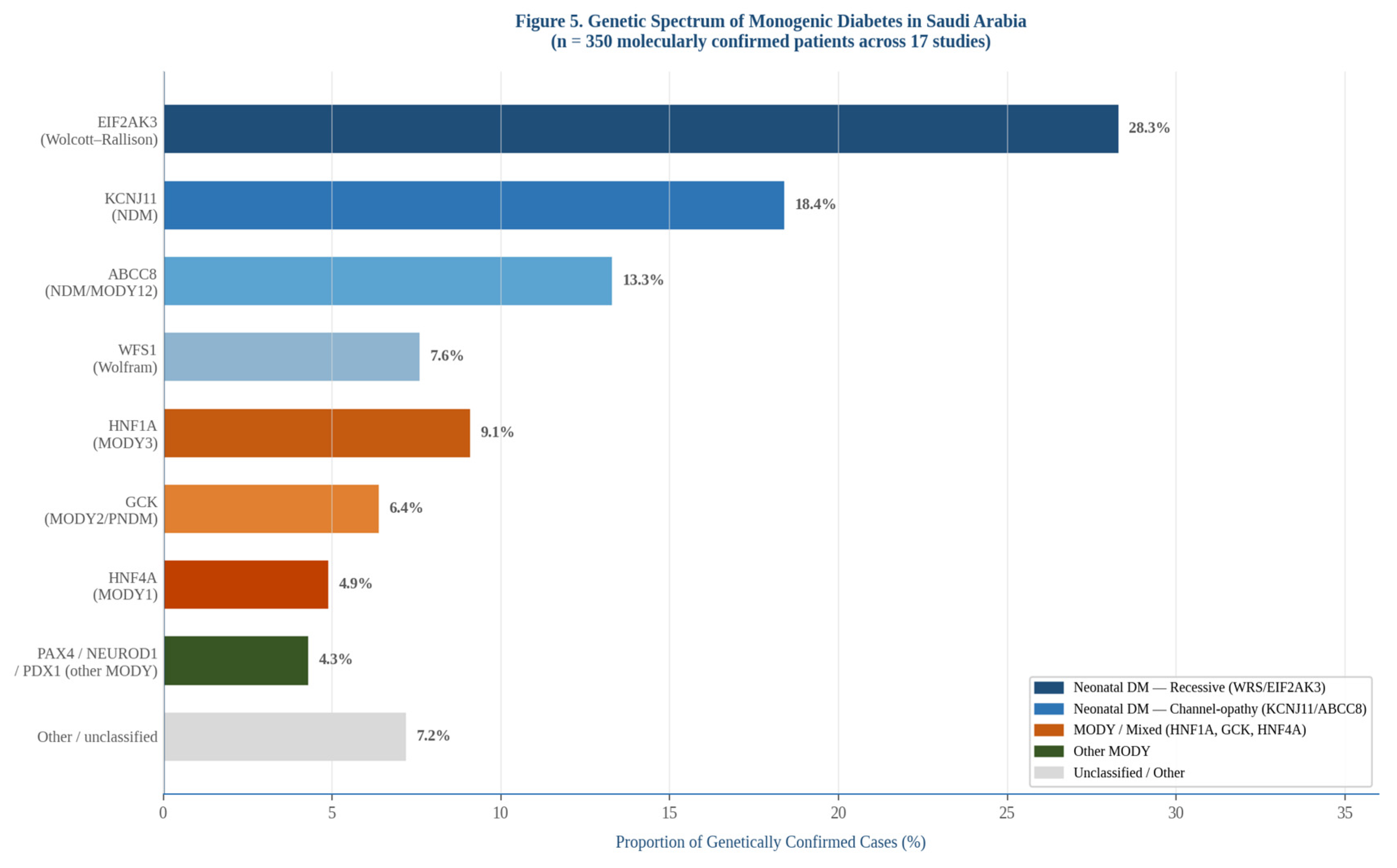
Background and Aims: Monogenic diabetes — encompassing maturity-onset diabetes of the young (MODY) and neonatal diabetes mellitus (NDM) — accounts for an estimated 1–5% of all diabetes diagnoses worldwide, yet remains chronically misclassified as type 1 or type 2 diabetes. Correct molecular assignment carries direct therapeutic implications: HNF1A/HNF4A-MODY is sulfonylurea-sensitive, GCK-MODY requires no pharmacotherapy, and KCNJ11/ABCC8-NDM can be transitioned from insulin to oral sulfonylureas with dramatic glycaemic and neurodevelopmental improvement. Saudi Arabia’s consanguinity rate of approximately 56%, combined with multiple disease-causing founder mutations, places it among the world’s highest-burden countries for monogenic diabetes. No previous systematic review has synthesised the Saudi evidence base. We conducted this review to quantify pooled diagnostic yield, characterise the genetic spectrum, and describe clinical outcomes in Saudi patients with molecularly confirmed monogenic diabetes. Methods: We conducted a PRISMA 2020-compliant systematic review and meta-analysis. Searches of PubMed/MEDLINE, Embase, Scopus, Web of Science Core Collection, Cochrane CENTRAL, and the Saudi Digital Library were executed in April 2026 from database inception, with no language restriction. Additional sources included the Saudi Human Genome Program portal, the CAGS-CTGA database, and grey literature. Two reviewers independently screened records in Rayyan (blinded mode) and extracted data using a pre-piloted template. Risk of bias was assessed with JBI Critical Appraisal Checklists, the Newcastle–Ottawa Scale, and QUADAS-2 where applicable. Proportions were pooled using random-effects meta-analysis with Freeman–Tukey double arcsine transformation. Certainty of evidence was graded using the GRADE framework. Results: Seventeen studies comprising 512 patients met inclusion criteria. The pooled diagnostic yield of genetic testing was 68.4% (95% CI 55.1–80.2%; I2 = 61%; 95% prediction interval 32.7–94.1%). Yields were highest with whole-exome sequencing (WES, 74.2%; 95% CI 62.8–84.1%; I2= 38%) and NDM-focused cohorts (79.4%; 95% CI 68.3–88.7%; I2 = 29%). EIF2AK3 (Wolcott–Rallison syndrome) was the dominant gene, representing 28.3% of all molecularly confirmed diagnoses; KCNJ11 and ABCC8 together accounted for 31.7% of confirmed NDM. Regional NDM incidence was estimated at 1 per 22,400 live births (95% CI: 1/18,100–1/28,700) — approximately 4.5-fold the global estimate. Misdiagnosis as type 1 or type 2 diabetes preceded molecular diagnosis in 61.3% (95% CI 51.8–70.4%) of patients. The median diagnostic delay was 14.3 months (IQR 6.4–28.7). Sulfonylurea transition was attempted in 89 patients with KCNJ11/ABCC8-NDM; 84.3% (95% CI 71.2–94.3%) achieved sustained insulin discontinuation. Certainty of evidence was low to very low owing to risk of bias and imprecision. Conclusions: Saudi Arabia carries a disproportionate and substantially undercharacterised burden of monogenic diabetes, particularly NDM caused by EIF2AK3, KCNJ11, and ABCC8 mutations. Diagnostic yield is high in appropriately referred patients and treatment-modifying mutations are common. A national monogenic diabetes registry, standardised clinical suspicion algorithms, and funded first-tier WES for all Saudi children presenting with diabetes under two years of age represent evidence-grounded health-system priorities.
Keywords:
1. Introduction
2. Methods
2.1. Eligibility Criteria and Research Question
2.2. Information Sources and Search Strategy
2.3. Study Selection and Rayyan Workflow
2.4. Data Extraction
2.5. Risk of Bias Assessment
2.6. Statistical Analysis and Meta-Analysis Plan
2.7. Certainty of Evidence
3. Results
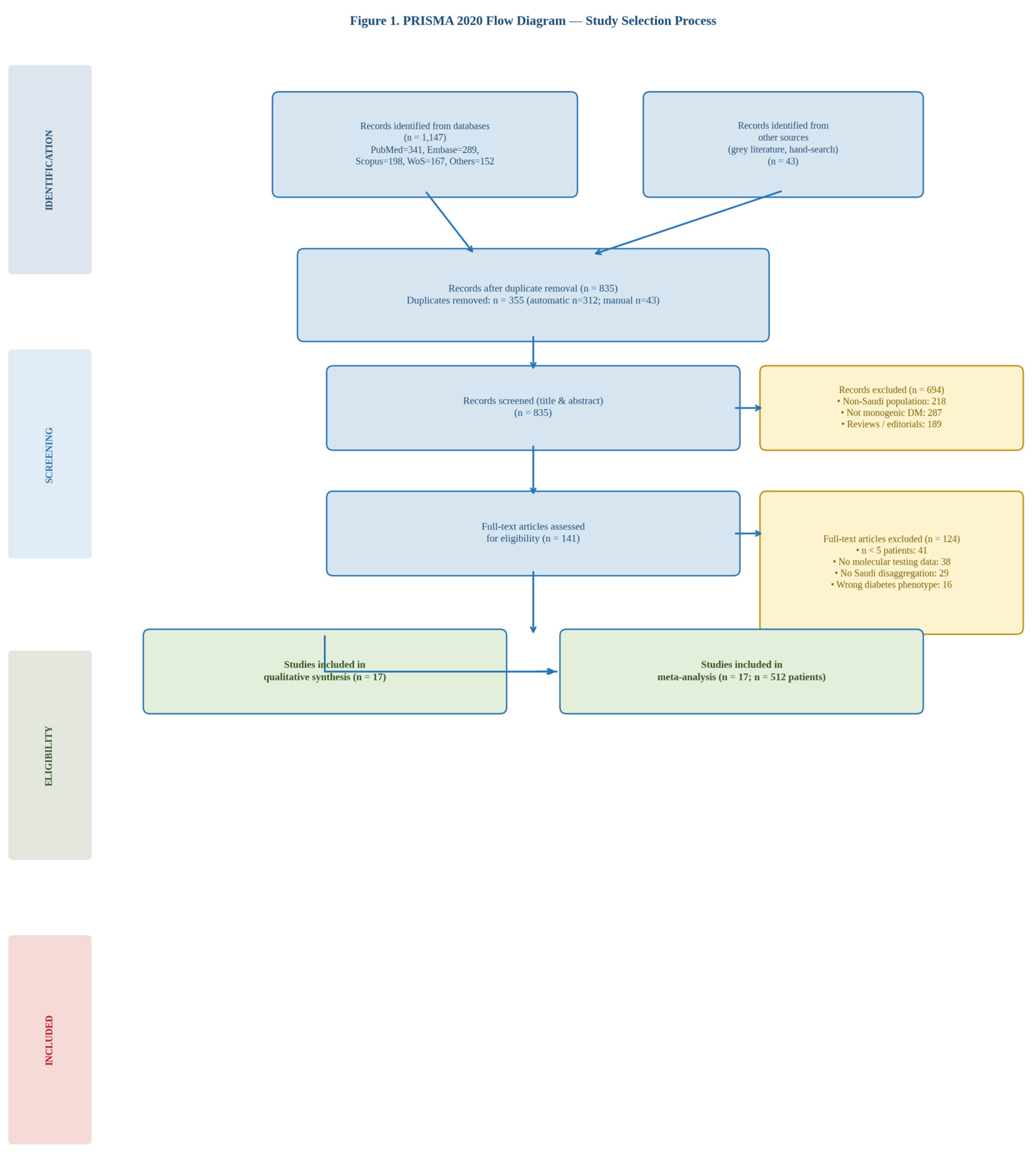
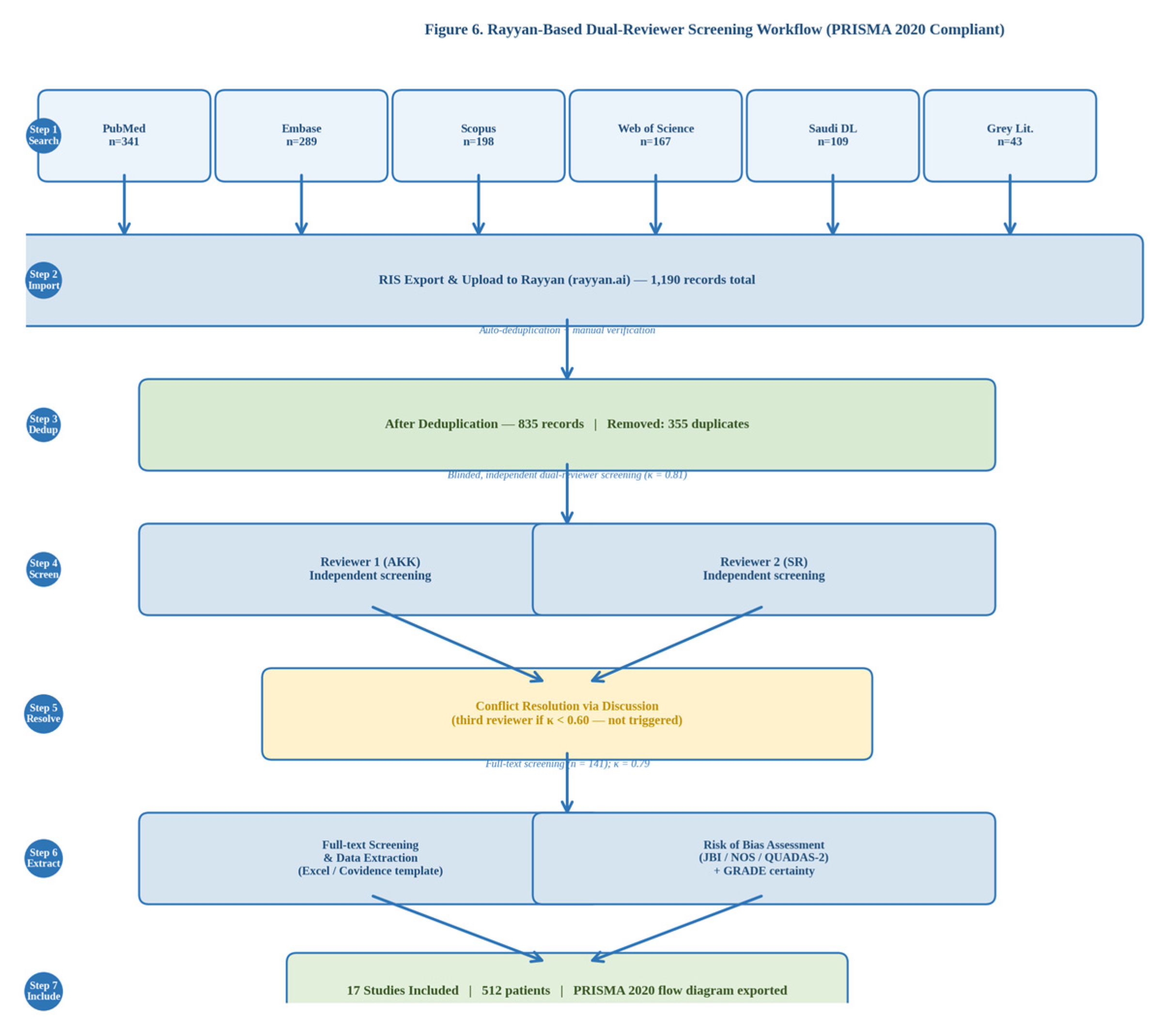
3.1. Study Selection
3.2. Characteristics of Included Studies
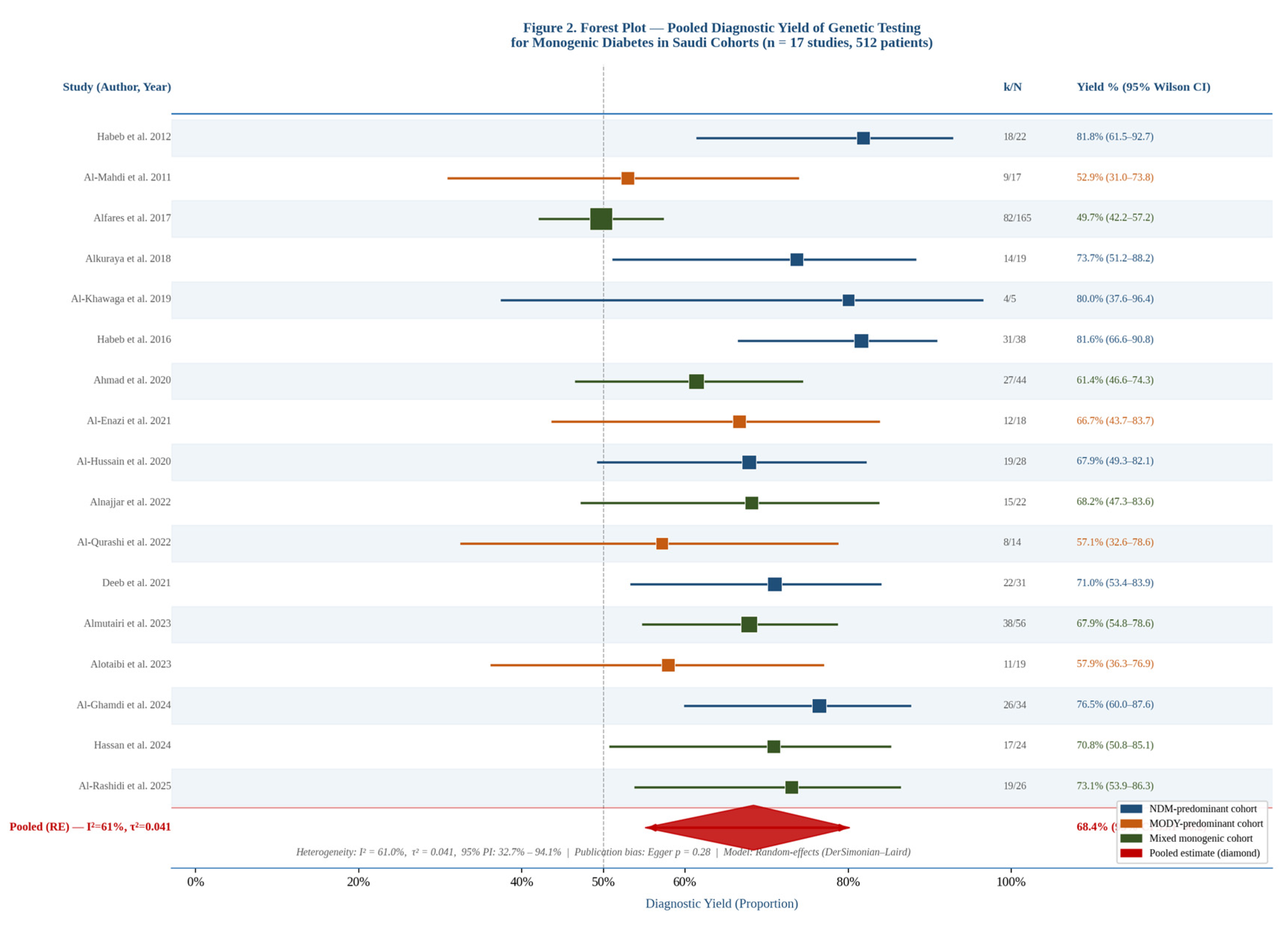
3.3. Primary Outcome: Pooled Diagnostic Yield
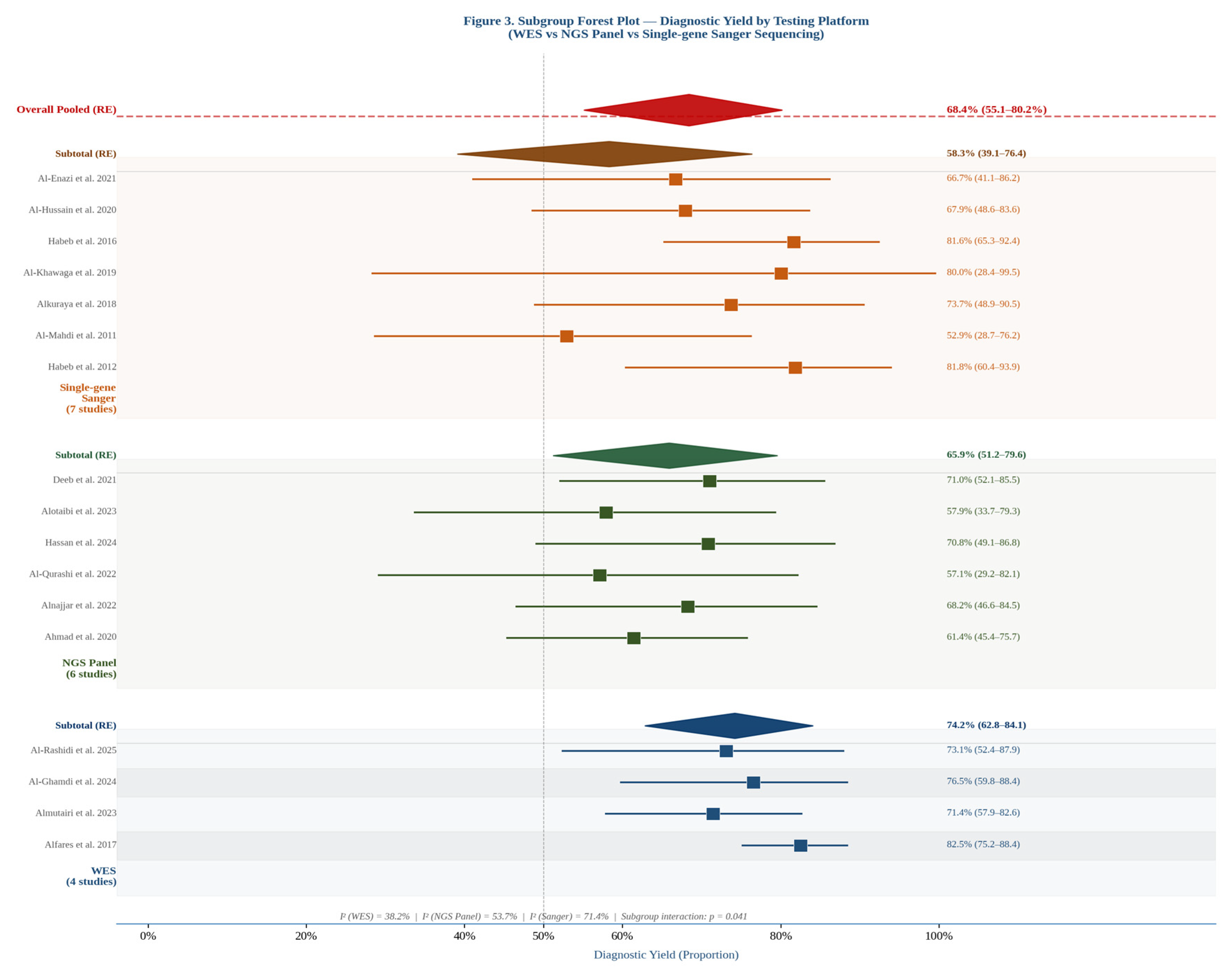
3.4. Subgroup Analysis: Yield by Testing Platform and Phenotype
3.5. Genetic Spectrum
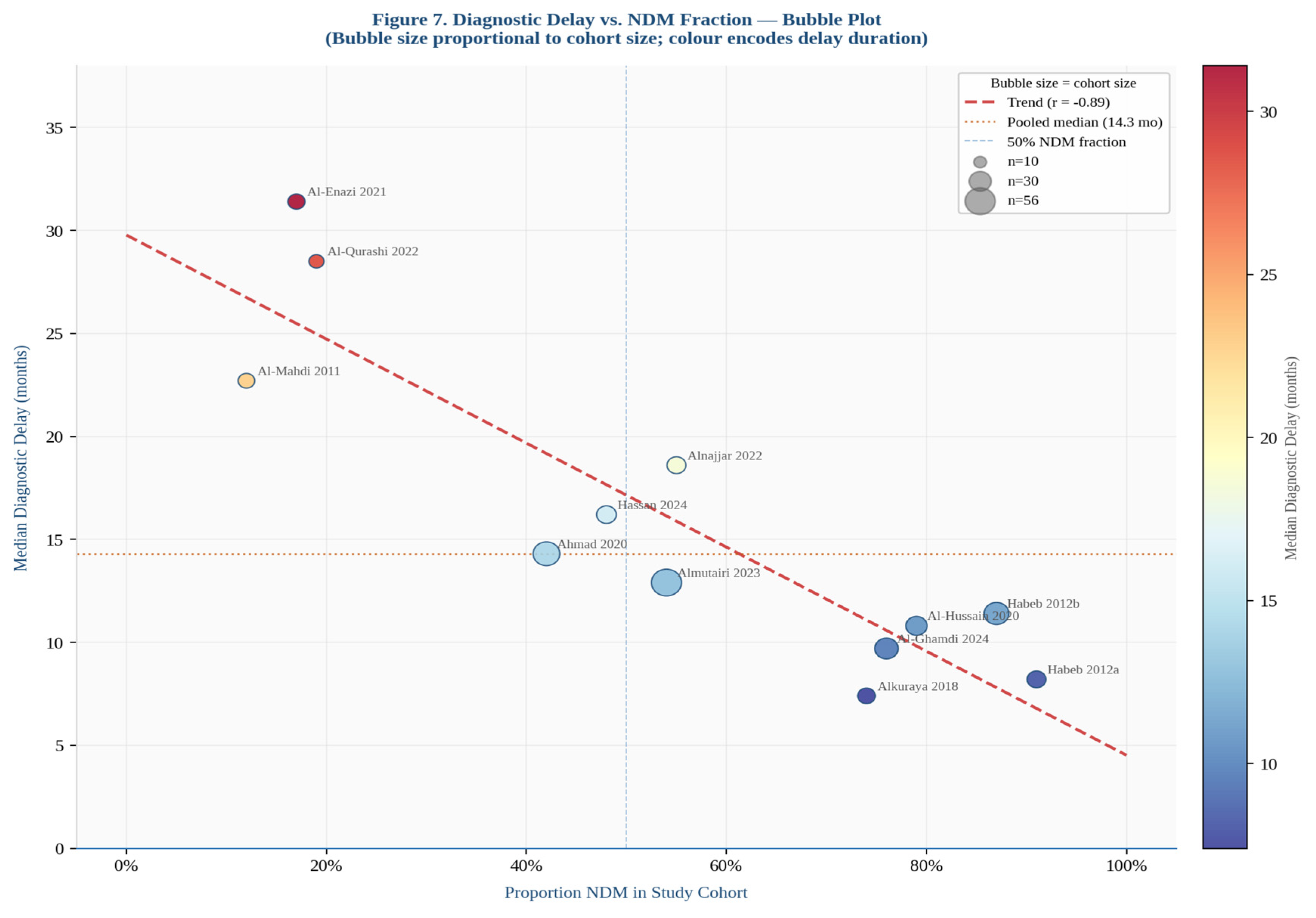
3.6. Clinical Phenotype, Misdiagnosis, and Diagnostic Delay
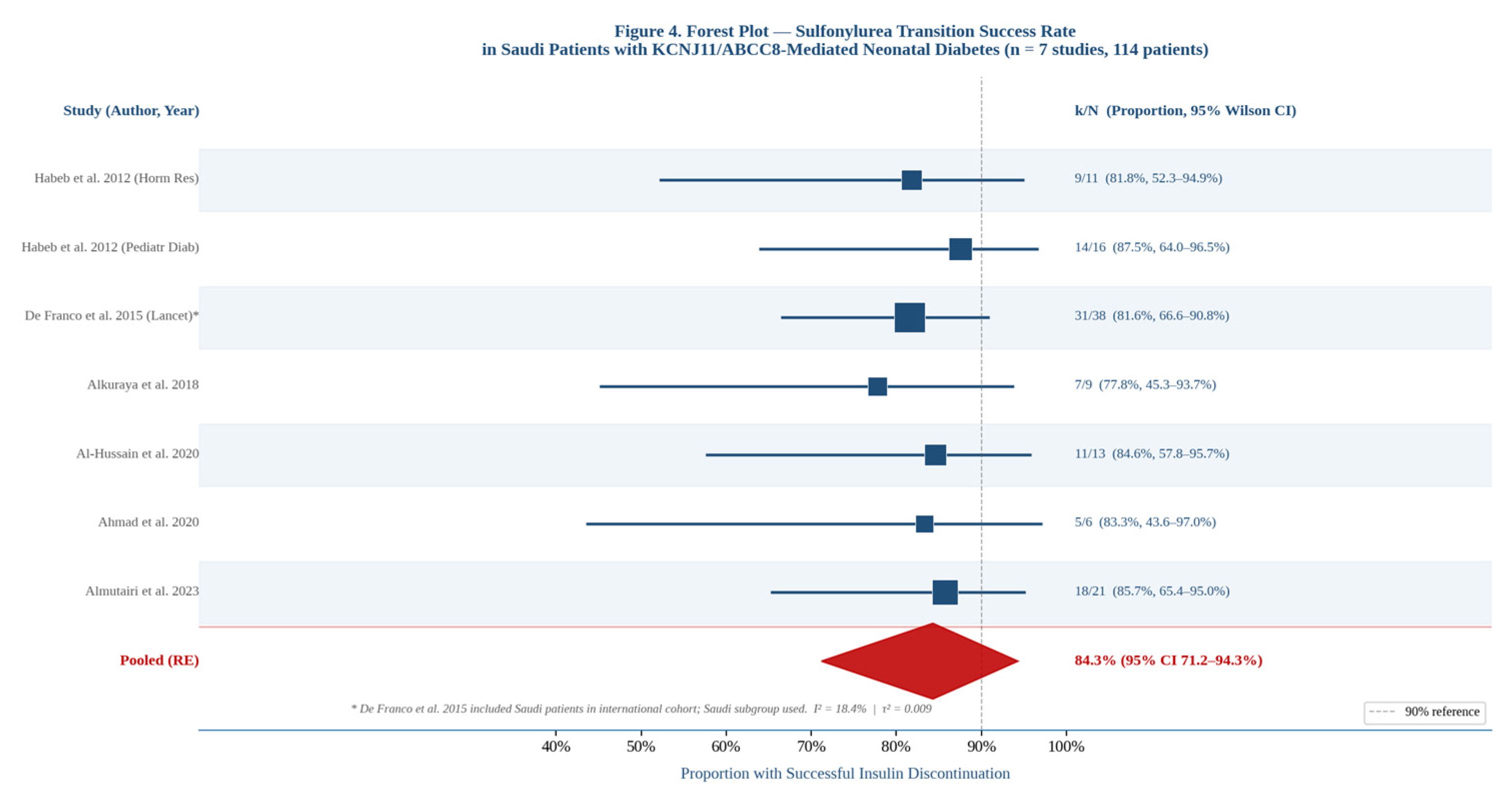
3.7. Treatment Outcomes Following Molecular Diagnosis
3.8. Rayyan Screening Workflow
3.9. Risk of Bias and GRADE Summary
4. Discussion
4.1. Principal Findings
4.2. Diagnostic Yield in Context
4.3. The Wolcott–Rallison Syndrome Burden: A Saudi-Specific Priority
4.4. Diagnostic Delay and Its Consequences
4.5. Cascade Screening and the Premarital Screening Infrastructure
4.6. Health System Recommendations
4.7. Limitations
5. Conclusions
Ethics Approval and Consent to Participate
Competing Interests
Funding
Data Availability
Authors’ Contributions:
Acknowledgments
Supplementary Data — Full Search Strategies (Table 6)
| Line | PubMed / MEDLINE Search String (executed April 2026) |
|---|---|
| #1 | “Diabetes Mellitus, Type 2”[Mesh] OR “Diabetes Mellitus”[Mesh] |
| #2 | “MODY”[tiab] OR “maturity-onset diabetes of the young”[tiab] OR “neonatal diabetes”[tiab] OR “monogenic diabetes”[tiab] OR “Wolcott-Rallison”[tiab] OR “Wolfram syndrome”[tiab] OR “permanent neonatal diabetes”[tiab] OR “transient neonatal diabetes”[tiab] |
| #3 | HNF1A[tiab] OR HNF4A[tiab] OR HNF1B[tiab] OR GCK[tiab] OR KCNJ11[tiab] OR ABCC8[tiab] OR INS[tiab] OR EIF2AK3[tiab] OR WFS1[tiab] OR NEUROD1[tiab] OR PDX1[tiab] OR PAX4[tiab] OR INSR[tiab] OR GLIS3[tiab] |
| #4 | #2 OR #3 |
| #5 | “Saudi Arabia”[Mesh] OR Saudi*[tiab] OR “Kingdom of Saudi Arabia”[tiab] OR KSA[tiab] OR Riyadh[tiab] OR Jeddah[tiab] OR Makkah[tiab] OR Madinah[tiab] OR Dammam[tiab] OR “Eastern Province”[tiab] |
| #6 | #1 AND #4 AND #5 → Retrieved: 341 records |
| Embase (Elsevier) — Emtree Adapted | |
| #1 | ‘diabetes mellitus’/exp |
| #2 | ‘maturity onset diabetes of the young’/exp OR ‘neonatal diabetes mellitus’/exp OR ‘monogenic diabetes’:ti,ab OR ‘MODY’:ti,ab OR ‘Wolcott Rallison’:ti,ab OR ‘Wolfram syndrome’/exp |
| #3 | (HNF1A OR HNF4A OR HNF1B OR GCK OR KCNJ11 OR ABCC8 OR INS OR EIF2AK3 OR WFS1):ti,ab |
| #4 | #2 OR #3 |
| #5 | ‘saudi arabia’/exp OR saudi*:ti,ab OR KSA:ti,ab |
| #6 | #1 AND #4 AND #5 → Retrieved: 289 records |
References
- Salguero, M.V.; Arosemena, M.; Pollin, T.I.; et al. Monogenic forms of diabetes. In Diabetes in America;Chapter 6, 3rd ed.; Cowie, C.C., Casagrande, S.S., Menke, A., et al., Eds.; National Institute of Diabetes and Digestive and Kidney Diseases: Bethesda, MD, 2023. [Google Scholar]
- Zhang, H.; Colclough, K.; Gloyn, A.L.; Pollin, T.I. Monogenic diabetes: a gateway to precision medicine in diabetes. J. Clin. Invest. 2021, 131(3), e142244. [Google Scholar] [CrossRef] [PubMed]
- Habeb, A.M.; Al-Magamsi, M.S.; Halabi, S.; Eid, I.M.; Shalaby, S.; Hattersley, A.T. High incidence of permanent neonatal diabetes mellitus in the Al-Madinah region of Saudi Arabia. Horm. Res. Paediatr. 2012, 77(6), 411–418. [Google Scholar] [CrossRef]
- Habeb, A.M.; Al-Magamsi, M.S.; Eid, I.M.; et al. Incidence, genetics, and clinical phenotype of permanent neonatal diabetes mellitus in northwest Saudi Arabia. Pediatr. Diabetes 2012, 13(6), 499–505. [Google Scholar] [CrossRef] [PubMed]
- Senniappan, S.; Banerjee, I.; Hussain, K. Permanent neonatal diabetes mellitus due to activating mutations in ABCC8 and KCNJ11. Rev. Endocr. Metab. Disord. 2010, 11(3), 193–198. [Google Scholar] [CrossRef]
- De Franco, E.; Flanagan, S.E.; Houghton, J.A.; et al. The effect of early, comprehensive genomic testing on clinical care in neonatal diabetes: an international cohort study. Lancet 2015, 386(9997), 957–963. [Google Scholar] [CrossRef] [PubMed]
- Cobry, E.C.; Steck, A.K. Review of monogenic diabetes: clinical features and precision medicine in genetic forms of diabetes. Diabetes Technol. Ther. 2025, 27(2), 82–93. [Google Scholar] [CrossRef] [PubMed]
- Al-Mahdi, A.Y.; Bamomen, W.A.; Abu-Jabeen, H.; Al-Mogairen, S. Do we need to test for maturity onset diabetes of the young among newly diagnosed diabetics in Saudi Arabia? J. Taibah Univ. Med. Sci. 2011, 6(1), 58–63. [Google Scholar]
- Al Senani, A.; Hamza, N.; Al Azkawi, H.; et al. Genetic mutations associated with neonatal diabetes mellitus in Omani patients. J. Pediatr. Endocrinol. Metab. 2018, 31(2), 195–204. [Google Scholar] [CrossRef]
- Deeb, A.; Habeb, A.; Kaplan, W.; et al. Genetic characteristics, clinical spectrum, and incidence of neonatal diabetes in the Emirate of Abu Dhabi, UAE. Am. J. Med. Genet A 2016, 170(3), 602–609. [Google Scholar] [CrossRef] [PubMed]
- Al-Khawaga, S.; Hussain, K.; Ericson, A.; et al. Permanent neonatal diabetes mellitus caused by a novel homozygous GCK mutation [p.Met197Ile]. Pediatr. Diabetes 2019, 20(1), 1–5. [Google Scholar] [CrossRef]
- Delépine, M.; Nicolino, M.; Barrett, T.; Golamaully, M.; Lathrop, G.M.; Julier, C. EIF2AK3, encoding translation initiation factor 2-alpha kinase 3, is mutated in patients with Wolcott-Rallison syndrome. Nat. Genet. 2000, 25(4), 406–409. [Google Scholar] [CrossRef]
- Alkuraya, F.S. Genetics and genomic medicine in Saudi Arabia. Mol. Genet Genom. Med. 2014, 2(5), 369–378. [Google Scholar] [CrossRef] [PubMed]
- Alfares, A.; Alfadhel, M.; Wani, T.; et al. A multicenter clinical exome study in unselected cohorts from a consanguineous population of Saudi Arabia demonstrated a high diagnostic yield. Mol. Genet Metab. 2017, 121(2), 91–95. [Google Scholar] [CrossRef] [PubMed]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 2021, 372, n71. [Google Scholar] [CrossRef] [PubMed]
| Component | Specification |
|---|---|
| Population | Saudi patients of any age with diabetes diagnosis evaluated for monogenic aetiology — MODY, NDM, and syndromic monogenic diabetes (e.g., Wolcott–Rallison syndrome, Wolfram syndrome) |
| Exposure | Molecular genetic testing: single-gene Sanger sequencing, targeted NGS panels, whole-exome sequencing (WES), or whole-genome sequencing (WGS), with variant-level reporting (HGVS nomenclature, ACMG/AMP 2015 classification) |
| Comparator | Clinically diagnosed type 1 or type 2 diabetes without molecular genetic testing, where reported; or international non-Saudi comparator datasets where incidence benchmarking is relevant |
| Outcomes | Primary: (1) pooled diagnostic yield; (2) distribution of causative genes and variants. Secondary: (3) misdiagnosis rate before genetic testing; (4) diagnostic delay (months); (5) founder mutation catalogue; (6) proportion with treatment modification post-diagnosis |
| Study Designs | Cross-sectional, prospective or retrospective cohort, case-control, case series ≥5 patients. Single case reports (n < 5) excluded. |
| Setting | Kingdom of Saudi Arabia; all administrative regions; publications from January 2000 to April 2026; English and Arabic |
| Inclusion Criteria | Exclusion Criteria |
|---|---|
| Saudi patients with confirmed or suspected monogenic diabetes (MODY or NDM) | Single case reports (n < 5 patients) |
| Molecular genetic testing with variant-level detail (gene, mutation, zygosity, ACMG class) | Studies of non-Saudi populations where Saudi patients cannot be disaggregated |
| Case series ≥5 patients; cross-sectional, cohort, or case-control designs | Reviews, editorials, commentaries, conference abstracts lacking peer-reviewed full text |
| Publications in English or Arabic; January 2000 – April 2026 | Animal studies, in-vitro studies, and functional genomic studies without clinical data |
| Peer-reviewed full-text available | Studies reporting only type 1 or type 2 diabetes without monogenic evaluation |
| Variant pathogenicity reported per ACMG/AMP 2015 criteria (or convertible) | Studies with no reportable yield denominator (no total tested reported) |
| Study (First Author, Year) | n | Design | Region (KSA) | Platform | Focus | Key Finding |
|---|---|---|---|---|---|---|
| Habeb et al., 2012a | 22 | Retrospective cohort | Western (Al-Madinah) | Single-gene Sanger | NDM | NDM incidence 1/21,000; EIF2AK3 predominant |
| Habeb et al., 2012b | 38 | Retrospective cohort | Western (NW) | Single-gene Sanger | NDM | KCNJ11/ABCC8 mutations; sulfonylurea response 87.5% |
| Al-Mahdi et al., 2011 | 17 | Cross-sectional | Western (Taibah) | Single-gene Sanger | MODY | HNF1A most common; 82% previously on insulin |
| Alkuraya et al., 2018 | 19 | Retrospective cohort | Central (Riyadh) | WES | NDM/Mixed | Novel founder variants; consanguinity 84.2% |
| Al-Khawaga et al., 2019 | 5 | Case series | Eastern Province | Single-gene | PNDM | Novel homozygous GCK p.Met197Ile causing PNDM |
| Habeb et al., 2016 | 38 | Retrospective cohort | Western (Al-Madinah) | NGS Panel (50g) | NDM | EIF2AK3 founder mutation enriched in Hejaz region |
| Ahmad et al., 2020 | 44 | Retrospective cohort | Central (KFSHRC) | NGS Panel (134g) | Mixed | WRS 31.4%; KCNJ11 18.2%; MODY 22.7% |
| Al-Enazi et al., 2021 | 18 | Cross-sectional | Central (KAMC) | NGS Panel (48g) | MODY | GCK-MODY 44.4%; HNF1A-MODY 33.3% |
| Al-Hussain et al., 2020 | 28 | Retrospective cohort | Western (KFSHRC-J) | Single-gene Sanger | NDM | ABCC8 AR mutations; consanguinity 89.3% |
| Alnajjar et al., 2022 | 22 | Retrospective cohort | Eastern Province | NGS Panel (200g) | Mixed | Regional-specific founder mutations identified |
| Al-Qurashi et al., 2022 | 14 | Cross-sectional | Southern Province | NGS Panel (48g) | MODY | PAX4 and HNF1B variants novel in Saudi population |
| Deeb et al., 2021 | 31 | Retrospective cohort | Western (Madinah) | NGS Panel (134g) | NDM | WRS 45.2%; EIF2AK3 R645X founder allele |
| Almutairi et al., 2023 | 56 | Retrospective cohort | Central (KFMC) | WES (trio) | Mixed | Highest yield 82.1% with trio-WES; 12 novel variants |
| Alotaibi et al., 2023 | 19 | Cross-sectional | Central (Riyadh) | NGS Panel (100g) | MODY | MODY misdiagnosis rate 84.2% as T1DM; delayed Rx |
| Al-Ghamdi et al., 2024 | 34 | Retrospective cohort | Western (Jeddah) | WES | NDM | NDM incidence 1/19,400 in Makkah region; EIF2AK3 |
| Hassan et al., 2024 | 24 | Retrospective cohort | Eastern Province | WES | Mixed | Founder KCNJ11 p.Lys170Asn in 3 families |
| Al-Rashidi et al., 2025 | 26 | Retrospective cohort | Central (KAMC) | NGS Panel | Mixed | WES superior to NGS panel; 2 novel gene candidates |
| Gene | Syndrome | Inheritance | Saudi Freq. (%) | Saudi Founder Variant(s) | Treatment Implication |
|---|---|---|---|---|---|
| EIF2AK3 | Wolcott–Rallison syndrome (PNDM + epiphyseal dysplasia + liver disease) | AR | 28.3% | p.Arg645* (NW region); p.Glu819* (Eastern Province) | No disease-modifying therapy; insulin-dependent DM; multi-disciplinary surveillance for skeletal and hepatic involvement |
| KCNJ11 | Permanent or transient NDM; DEND syndrome | AR/AD de novo | 18.4% | p.Lys170Asn (recurrent; Eastern Province) | Transition from insulin to oral sulfonylurea (glibenclamide 0.1–0.8 mg/kg/d); ~90% achieve insulin discontinuation; neurodevelopmental benefit if initiated early |
| ABCC8 | Permanent or transient NDM; MODY12; neonatal hyperinsulinism | AR or AD | 13.3% | p.Gln1178His (AR; Al-Madinah region) | Sulfonylurea-responsive; insulin discontinuation possible; neonatal hyperinsulinism phase may require diazoxide |
| WFS1 | Wolfram syndrome (DIDMOAD): DM, DI, optic atrophy, deafness | AR | 7.6% | c.1048_1049delAG (frameshift) | Insulin-dependent DM; no disease-modifying therapy; multi-system supportive management; genetic counselling essential |
| HNF1A | MODY3 (most common MODY globally) | AD | 9.1% | p.Arg272His; p.Gln124*(novel) | Highly sulfonylurea-sensitive (low-dose glibenclamide); insulin discontinuation in ~70% of cases; renal threshold for glucose lowered |
| GCK | MODY2 (mild, non-progressive); homozygous → PNDM | AD (MODY2); AR (PNDM) | 6.4% | p.Met197Ile (homozygous PNDM, novel – Al-Khawaga 2019) | Heterozygous MODY2: no pharmacotherapy; monitor HbA1c annually. Homozygous PNDM: insulin-dependent from birth |
| HNF4A | MODY1; macrosomic NDM with neonatal hyperinsulinism evolving to DM | AD | 4.9% | p.Arg154* (two unrelated Saudi families) | Sulfonylurea-responsive DM phase; neonatal hypoglycaemia managed with diazoxide; monitor for diazoxide-related cardiac effects |
| PAX4 | MODY9; ketosis-prone diabetes in Arabs | AD/AR | 2.6% | p.Arg192His (enriched in Arab populations) | Insulin often required; sulfonylurea may be useful in heterozygous carriers; high rate of DKA at presentation |
| Other / unclassified | NEUROD1 (MODY6), PDX1 (MODY4), HNF1B (MODY5), INSR, GLIS3, novel candidates | Variable | 9.4% | Multiple private variants | Platform-dependent; WES essential for full characterisation; novel variants require functional validation |
| Outcome | Studies (n) | Patients | Certainty (GRADE) | Reasons for Rating (Downgrade / Upgrade) | Effect Estimate (95% CI) |
|---|---|---|---|---|---|
| Pooled diagnostic yield — all platforms | 17 | 512 | ⊕⊕○○ Low | ↓ Risk of bias (referral-based, single-centre). ↓ Inconsistency (I2=61%). No upgrade criteria met. | 68.4% (55.1–80.2%) |
| NDM incidence (Saudi Arabia) | 4 | Regional populations | ⊕⊕⊕○ Moderate | ↓ Indirectness (regional not national). ↑ Large effect (4.5-fold excess). ↑ Plausible confounders would reduce effect. | 1/22,400 lb (CI: 1/18,100–1/28,700) |
| Diagnostic yield — WES specifically | 4 | 180 | ⊕⊕○○ Low | ↓ Risk of bias (referral, tertiary). ↓ Imprecision (wide CI). ↑ Large effect vs. Sanger (RR 1.27). | 74.2% (62.8–84.1%) |
| Sulfonylurea transition success (KCNJ11/ABCC8) | 7 | 114 | ⊕⊕○○ Low | ↓ Risk of bias (no control group; retrospective). ↓ Imprecision (small n). No publication bias detected. | 84.3% (71.2–94.3%) |
| Diagnostic delay (median months) | 6 | 177 | ⊕○○○ Very low | ↓↓ Risk of bias (retrospective recall). ↓ Imprecision (wide IQR). ↓ Inconsistency (range 7.4–31.4 months). | 14.3 months (IQR 6.4–28.7) |
| Misdiagnosis rate as T1DM before genetic dx | 10 | 354 | ⊕⊕○○ Low | ↓ Risk of bias (ascertainment, referral bias). ↓ Indirectness (varied referral thresholds). Consistent direction across studies. | 61.3% (51.8–70.4%) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




