Background/Objectives: Parvovirus infection cause sever diseases in both feline and ca-nine species, mostly affects adult cats and dogs, but cause higher risk in the kittens and puppies. This virus is known to be contagious; the simple way of spreading occurs through food and shelter as well as hands and clothing of people. The recovered species may continue to shed parvovirus in their feces for an extended period, leading to severe environmental contamination. There is no universal vaccine available that protect dogs and cats against parvovirus infections. The main goal of this study is to design a pan parvovirus multiepitope DNA based vaccine that could be administered to dog and cats.
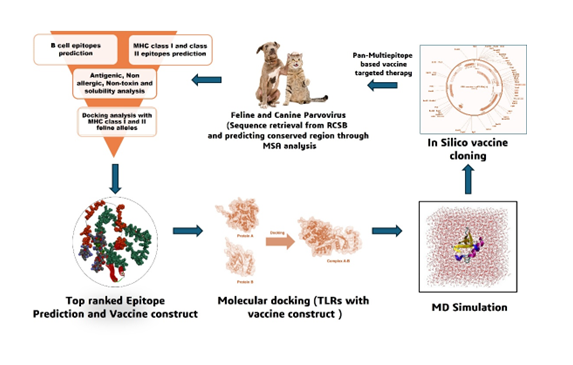
Methods: We utilized AI-machine learning incorporated server tools such as IEDB and NetMHCpan to predict B-cell and T-cell epitopes. VaxiJen and ToxinPred were used to analyze immune characteristic features and docking with feline alleles using HADdock server. Following, the immune response and stability of vaccine construct was confirmed with disulfide engineering, Normal mode analysis and molecular docking, and dynamic simulations were done with Toll like receptors of both feline and canine (TLR4 and TLR5) for 50ns. The triggered immune response was determined with immuno-simulation (ImmSim) and their activity in biological environment was reinforced with in silico cloning.
Results: The B cell epitopes (NS1 - 9, NS2 - 4, VP1- 12 and VP2- 9) predicted with IEDB database were subjected to antigenicity prediction. MHC class I and IFN prediction and MHC class II and IL-4 prediction were done with IEDB and NetMHCpan. The T cell epitopes showed high binding affinities with the feline alleles. The final vaccine was de-signed by combining the top-ranked B-cell epitopes T- cell epitopes, filtered from high antigenic, non-allergic, non-toxic and good solubility, and with the better binding affinity score of the structural and non- structural proteins (NS1, NS2, VP1 and VP2) of feline and canine parvoviruses through linkers and adjuvants. The disulfide bond prediction and Normal mode analysis showed our vaccine construct are stable and flexible. The molecular docking analysis was performed between the designed vaccine epitopes and the TLRs (TLR4 – feline and TLR5 – canine) with Biovia Discovery Studio using Zdocker, it showed the better binding interaction with value of 22.26 (Zdock score), -47.409 (Zrank score) for feline and 16.54 (Zdock score), -134.295 (Zrank score) for canine.
Conclusions: The pan multi-epitope DNA based vaccine combining the four major structural proteins (NS1, NS2, VP1 and VP2) possess dual purpose to protect both feline and canine species against the parvovirus were designed and constructed. The molecular docking and dynamic simulation analysis showed higher binding affinities and stable conformation with canine (TLR5) and feline (TLR4) toll like receptors. Though computational analysis will support us to predict the more precise top-ranked epitopes and their immuno-antigenic properties, further experimental validation will be required to be used against those viruses.